

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121241.10 + phase: 0 /pseudo
(330 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
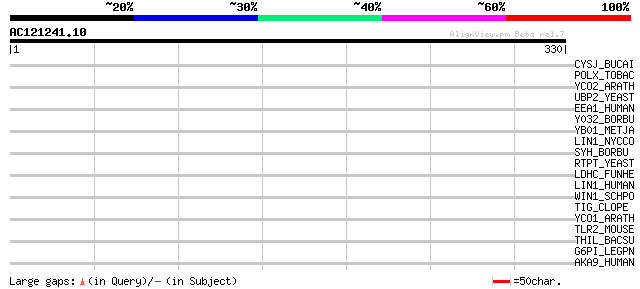
Score E
Sequences producing significant alignments: (bits) Value
CYSJ_BUCAI (P57503) Sulfite reductase [NADPH] flavoprotein alpha... 39 0.014
POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from tran... 36 0.15
YCO2_ARATH (Q9LME2) Hypothetical protein At1g22260 33 0.75
UBP2_YEAST (Q01476) Ubiquitin carboxyl-terminal hydrolase 2 (EC ... 33 0.98
EEA1_HUMAN (Q15075) Early endosome antigen 1 (Endosome-associate... 33 0.98
Y032_BORBU (O51063) Hypothetical protein BB0032 33 1.3
YB01_METJA (Q58501) Hypothetical acetyltransferase MJ1101 (EC 2.... 32 1.7
LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog 32 2.9
SYH_BORBU (O51160) Histidyl-tRNA synthetase (EC 6.1.1.21) (Histi... 31 3.7
RTPT_YEAST (P17558) 40S ribosomal protein PET123, mitochondrial ... 31 3.7
LDHC_FUNHE (Q06176) L-lactate dehydrogenase C chain (EC 1.1.1.27... 31 3.7
LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog 31 4.9
WIN1_SCHPO (O74304) MAP kinase kinase kinase win1 (EC 2.7.1.-) 30 6.4
TIG_CLOPE (Q8XKK0) Trigger factor (TF) 30 6.4
YCO1_ARATH (P61430) Hypothetical protein At1g22275 30 8.3
TLR2_MOUSE (Q9QUN7) Toll-like receptor 2 precursor 30 8.3
THIL_BACSU (O05514) Thiamine-monophosphate kinase (EC 2.7.4.16) ... 30 8.3
G6PI_LEGPN (Q9RDY2) Glucose-6-phosphate isomerase (EC 5.3.1.9) (... 30 8.3
AKA9_HUMAN (Q99996) A-kinase anchor protein 9 (Protein kinase A ... 30 8.3
>CYSJ_BUCAI (P57503) Sulfite reductase [NADPH] flavoprotein
alpha-component (EC 1.8.1.2) (SIR-FP)
Length = 601
Score = 39.3 bits (90), Expect = 0.014
Identities = 45/219 (20%), Positives = 98/219 (44%), Gaps = 26/219 (11%)
Query: 14 KFDGHYDHWAMFMENFLRSKE--------YWDLIENEILVVEAGVEQTEAQRKLIKEQKL 65
+++ +Y+ W+ + + SKE Y D IL ++ A+ ++ QK+
Sbjct: 190 EYEDNYNKWSQDLLQSINSKEKIYKSSVSYIDQENTLILSKNHYTKKNPAEGIILTNQKI 249
Query: 66 MNLKTKNYLFQAISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILH 125
+K + H + I N + + D++ ++ + +++ L E+L
Sbjct: 250 TGRNSKKDV-----HHIEIDISNLNIKYSPGDALGVWYKNDSNLVKNIL-------ELLS 297
Query: 126 MKEGETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMV--SKIDYVVCSIEESNNL 183
+ ETI T + +TI + ++ H E+ + T + I++S SK ++ I ++L
Sbjct: 298 INISETI-TIKNDVITIFDALQNHFELTNNT---KNIIKSYANFSKNKFLKDIISNDSDL 353
Query: 184 DTMTIDELQSSLVHEQRIKSNGEEENGGDNPTTKLLWSI 222
+ TI+ ++H+ +K + ++ G P T L+SI
Sbjct: 354 ENYTINTPLIKMIHDHPLKLSSQQLIGLLRPLTPRLYSI 392
>POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from
transposon TNT 1-94 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1328
Score = 35.8 bits (81), Expect = 0.15
Identities = 28/134 (20%), Positives = 66/134 (48%), Gaps = 11/134 (8%)
Query: 78 ISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGETINTY-- 135
