

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
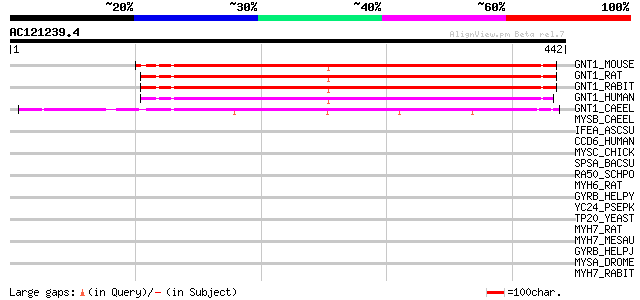
Query= AC121239.4 - phase: 0
(442 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
GNT1_MOUSE (P27808) Alpha-1,3-mannosyl-glycoprotein 2-beta-N-ace... 277 4e-74
GNT1_RAT (Q09325) Alpha-1,3-mannosyl-glycoprotein 2-beta-N-acety... 276 8e-74
GNT1_RABIT (P27115) Alpha-1,3-mannosyl-glycoprotein 2-beta-N-ace... 272 1e-72
GNT1_HUMAN (P26572) Alpha-1,3-mannosyl-glycoprotein 2-beta-N-ace... 266 6e-71
GNT1_CAEEL (Q11068) Putative alpha-1,3-mannosyl-glycoprotein 2-b... 238 2e-62
MYSB_CAEEL (P02566) Myosin heavy chain B (MHC B) 35 0.39
IFEA_ASCSU (P23730) Intermediate filament protein A (IF-A) (Frag... 34 0.86
CCD6_HUMAN (Q16204) Coiled-coil domain containing protein 6 (H4 ... 33 1.1
MYSC_CHICK (P29616) Myosin heavy chain, cardiac muscle isoform (... 33 1.5
SPSA_BACSU (P39621) Spore coat polysaccharide biosynthesis prote... 33 1.9
RA50_SCHPO (Q9UTJ8) DNA repair protein rad50 33 1.9
MYH6_RAT (P02563) Myosin heavy chain, cardiac muscle alpha isofo... 33 1.9
GYRB_HELPY (P55992) DNA gyrase subunit B (EC 5.99.1.3) 33 1.9
YC24_PSEPK (P43037) Hypothetical protein PP1224 precursor (ORF2) 32 2.5
TP20_YEAST (P33891) Protein transport protein TIP20 32 2.5
MYH7_RAT (P02564) Myosin heavy chain, cardiac muscle beta isofor... 32 2.5
MYH7_MESAU (P13540) Myosin heavy chain, cardiac muscle beta isof... 32 2.5
GYRB_HELPJ (Q9ZLX3) DNA gyrase subunit B (EC 5.99.1.3) 32 3.3
MYSA_DROME (P05661) Myosin heavy chain, muscle 32 4.3
MYH7_RABIT (P04461) Myosin heavy chain, cardiac muscle beta isof... 32 4.3
>GNT1_MOUSE (P27808) Alpha-1,3-mannosyl-glycoprotein
2-beta-N-acetylglucosaminyltransferase (EC 2.4.1.101)
(N-glycosyl-oligosaccharide-glycoprotein
N-acetylglucosaminyltransferase I) (GNT-I) (GlcNAc-T I)
Length = 447
Score = 277 bits (708), Expect = 4e-74
Identities = 142/339 (41%), Positives = 210/339 (61%), Gaps = 10/339 (2%)
Query: 101 VQVPVAAVVIMACNRADYLERTINSVLKYQRPISSRFPLFVSQDGSNSDVKRKALSYDE- 159
VQ+P+ +++AC+R+ + R ++ +L Y RP + RFP+ VSQD + + + SY
Sbjct: 106 VQIPI---LVIACDRST-VRRCLDKLLHY-RPSAERFPIIVSQDCGHEETAQVIASYGTA 160
Query: 160 LSHMQHLDFEPVQTERPGELI-AYYKIARHYKWALGQLFDKHNFNRVIILEDDMEIAPDF 218
