

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121245.9 + phase: 2 /pseudo/partial
(223 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
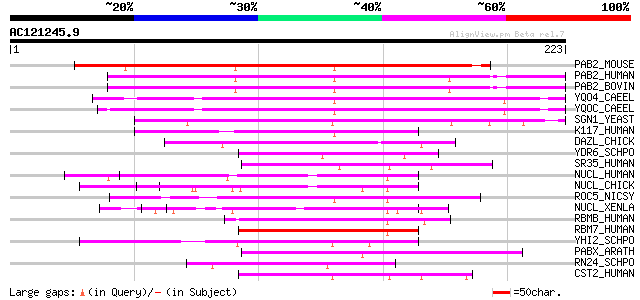
Score E
Sequences producing significant alignments: (bits) Value
PAB2_MOUSE (Q8CCS6) Polyadenylate-binding protein 2 (Poly(A)-bin... 141 1e-33
PAB2_HUMAN (Q86U42) Polyadenylate-binding protein 2 (Poly(A)-bin... 139 4e-33
PAB2_BOVIN (Q28165) Polyadenylate-binding protein 2 (Poly(A)-bin... 139 4e-33
YQO4_CAEEL (Q09295) Hypothetical RNA-binding protein EEED8.4 in ... 105 1e-22
YQOC_CAEEL (Q09301) Hypothetical RNA-binding protein EEED8.12 in... 104 2e-22
SGN1_YEAST (P40561) RNA-binding protein SGN1 97 3e-20
K117_HUMAN (P42696) Probable RNA-binding protein KIAA0117 (HAL845) 67 4e-11
DAZL_CHICK (Q804A9) Deleted in azoospermia-like (DAZ-like protein) 59 1e-08
YDR6_SCHPO (O13741) Hypothetical RNA-binding protein C16E8.06c i... 56 6e-08
SR35_HUMAN (Q8WXF0) 35 kDa SR repressor protein (SRrp35) 55 1e-07
NUCL_HUMAN (P19338) Nucleolin (Protein C23) 55 1e-07
NUCL_CHICK (P15771) Nucleolin (Protein C23) 54 4e-07
ROC5_NICSY (P19684) 33 kDa ribonucleoprotein, chloroplast precursor 53 7e-07
NUCL_XENLA (P20397) Nucleolin (Protein C23) 52 9e-07
RBMB_HUMAN (P57052) Putative RNA-binding protein 11 (RNA binding... 52 1e-06
RBM7_HUMAN (Q9Y580) RNA-binding protein 7 (RNA binding motif pro... 51 2e-06
YHI2_SCHPO (O13620) Probable RNA-binding protein P22H7.02c 50 4e-06
PABX_ARATH (Q9ZQA8) Probable polyadenylate-binding protein At2g3... 50 6e-06
RN24_SCHPO (Q09100) RNA-binding protein rnp24 49 7e-06
CST2_HUMAN (P33240) Cleavage stimulation factor, 64 kDa subunit ... 49 7e-06
>PAB2_MOUSE (Q8CCS6) Polyadenylate-binding protein 2
(Poly(A)-binding protein 2) (PolyA binding protein II)
(PABII) (Polyadenylate-binding nuclear protein 1)
(Nuclear poly(A)-binding protein 1)
Length = 301
Score = 141 bits (356), Expect = 1e-33
Identities = 80/176 (45%), Positives = 111/176 (62%), Gaps = 12/176 (6%)
Query: 27 ENDDVDMIGADNNEADLDD-----MKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANAS 81
E ++ ++ AD + ++D +K R++EME+EA LKE+Q +VEK+M P NA
Sbjct: 93 EEEEPGLVEADPGDGAIEDPELEAIKARVREMEEEAEKLKELQNEVEKQMNMSPPPGNAG 152
Query: 82 ASQINREE---VDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKF-GQPKGYAY 137
++ EE DARSI+VGNVDY T E+++ HF CG+VNRVTI DKF G PKG+AY
Sbjct: 153 PVIMSLEEKMEADARSIYVGNVDYGATAEELEAHFHGCGSVNRVTILCDKFSGHPKGFAY 212
Query: 138 VEFVEVEAVQEALLLNESELHGRQLKVTAKRTNVPGMKQFRPRRPSNPYMGFRGRT 193
+EF + E+V+ +L L+ES GRQ+KV KRTN PG+ P + Y R RT
Sbjct: 213 IEFSDKESVRTSLALDESLFRGRQIKVIPKRTNRPGISTTDRGFPRSRY---RART 265
>PAB2_HUMAN (Q86U42) Polyadenylate-binding protein 2
(Poly(A)-binding protein 2) (PolyA binding protein II)