+S DV+ I+++DT + IW +++ + ++ + +L L++ LHM EG ++
Sbjct: 67 LSDDVVNNIIDEDTARGIWTRLESLY--MSKTLTNKLY-LKKQLYALHMSEGTNFLSHLN 123
Query: 136 -FSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMTIDELQSS 194
F+ +T + E + I+ +L S+ S D + +I T+ + ++ S+
Sbjct: 124 VFNGLITQLANLGVKIEEEDKAIL---LLNSLPSSYDNLATTILHGKT--TIELKDVTSA 178
Query: 195 LVHEQRIKSNGEEE 208
L+ ++++ E +
Sbjct: 179 LLLNEKMRKKPENQ 192
>YCO2_ARATH (Q9LME2) Hypothetical protein At1g22260
Length = 871
Score = 33.5 bits (75), Expect = 0.75
Identities = 22/85 (25%), Positives = 38/85 (43%)
Query: 125 HMKEGETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLD 184
H+KE N + ++ H I S+ EKI++ + +K D + +E +
Sbjct: 551 HLKELSQRNDQAINEIRRKYDVEKHEIINSEKDKVEKIIKDLSNKFDKELSDCKEESKRQ 610
Query: 185 TMTIDELQSSLVHEQRIKSNGEEEN 209
+TI E SSL+ R + +E N
Sbjct: 611 LLTIQEEHSSLILSLREEHESKELN 635
>UBP2_YEAST (Q01476) Ubiquitin carboxyl-terminal hydrolase 2 (EC
3.1.2.15) (Ubiquitin thiolesterase 2)
(Ubiquitin-specific processing protease 2)
(Deubiquitinating enzyme 2)
Length = 1272
Score = 33.1 bits (74), Expect = 0.98
Identities = 26/91 (28%), Positives = 47/91 (51%), Gaps = 7/91 (7%)
Query: 119 RDFEILHMKEGETINTYFSRTLTIANKMKAHSEIM---SQTIITEKILRS--MVSKIDYV 173
R++E L + E I YF IAN+ A+ I Q II ++ L + ++ KI+
Sbjct: 464 RNYESLSSLDPENIGLYFDALTYIANRKGAYQLIAYCGKQDIIGQEALENALLMFKINPK 523
Query: 174 VCSIEESNNLDTMTIDELQSSLVHEQRIKSN 204
C+I E N ++I + ++S ++ ++ SN
Sbjct: 524 ECNISELNEATLLSIYKYETS--NKSQVTSN 552
>EEA1_HUMAN (Q15075) Early endosome antigen 1 (Endosome-associated
protein p162) (Zinc finger FYVE domain containing
protein 2)
Length = 1411
Score = 33.1 bits (74), Expect = 0.98
Identities = 38/183 (20%), Positives = 70/183 (37%), Gaps = 16/183 (8%)
Query: 2 NDSSNFVHSIIPKFDGHYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQR---K 58
N + + + K +H + + KE + +E + +E +++ EA K
Sbjct: 684 NKITTQLDQVTAKLQDKQEHCSQLESHLKEYKEKYLSLEQKTEELEGQIKKLEADSLEVK 743
Query: 59 LIKEQKLMNLKTKNYLFQAISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRA------ 112
KEQ L +L+ + L + E + K I S + + + +
Sbjct: 744 ASKEQALQDLQQQRQLNTDLELRATELSKQLEMEKEIVSSTRLDLQKKSEALESIKQKLT 803
Query: 113 ----QLQALRRDFEILHMK---EGETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRS 165
+ Q L++DFE L + + E +N T+T K+K E + + T K S
Sbjct: 804 KQEEEKQILKQDFETLSQETKIQHEELNNRIQTTVTELQKVKMEKEALMTELSTVKDKLS 863
Query: 166 MVS 168
VS
Sbjct: 864 KVS 866
>Y032_BORBU (O51063) Hypothetical protein BB0032
Length = 490
Score = 32.7 bits (73), Expect = 1.3
Identities = 31/124 (25%), Positives = 58/124 (46%), Gaps = 4/124 (3%)
Query: 93 KNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGETINTYFSRTLTIAN--KMKAHS 150
+ IW+S+KT F NR I L+ L +D +IL K + T ++TL N + +
Sbjct: 208 EEIWNSLKTNFDLENRAIFWFLENLDKDLKIL-TKNAPWVKT-LAKTLDNENIQLISKNE 265
Query: 151 EIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMTIDELQSSLVHEQRIKSNGEEENG 210