++H++ D + + YYKIARHY+WALGQ+F+K F +++EDD+E+APDF
Sbjct: 161 VTHIRQPDLSNIAVQPDHRKFQGYYKIARHYRWALGQIFNKFKFPAAVVVEDDLEVAPDF 220
Query: 219 FDYFEAMATLLDKDKSIMAVSSWNDNGQKQFVHD--PYELYRSDFFPGLGWMLARSTWDE 276
F+YF+A LL D S+ VS+WNDNG++Q V P LYR+DFFPGLGW+L W E
Sbjct: 221 FEYFQATYPLLRTDPSLWCVSAWNDNGKEQMVDSSKPELLYRTDFFPGLGWLLLADLWAE 280
Query: 277 LSPKWPKAYWDDWLRLKENHKGRQFIRPEVCRTYNFGEHGSSLGQFFKQYLEPIKLNEVQ 336
L PKWPKA+WDDW+R E KGR IRPE+ RT FG G S GQFF Q+L+ IKLN+
Sbjct: 281 LEPKWPKAFWDDWMRRPEQRKGRACIRPEISRTMTFGRKGVSHGQFFDQHLKFIKLNQQF 340
Query: 337 VDWKSMDLSYLLEDKYVIHFANIIKKAKPVSGADFAQKARHIDGDVRIKYKDQWDFENIA 396
V + +DLSYL ++ Y F + A + + G+VR++Y + F+ A
Sbjct: 341 VPFTQLDLSYLQQEAYDRDFLAQVYGAPQLQVEKVRTNDQKELGEVRVQYTSRDSFKAFA 400
Query: 397 QQFGIFQEWKDGVPRTAYKGVVVFRYQTTKRIFLVGPES 435
+ G+ + K GVPR Y+G+V F+++ +R+ L P++
Sbjct: 401 KALGVMDDLKSGVPRAGYRGIVTFQFR-GRRVHLAPPQT 438
>GNT1_RAT (Q09325) Alpha-1,3-mannosyl-glycoprotein
2-beta-N-acetylglucosaminyltransferase (EC 2.4.1.101)
(N-glycosyl-oligosaccharide-glycoprotein
N-acetylglucosaminyltransferase I) (GNT-I) (GlcNAc-T I)
Length = 447
Score = 276 bits (706), Expect = 8e-74
Identities = 141/335 (42%), Positives = 205/335 (61%), Gaps = 7/335 (2%)
Query: 105 VAAVVIMACNRADYLERTINSVLKYQRPISSRFPLFVSQDGSNSDVKRKALSYDE-LSHM 163
V ++++AC+R+ + R ++ +L Y RP + FP+ VSQD + + + SY ++H+
Sbjct: 107 VIPILVIACDRST-VRRCLDKLLHY-RPSAEHFPIIVSQDCGHEETAQVIASYGTAVTHI 164
Query: 164 QHLDFEPVQTERPGELI-AYYKIARHYKWALGQLFDKHNFNRVIILEDDMEIAPDFFDYF 222
+ D + + YYKIARHY+WALGQ+F+K F +++EDD+E+APDFF+YF
Sbjct: 165 RQPDLSNIAVQPDHRKFQGYYKIARHYRWALGQIFNKFKFPAAVVVEDDLEVAPDFFEYF 224
Query: 223 EAMATLLDKDKSIMAVSSWNDNGQKQFVHD--PYELYRSDFFPGLGWMLARSTWDELSPK 280
+A LL D S+ VS+WNDNG++Q V P LYR+DFFPGLGW+L W EL PK
Sbjct: 225 QATYPLLKADPSLWCVSAWNDNGKEQMVDSSKPELLYRTDFFPGLGWLLLADLWAELEPK 284
Query: 281 WPKAYWDDWLRLKENHKGRQFIRPEVCRTYNFGEHGSSLGQFFKQYLEPIKLNEVQVDWK 340
WPKA+WDDW+R E KGR IRPE+ RT FG G S GQFF Q+L+ IKLN+ V +
Sbjct: 285 WPKAFWDDWMRRPEQRKGRACIRPEISRTMTFGRKGVSHGQFFDQHLKFIKLNQQFVPFT 344
Query: 341 SMDLSYLLEDKYVIHFANIIKKAKPVSGADFAQKARHIDGDVRIKYKDQWDFENIAQQFG 400
+DLSYL + Y F + A + R G+VR++Y + F+ A+ G
