

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121233.1 - phase: 0 /pseudo
(1172 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
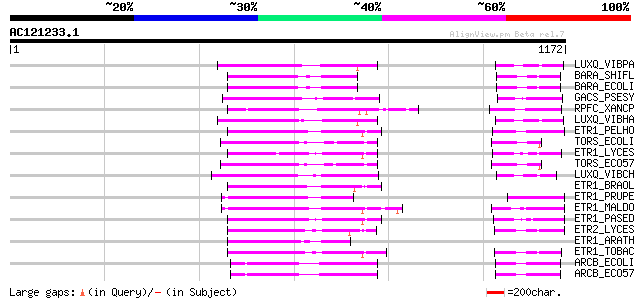
Score E
Sequences producing significant alignments: (bits) Value
LUXQ_VIBPA (Q87GU5) Autoinducer 2 sensor kinase/phosphatase luxQ... 152 4e-36
BARA_SHIFL (P59342) Sensor protein barA (EC 2.7.3.-) 150 2e-35
BARA_ECOLI (P26607) Sensor protein barA (EC 2.7.3.-) 150 2e-35
GACS_PSESY (P48027) Sensor protein gacS (EC 2.7.3.-) 146 4e-34
RPFC_XANCP (P49246) Sensory/regulatory protein rpfC (EC 2.7.3.-) 145 6e-34
LUXQ_VIBHA (P54302) Autoinducer 2 sensor kinase/phosphatase luxQ... 142 7e-33
ETR1_PELHO (Q9XH58) Ethylene receptor 1 (EC 2.7.3.-) (PhETR1) 138 8e-32
TORS_ECOLI (P39453) Sensor protein torS (EC 2.7.3.-) 137 2e-31
ETR1_LYCES (Q41342) Ethylene receptor 1 (EC 2.7.3.-) (LeETR1) 135 5e-31
TORS_ECO57 (P58356) Sensor protein torS (EC 2.7.3.-) 134 1e-30
LUXQ_VIBCH (Q9KLK7) Autoinducer 2 sensor kinase/phosphatase luxQ... 134 2e-30
ETR1_BRAOL (O49230) Ethylene receptor (EC 2.7.3.-) 132 5e-30
ETR1_PRUPE (Q9M7M1) Ethylene receptor (EC 2.7.3.-) 132 7e-30
ETR1_MALDO (O81122) Ethylene receptor (EC 2.7.3.-) 131 9e-30
ETR1_PASED (Q9ZWL6) Ethylene receptor (EC 2.7.3.-) (PE-ETR1) 130 3e-29
ETR2_LYCES (O49187) Ethylene receptor 2 (EC 2.7.3.-) (LeETR2) 128 1e-28
ETR1_ARATH (P49333) Ethylene receptor (EC 2.7.3.-) 126 3e-28
ETR1_TOBAC (O48929) Ethylene receptor (EC 2.7.3.-) (NT-ETR1) 122 7e-27
ARCB_ECOLI (P22763) Aerobic respiration control sensor protein a... 111 1e-23
ARCB_ECO57 (P58363) Aerobic respiration control sensor protein a... 111 1e-23
>LUXQ_VIBPA (Q87GU5) Autoinducer 2 sensor kinase/phosphatase luxQ
(EC 2.7.3.-) (EC 3.1.3.-)
Length = 858
Score = 152 bits (385), Expect = 4e-36
Identities = 105/342 (30%), Positives = 175/342 (50%), Gaps = 43/342 (12%)
Query: 440 ILTNGVSKEMNLRAELISHLEARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLS 499
I+ G + AE S+L ARR+AE S+ ++ FLA MSHE+RTP+ ++G+ LL
Sbjct: 451 IIVQGQDITTLIEAEKQSNL-ARREAEKSAQARADFLAKMSHEIRTPINGILGVAQ-LLK 508
Query: 500 DDRLTNEQCATVTQIRKCSTALLHLLNNILDISKVESGKLVLEEAEFDFGRELEGLVDMF 559
D EQ + +R LL +LN+ILD SK+E GK +++ F F + L +++
Sbjct: 509 DSVEAEEQKNQIDVLRHSGEHLLAVLNDILDFSKIEQGKFNIQKHPFSFADTMRTLENIY 568
Query: 560 SVQCINHNVEIILDLSDDMPKLVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPN 619
C N VE++++ D + D R+ QI NL++N++KFT SG + L E
Sbjct: 569 RPICENKGVELVIENQLDGNVEIFTDQVRLNQILFNLVSNAVKFTPSGCVRLHAELEQ-- 626
Query: 620 SYGDSENFTLEQKPFGCSRKTKMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDS 679
YG DN +++ E+ DTG GI+ K D
Sbjct: 627 FYG--------------------------------ADNSVLV-VEISDTGIGIESDKLDE 653