+I + +KI + + KI + +S N+ + ++ L+ EQ + ++
Sbjct: 266 KINIKLPGFKKINSNKIEKIQNYELNSLDSKNISLKSTYNVKEFLIQEQNVTKYEYQDFL 325
Query: 211 GDNP 214
+NP
Sbjct: 326 KENP 329
>YB01_METJA (Q58501) Hypothetical acetyltransferase MJ1101 (EC
2.7.3.-)
Length = 408
Score = 32.3 bits (72), Expect = 1.7
Identities = 22/73 (30%), Positives = 34/73 (46%), Gaps = 16/73 (21%)
Query: 1 ENDSSNFVHSIIPKFDGHYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLI 60
EN SN +++ I KFD K+ ++LIE + E T+A + LI
Sbjct: 148 ENPKSNLINAGIYKFD----------------KKIFELIEKTKISERGERELTDAIKHLI 191
Query: 61 KEQKLMNLKTKNY 73
KE+K+ +K Y
Sbjct: 192 KEEKVKGIKLNGY 204
>LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog
Length = 1260
Score = 31.6 bits (70), Expect = 2.9
Identities = 42/167 (25%), Positives = 78/167 (46%), Gaps = 23/167 (13%)
Query: 61 KEQKLMNLKTKN-YLFQAISHDV---LETILNKDT-YKNIWDSMKTFF*GSNRVIRAQLQ 115
K KL NL K+ ++ I ++ LE N+DT Y+N+WD+ K V+R +
Sbjct: 247 KTWKLNNLMLKDTWVIDEIKKEITKFLEQNNNQDTNYQNLWDTAKA-------VLRGKFI 299
Query: 116 ALRRDFEILHMKEGETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKID--YV 173
AL+ L E E +N + + ++ + + IT+ +R+ +++I+ +
Sbjct: 300 ALQ---AFLKKTEREEVNNLMGHLKQLEKEEHSNPKPSRRKEITK--IRAELNEIENKRI 354
Query: 174 VCSIEESNNL---DTMTIDELQSSLVHEQRIKS-NGEEENGGDNPTT 216
+ I +S + ID+ ++L ++R+KS NG D TT
Sbjct: 355 IQQINKSKSWFFEKINKIDKPLANLTRKKRVKSLISSIRNGNDEITT 401
>SYH_BORBU (O51160) Histidyl-tRNA synthetase (EC 6.1.1.21)
(Histidine--tRNA ligase) (HisRS)
Length = 457
Score = 31.2 bits (69), Expect = 3.7
Identities = 35/156 (22%), Positives = 66/156 (41%), Gaps = 20/156 (12%)
Query: 6 NFVHSIIPKFDGHYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKL 65
NF+ I KF HY H + F E L E I ++ R + K K+
Sbjct: 154 NFIEGINKKFIIHYSHIGILNSFF----EKLGLKEKSIFIL----------RNIDKIDKI 199
Query: 66 MNLKTKNYLFQAISHDVLETILN----KDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDF 121
K K L I + +++IL+ + T+K+ ++K+ G N I+ +++ + +
Sbjct: 200 GIDKVKEALLLEIEKEAVDSILSLVSLQGTFKDKIQALKSIL-GDNESIK-RVEDVFQHL 257
Query: 122 EILHMKEGETINTYFSRTLTIANKMKAHSEIMSQTI 157
+L +++ +N SR L + SE+ +
Sbjct: 258 SLLKIQDSFNLNLKISRGLDYYTGIVFESEVFGSNM 293
>RTPT_YEAST (P17558) 40S ribosomal protein PET123, mitochondrial
precursor
Length = 318
Score = 31.2 bits (69), Expect = 3.7
Identities = 38/179 (21%), Positives = 75/179 (41%), Gaps = 16/179 (8%)
Query: 43 ILVVEAGVEQTEAQRKLIKEQKLMNLKTKNYLFQAISHDVLETILNKDTYKNIWDSMKTF 102
I+ ++ + Q + + +E +L+ LK +NY ++LN +K + +
Sbjct: 153 IMTLDKMMSQPLLRNRSPEESELLKLK-RNYN---------RSLLNFQAHKKKLNELLNL 202
Query: 103 F*GSNRVIRAQLQALRRDFEILHMKEGETINTY--FSRTLTIAN-KMKAHSEIMSQTIIT 159
+ +N I + Q L++ ++ + + E + Y S N K+ QT I
Sbjct: 203 YHVANEFIVTESQLLKKIDKVFNDETEEFTDAYDVTSNFTQFGNRKLLLSGNTTLQTQIN 262
Query: 160 EKILRSMVSKIDYVVCSIEESNNLDTMTIDELQSSLVHEQRIKSNGEEENGGDNPTTKL 218
I+ S+ ++ + + ++ N D I S ++ SNG NG +N TT L