Sbjct: 345 QLDLSYLQREAYDRDFLAQVYGAPQLQVEKVRTNDRKELGEVRVQYTSRDSFKAFAKALG 404
Query: 401 IFQEWKDGVPRTAYKGVVVFRYQTTKRIFLVGPES 435
+ + K GVPR Y+G+V F+++ +R+ L PE+
Sbjct: 405 VMDDLKSGVPRAGYRGIVTFQFR-GRRVHLAPPET 438
>GNT1_RABIT (P27115) Alpha-1,3-mannosyl-glycoprotein
2-beta-N-acetylglucosaminyltransferase (EC 2.4.1.101)
(N-glycosyl-oligosaccharide-glycoprotein
N-acetylglucosaminyltransferase I) (GNT-I) (GlcNAc-T I)
Length = 447
Score = 272 bits (695), Expect = 1e-72
Identities = 138/335 (41%), Positives = 204/335 (60%), Gaps = 7/335 (2%)
Query: 105 VAAVVIMACNRADYLERTINSVLKYQRPISSRFPLFVSQDGSNSDVKRKALSYDE-LSHM 163
V ++++AC+R+ + R ++ +L Y RP + FP+ VSQD + + + SY ++H+
Sbjct: 107 VIPILVIACDRST-VRRCLDKLLHY-RPSAELFPIIVSQDCGHEETAQVIASYGSAVTHI 164
Query: 164 QHLDFEPVQTERPGELI-AYYKIARHYKWALGQLFDKHNFNRVIILEDDMEIAPDFFDYF 222
+ D + + YYKIARHY+WALGQ+F N+ +++EDD+E+APDFF+YF
Sbjct: 165 RQPDLSNIAVQPDHRKFQGYYKIARHYRWALGQIFHNFNYPAAVVVEDDLEVAPDFFEYF 224
Query: 223 EAMATLLDKDKSIMAVSSWNDNGQKQFVHD--PYELYRSDFFPGLGWMLARSTWDELSPK 280
+A LL D S+ VS+WNDNG++Q V P LYR+DFFPGLGW+L W EL PK
Sbjct: 225 QATYPLLKADPSLWCVSAWNDNGKEQMVDSSKPELLYRTDFFPGLGWLLLAELWAELEPK 284
Query: 281 WPKAYWDDWLRLKENHKGRQFIRPEVCRTYNFGEHGSSLGQFFKQYLEPIKLNEVQVDWK 340
WPKA+WDDW+R E KGR +RPE+ RT FG G S GQFF Q+L+ IKLN+ V +
Sbjct: 285 WPKAFWDDWMRRPEQRKGRACVRPEISRTMTFGRKGVSHGQFFDQHLKFIKLNQQFVPFT 344
Query: 341 SMDLSYLLEDKYVIHFANIIKKAKPVSGADFAQKARHIDGDVRIKYKDQWDFENIAQQFG 400
+DLSYL ++ Y F + A + R G+VR++Y + F+ A+ G
Sbjct: 345 QLDLSYLQQEAYDRDFLARVYGAPQLQVEKVRTNDRKELGEVRVQYTGRDSFKAFAKALG 404
Query: 401 IFQEWKDGVPRTAYKGVVVFRYQTTKRIFLVGPES 435
+ + K GVPR Y+G+V F ++ +R+ L P++
Sbjct: 405 VMDDLKSGVPRAGYRGIVTFLFR-GRRVHLAPPQT 438
>GNT1_HUMAN (P26572) Alpha-1,3-mannosyl-glycoprotein
2-beta-N-acetylglucosaminyltransferase (EC 2.4.1.101)
(N-glycosyl-oligosaccharide-glycoprotein
N-acetylglucosaminyltransferase I) (GNT-I) (GlcNAc-T I)
Length = 445
Score = 266 bits (681), Expect = 6e-71
Identities = 138/333 (41%), Positives = 200/333 (59%), Gaps = 7/333 (2%)
Query: 105 VAAVVIMACNRADYLERTINSVLKYQRPISSRFPLFVSQDGSNSDVKRKALSYDE-LSHM 163
V ++++AC+R+ + R ++ +L Y RP + FP+ VSQD + + + SY ++H+
Sbjct: 105 VIPILVIACDRST-VRRCLDKLLHY-RPSAELFPIIVSQDCGHEETAQAIASYGSAVTHI 162
