

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
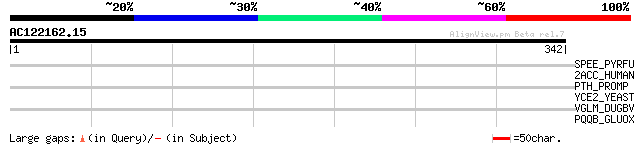
Query= AC122162.15 - phase: 0
(342 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
SPEE_PYRFU (Q8U4G1) Probable spermidine synthase (EC 2.5.1.16) (... 33 1.0
2ACC_HUMAN (Q9Y5P8) Serine/threonine protein phosphatase 2A, 48 ... 32 1.8
PTH_PROMP (Q7V342) Peptidyl-tRNA hydrolase (EC 3.1.1.29) (PTH) 31 5.2
YCE2_YEAST (P25572) Hypothetical protein YCL042W 30 6.7
VGLM_DUGBV (Q02004) M polyprotein precursor [Contains: Glycoprot... 30 6.7
PQQB_GLUOX (Q9L3B3) Coenzyme PQQ synthesis protein B (Pyrroloqui... 30 6.7
>SPEE_PYRFU (Q8U4G1) Probable spermidine synthase (EC 2.5.1.16)
(Putrescine aminopropyltransferase) (SPDSY)
Length = 281
Score = 33.1 bits (74), Expect = 1.0
Identities = 33/124 (26%), Positives = 51/124 (40%), Gaps = 19/124 (15%)
Query: 232 GRFYVLDQSGQTVTVGPDSSTE------LAAHPLYQRCV------RGKNRKLLVESEGEL 279
GR LD + Q VT+G S E + AHP +R + G R++L E+
Sbjct: 42 GRLLALDGTVQLVTLGERSYHEPLVHPAMLAHPKPKRVLVIGGGDGGTVREVLQHDVDEV 101
Query: 280 LLLDIHQTFFQFSMKIFKLDEN------RKKWEKLK-KLGDRILFVGSGCSFSASALDLC 332
++++I + S + K+D K EK K +GD F+ + F D
Sbjct: 102 IMVEIDEDVIMVSKDLIKIDNGLLEAMLNGKHEKAKLTIGDGFEFIKNNRGFDVIIADST 161
Query: 333 LPKG 336
P G
Sbjct: 162 DPVG 165
>2ACC_HUMAN (Q9Y5P8) Serine/threonine protein phosphatase 2A, 48 kDa
regulatory subunit B (PP2A, subunit B, PR48 isoform)
Length = 414
Score = 32.3 bits (72), Expect = 1.8
Identities = 44/191 (23%), Positives = 76/191 (39%), Gaps = 23/191 (12%)
Query: 138 FSHLPNAVNFIKFSAQHLRTDFIIDEDDFTFQNQH--GSYLYPQTVLAVTCHGKKPLVLG 195
FS+ V + KF D +ID DD N H + + + G+K G
Sbjct: 150 FSYEHFYVIYCKFWELDTDHDLLIDADDLARHNDHALSTKMIDRIFSGAVTRGRKVQKEG 209
Query: 196 TLSYCSSKPVLFHDQDKR-------WTPVSNLSTANGDICLFKGRFYVLDQSGQTVTVGP 248
+SY L ++DK+ W +L +G + +F+ ++ +Q +
Sbjct: 210 KISYADFVWFLISEEDKKTPTSIEYWFRCMDLD-GDGALSMFELEYFYEEQCRRL----- 263
Query: 249 DSSTELAAHPLYQRCVRGKNRKLLVESEGELLLLDIHQ-----TFFQFSMKIFK-LDENR 302
S + A P +Q C+ + +EG++ L D+ + FF I K LD +
Sbjct: 264 -DSMAIEALP-FQDCLCQMLDLVKPRTEGKITLQDLKRCKLANVFFDTFFNIEKYLDHEQ 321
Query: 303 KKWEKLKKLGD 313
K+ L + GD
Sbjct: 322 KEQISLLRDGD 332
>PTH_PROMP (Q7V342) Peptidyl-tRNA hydrolase (EC 3.1.1.29) (PTH)
Length = 203
Score = 30.8 bits (68), Expect = 5.2
Identities = 22/77 (28%), Positives = 38/77 (48%), Gaps = 1/77 (1%)
