

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122162.2 - phase: 0 /pseudo
(547 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
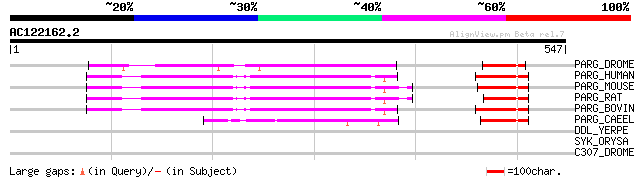
Score E
Sequences producing significant alignments: (bits) Value
PARG_DROME (O46043) Poly(ADP-ribose) glycohydrolase (EC 3.2.1.143) 194 6e-49
PARG_HUMAN (Q86W56) Poly(ADP-ribose) glycohydrolase (EC 3.2.1.143) 172 2e-42
PARG_MOUSE (O88622) Poly(ADP-ribose) glycohydrolase (EC 3.2.1.143) 171 4e-42
PARG_RAT (Q9QYM2) Poly(ADP-ribose) glycohydrolase (EC 3.2.1.143) 167 6e-41
PARG_BOVIN (O02776) Poly(ADP-ribose) glycohydrolase (EC 3.2.1.143) 167 8e-41
PARG_CAEEL (Q867X0) Poly(ADP-ribose) glycohydrolase pme-3 (EC 3.... 106 2e-22
DDL_YERPE (Q8ZIE7) D-alanine--D-alanine ligase (EC 6.3.2.4) (D-a... 32 5.6
SYK_ORYSA (Q6F2U9) Lysyl-tRNA synthetase (EC 6.1.1.6) (Lysine--t... 31 7.3
C307_DROME (Q9VRM7) Probable cytochrome P450 307a1 (EC 1.14.-.-)... 31 7.3
>PARG_DROME (O46043) Poly(ADP-ribose) glycohydrolase (EC 3.2.1.143)
Length = 768
Score = 194 bits (492), Expect = 6e-49
Identities = 124/321 (38%), Positives = 176/321 (54%), Gaps = 51/321 (15%)
Query: 78 DELISREECRKWFEEVLPSLGDLLLRLPSLLEA------HYENADMVIDGKGATVRTGLR 131
DE + E R +FE++LP + L LRLP L+++ H++NA +
Sbjct: 197 DEELDESETRVFFEDLLPRIIRLALRLPDLIQSPVPLLKHHKNASLS------------- 243
Query: 132 MLDSQEAGIVFLTQELIAALLACSFLCLFPVHDRYEKQLQPVNFDEL-FASLYNDYTQKQ 190
L+Q+ I+ LLA +FLC FP + +++ + F ++ F LY
Sbjct: 244 -----------LSQQQISCLLANAFLCTFPRRNTLKRKSEYSTFPDINFNRLYQSTGPAV 292
Query: 191 EDKIWCIIHYFQRI------TSNMPKGVVSFERKV-LPWEDDCIHISYPNASFWSTSVKP 243
+K+ CI+HYF+R+ SN+P GVV+F R+ LP + WS S P
Sbjct: 293 LEKLKCIMHYFRRVCPTERDASNVPTGVVTFVRRSGLP----------EHLIDWSQSAAP 342
Query: 244 L--CRFEVKSSGLIEDHSSETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIAGMIFL 301
L V + G IED ++VDFAN+YLGGG L GCVQEEIRF+I PEL+ G +F
Sbjct: 343 LGDVPLHVDAEGTIEDEGIGLLQVDFANKYLGGGVLGHGCVQEEIRFVICPELLVGKLFT 402
Query: 302 PSMADNEAIDIVGVERFSSYTGYASSFRFSGDYVDDKEVDPFGRRKTRIVAIDAL-CGPG 360
+ EA+ ++G ER+S+YTGYA SF +SG++ D D GRR+T IVAIDAL
Sbjct: 403 ECLRPFEALVMLGAERYSNYTGYAGSFEWSGNFEDSTPRDSSGRRQTAIVAIDALHFAQS 462
Query: 361 MRQYREKFLLREINKAFCGFL 381
QYRE + RE+NKA+ GF+
Sbjct: 463 HHQYREDLMERELNKAYIGFV 483
Score = 63.2 bits (152), Expect = 2e-09
Identities = 27/42 (64%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 467 GVATGNWGCGAFGGDPEVKTIIQWLAASQAGRPFIAYYTFGS 508