Query: 164 QHLDFEPVQTERPGELI-AYYKIARHYKWALGQLFDKHNFNRVIILEDDMEIAPDFFDYF 222
+ D + YYKIARHY+WALGQ+F + F +++EDD+E+APDFF+YF
Sbjct: 163 RQPDLSSIAVPPDHRKFQGYYKIARHYRWALGQVFRQFRFPAAVVVEDDLEVAPDFFEYF 222
Query: 223 EAMATLLDKDKSIMAVSSWNDNGQKQFVHD--PYELYRSDFFPGLGWMLARSTWDELSPK 280
A LL D S+ VS+WNDNG++Q V P LYR+DFFPGLGW+L W EL PK
Sbjct: 223 RATYPLLKADPSLWCVSAWNDNGKEQMVDASRPELLYRTDFFPGLGWLLLAELWAELEPK 282
Query: 281 WPKAYWDDWLRLKENHKGRQFIRPEVCRTYNFGEHGSSLGQFFKQYLEPIKLNEVQVDWK 340
WPKA+WDDW+R E +GR IRPE+ RT FG G S GQFF Q+L+ IKLN+ V +
Sbjct: 283 WPKAFWDDWMRRPEQRQGRACIRPEISRTMTFGRKGVSHGQFFDQHLKFIKLNQQFVHFT 342
Query: 341 SMDLSYLLEDKYVIHFANIIKKAKPVSGADFAQKARHIDGDVRIKYKDQWDFENIAQQFG 400
+DLSYL + Y F + A + R G+VR++Y + F+ A+ G
Sbjct: 343 QLDLSYLQREAYDRDFLARVYGAPQLQVEKVRTNDRKELGEVRVQYTGRDSFKAFAKALG 402
Query: 401 IFQEWKDGVPRTAYKGVVVFRYQTTKRIFLVGP 433
+ + K GVPR Y+G+V F+++ +R+ L P
Sbjct: 403 VMDDLKSGVPRAGYRGIVTFQFR-GRRVHLAPP 434
>GNT1_CAEEL (Q11068) Putative alpha-1,3-mannosyl-glycoprotein
2-beta-N-acetylglucosaminyltransferase (EC 2.4.1.101)
(N-glycosyl-oligosaccharide-glycoprotein
N-acetylglucosaminyltransferase I) (GNT-I) (GlcNAc-T I)
Length = 449
Score = 238 bits (608), Expect = 2e-62
Identities = 142/443 (32%), Positives = 244/443 (55%), Gaps = 28/443 (6%)
Query: 8 FRFLLFVAALVFIYIQMRLFASQSQYADRLAAAIEAENHCTAQIRSLIDQISLQQGRILD 67
F +FV L +Y++ + ++++ D + +E+ N ++ +I+ +
Sbjct: 8 FIIFIFVFILWTLYVENDI-TNRTRNTDNIDDLLESANRLERLLKFEAKKIAALAEDVHK 66
Query: 68 LQQERNRREQECSQIKSLVQDLERKDVRRLIDKVQVPVAAVVIMACNRADYLERTINSVL 127
++ R + +++++ +D+++ D + V ++ +CNRA + + ++
Sbjct: 67 IRANRKGKHV-------IMEEMVSQDLKQWKDPIPV-----LVFSCNRAMAVRDHVEKLI 114
Query: 128 KYQRPISSRFPLFVSQDGSNSDVKRKALSY-DELSHMQHLDFEPVQTERPG---ELIAYY 183
+Y RP +FP+ V+QD N +VK + + D++ +++HL + P + AYY
Sbjct: 115 RY-RPSQEKFPIIVTQDCDNENVKNEVKKFGDKVEYIKHLAGDKANITIPPSHRQYTAYY 173
Query: 184 KIARHYKWALGQLFDKHNFNRVIILEDDMEIAPDFFDYFEAMATLLDKDKSIMAVSSWND 243
+IARHYK AL +F ++ VII EDD++I+PDFF YF + LL+ D+ + V++WND
Sbjct: 174 RIARHYKLALNHVFVDKGYSSVIITEDDLDISPDFFSYFSSTRYLLENDEKLWCVTAWND 233
Query: 244 NGQKQFVH--DPYELYRSDFFPGLGWMLARSTWDELSPKWPKAYWDDWLRLKENHKGRQF 301
NG+++ + LYRSDFF GLGWM++ TW EL P WP +WDDW+R K RQ