Query: 2 ASVDWTELPRDILFLISQRLDVELDLIRFRSVCSTWHSSPIPNDHNILPFQ-FPLLKYVP 60
A VDW ++ D LF+I +D+ L IRFR S+ + I + N L + F +K
Sbjct: 79 AIVDWYKIDLDRLFIIVDDIDLPLGKIRFRKKGSSGGHNGIKDIINKLQTENFNRIKIGI 138
Query: 61 APDSINNNNEIIDNINN 77
+N + ++ I++
Sbjct: 139 GSPPVNERKKTLNTISH 155
>YCE2_YEAST (P25572) Hypothetical protein YCL042W
Length = 119
Score = 30.4 bits (67), Expect = 6.7
Identities = 18/62 (29%), Positives = 27/62 (43%), Gaps = 8/62 (12%)
Query: 50 PFQFPLLKYVPAPDSINNNNEIIDNINNANTSFCYLSKRSFFLVKPPQEQQQQETLIRCR 109
P+++PL P P S+ N+ + N NT+ Y S KP +E+ R
Sbjct: 50 PYKYPLSTEPPTPPSVPNSASVNHN-TTTNTTLSYTRCHSTTYTKPLRERSS-------R 101
Query: 110 PW 111
PW
Sbjct: 102 PW 103
>VGLM_DUGBV (Q02004) M polyprotein precursor [Contains: Glycoprotein
G2; Nonstructural protein NS-M; Glycoprotein G1]
Length = 1551
Score = 30.4 bits (67), Expect = 6.7
Identities = 21/89 (23%), Positives = 36/89 (39%), Gaps = 4/89 (4%)
Query: 62 PDSINNNNEIIDNINNANTSFCYLSKRSFFLVKPPQEQQQQETLIRCRPWLIQITKNSSG 121
P + + + I T + +S + K Q ++ET+I P ++ +S
Sbjct: 49 PQTTTTSTAVSTTITATTTPTASWTTQSQYFNKTTQHHWREETMISRNPTVLDRQSRASS 108
Query: 122 KTQLLNPPFVSL----KSTAFSHLPNAVN 146
+LLN F+ L +HL NA N
Sbjct: 109 VRELLNTKFLMLLGFIPKGEVNHLENACN 137
>PQQB_GLUOX (Q9L3B3) Coenzyme PQQ synthesis protein B
(Pyrroloquinoline quinone biosynthesis protein B)
Length = 304
Score = 30.4 bits (67), Expect = 6.7
Identities = 22/96 (22%), Positives = 40/96 (40%), Gaps = 4/96 (4%)
Query: 169 QNQHGSYLYPQTVLAVTCHGKKPLVLGTLSYC----SSKPVLFHDQDKRWTPVSNLSTAN 224
+ + G+ Q LAV+ GK+ +L S P L H R TP+ +
Sbjct: 29 RTRQGAKAGTQASLAVSADGKRWFILNASPDLRQNHSDAPALHHQGSLRGTPIQGVVLTC 88
Query: 225 GDICLFKGRFYVLDQSGQTVTVGPDSSTELAAHPLY 260
G+I G + ++ T+ + +LA +P++
Sbjct: 89 GEIDAITGLLTLREREPFTLMGSESTLQQLADNPIF 124
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.138 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 41,883,695
Number of Sequences: 164201
Number of extensions: 1805352
Number of successful extensions: 3985
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 3985
Number of HSP's gapped (non-prelim): 6
length of query: 342
length of database: 59,974,054
effective HSP length: 111
effective length of query: 231
effective length of database: 41,747,743
effective search space: 9643728633
effective search space used: 9643728633
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 66 (30.0 bits)
Medicago: description of AC122162.15