GVATGNWGCGAFGGD +K ++Q + +Q GRP +AYYTFG+
Sbjct: 492 GVATGNWGCGAFGGDSYLKALLQLMVCAQLGRP-LAYYTFGN 532
>PARG_HUMAN (Q86W56) Poly(ADP-ribose) glycohydrolase (EC 3.2.1.143)
Length = 976
Score = 172 bits (436), Expect = 2e-42
Identities = 107/311 (34%), Positives = 166/311 (52%), Gaps = 36/311 (11%)
Query: 76 FFDELISREECRKWFEEVLPSLGDLLLRLPSLLEAHYENADMVIDGKGATVRTGLRMLDS 135
F+D+++ E + ++ +LP + + L LP++ + +L
Sbjct: 576 FWDKVLEEAEAQHLYQSILPDMVKIALCLPNICTQP------------------IPLLKQ 617
Query: 136 QEAGIVFLTQELIAALLACSFLCLFPVHDRYEKQLQPVNFDELFASLYNDYTQKQEDKIW 195
+ + ++QE IA+LLA +F C FP + K D F L+ + ++ +K+
Sbjct: 618 KMNHSITMSQEQIASLLANAFFCTFPRRNAKMKSEYSSYPDINFNRLFEGRSSRKPEKLK 677
Query: 196 CIIHYFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASFWSTSVKPLCRFEVKSSGLI 255
+ YF+R+T P G+V+F R+ L ED +P W KPL R V G I
Sbjct: 678 TLFCYFRRVTEKKPTGLVTFTRQSL--ED------FPE---WERCEKPLTRLHVTYEGTI 726
Query: 256 EDHSSETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIAGMIFLPSMADNEAIDIVGV 315
E++ ++VDFAN ++GGG G VQEEIRF+I+PELI +F + NE + I G
Sbjct: 727 EENGQGMLQVDFANRFVGGGVTSAGLVQEEIRFLINPELIISRLFTEVLDHNECLIITGT 786
Query: 316 ERFSSYTGYASSFRFSGDYVDDKEVDPFGRRKTRIVAIDALCGPGMRQYREKF----LLR 371
E++S YTGYA ++R+S + D E D + RR T IVAIDAL R+Y ++F + R
Sbjct: 787 EQYSEYTGYAETYRWSRSHEDGSERDDWQRRCTEIVAIDAL---HFRRYLDQFVPEKMRR 843
Query: 372 EINKAFCGFLQ 382
E+NKA+CGFL+
Sbjct: 844 ELNKAYCGFLR 854
Score = 58.2 bits (139), Expect = 6e-08
Identities = 28/52 (53%), Positives = 35/52 (66%), Gaps = 1/52 (1%)
Query: 460 MENSNDIGVATGNWGCGAFGGDPEVKTIIQWLAASQAGRPFIAYYTFGSGAL 511
+ + N VATGNWGCGAFGGD +K +IQ LAA+ A R + Y+TFG L
Sbjct: 857 VSSENLSAVATGNWGCGAFGGDARLKALIQILAAAAAERD-VVYFTFGDSEL 907
>PARG_MOUSE (O88622) Poly(ADP-ribose) glycohydrolase (EC 3.2.1.143)
Length = 969
Score = 171 bits (433), Expect = 4e-42
Identities = 110/326 (33%), Positives = 170/326 (51%), Gaps = 43/326 (13%)
Query: 76 FFDELISREECRKWFEEVLPSLGDLLLRLPSLLEAHYENADMVIDGKGATVRTGLRMLDS 135
F+D+++ E + ++ +LP + + L LP++ + +L
Sbjct: 569 FWDKVLEEAEAQHLYQSILPDMVKIALCLPNICTQP------------------IPLLKQ 610
Query: 136 QEAGIVFLTQELIAALLACSFLCLFPVHDRYEKQLQPVNFDELFASLYNDYTQKQEDKIW 195
+ V ++QE IA+LLA +F C FP + K D F L+ + ++ +K+
Sbjct: 611 KMNHSVTMSQEQIASLLANAFFCTFPRRNAKMKSEYSSYPDINFNRLFEGRSSRKPEKLK 670
Query: 196 CIIHYFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASFWSTSVKPLCRFEVKSSGLI 255
+ YF+R+T P G+V+F R+ L ED +P W KPL R V G I