(PABII) (Polyadenylate-binding nuclear protein 1)
(Nuclear poly(A)-binding protein 1)
Length = 305
Score = 139 bits (351), Expect = 4e-33
Identities = 86/195 (44%), Positives = 116/195 (59%), Gaps = 15/195 (7%)
Query: 40 EADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINREE---VDARSIF 96
+ +L+ +K R++EME+EA LKE+Q +VEK+M P NA ++ EE DARSI+
Sbjct: 115 DPELEAIKARVREMEEEAEKLKELQNEVEKQMNMSPPPGNAGPVIMSIEEKMEADARSIY 174
Query: 97 VGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKF-GQPKGYAYVEFVEVEAVQEALLLNES 155
VGNVDY T E+++ HF CG+VNRVTI DKF G PKG+AY+EF + E+V+ +L L+ES
Sbjct: 175 VGNVDYGATAEELEAHFHGCGSVNRVTILCDKFSGHPKGFAYIEFSDKESVRTSLALDES 234
Query: 156 ELHGRQLKVTAKRTNVPGMK-------QFRPRRPSNPYMGFRGRTPYAPPFAYAPAPYGY 208
GRQ+KV KRTN PG+ + R R + Y R R Y+ + P G
Sbjct: 235 LFRGRQIKVIPKRTNRPGISTTDRGFPRARYRARTTNYNSSRSRF-YS---GFNSRPRGR 290
Query: 209 GKIPRFRMGMRYSPY 223
R R YSPY
Sbjct: 291 VYRGRARATSWYSPY 305
>PAB2_BOVIN (Q28165) Polyadenylate-binding protein 2
(Poly(A)-binding protein 2) (PolyA binding protein II)
(PABII) (Polyadenylate-binding nuclear protein 1)
(Nuclear poly(A)-binding protein 1)
Length = 305
Score = 139 bits (351), Expect = 4e-33
Identities = 86/195 (44%), Positives = 116/195 (59%), Gaps = 15/195 (7%)
Query: 40 EADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINREE---VDARSIF 96
+ +L+ +K R++EME+EA LKE+Q +VEK+M P NA ++ EE DARSI+
Sbjct: 115 DPELEAIKARVREMEEEAEKLKELQNEVEKQMNMSPPPGNAGPVIMSIEEKMEADARSIY 174
Query: 97 VGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKF-GQPKGYAYVEFVEVEAVQEALLLNES 155
VGNVDY T E+++ HF CG+VNRVTI DKF G PKG+AY+EF + E+V+ +L L+ES
Sbjct: 175 VGNVDYGATAEELEAHFHGCGSVNRVTILCDKFSGHPKGFAYIEFSDKESVRTSLALDES 234
Query: 156 ELHGRQLKVTAKRTNVPGMK-------QFRPRRPSNPYMGFRGRTPYAPPFAYAPAPYGY 208
GRQ+KV KRTN PG+ + R R + Y R R Y+ + P G
Sbjct: 235 LFRGRQIKVIPKRTNRPGISTTDRGFPRARYRARTTNYNSSRSRF-YS---GFNSRPRGR 290
Query: 209 GKIPRFRMGMRYSPY 223
R R YSPY
Sbjct: 291 VYRGRARATSWYSPY 305
>YQO4_CAEEL (Q09295) Hypothetical RNA-binding protein EEED8.4 in
chromosome II
Length = 237
Score = 105 bits (261), Expect = 1e-22
Identities = 67/199 (33%), Positives = 107/199 (53%), Gaps = 20/199 (10%)
Query: 34 IGADNNEADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINREEVDAR 93
I ADNN +M E+E E+A L+++Q K+ K + S A ++ ++ +DA+
Sbjct: 50 ITADNNSNSHFEM-----EIEAESAILQQIQNKMAKHLESA---AYVPPTEEEQKAIDAK 101
Query: 94 SIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKF-GQPKGYAYVEFVEVEAVQEALLL 152
S+F+GNVD+ T E++++HF+ CG + + TI DKF + K +AY+EF + +++ AL++
Sbjct: 102 SVFIGNVDFNSTIEEIEEHFKGCGQIVKTTIPKDKFTKKQKNFAYIEFDDSSSIENALVM 161
Query: 153 NESELHGRQLKVTAKRTNVPGMKQFRPRRPSNPYMGFRGRTPYAP--------PFAYAPA 204
N S R + VTAKRTN+PGM + RG AP + Y
Sbjct: 162 NGSLFRSRPIVVTAKRTNIPGMGHGVRGSSRGTFGRGRGAARGAPGRQQTVVVKYVYVNG 221