Sbjct: 234 NGKQENIDMTAASTLYRSDFFAGLGWMMSSKTWHELEPIWPVGFWDDWMRDPARRKDRQC 293
Query: 302 IRPEVCRT--YNFGEHGSSLGQFFKQYLEPIKLNEVQVDWKSMDLSYLLEDKYVIHF-AN 358
IRPE+ RT ++G+ G+S GQFF ++L IK+N+ +++ +DL YLL +
Sbjct: 294 IRPEISRTGMMSYGKEGASKGQFFSKHLAKIKVNDKYINFGKIDLDYLLPANFAKKTNLE 353
Query: 359 IIKKAKPVS---GADFAQKARHIDGDVRIKYKDQWDFENIAQQFGIFQEWKDGVPRTAYK 415
++K+A +S A F + + VR+ Y D+ A + I ++K GVPRTAY
Sbjct: 354 VMKEAVELSIDNVASFVLSSENKGKSVRVMYDGNIDYIRKADKLHIMHDFKAGVPRTAYD 413
Query: 416 GVVVFRYQTTKRIFLVGPESLKL 438
G+V + RI+LV P+ K+
Sbjct: 414 GIVTC-FINGIRIYLV-PDRTKV 434
>MYSB_CAEEL (P02566) Myosin heavy chain B (MHC B)
Length = 1966
Score = 35.0 bits (79), Expect = 0.39
Identities = 18/60 (30%), Positives = 31/60 (51%)
Query: 31 SQYADRLAAAIEAENHCTAQIRSLIDQISLQQGRILDLQQERNRREQECSQIKSLVQDLE 90
S +RLA + + Q+ L DQ++ + R D+Q+ + + E E +K +QDLE
Sbjct: 911 SDAEERLAKLEAQQKDASKQLSELNDQLADNEDRTADVQRAKKKIEAEVEALKKQIQDLE 970
>IFEA_ASCSU (P23730) Intermediate filament protein A (IF-A)
(Fragment)
Length = 497
Score = 33.9 bits (76), Expect = 0.86
Identities = 26/78 (33%), Positives = 37/78 (47%), Gaps = 7/78 (8%)
Query: 31 SQYADRLAAAIEAENHCTAQIRSLIDQISLQQGRI-LDLQQERNRREQECSQIKSLVQDL 89
S DRLA+ IE AQ R L + L +GR D R E E + + LV D
Sbjct: 2 SDLNDRLASYIEKVRFLEAQNRKLAADLDLLRGRWGKDTLSVRAMYEGELQEARKLVNDT 61
Query: 90 ER------KDVRRLIDKV 101
R K++R+L+D++
Sbjct: 62 NRQREELEKEIRKLLDEL 79
>CCD6_HUMAN (Q16204) Coiled-coil domain containing protein 6 (H4
protein)
Length = 585
Score = 33.5 bits (75), Expect = 1.1
Identities = 22/74 (29%), Positives = 37/74 (49%), Gaps = 10/74 (13%)
Query: 22 IQMRLFASQSQYADRLAAAIEAENHCTAQIRSLIDQISLQQGRILDLQQERNRREQECSQ 81
++ +L A+Q Q+++++A +E E H + + LQ+ LQ+E RRE C Q
Sbjct: 272 LKKQLRAAQLQHSEKMAQYLEEERHMREE------NLRLQR----KLQREMERREALCRQ 321
Query: 82 IKSLVQDLERKDVR 95
+ LE D R
Sbjct: 322 LSESESSLEMDDER 335
>MYSC_CHICK (P29616) Myosin heavy chain, cardiac muscle isoform
(Fragment)
Length = 1102
Score = 33.1 bits (74), Expect = 1.5
Identities = 18/70 (25%), Positives = 38/70 (53%), Gaps = 3/70 (4%)
Query: 24 MRLFASQSQYAD---RLAAAIEAENHCTAQIRSLIDQISLQQGRILDLQQERNRREQECS 80
++L A Q AD R I+++ A+++ L +++ ++ +L ++ + E ECS
Sbjct: 55 LQLQAEQDTLADAEERCDLLIKSKIQLEAKVKELTERVEDEEEMNSELTSKKRKLEDECS 114