Sbjct: 671 TLFCYFRRVTEKKPTGLVTFTRQSL--ED------FPE---WERCEKPLTRLHVTYEGTI 719
Query: 256 EDHSSETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIAGMIFLPSMADNEAIDIVGV 315
E + ++VDFAN ++GGG G VQEEIRF+I+PELI +F + NE + I G
Sbjct: 720 EGNGRGMLQVDFANRFVGGGVTGAGLVQEEIRFLINPELIVSRLFTEVLDHNECLIITGT 779
Query: 316 ERFSSYTGYASSFRFSGDYVDDKEVDPFGRRKTRIVAIDALCGPGMRQYREKF----LLR 371
E++S YTGYA ++R++ + D E D + RR T IVAIDAL R+Y ++F + R
Sbjct: 780 EQYSEYTGYAETYRWARSHEDGSEKDDWQRRCTEIVAIDAL---HFRRYLDQFVPEKVRR 836
Query: 372 EINKAFCGFLQQSQYQRDQKIPQENV 397
E+NKA+CGFL+ +P EN+
Sbjct: 837 ELNKAYCGFLRPG-------VPSENL 855
Score = 57.8 bits (138), Expect = 7e-08
Identities = 28/50 (56%), Positives = 34/50 (68%), Gaps = 1/50 (2%)
Query: 462 NSNDIGVATGNWGCGAFGGDPEVKTIIQWLAASQAGRPFIAYYTFGSGAL 511
+ N VATGNWGCGAFGGD +K +IQ LAA+ A R + Y+TFG L
Sbjct: 852 SENLSAVATGNWGCGAFGGDARLKALIQILAAAAAERD-VVYFTFGDSEL 900
>PARG_RAT (Q9QYM2) Poly(ADP-ribose) glycohydrolase (EC 3.2.1.143)
Length = 972
Score = 167 bits (423), Expect = 6e-41
Identities = 109/326 (33%), Positives = 169/326 (51%), Gaps = 43/326 (13%)
Query: 76 FFDELISREECRKWFEEVLPSLGDLLLRLPSLLEAHYENADMVIDGKGATVRTGLRMLDS 135
F+D+++ E + ++ +LP + + L LP++ + +L
Sbjct: 572 FWDKVLEEAEAQHLYQSILPDMVKIALCLPNICTQP------------------IPLLKQ 613
Query: 136 QEAGIVFLTQELIAALLACSFLCLFPVHDRYEKQLQPVNFDELFASLYNDYTQKQEDKIW 195
+ V ++QE IA+LLA +F C FP + K D F L+ + ++ +K+
Sbjct: 614 KMNHSVTMSQEQIASLLANAFFCTFPRRNAKMKSEYSSYPDINFNRLFEGRSSRKPEKLK 673
Query: 196 CIIHYFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASFWSTSVKPLCRFEVKSSGLI 255
+ YF+R+T P G+V+F R+ L ED +P W KPL R V G I
Sbjct: 674 TLFCYFRRVTEKKPTGLVTFTRQSL--ED------FPE---WERCDKPLTRLHVTYEGTI 722
Query: 256 EDHSSETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIAGMIFLPSMADNEAIDIVGV 315
E + ++VDFAN ++GGG G VQEEIRF+I+PELI +F + NE + I G
Sbjct: 723 EGNGRGMLQVDFANRFVGGGVTGAGLVQEEIRFLINPELIVSRLFTEVLDHNECLIITGT 782
Query: 316 ERFSSYTGYASSFRFSGDYVDDKEVDPFGRRKTRIVAIDALCGPGMRQYREKF----LLR 371
E++S YTGYA ++R++ + D E D + R T IVAIDAL R+Y ++F + R
Sbjct: 783 EQYSEYTGYAETYRWARSHEDGSEKDDWQRCCTEIVAIDAL---HFRRYLDQFVPEKVRR 839
Query: 372 EINKAFCGFLQQSQYQRDQKIPQENV 397
E+NKA+CGFL+ +P EN+
Sbjct: 840 ELNKAYCGFLRPG-------VPPENL 858
Score = 57.8 bits (138), Expect = 7e-08
Identities = 27/44 (61%), Positives = 32/44 (72%), Gaps = 1/44 (2%)
Query: 468 VATGNWGCGAFGGDPEVKTIIQWLAASQAGRPFIAYYTFGSGAL 511