Query: 205 PYGYGKIPRFRMGMRYSPY 223
P G+ R G R++PY
Sbjct: 222 PNRGGRGGR---GRRFNPY 237
>YQOC_CAEEL (Q09301) Hypothetical RNA-binding protein EEED8.12 in
chromosome II
Length = 197
Score = 104 bits (259), Expect = 2e-22
Identities = 67/197 (34%), Positives = 106/197 (53%), Gaps = 16/197 (8%)
Query: 36 ADNNEADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINREEVDARSI 95
ADNN + + E+E E+A L+++Q K+ K + S A ++ ++ +DA+S+
Sbjct: 8 ADNN-CNEESSNSFEMEIEAESAILEQIQNKMAKNLESA---AYVPPTEEEQKAIDAKSV 63
Query: 96 FVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKF-GQPKGYAYVEFVEVEAVQEALLLNE 154
F+GNVD+ T E+V++HF+ CG + R TI DKF + K +AY+EF + +++ AL++N
Sbjct: 64 FIGNVDFNSTIEEVEEHFKGCGHIVRTTIPKDKFTKKQKNFAYIEFDDSSSIENALVMNG 123
Query: 155 SELHGRQLKVTAKRTNVPGMKQFRPRRPSNPYMGFRGRTPYAP--------PFAYAPAPY 206
S R + VTAKRTN+PGM + RG AP + Y P
Sbjct: 124 SLFRSRPIVVTAKRTNIPGMGHGVRGSSRGTFGRGRGAARGAPGRQQTVVVKYVYVNGPN 183
Query: 207 GYGKIPRFRMGMRYSPY 223
G+ R G R++PY
Sbjct: 184 RGGRGGR---GRRFNPY 197
>SGN1_YEAST (P40561) RNA-binding protein SGN1
Length = 250
Score = 97.1 bits (240), Expect = 3e-20
Identities = 62/188 (32%), Positives = 96/188 (50%), Gaps = 20/188 (10%)
Query: 51 KEMEDEAAALKEMQAKVEKE-MGSVQDPANASASQINREEVDARSIFVGNVDYACTPEDV 109
K+ + + L E+ K+ E V + ++ E D+RSIFVGN+ TPE +
Sbjct: 21 KKNDKQELELDELVGKLSIEGTPQVSQKLSKEEKHAHQLEADSRSIFVGNITPDVTPEQI 80
Query: 110 QQHFQSCGTVNRVTIRTDK-FGQPKGYAYVEFVEVEAVQEALLLNESELHGRQLKVTAKR 168
+ HF+ CG + R+T+ D+ G PKGY Y+EF ++AL LN EL G+++ V+ KR
Sbjct: 81 EDHFKDCGQIKRITLLYDRNTGTPKGYGYIEFESPAYREKALQLNGGELKGKKIAVSRKR 140
Query: 169 TNVPGMKQ--------FRPRRPSNPYMGFRG--RTPYAPPFAYAPAP---YGYGKIPRFR 215
TN+PG + F+ + + P M + PY PP+ +P +GY K +R
Sbjct: 141 TNIPGFNRHYNSQNQYFQQWQWNYPLMAYPNPDTFPYYPPYPPNQSPNQNFGYNKNNYYR 200
Query: 216 MGMRYSPY 223
SPY
Sbjct: 201 -----SPY 203
>K117_HUMAN (P42696) Probable RNA-binding protein KIAA0117 (HAL845)
Length = 430
Score = 66.6 bits (161), Expect = 4e-11
Identities = 43/115 (37%), Positives = 61/115 (52%), Gaps = 7/115 (6%)
Query: 51 KEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINREEVDARSIFVGNVDYACTPEDVQ 110
KE ALK A++ D A+ ++S+ D RS+FVGN+ Y ++
Sbjct: 251 KEESAATQALKRNGAQIADGFRIRVDLASETSSR------DKRSVFVGNLPYKVEESAIE 304
Query: 111 QHFQSCGTVNRVTIRTDKF-GQPKGYAYVEFVEVEAVQEALLLNESELHGRQLKV 164
+HF CG++ V I DK G KG+ YV F ++V AL LN SEL GR+L+V
Sbjct: 305 KHFLDCGSIMAVRIVRDKMTGIGKGFGYVLFENTDSVHLALKLNNSELMGRKLRV 359
Score = 31.6 bits (70), Expect = 1.6
Identities = 15/46 (32%), Positives = 26/46 (55%), Gaps = 3/46 (6%)
Query: 84 QINREEV---DARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRT 126
QIN+EE + R++FVGN+ C + ++ F+ G + V R+
Sbjct: 173 QINQEEERLKNERTVFVGNLPVTCNKKKLKSFFKEYGQIESVRFRS 218
>DAZL_CHICK (Q804A9) Deleted in azoospermia-like (DAZ-like protein)