Sbjct: 263 NAIMGSLSNEKFFDISLVDSYLNKDLKNISNKIDSKLNPT---SNGAGNNGNNNNTTNL 318
>LDHC_FUNHE (Q06176) L-lactate dehydrogenase C chain (EC 1.1.1.27)
(LDH-C)
Length = 333
Score = 31.2 bits (69), Expect = 3.7
Identities = 16/50 (32%), Positives = 30/50 (60%), Gaps = 2/50 (4%)
Query: 21 HWAMFMEN--FLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNL 68
H ++F++ + K+Y + I+VV AGV Q E +R+L +Q+ +N+
Sbjct: 67 HGSLFLKTPKIVADKDYSVTSNSRIVVVTAGVRQQEGERRLNLDQRNVNI 116
>LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog
Length = 1259
Score = 30.8 bits (68), Expect = 4.9
Identities = 39/147 (26%), Positives = 61/147 (40%), Gaps = 22/147 (14%)
Query: 64 KLMNLKTKNYL----FQAISHDVLETILNKDT-YKNIWDSMKTFF*GSNRVIRAQLQALR 118
KL NL +Y +A ET NKDT Y+N+WD+ K G + A +
Sbjct: 250 KLNNLLLNDYWVHNEMKAEIKKFFETNENKDTTYQNLWDTAKAVCRGKFIALNAHKRKQE 309
Query: 119 RD------FEILHMKEGETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRS------M 166
R ++ +++ E N+ SR I EI +Q + +KI S
Sbjct: 310 RSKIDTLISQLKELEKQEQTNSKASRRQEIIKIRAELKEIETQKTL-QKINESRSWFFEK 368
Query: 167 VSKIDYVVCSI----EESNNLDTMTID 189
++KID + + E N +DT+ D
Sbjct: 369 INKIDRPLARLIKKKREKNQIDTIKND 395
>WIN1_SCHPO (O74304) MAP kinase kinase kinase win1 (EC 2.7.1.-)
Length = 1436
Score = 30.4 bits (67), Expect = 6.4
Identities = 25/96 (26%), Positives = 45/96 (46%), Gaps = 16/96 (16%)
Query: 107 NRVIRAQLQALRRDFEIL----HMKEGETINTYFSRTLTIANKMKAHSEIMSQTIITEKI 162
NR ++ QLQ D L ++K + + F + ++ +KAH E + +I E
Sbjct: 694 NRYLKDQLQGPDTDDSALITSFYIKVLDCVRIRFRKLMSFTRILKAHLENSCEYVIKENS 753
Query: 163 LRSMVSKIDYVVCSIEESNNLDTMTIDELQSSLVHE 198
L ++ + +EESN++ T T +S+ HE
Sbjct: 754 LSLLIQR-------LEESNHVLTYT-----ASIEHE 777
>TIG_CLOPE (Q8XKK0) Trigger factor (TF)
Length = 428
Score = 30.4 bits (67), Expect = 6.4
Identities = 21/99 (21%), Positives = 51/99 (51%), Gaps = 11/99 (11%)
Query: 109 VIRAQLQALRRDFEILHMKEGETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVS 168
+I +++ + RD E+ ++G ++ Y+ T + +KMK++ + ++ +
Sbjct: 307 MINREVENMVRDLEMRLGQQGLSLEQYYEFTGSTEDKMKSYMKENAERKV---------- 356
Query: 169 KIDYVVCSIEESNNLDTMTIDELQSSLVHEQRIKSNGEE 207
K D V+ ++ E+ ++ T +EL++ ++ SNG E
Sbjct: 357 KTDLVMSAVTEAEAIEA-TEEELKAKAEEVAKMYSNGAE 394
>YCO1_ARATH (P61430) Hypothetical protein At1g22275
Length = 856
Score = 30.0 bits (66), Expect = 8.3
Identities = 21/85 (24%), Positives = 37/85 (42%)
Query: 125 HMKEGETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLD 184
H+KE N + ++ H I S+ EKI++ + +K D + +E +
Sbjct: 551 HLKELSQRNDQAINEIRRKYDVEKHEIINSEKDKVEKIIKELSTKYDKGLSDCKEESKRQ 610
Query: 185 TMTIDELQSSLVHEQRIKSNGEEEN 209
+TI E SS + R + +E N
Sbjct: 611 LLTIQEEHSSRILNIREEHESKELN 635
>TLR2_MOUSE (Q9QUN7) Toll-like receptor 2 precursor
Length = 784
Score = 30.0 bits (66), Expect = 8.3
Identities = 20/67 (29%), Positives = 33/67 (48%), Gaps = 10/67 (14%)
Query: 4 SSNFVHSIIPKFDGHYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQ 63