Query: 81 QIKSLVQDLE 90
++K + DLE
Sbjct: 115 ELKKDIDDLE 124
>SPSA_BACSU (P39621) Spore coat polysaccharide biosynthesis protein
spsA
Length = 256
Score = 32.7 bits (73), Expect = 1.9
Identities = 15/47 (31%), Positives = 29/47 (60%), Gaps = 3/47 (6%)
Query: 103 VPVAAVVIMACNRADYLERTINSVLKYQRPISSRFPLFVSQDGSNSD 149
+P +V++ + N++DY+ ++I+S+L S F LF+ D SN +
Sbjct: 1 MPKVSVIMTSYNKSDYVAKSISSILS---QTFSDFELFIMDDNSNEE 44
>RA50_SCHPO (Q9UTJ8) DNA repair protein rad50
Length = 1290
Score = 32.7 bits (73), Expect = 1.9
Identities = 28/104 (26%), Positives = 51/104 (48%), Gaps = 7/104 (6%)
Query: 68 LQQERNRREQECSQ--IKSLVQDLERKDVRRLIDKVQVPVAAVVIMACNRADYLERTINS 125
LQ+E ++ E S IKSL Q+ E K++ L+DK Q +++V NR D + ++S
Sbjct: 461 LQREDFTKDVEKSDLWIKSLRQEYESKNLLELLDKHQTALSSVE----NRLDEISEIVDS 516
Query: 126 VLKYQRPISSRFPLFVSQDGSNSDVKRKALSYDELSHMQHLDFE 169
KY + ++ +F + S + L + S + + +E
Sbjct: 517 YHKYS-GVRTKLQVFEENKTNKSAILANQLMTLKSSFSEVMSYE 559
>MYH6_RAT (P02563) Myosin heavy chain, cardiac muscle alpha isoform
(MyHC-alpha)
Length = 1938
Score = 32.7 bits (73), Expect = 1.9
Identities = 17/72 (23%), Positives = 39/72 (53%), Gaps = 3/72 (4%)
Query: 22 IQMRLFASQSQYAD---RLAAAIEAENHCTAQIRSLIDQISLQQGRILDLQQERNRREQE 78
+Q+++ A Q AD R I+ + A+++ + +++ ++ +L ++ + E E
Sbjct: 888 LQLQVQAEQDNLADAEERCDQLIKNKIQLEAKVKEMTERLEDEEEMNAELTAKKRKLEDE 947
Query: 79 CSQIKSLVQDLE 90
CS++K + DLE
Sbjct: 948 CSELKKDIDDLE 959
>GYRB_HELPY (P55992) DNA gyrase subunit B (EC 5.99.1.3)
Length = 773
Score = 32.7 bits (73), Expect = 1.9
Identities = 26/155 (16%), Positives = 68/155 (43%), Gaps = 11/155 (7%)
Query: 62 QGRILDLQQERNRREQECSQIKSLVQ----------DLERKDVRRLIDKVQVPVAAVVIM 111
+G+IL++++ + + +IK+++ D+ER ++I V I
Sbjct: 445 KGKILNVEKSHLSKILKSEEIKNMITAFGCGIQESFDIERLRYHKIIIMTDADVDGSHIQ 504
Query: 112 ACNRADYLERTINSVLKYQRPISSRFPLFVSQDGSNSDVKRKALSYDELSHMQHLDFEPV 171
+ R + +++ ++ PL+ + G + +++ D ++ +
Sbjct: 505 TLLMT-FFYRYLRPLIEQGHVYIAQAPLYKYKKGKTEIYLKDSVALDHFLIEHGINSVDI 563
Query: 172 QTERPGELIAYYKIARHYKWALGQLFDKHNFNRVI 206
+ +L+ K+ARHY++AL +L ++N ++
Sbjct: 564 EGIGKNDLMNLLKVARHYRYALLELEKRYNLLEIL 598
>YC24_PSEPK (P43037) Hypothetical protein PP1224 precursor (ORF2)
Length = 268