Length = 289
Score = 58.5 bits (140), Expect = 1e-08
Identities = 35/125 (28%), Positives = 66/125 (52%), Gaps = 9/125 (7%)
Query: 63 MQAKVEKEMGSVQDPANASASQ-----INREEVDARSIFVGNVDYACTPEDVQQHFQSCG 117
M A E + GS+ + S++ + ++ ++FVG +D +++ +F+ G
Sbjct: 1 MSANAEAQCGSISEDNTHSSTTCQGYVLPEGKIMPNTVFVGGIDIRMNEAEIRSYFEQYG 60
Query: 118 TVNRVTIRTDKFGQPKGYAYVEFVEVEAVQEALLLNESELHGRQLKV---TAKRTNVPGM 174
TV V I TD+ G KGY +V F++ VQ+ ++ ++ +HG++LK+ K+ N+
Sbjct: 61 TVKEVKIITDRTGVSKGYGFVSFLDNVDVQK-IVESQISVHGKRLKLGPAIRKQQNLCSY 119
Query: 175 KQFRP 179
Q RP
Sbjct: 120 MQPRP 124
>YDR6_SCHPO (O13741) Hypothetical RNA-binding protein C16E8.06c in
chromosome I
Length = 438
Score = 56.2 bits (134), Expect = 6e-08
Identities = 29/82 (35%), Positives = 49/82 (59%), Gaps = 2/82 (2%)
Query: 93 RSIFVGNVDYACTPEDVQQHFQSCGTVNRVTI-RTDKFGQPKGYAYVEFVEVEAVQEALL 151
R +FVGN+ + E + ++F CG+++ V I R K KG+AY++F + V +ALL
Sbjct: 270 RCVFVGNLAFEAEEEPLWRYFGDCGSIDYVRIVRDPKTNLGKGFAYIQFKDTMGVDKALL 329
Query: 152 LNESEL-HGRQLKVTAKRTNVP 172
LNE ++ GR L++ ++ P
Sbjct: 330 LNEKKMPEGRTLRIMRAKSTKP 351
>SR35_HUMAN (Q8WXF0) 35 kDa SR repressor protein (SRrp35)
Length = 261
Score = 55.5 bits (132), Expect = 1e-07
Identities = 35/105 (33%), Positives = 58/105 (54%), Gaps = 4/105 (3%)
Query: 94 SIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQ-PKGYAYVEFVEVEAVQEALL- 151
S+F+ NV A PED+++ F G + V I D + + P+G+AYV+F +V ++AL
Sbjct: 11 SLFIRNVADATRPEDLRREFGRYGPIVDVYIPLDFYTRRPRGFAYVQFEDVRDAEDALYN 70
Query: 152 LNESELHGRQLKVTAKR--TNVPGMKQFRPRRPSNPYMGFRGRTP 194
LN + GRQ+++ + PG + + R P +P R R+P
Sbjct: 71 LNRKWVCGRQIEIQFAQGDRKTPGQMKSKERHPCSPSDHRRSRSP 115
>NUCL_HUMAN (P19338) Nucleolin (Protein C23)
Length = 706
Score = 55.5 bits (132), Expect = 1e-07
Identities = 37/144 (25%), Positives = 70/144 (47%), Gaps = 2/144 (1%)
Query: 23 D*DMENDDVDMIGADN-NEADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANAS 81
D D E DD D D+ ++ D DD +E E+E +KE K +KEM + A
Sbjct: 234 DEDEEEDDEDEDDDDDEDDEDDDDEDDEEEEEEEEEEPVKEAPGKRKKEMAKQKAAPEAK 293
Query: 82 ASQIN-REEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEF 140
++ E A ++FVGN+++ + +++ N + + + G + + YV+F
Sbjct: 294 KQKVEGTEPTTAFNLFVGNLNFNKSAPELKTGISDVFAKNDLAVVDVRIGMTRKFGYVDF 353
Query: 141 VEVEAVQEALLLNESELHGRQLKV 164
E +++AL L ++ G ++K+
Sbjct: 354 ESAEDLEKALELTGLKVFGNEIKL 377
Score = 45.1 bits (105), Expect = 1e-04
Identities = 30/121 (24%), Positives = 59/121 (47%), Gaps = 7/121 (5%)
Query: 45 DMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINREEVDARSIFVGNVDYAC 104
D +K +E + + + E G QD S + E ++++ + N+ Y+
Sbjct: 440 DAEKTFEEKQGTEIDGRSISLYYTGEKGQNQDYRGGKNSTWSGE---SKTLVLSNLSYSA 496
Query: 105 TPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAVQEAL-LLNESELHGRQLK 163
T E +Q+ F+ + + ++ G+ KGYA++EF E +EAL N+ E+ GR ++
