

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
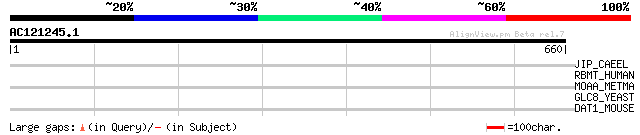
Query= AC121245.1 - phase: 0 /pseudo
(660 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
JIP_CAEEL (P34609) JNK-interacting protein (JIP) (JNK MAP kinase... 32 5.3
RBMT_HUMAN (Q9NW13) RNA-binding protein 28 (RNA binding motif pr... 31 9.1
MOAA_METMA (Q8PX29) Probable molybdenum cofactor biosynthesis pr... 31 9.1
GLC8_YEAST (P41818) GLC8 protein 31 9.1
DAT1_MOUSE (Q8C9B9) Death associated transcription factor 1 (Dea... 31 9.1
>JIP_CAEEL (P34609) JNK-interacting protein (JIP) (JNK MAP kinase
scaffold protein) (Uncoordinated protein 16)
Length = 1157
Score = 32.0 bits (71), Expect = 5.3
Identities = 21/93 (22%), Positives = 43/93 (45%), Gaps = 3/93 (3%)
Query: 151 ITQATSKQMSKRDSLRPYMNIIDKMLVQMLNEDPSFKPQGGRLGESDMDYDTDEKSMASL 210
+ + S+ M RD + +KM Q+ + K +L E + + + D+ MA
Sbjct: 364 VDELNSENMILRDENLSRQMVSEKMQEQITKHEEEIKTLKQKLMEKENEQEEDDVPMAMR 423
Query: 211 RRRLRRAREQYYRYRSEYTKFVLEALELEDTVK 243
+R R ++ R+ Y + + +ELE+++K
Sbjct: 424 KRFTRSEMQRVLMDRNAYKE---KLMELEESIK 453
>RBMT_HUMAN (Q9NW13) RNA-binding protein 28 (RNA binding motif
protein 28)
Length = 758
Score = 31.2 bits (69), Expect = 9.1
Identities = 22/79 (27%), Positives = 35/79 (43%), Gaps = 2/79 (2%)
Query: 65 ALFGLILLITLNKFWSGSVRGFAMACSNTFGLVTGAFLLGFGMSEIPKGIWLNADWAIQQ 124
A FG +L + + + G +RGF L G L G M EI KG + DWA+ +
Sbjct: 134 AQFGAVLEVNIPRKPDGKMRGFGFVQFKNL-LEAGKALKGMNMKEI-KGRTVAVDWAVAK 191
Query: 125 KFLSHKVAKMAVKLDDAHQ 143
+ A+ + +H+
Sbjct: 192 DKYKDTQSVSAIGEEKSHE 210
>MOAA_METMA (Q8PX29) Probable molybdenum cofactor biosynthesis
protein A
Length = 334
Score = 31.2 bits (69), Expect = 9.1
Identities = 21/77 (27%), Positives = 37/77 (47%), Gaps = 13/77 (16%)
Query: 163 DSLRPYMNIIDKMLVQMLNEDPSFKPQGGRLGESDMDYDTDEKSMASLRRRLRRAREQYY 222
D +RPY + L++++N DP L + +D T EKS+ RA E
Sbjct: 195 DFIRPYKGKVILQLIELMNIDP-------ELSKYTIDSKTLEKSLE------ERASEVRV 241
Query: 223 RYRSEYTKFVLEALELE 239
R+ K++++ +E+E
Sbjct: 242 RHLHHRKKYMIDGVEVE 258
>GLC8_YEAST (P41818) GLC8 protein
Length = 229
Score = 31.2 bits (69), Expect = 9.1
Identities = 23/87 (26%), Positives = 41/87 (46%), Gaps = 12/87 (13%)
Query: 173 DKMLVQMLNED--------PSFKPQGGRLGESDMDYDTDEKSMASLRRRLRRAREQYYRY 224
DK Q+ N+D P F+ + + + + + + DE S + ++ R+++Y
Sbjct: 143 DKKPCQVANDDIDDLSLGEPEFEIKENKQPDFETNDENDEDSPEARHKKFEEMRKKHYDV 202
Query: 225 RSEYTKFVLEALELEDTVKNYDRRDST 251
R+ + K EAL+ ED D DST
Sbjct: 203 RAIFNKKSREALKDEDE----DEDDST 225
>DAT1_MOUSE (Q8C9B9) Death associated transcription factor 1 (Death
inducer-obliterator-1) (DIO-1)
Length = 614
Score = 31.2 bits (69), Expect = 9.1
Identities = 16/36 (44%), Positives = 21/36 (57%)
Query: 185 SFKPQGGRLGESDMDYDTDEKSMASLRRRLRRAREQ 220
S K +GG E D D+D ++ L+ RLRR REQ
Sbjct: 136 SEKAKGGEEEEDTSDSDSDGLTLKELQNRLRRKREQ 171
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.348 0.152 0.521
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 67,928,679
Number of Sequences: 164201
Number of extensions: 2642512
Number of successful extensions: 11612
Number of sequences better than 10.0: 5
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 11611
Number of HSP's gapped (non-prelim): 5
length of query: 660
length of database: 59,974,054
effective HSP length: 117
effective length of query: 543
effective length of database: 40,762,537
effective search space: 22134057591
effective search space used: 22134057591
T: 11
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.8 bits)
S2: 69 (31.2 bits)
Medicago: description of AC121245.1