Score = 32.3 bits (72), Expect = 2.5
Identities = 22/67 (32%), Positives = 37/67 (54%), Gaps = 4/67 (5%)
Query: 27 FASQSQYADRLAAA-IEAENHCTAQIRSLIDQISLQQGRILDLQQERNRREQECSQIKSL 85
+ + YA A+A A+ Q++ + DQ+S QQG I +LQ + +R +QE +
Sbjct: 38 YGTSGAYAGSGASAPASAQGQLFMQLQQMQDQLSRQQGIIEELQNDVSRMKQENLE---R 94
Query: 86 VQDLERK 92
QDL+R+
Sbjct: 95 YQDLDRR 101
>TP20_YEAST (P33891) Protein transport protein TIP20
Length = 701
Score = 32.3 bits (72), Expect = 2.5
Identities = 28/103 (27%), Positives = 47/103 (45%), Gaps = 14/103 (13%)
Query: 233 KSIMAVSSWNDNGQKQFVHDPYELYRSDFFPGLGWMLARSTWDELSPKWPKAYWDDWLRL 292
+SI+ ++ +N NG QF+HD L P + EL L+L
Sbjct: 601 ESILKLNKFNQNGLNQFLHDFKSLSSILSLPSHATNYKCMSLHELV---------KILKL 651
Query: 293 KENHKGRQFIRPEVCRTYNFGEHGSSLGQFFK-QYLEPIKLNE 334
K + +QF+ PE +T NF +SL + + +YL+ K+ +
Sbjct: 652 KYDPNNQQFLNPEYIKTGNF----TSLKEAYSIKYLKDTKIQD 690
>MYH7_RAT (P02564) Myosin heavy chain, cardiac muscle beta isoform
(MyHC-beta)
Length = 1935
Score = 32.3 bits (72), Expect = 2.5
Identities = 17/72 (23%), Positives = 39/72 (53%), Gaps = 3/72 (4%)
Query: 22 IQMRLFASQSQYAD---RLAAAIEAENHCTAQIRSLIDQISLQQGRILDLQQERNRREQE 78
+Q+++ A Q AD R I+ + A+++ + +++ ++ +L ++ + E E
Sbjct: 887 LQLQVQAEQDNLADAEERCDQLIKNKIQLEAKVKEMTERLEDEEEMNAELTAKKRKLEDE 946
Query: 79 CSQIKSLVQDLE 90
CS++K + DLE
Sbjct: 947 CSELKRDIDDLE 958
>MYH7_MESAU (P13540) Myosin heavy chain, cardiac muscle beta isoform
(MyHC-beta)
Length = 1934
Score = 32.3 bits (72), Expect = 2.5
Identities = 17/72 (23%), Positives = 39/72 (53%), Gaps = 3/72 (4%)
Query: 22 IQMRLFASQSQYAD---RLAAAIEAENHCTAQIRSLIDQISLQQGRILDLQQERNRREQE 78
+Q+++ A Q AD R I+ + A+++ + +++ ++ +L ++ + E E
Sbjct: 886 LQLQVQAEQDNLADAEERCDQLIKNKIQLEAKVKEMTERLEDEEEMNAELTAKKRKLEDE 945
Query: 79 CSQIKSLVQDLE 90
CS++K + DLE
Sbjct: 946 CSELKRDIDDLE 957
>GYRB_HELPJ (Q9ZLX3) DNA gyrase subunit B (EC 5.99.1.3)
Length = 773
Score = 32.0 bits (71), Expect = 3.3
Identities = 26/155 (16%), Positives = 67/155 (42%), Gaps = 11/155 (7%)
Query: 62 QGRILDLQQERNRREQECSQIKSLVQ----------DLERKDVRRLIDKVQVPVAAVVIM 111
+G+IL++++ + + +IK+++ D+ER ++I V I
Sbjct: 445 KGKILNVEKSHLSKILKSEEIKNMITAFGCGIQESFDIERLRYHKIIIMTDADVDGSHIQ 504
Query: 112 ACNRADYLERTINSVLKYQRPISSRFPLFVSQDGSNSDVKRKALSYDELSHMQHLDFEPV 171