Sbjct: 497 TEETLQEVFEKATFIK---VPQNQNGKSKGYAFIEFASFEDAKEALNSCNKREIEGRAIR 553
Query: 164 V 164
+
Sbjct: 554 L 554
Score = 32.0 bits (71), Expect = 1.2
Identities = 34/138 (24%), Positives = 55/138 (39%), Gaps = 7/138 (5%)
Query: 54 EDEAAALKEMQAKVEKEMGSVQDPANASASQINREEVDARSIFVGNVDYACTPEDVQQHF 113
ED AL K E E +++ N ++++FV + T E +++ F
Sbjct: 533 EDAKEALNSCN-KREIEGRAIRLELQGPRGSPNARSQPSKTLFVKGLSEDTTEETLKESF 591
Query: 114 QSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAVQEALLLNESELHGRQLKVTAKRTNVPG 173
G+V + + G KG+ +V+F E +EA + + E+ G KVT G
Sbjct: 592 D--GSVRARIVTDRETGSSKGFGFVDFNSEEDAKEA--MEDGEIDGN--KVTLDWAKPKG 645
Query: 174 MKQFRPRRPSNPYMGFRG 191
F R G RG
Sbjct: 646 EGGFGGRGGGRGGFGGRG 663
>NUCL_CHICK (P15771) Nucleolin (Protein C23)
Length = 694
Score = 53.5 bits (127), Expect = 4e-07
Identities = 40/140 (28%), Positives = 70/140 (49%), Gaps = 11/140 (7%)
Query: 29 DDVDMIGADNNEADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINRE 88
D+ D D +E D DD ++ +E EDE +KE K +KEM AN SA + ++
Sbjct: 220 DEEDEEDEDEDEEDEDDEEEDEEESEDEKP-VKEAPGKRKKEM------ANKSAPEAKKK 272
Query: 89 EVD----ARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVE 144
+ + A S+FV N+ E+++ + + + + G K + YV+F+ E
Sbjct: 273 KTETPASAFSLFVKNLTPTKDYEELRTAIKEFFGKKNLQVSEVRIGSSKRFGYVDFLSAE 332
Query: 145 AVQEALLLNESELHGRQLKV 164
+ +AL LN +L G ++K+
Sbjct: 333 DMDKALQLNGKKLMGLEIKL 352
Score = 51.2 bits (121), Expect = 2e-06
Identities = 35/120 (29%), Positives = 65/120 (54%), Gaps = 10/120 (8%)
Query: 52 EMEDEAAALKEMQAKVEKEMGS---VQDPANASASQINRE---EVDARSIFVGNVDYACT 105
E + EA A K ++ K E+ V D + Q +++ E +++++ V N+ YA +
Sbjct: 414 EFKTEAEAEKALEEKQGTEVDGRAMVIDYTGEKSQQESQKGGGERESKTLIVNNLSYAAS 473
Query: 106 PEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAVQEAL-LLNESELHGRQLKV 164
E +Q+ F+ ++ + + G+PKGYA+VEF E +EAL N +E+ GR +++
Sbjct: 474 EETLQELFKKATSIK---MPQNNQGRPKGYAFVEFPTAEDAKEALNSCNNTEIEGRAIRL 530
Score = 45.1 bits (105), Expect = 1e-04
Identities = 32/108 (29%), Positives = 57/108 (52%), Gaps = 7/108 (6%)
Query: 61 KEMQAKVEKEMG---SVQDPANASASQINREEVDARSIFVGNVDYACTPEDVQQHFQSCG 117
K +Q +K MG ++ + + + N++E DAR++FV N+ Y T ++++ F++
Sbjct: 336 KALQLNGKKLMGLEIKLEKAKSKESLKENKKERDARTLFVKNLPYRVTEDEMKNVFENAL 395
Query: 118 TVNRVTIRTDKFGQPKGYAYVEF-VEVEAVQEALLLNESELHGRQLKV 164
V V +K G KG AY+EF E EA + +E+ GR + +
Sbjct: 396 EVRLV---LNKEGSSKGMAYIEFKTEAEAEKALEEKQGTEVDGRAMVI 440
>ROC5_NICSY (P19684) 33 kDa ribonucleoprotein, chloroplast precursor
Length = 324
Score = 52.8 bits (125), Expect = 7e-07
Identities = 39/151 (25%), Positives = 75/151 (48%), Gaps = 12/151 (7%)
Query: 41 ADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINREEVDARSIFVGNV 100
A + D + ++E ++E AL A+ E+E+ ++ ++ E V+ ++VGN+
Sbjct: 72 ASVSDGVEVVQEDDEEEVALS---AEEEEEIEEKEE-------RVESESVEGGRLYVGNL 121