S NFV S K++ + H+ +F EN + ILV+ +E+ ++ K +
Sbjct: 704 SENFVRSEWCKYELDFSHFRLFDENN----------DAAILVLLEPIERKAIPQRFCKLR 753
Query: 64 KLMNLKT 70
K+MN KT
Sbjct: 754 KIMNTKT 760
>THIL_BACSU (O05514) Thiamine-monophosphate kinase (EC 2.7.4.16)
(Thiamine-phosphate kinase)
Length = 325
Score = 30.0 bits (66), Expect = 8.3
Identities = 21/78 (26%), Positives = 39/78 (49%), Gaps = 12/78 (15%)
Query: 8 VHSIIPKFDGHYDHWAMF-MENF----LRSKEYWDLIENEILVVEAGV-------EQTEA 55
+HS +PK ++ WA+F E+F S E W++++ E + + E+T++
Sbjct: 244 IHSDLPKLHPNWKEWALFGGEDFELTGTVSNEEWEVLKQECAALHLPITKIGYVREKTKS 303
Query: 56 QRKLIKEQKLMNLKTKNY 73
+ L +Q M L+ K Y
Sbjct: 304 KVILKTDQTSMILEKKGY 321
>G6PI_LEGPN (Q9RDY2) Glucose-6-phosphate isomerase (EC 5.3.1.9)
(GPI) (Phosphoglucose isomerase) (PGI) (Phosphohexose
isomerase) (PHI)
Length = 497
Score = 30.0 bits (66), Expect = 8.3
Identities = 13/44 (29%), Positives = 28/44 (63%), Gaps = 1/44 (2%)
Query: 142 IANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDT 185
I N MK+H+E++S ++ ++ R ++ D + C + +SNN ++
Sbjct: 2 IRNSMKSHTELLSWNLLQKEADRVRLNS-DSLTCVVPDSNNYES 44
>AKA9_HUMAN (Q99996) A-kinase anchor protein 9 (Protein kinase A
anchoring protein 9) (PRKA9) (A-kinase anchor protein 450
kDa) (AKAP 450) (A-kinase anchor protein 350 kDa) (AKAP
350) (hgAKAP 350) (AKAP 120 like protein) (Hyperion
protein) (Yotiao protein
Length = 3911
Score = 30.0 bits (66), Expect = 8.3
Identities = 35/194 (18%), Positives = 79/194 (40%), Gaps = 12/194 (6%)
Query: 16 DGHYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNLKTKNYLF 75
DG+ D +F + D +E E+L +++ EA+++ I+E++ + L +
Sbjct: 1937 DGYADEKTLFERQIQEKTDIIDRLEQELLCASNRLQELEAEQQQIQEEREL-LSRQKEAM 1995
Query: 76 QAISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGETINTY 135
+A + V + +L ++T K + + ++ ++ Q + +R D + +
Sbjct: 1996 KAEAGPVEQQLL-QETEKLMKEKLE---------VQCQAEKVRDDLQKQVKALEIDVEEQ 2045
Query: 136 FSRTLTIANKMKAH-SEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMTIDELQSS 194
SR + + + ++ Q EK L M +D E ++ I +L+
Sbjct: 2046 VSRFIELEQEKNTELMDLRQQNQALEKQLEKMRKFLDEQAIDREHERDVFQQEIQKLEQQ 2105
Query: 195 LVHEQRIKSNGEEE 208
L R + E +
Sbjct: 2106 LKVVPRFQPISEHQ 2119
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.334 0.144 0.439
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 33,497,822
Number of Sequences: 164201
Number of extensions: 1252987
Number of successful extensions: 5313
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 18
Number of HSP's that attempted gapping in prelim test: 5305
Number of HSP's gapped (non-prelim): 25
length of query: 330
length of database: 59,974,054
effective HSP length: 111
effective length of query: 219
effective length of database: 41,747,743
effective search space: 9142755717
effective search space used: 9142755717
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 66 (30.0 bits)
Medicago: description of AC121241.10