+ R + +++ ++ PL+ + G + +++ D ++ +
Sbjct: 505 TLLMT-FFYRYLRPLIEQGHVFIAQAPLYKYKKGKTEIYLKDSVALDHFLIEHGINSVDI 563
Query: 172 QTERPGELIAYYKIARHYKWALGQLFDKHNFNRVI 206
+ +L+ K+ARHY++ L +L ++N V+
Sbjct: 564 EGIGKNDLMNLLKVARHYRYTLLELEKRYNLLEVL 598
>MYSA_DROME (P05661) Myosin heavy chain, muscle
Length = 1962
Score = 31.6 bits (70), Expect = 4.3
Identities = 27/143 (18%), Positives = 61/143 (41%), Gaps = 15/143 (10%)
Query: 33 YADRLAAAIEAENHCTAQIRSLIDQISLQQGRILDLQQERNRREQECSQIKSLVQDLERK 92
Y +R A +N Q+R + ++++ ++ L Q++ + +QE S +K ++DLE
Sbjct: 900 YQERNAKLTAQKNDLENQLRDIQERLTQEEDARNQLFQQKKKADQEISGLKKDIEDLELN 959
Query: 93 DVRRLIDKVQVPVAAVVIMACNRADYLERTINSVLKYQRPISSRFPLFVSQDGSNSDVKR 152
+ DK D+ R +N + +Q + ++ G +
Sbjct: 960 VQKAEQDKA-------------TKDHQIRNLNDEIAHQDELINKLNKEKKMQGETNQKTG 1006
Query: 153 KAL--SYDELSHMQHLDFEPVQT 173
+ L + D+++H+ + + QT
Sbjct: 1007 EELQAAEDKINHLNKVKAKLEQT 1029
>MYH7_RABIT (P04461) Myosin heavy chain, cardiac muscle beta isoform
(MyHC-beta) (Beta isomyosin) (Fragment)
Length = 736
Score = 31.6 bits (70), Expect = 4.3
Identities = 17/72 (23%), Positives = 39/72 (53%), Gaps = 3/72 (4%)
Query: 22 IQMRLFASQSQYAD---RLAAAIEAENHCTAQIRSLIDQISLQQGRILDLQQERNRREQE 78
+Q+++ A Q AD R I+ + A+++ + +++ ++ +L ++ + E E
Sbjct: 451 LQLQVQAEQDNLADAEERCDQLIKNKIQLEAKVKEMNERLEDEEEMNAELTAKKRKLEDE 510
Query: 79 CSQIKSLVQDLE 90
CS++K + DLE
Sbjct: 511 CSELKRDIDDLE 522
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.324 0.139 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 51,566,111
Number of Sequences: 164201
Number of extensions: 2209837
Number of successful extensions: 5935
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 12
Number of HSP's successfully gapped in prelim test: 24
Number of HSP's that attempted gapping in prelim test: 5869
Number of HSP's gapped (non-prelim): 70
length of query: 442
length of database: 59,974,054
effective HSP length: 113
effective length of query: 329
effective length of database: 41,419,341
effective search space: 13626963189
effective search space used: 13626963189
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 67 (30.4 bits)
Medicago: description of AC121239.4