Query: 101 DYACTPEDVQQHFQSCGTVNRVTIRTDKF-GQPKGYAYVEFVEVEAVQEAL-LLNESELH 158
++ T + + F GTV V I D+ + +G+A+V VE +EA+ L + S++
Sbjct: 122 PFSMTSSQLSEIFAEAGTVANVEIVYDRVTDRSRGFAFVTMGSVEEAKEAIRLFDGSQVG 181
Query: 159 GRQLKVTAKRTNVPGMKQFRPRRPSNPYMGF 189
GR +KV G ++ + + Y GF
Sbjct: 182 GRTVKVNFPEVPRGGEREVMSAKIRSTYQGF 212
>NUCL_XENLA (P20397) Nucleolin (Protein C23)
Length = 650
Score = 52.4 bits (124), Expect = 9e-07
Identities = 38/116 (32%), Positives = 65/116 (55%), Gaps = 11/116 (9%)
Query: 64 QAKVEKEMGSVQDPANASASQINREEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVT 123
+ K+EK M + N +A N++E D+R++FV N+ Y+ T E++Q+ F++ +
Sbjct: 300 EVKIEKAMAFDK---NKTAE--NKKERDSRTLFVKNIPYSTTVEELQEIFEN---AKDIR 351
Query: 124 IRTDKFGQPKGYAYVEFVEVEAVQEALLLNE-SELHGRQLKV--TAKRTNVPGMKQ 176
I T K G KG AYVEF + +AL + +E+ GR + V T +++ G K+
Sbjct: 352 IPTGKDGSNKGIAYVEFSNEDEANKALEEKQGAEIEGRSIFVDFTGEKSQNSGNKK 407
Score = 46.6 bits (109), Expect = 5e-05
Identities = 33/128 (25%), Positives = 63/128 (48%), Gaps = 9/128 (7%)
Query: 37 DNNEADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINREEVDARSIF 96
D+ E D DD E +D+ ++ AK +KEM A + + E + SIF
Sbjct: 185 DDEEEDDDD------EEDDDDEEEQQGSAKRKKEMPKTIPEAKKTKTDTASEGL---SIF 235
Query: 97 VGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAVQEALLLNESE 156
+GN++ ++++ + + +TI+ + G K + YV+F E V++AL L +
Sbjct: 236 IGNLNSTKEFDELKDALREFFSKKNLTIQDIRIGNSKKFGYVDFSSEEEVEKALKLTGKK 295
Query: 157 LHGRQLKV 164
+ G ++K+
Sbjct: 296 ILGTEVKI 303
Score = 43.9 bits (102), Expect = 3e-04
Identities = 32/116 (27%), Positives = 62/116 (52%), Gaps = 9/116 (7%)
Query: 54 EDEA-AALKEMQ-AKVEKEMGSVQDPANASASQINRE--EVDARSIFVGNVDYACTPEDV 109
EDEA AL+E Q A++E V S + N++ E D++ + V N+ Y+ T + +
Sbjct: 371 EDEANKALEEKQGAEIEGRSIFVDFTGEKSQNSGNKKGPEGDSKVLVVNNLSYSATEDSL 430
Query: 110 QQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAVQEAL-LLNESELHGRQLKV 164
++ F+ ++ G+ KG+A++EF E ++A+ N +E+ GR +++
Sbjct: 431 REVFEKATSIRI----PQNQGRAKGFAFIEFSSAEDAKDAMDSCNNTEIEGRSIRL 482
Score = 30.4 bits (67), Expect = 3.5
Identities = 27/101 (26%), Positives = 44/101 (42%), Gaps = 5/101 (4%)
Query: 92 ARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAVQEAL- 150
++++FV + T E +++ F G+VN + G KG+ +V+F E + A
Sbjct: 501 SKTLFVRGLSEDTTEETLKEAFD--GSVNARIVTDRDTGASKGFGFVDFSTAEDAKAAKE 558
Query: 151 LLNESELHGRQLKVTAKRTNVPGMKQFRPRRPSNPYMGFRG 191
+ + E+ G KVT G Q R GFRG
Sbjct: 559 AMEDGEIDGN--KVTLDFAKPKGDSQRGGRGGFGRGGGFRG 597
>RBMB_HUMAN (P57052) Putative RNA-binding protein 11 (RNA binding
motif protein 11)
Length = 281
Score = 52.0 bits (123), Expect = 1e-06
Identities = 33/96 (34%), Positives = 54/96 (55%), Gaps = 6/96 (6%)
Query: 87 REEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAV 146