Query: 680 VFESFEQADLSTTRLHGGTGLGLCIVRSLVNKMGGEIKIVKKEGPGTLMRLYM------Q 733
+FE F Q + +TTR +GG+GLGL IV++LV+ + G++++ ++G GT + + +
Sbjct: 654 MFEPFVQEEATTTREYGGSGLGLTIVKNLVDMLDGDVQVRSQKGQGTTFVVTLPVKDRER 713
Query: 734 LTAPVDATEQHCQVDFANNNLMVLLALHGNMSRLITSKWLQK 775
+ AP+D++++ + + +L VLL + + I + K
Sbjct: 714 VLAPLDSSQRVKPAELFDESLKVLLVEDNHTNAFILKAFCTK 755
Score = 53.1 bits (126), Expect = 4e-06
Identities = 44/143 (30%), Positives = 60/143 (41%), Gaps = 23/143 (16%)
Query: 1027 EGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEIL 1086
E L++LL ED + K V DG +A+ E L
Sbjct: 732 ESLKVLLVEDNHTNAFILKAFCTKYKMQVDWAKDGLEAM------------------EFL 773
Query: 1087 SFPSYDLILMDCQMPKMDGYEATKEIRKS-EIGTSFHIPIVALTAHAMSCDEAKCLEVGM 1145
SYDLILMD Q+P + G E TKEIR++ ++GT PI A TA +E G
Sbjct: 774 KDHSYDLILMDNQLPHLGGIETTKEIRQNLKLGT----PIYACTADTAQETSDAFMEAGA 829
Query: 1146 DAYLTKPIDFKLMESTILSLTKR 1168
+ L KPI + + +R
Sbjct: 830 NYVLLKPIKENALHEAFVDFKQR 852
>BARA_SHIFL (P59342) Sensor protein barA (EC 2.7.3.-)
Length = 918
Score = 150 bits (379), Expect = 2e-35
Identities = 91/274 (33%), Positives = 146/274 (53%), Gaps = 37/274 (13%)
Query: 461 ARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTA 520
A+++A+ ++ KS+FLANMSHELRTP+ VIG + L + LT Q + I + +
Sbjct: 282 AKKRAQEAARIKSEFLANMSHELRTPLNGVIGFTRLTLKTE-LTPTQRDHLNTIERSANN 340
Query: 521 LLHLLNNILDISKVESGKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPK 580
LL ++N++LD SK+E+GKL+LE F L+ +V + + + +E+ L++ D+P
Sbjct: 341 LLAIINDVLDFSKLEAGKLILESIPFPLRSTLDEVVTLLAHSSHDKGLELTLNIKSDVPD 400
Query: 581 LVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGDSENFTLEQKPFGCSRKT 640
V GD R+ QI NL+ N+IKFT +G+I + +E++
Sbjct: 401 NVIGDPLRLQQIITNLVGNAIKFTENGNI----------------DILVEKRALS----- 439
Query: 641 KMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHGGTGL 700
+ K+ + ++ DTG GI +F++F QAD S +R HGGTGL
Sbjct: 440 ---------------NTKVQIEVQIRDTGIGIPERDQSRLFQAFRQADASISRRHGGTGL 484
Query: 701 GLCIVRSLVNKMGGEIKIVKKEGPGTLMRLYMQL 734
GL I + LVN+MGG+I + G+ ++ L
Sbjct: 485 GLVITQKLVNEMGGDISFHSQPNRGSTFWFHINL 518
Score = 64.3 bits (155), Expect = 2e-09
Identities = 42/135 (31%), Positives = 64/135 (47%), Gaps = 20/135 (14%)
Query: 1029 LRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEILSF 1088
+ ++ +D ++ +LE M V G QAV+ MP
Sbjct: 668 MTVMAVDDNPANLKLIGALLEDMVQHVELCDSGHQAVERAKQMP---------------- 711
Query: 1089 PSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTAHAMSCDEAKCLEVGMDAY 1148
+DLILMD QMP MDG A + I ++ P++A+TAHAM+ + K L GM Y
Sbjct: 712 --FDLILMDIQMPDMDGIRACELIH--QLPHQRQTPVIAVTAHAMAGQKEKLLGAGMSDY 767
Query: 1149 LTKPIDFKLMESTIL 1163
L KPI+ + + + +L
Sbjct: 768 LAKPIEEERLHNLLL 782
>BARA_ECOLI (P26607) Sensor protein barA (EC 2.7.3.-)
Length = 918