VATGNWGCGAFGGD +K +IQ LAA+ A R + Y+TFG L
Sbjct: 861 VATGNWGCGAFGGDARLKALIQLLAAAAAERD-VVYFTFGDSEL 903
>PARG_BOVIN (O02776) Poly(ADP-ribose) glycohydrolase (EC 3.2.1.143)
Length = 977
Score = 167 bits (422), Expect = 8e-41
Identities = 105/311 (33%), Positives = 164/311 (51%), Gaps = 36/311 (11%)
Query: 76 FFDELISREECRKWFEEVLPSLGDLLLRLPSLLEAHYENADMVIDGKGATVRTGLRMLDS 135
F+D+++ E + ++ +LP + + L LP++ + +L
Sbjct: 577 FWDKVLEEAEAQHLYQSILPDMVKIALCLPNICTQP------------------IPLLKQ 618
Query: 136 QEAGIVFLTQELIAALLACSFLCLFPVHDRYEKQLQPVNFDELFASLYNDYTQKQEDKIW 195
+ + ++QE IA+LLA +F C FP + K D F L+ + ++ +K+
Sbjct: 619 KMNHSITMSQEQIASLLANAFFCTFPRRNAKMKSEYSSYPDINFNRLFEGRSSRKPEKLK 678
Query: 196 CIIHYFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASFWSTSVKPLCRFEVKSSGLI 255
+ YF+R+T P G+V+F R+ L ED +P W K L R V G I
Sbjct: 679 TLFCYFRRVTEKKPTGLVTFTRQSL--ED------FPE---WERCEKLLTRLHVTYEGTI 727
Query: 256 EDHSSETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIAGMIFLPSMADNEAIDIVGV 315
E + ++VDFAN ++GGG G VQEEIRF+I+PELI +F + NE + I G
Sbjct: 728 EGNGQGMLQVDFANRFVGGGVTSAGLVQEEIRFLINPELIVSRLFTEVLDHNECLIITGT 787
Query: 316 ERFSSYTGYASSFRFSGDYVDDKEVDPFGRRKTRIVAIDALCGPGMRQYREKF----LLR 371
E++S YTGYA ++R++ + D E D + RR T IVAIDAL R+Y ++F + R
Sbjct: 788 EQYSEYTGYAETYRWARSHEDRSERDDWQRRTTEIVAIDAL---HFRRYLDQFVPEKIRR 844
Query: 372 EINKAFCGFLQ 382
E+NKA+CGFL+
Sbjct: 845 ELNKAYCGFLR 855
Score = 57.8 bits (138), Expect = 7e-08
Identities = 28/52 (53%), Positives = 35/52 (66%), Gaps = 1/52 (1%)
Query: 460 MENSNDIGVATGNWGCGAFGGDPEVKTIIQWLAASQAGRPFIAYYTFGSGAL 511
+ + N VATGNWGCGAFGGD +K +IQ LAA+ A R + Y+TFG L
Sbjct: 858 VSSENLSAVATGNWGCGAFGGDARLKALIQILAAAVAERD-VVYFTFGDSEL 908
>PARG_CAEEL (Q867X0) Poly(ADP-ribose) glycohydrolase pme-3 (EC
3.2.1.143) (Poly ADP-ribose metabolism enzyme-3)
Length = 781
Score = 106 bits (264), Expect = 2e-22
Identities = 71/204 (34%), Positives = 107/204 (51%), Gaps = 20/204 (9%)
Query: 192 DKIWCIIHYFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASFWSTSVKPLCRFEVKS 251
+K+ + YF +++ + P G VSF ++ + D N + ++ L E
Sbjct: 470 EKLKFLFTYFDKMSMDPPDGAVSF--RLTKMDKDTF-----NEEWKDKKLRSLPEVEFFD 522
Query: 252 SGLIEDHSSETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIAGMIFLPSMADNEAID 311
LIED + T +VDFANE+LGGG L G VQEEIRF++ PE++ GM+ M EAI
Sbjct: 523 EMLIEDTALCT-QVDFANEHLGGGVLNHGSVQEEIRFLMCPEMMVGMLLCEKMKQLEAIS 581