+EE D R++FVGN++ E + + F G + +VTI D+ G+PK + +V F E+V
Sbjct: 5 QEEAD-RTVFVGNLEARVREEILYELFLQAGPLTKVTICKDREGKPKSFGFVCFKHPESV 63
Query: 147 QEAL-LLNESELHGRQLKVT----AKRTNVPGMKQF 177
A+ LLN L+GR + V + R++ P + F
Sbjct: 64 SYAIALLNGIRLYGRPINVQYRFGSSRSSEPANQSF 99
>RBM7_HUMAN (Q9Y580) RNA-binding protein 7 (RNA binding motif
protein 7)
Length = 266
Score = 51.2 bits (121), Expect = 2e-06
Identities = 29/73 (39%), Positives = 45/73 (60%), Gaps = 1/73 (1%)
Query: 93 RSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAVQEAL-L 151
R++FVGN++ T E + + F G V +V I DK G+PK +A+V F +V A+ L
Sbjct: 10 RTLFVGNLETKVTEELLFELFHQAGPVIKVKIPKDKDGKPKQFAFVNFKHEVSVPYAMNL 69
Query: 152 LNESELHGRQLKV 164
LN +L+GR +K+
Sbjct: 70 LNGIKLYGRPIKI 82
>YHI2_SCHPO (O13620) Probable RNA-binding protein P22H7.02c
Length = 833
Score = 50.1 bits (118), Expect = 4e-06
Identities = 35/139 (25%), Positives = 69/139 (49%), Gaps = 12/139 (8%)
Query: 29 DDVDMIGADNNEADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINRE 88
++V ++G + D ++ + L+ + ++ A M K E N S + N +
Sbjct: 267 EEVSVVGDELKSFDKENNDEHLERVTNDKIADASMLQKAEN---------NVSEQERNIQ 317
Query: 89 EV-DARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDK-FGQPKGYAYVEFVEV-EA 145
+ + + +F+ N+ Y+C +D++ F G + +V + DK PKG+AY++F + +A
Sbjct: 318 LISETKRLFLRNLTYSCAEDDLKSLFGPFGQLEQVHMPIDKKTNNPKGFAYIDFHDADDA 377
Query: 146 VQEALLLNESELHGRQLKV 164
V+ L L+ GR L V
Sbjct: 378 VRAYLELDAKPFQGRLLHV 396
Score = 36.2 bits (82), Expect = 0.063
Identities = 30/121 (24%), Positives = 51/121 (41%), Gaps = 10/121 (8%)
Query: 52 EMEDEAAALKEMQA---------KVEKEMGSVQDPANASASQINREEVDARSIFVGNVDY 102
E +D+A+A+ M A K+E ++ A A + + + I + N+ +
Sbjct: 671 EFKDKASAVAAMHAMNGFVLDGHKLEIKLSHQGVDAAAEVRKQDSSKPKGTKILIKNLPF 730
Query: 103 ACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEV-EAVQEALLLNESELHGRQ 161
T +DVQ + G + V + +G+A+ EFV EA L + L GR
Sbjct: 731 EATKKDVQSLLGAYGQLRSVRVPKKFDRSARGFAFAEFVTAREAANAMRALKNTHLLGRH 790
Query: 162 L 162
L
Sbjct: 791 L 791
>PABX_ARATH (Q9ZQA8) Probable polyadenylate-binding protein
At2g36660 (Poly(A)-binding protein At2g36660) (PABP)
Length = 609
Score = 49.7 bits (117), Expect = 6e-06
Identities = 30/114 (26%), Positives = 51/114 (44%), Gaps = 1/114 (0%)
Query: 94 SIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEF-VEVEAVQEALLL 152
+I+V NV+ A T E++++HF CGT+ + D+ G+ KG+ +V F EA+
Sbjct: 305 NIYVKNVNVAVTEEELRKHFSQCGTITSTKLMCDEKGKSKGFGFVCFSTPEEAIDAVKTF 364
Query: 153 NESELHGRQLKVTAKRTNVPGMKQFRPRRPSNPYMGFRGRTPYAPPFAYAPAPY 206
+ HG+ L V + Q + + + + P YAP Y
Sbjct: 365 HGQMFHGKPLYVAIAQKKEDRKMQLQVQFGNRVEARKSSSSASVNPGTYAPLYY 418
Score = 34.3 bits (77), Expect = 0.24
Identities = 24/80 (30%), Positives = 40/80 (50%), Gaps = 6/80 (7%)
Query: 91 DARSIFVGNVDYACTPEDV-----QQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEA 145