Score = 150 bits (379), Expect = 2e-35
Identities = 91/274 (33%), Positives = 146/274 (53%), Gaps = 37/274 (13%)
Query: 461 ARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTA 520
A+++A+ ++ KS+FLANMSHELRTP+ VIG + L + LT Q + I + +
Sbjct: 282 AKKRAQEAARIKSEFLANMSHELRTPLNGVIGFTRLTLKTE-LTPTQRDHLNTIERSANN 340
Query: 521 LLHLLNNILDISKVESGKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPK 580
LL ++N++LD SK+E+GKL+LE F L+ +V + + + +E+ L++ D+P
Sbjct: 341 LLAIINDVLDFSKLEAGKLILESIPFPLRSTLDEVVTLLAHSSHDKGLELTLNIKSDVPD 400
Query: 581 LVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGDSENFTLEQKPFGCSRKT 640
V GD R+ QI NL+ N+IKFT +G+I + +E++
Sbjct: 401 NVIGDPLRLQQIITNLVGNAIKFTENGNI----------------DILVEKRALS----- 439
Query: 641 KMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHGGTGL 700
+ K+ + ++ DTG GI +F++F QAD S +R HGGTGL
Sbjct: 440 ---------------NTKVQIEVQIRDTGIGIPERDQSRLFQAFRQADASISRRHGGTGL 484
Query: 701 GLCIVRSLVNKMGGEIKIVKKEGPGTLMRLYMQL 734
GL I + LVN+MGG+I + G+ ++ L
Sbjct: 485 GLVITQKLVNEMGGDISFHSQPNRGSTFWFHINL 518
Score = 64.3 bits (155), Expect = 2e-09
Identities = 42/135 (31%), Positives = 64/135 (47%), Gaps = 20/135 (14%)
Query: 1029 LRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEILSF 1088
+ ++ +D ++ +LE M V G QAV+ MP
Sbjct: 668 MTVMAVDDNPANLKLIGALLEDMVQHVELCDSGHQAVERAKQMP---------------- 711
Query: 1089 PSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTAHAMSCDEAKCLEVGMDAY 1148
+DLILMD QMP MDG A + I ++ P++A+TAHAM+ + K L GM Y
Sbjct: 712 --FDLILMDIQMPDMDGIRACELIH--QLPHQQQTPVIAVTAHAMAGQKEKLLGAGMSDY 767
Query: 1149 LTKPIDFKLMESTIL 1163
L KPI+ + + + +L
Sbjct: 768 LAKPIEEERLHNLLL 782
>GACS_PSESY (P48027) Sensor protein gacS (EC 2.7.3.-)
Length = 907
Score = 146 bits (368), Expect = 4e-34
Identities = 103/335 (30%), Positives = 163/335 (47%), Gaps = 42/335 (12%)
Query: 450 NLRAELISHLE---ARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNE 506
NL I ++E AR++A +S KS+FLANMSHE+RTP+ ++G LL LT
Sbjct: 250 NLETIEIQNIELDLARKEALEASRIKSEFLANMSHEIRTPLNGILGFTH-LLQKSELTPR 308
Query: 507 QCATVTQIRKCSTALLHLLNNILDISKVESGKLVLEEAEFDFGRELEGLVDMFSVQCINH 566
Q + I K + LL ++N ILD SK+E+GKLVL+ F+ L+ + + +
Sbjct: 309 QFDYLGTIEKSADNLLSIINEILDFSKIEAGKLVLDNIPFNLRDLLQDTLTILAPAAHAK 368
Query: 567 NVEIILDLSDDMPKLVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGDSEN 626
+E++ + D P + GD R+ QI NL++N+IKFT G I+ R E+
Sbjct: 369 QLELVSLVYRDTPLALSGDPLRLRQILTNLVSNAIKFTREGTIVARAMLED--------- 419
Query: 627 FTLEQKPFGCSRKTKMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESFEQ 686
+ E H++ L V DTG G+ ++F++F Q
Sbjct: 420 -----------------ETEEHAQ----------LRISVQDTGIGLSSQDVRALFQAFSQ 452
Query: 687 ADLSTTRLHGGTGLGLCIVRSLVNKMGGEIKIVKKEGPGTLMRLYMQLTAPVDATEQHCQ 746