Query: 312 IVGVERFSSYTGYASSFRFS------GDYVDDKEVDPFGRRKTRIVAIDALCGPGMR--- 362
IVG FSSYTGY + +++ ++ D FGR + +AIDA+ G +
Sbjct: 582 IVGAYVFSSYTGYGHTLKWAELQPNHSRQNTNEFRDRFGRLRVETIAIDAILFKGSKLDC 641
Query: 363 ---QYREKFLLREINKAFCGFLQQ 383
Q + ++RE+ KA GF+ Q
Sbjct: 642 QTEQLNKANIIREMKKASIGFMSQ 665
Score = 45.1 bits (105), Expect = 5e-04
Identities = 23/47 (48%), Positives = 30/47 (62%), Gaps = 1/47 (2%)
Query: 465 DIGVATGNWGCGAFGGDPEVKTIIQWLAASQAGRPFIAYYTFGSGAL 511
+I + TG WGCGAF GD +K IIQ +AA A RP + + +FG L
Sbjct: 671 NIPIVTGWWGCGAFNGDKPLKFIIQVIAAGVADRP-LHFCSFGEPEL 716
>DDL_YERPE (Q8ZIE7) D-alanine--D-alanine ligase (EC 6.3.2.4)
(D-alanylalanine synthetase) (D-Ala-D-Ala ligase)
Length = 306
Score = 31.6 bits (70), Expect = 5.6
Identities = 21/58 (36%), Positives = 29/58 (49%), Gaps = 2/58 (3%)
Query: 28 QVVGALRELGCGRVDSGQLL--FIFITELRNSLSLSSEPLAPSAAHGYALFFDELISR 83
Q AL G GRVD Q ++ E+ S ++S L P AA Y L F +L++R
Sbjct: 243 QAYHALDCSGWGRVDVMQDRDGHFYLLEVNTSPGMTSHSLVPMAARQYGLSFSQLVAR 300
>SYK_ORYSA (Q6F2U9) Lysyl-tRNA synthetase (EC 6.1.1.6) (Lysine--tRNA
ligase) (LysRS)
Length = 602
Score = 31.2 bits (69), Expect = 7.3
Identities = 14/34 (41%), Positives = 21/34 (61%)
Query: 336 DDKEVDPFGRRKTRIVAIDALCGPGMRQYREKFL 369
DD+E+DP + R+ A+D+L G+ Y KFL
Sbjct: 73 DDEEMDPTQYYENRLKALDSLKATGVNPYPHKFL 106
>C307_DROME (Q9VRM7) Probable cytochrome P450 307a1 (EC 1.14.-.-)
(CYPCCCVIIA1)
Length = 543
Score = 31.2 bits (69), Expect = 7.3
Identities = 18/64 (28%), Positives = 34/64 (53%), Gaps = 5/64 (7%)
Query: 138 AGIVFLTQELIAALLACSFLCLFPVHDRYEKQLQPV---NFDELFASLYNDYTQKQEDKI 194
A +++ ++ ++LA S++C+ ++ + LQPV N E+ + Y YTQ +
Sbjct: 3 AALIYTILAILLSVLATSYICI--IYGVKRRVLQPVKTKNSTEINHNAYQKYTQAPGPRP 60
Query: 195 WCII 198
W II
Sbjct: 61 WPII 64
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.324 0.140 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 63,838,862
Number of Sequences: 164201
Number of extensions: 2733422
Number of successful extensions: 7203
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 7162
Number of HSP's gapped (non-prelim): 19
length of query: 547
length of database: 59,974,054
effective HSP length: 115
effective length of query: 432
effective length of database: 41,090,939
effective search space: 17751285648
effective search space used: 17751285648
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 68 (30.8 bits)
Medicago: description of AC122162.2