DAR VGNV PE V Q F+ G + + T + G+ +GY +V+F + +A
Sbjct: 105 DARRNGVGNVFVKNLPESVTNAVLQDMFKKFGNIVSCKVATLEDGKSRGYGFVQFEQEDA 164
Query: 146 VQEAL-LLNESELHGRQLKV 164
A+ LN + + +++ V
Sbjct: 165 AHAAIQTLNSTIVADKEIYV 184
>RN24_SCHPO (Q09100) RNA-binding protein rnp24
Length = 369
Score = 49.3 bits (116), Expect = 7e-06
Identities = 26/88 (29%), Positives = 50/88 (56%), Gaps = 4/88 (4%)
Query: 72 GSVQDPANA---SASQINREEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRT-D 127
G PAN +AS + ++ + +FVGN+D+ T D+++HF G + RV + T +
Sbjct: 204 GRPSKPANTLSKTASIQSSKKEPSSILFVGNLDFETTDADLKEHFGQVGQIRRVRLMTFE 263
Query: 128 KFGQPKGYAYVEFVEVEAVQEALLLNES 155
G+ KG+ +V+F +++ +A+ L +
Sbjct: 264 DTGKCKGFGFVDFPDIDTCMKAMELGHN 291
Score = 38.9 bits (89), Expect = 0.010
Identities = 39/149 (26%), Positives = 68/149 (45%), Gaps = 29/149 (19%)
Query: 59 ALKEMQAKVEKEMGSVQDPANASASQINREEVDARS---IFVGNVDYACTPEDVQQHF-- 113
AL+E + K EK + + D + +E RS I+VGN+ + T E + F
Sbjct: 69 ALEEAKKKEEKRLKRL-DAKYGRKEEGESQEESKRSPWGIWVGNLSFHTTKEILTDFFVR 127
Query: 114 QSCGTVNRVT------------------IRTDKFGQPKGYAYVEFVEVEAVQEALLLNES 155
++ + V+ + +K Q KG+AYV+F +A++ AL +E
Sbjct: 128 ETSEMIKEVSEENIKPITTEQITRIHMPMSKEKRFQNKGFAYVDFATEDALKLALQCSEK 187
Query: 156 ELHGRQLKVTAKRTNVPGMKQFRPRRPSN 184
L+GR + + + T+ G RP +P+N
Sbjct: 188 ALNGRNILIKS-NTDFSG----RPSKPAN 211
>CST2_HUMAN (P33240) Cleavage stimulation factor, 64 kDa subunit
(CSTF 64 kDa subunit) (CF-1 64 kDa subunit)
Length = 577
Score = 49.3 bits (116), Expect = 7e-06
Identities = 34/100 (34%), Positives = 48/100 (48%), Gaps = 6/100 (6%)
Query: 93 RSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDK-FGQPKGYAYVEFVEVEAVQEALL 151
RS+FVGN+ Y T E ++ F G V + D+ G+PKGY + E+ + E A+
Sbjct: 16 RSVFVGNIPYEATEEQLKDIFSEVGPVVSFRLVYDRETGKPKGYGFCEYQDQETALSAMR 75
Query: 152 -LNESELHGRQLKV--TAKRTNVPGMKQFRPRRP--SNPY 186
LN E GR L+V A N +K P +PY
Sbjct: 76 NLNGREFSGRALRVDNAASEKNKEELKSLGTGAPVIESPY 115
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.137 0.408
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,986,315
Number of Sequences: 164201
Number of extensions: 1122166
Number of successful extensions: 5915
Number of sequences better than 10.0: 343
Number of HSP's better than 10.0 without gapping: 154
Number of HSP's successfully gapped in prelim test: 190
Number of HSP's that attempted gapping in prelim test: 5490
Number of HSP's gapped (non-prelim): 586
length of query: 223
length of database: 59,974,054
effective HSP length: 106
effective length of query: 117
effective length of database: 42,568,748
effective search space: 4980543516
effective search space used: 4980543516
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 64 (29.3 bits)
Medicago: description of AC121245.9