AD S +R GGTGLGL I + L+ +MGGEI + G G+ ++ L P ++
Sbjct: 453 ADNSLSRQPGGTGLGLVISKRLIEQMGGEIGVDSTPGEGS--EFWISLKLPKAREDKEES 510
Query: 747 VDFANNNLMVLLALHGNMSRLITSKWLQKNGVLTM 781
++ L + H +++R L+ G+ T+
Sbjct: 511 LNIPLGGLRAAVLEHHDLARQALEHQLEDCGLQTI 545
Score = 84.3 bits (207), Expect = 2e-15
Identities = 56/138 (40%), Positives = 76/138 (54%), Gaps = 19/138 (13%)
Query: 1030 RILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEILSFP 1089
R+L +D + +LE MGA VVAV G AV+A+ Q E
Sbjct: 658 RVLCVDDNPANLLLVQTLLEDMGAEVVAVEGGYAAVNAV-------------QQE----- 699
Query: 1090 SYDLILMDCQMPKMDGYEATKEIRKSEIGTS-FHIPIVALTAHAMSCDEAKCLEVGMDAY 1148
++DL+LMD QMP MDG +AT+ IR E + +PIVALTAHAM+ ++ L+ GMD Y
Sbjct: 700 AFDLVLMDVQMPGMDGRQATEAIRAWEAERNQSSLPIVALTAHAMANEKRSLLQSGMDDY 759
Query: 1149 LTKPIDFKLMESTILSLT 1166
LTKPI + + +L T
Sbjct: 760 LTKPISERQLAQVVLKWT 777
>RPFC_XANCP (P49246) Sensory/regulatory protein rpfC (EC 2.7.3.-)
Length = 677
Score = 145 bits (366), Expect = 6e-34
Identities = 123/434 (28%), Positives = 207/434 (47%), Gaps = 80/434 (18%)
Query: 461 ARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTA 520
A R+A ++ KS+FLANMSHE RTP+ + G+ ++L + RL EQ + I+ + +
Sbjct: 129 AVREARHANQAKSRFLANMSHEFRTPLNGLSGMTEVLATT-RLDAEQKECLNTIQASARS 187
Query: 521 LLHLLNNILDISKVESGKLVLEEAEFDFGRELEGLVDM-FSVQCINHNVEIILDLSDDMP 579
LL L+ +LDIS +E+GK+ ++ +F RE+ G V++ Q +E ++DD+P
Sbjct: 188 LLSLVEEVLDISAIEAGKIRIDRRDFSL-REMIGSVNLILQPQARGRRLEYGTQVADDVP 246
Query: 580 KLVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGDSENFTLEQKPFGCSRK 639
L++GD+A + Q+ NL+ N++KFT GH++LR
Sbjct: 247 DLLKGDTAHLRQVLLNLVGNAVKFTEHGHVLLR--------------------------- 279
Query: 640 TKMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHGGTG 699
++ + + ++ + L F+V+DTG G+ +FE+FEQAD+ +R + GTG
Sbjct: 280 --------VTRVSGSAEDAVRLRFDVEDTGIGVPMDMRPRLFEAFEQADVGLSRRYEGTG 331
Query: 700 LGLCIVRSLVNKMGGEIKIVKKEGPGTL--MRLYMQLTAP-------------VDATEQ- 743
LG I + LV MGG I + + G++ L M + P VDA E+
Sbjct: 332 LGTTIAKGLVEAMGGSIGFKENQPSGSVFWFELPMAIGEPLKSSTVRVPTGALVDAPEEL 391
Query: 744 --HCQVDFAN---------NNLMVLLALHGNMSRLITSKWLQKNG--VLTMEASEWNGLT 790
+ F+N ++ +L+A +R++ + L+K G VL + NG
Sbjct: 392 ESSNIIAFSNPFLRHRARVRSMRMLVADDHEANRMVLQRLLEKAGHKVLCV-----NGAE 446
Query: 791 QILKELFEAKTSTHNNDFDTHFPAPEGLNS-KFISIQELPNPTFVIVVDIDLLDLSTDIW 849
Q+L + A+ D H P GL+ K + + + + VV LS D+
Sbjct: 447 QVLDAM--AEEDYDAVIVDLHMPGMNGLDMLKQLRVMQASGMRYTPVV-----VLSADVT 499
Query: 850 KEQLNFLHKYYARA 863
E + + ARA
Sbjct: 500 PEAIRACEQAGARA 513
Score = 56.2 bits (134), Expect = 5e-07
Identities = 36/153 (23%), Positives = 69/153 (44%), Gaps = 18/153 (11%)
Query: 1013 VSGSSKAMNGKKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMP 1072
++ S+ + + + +R+L+A+D + V +LEK G V+ V +Q +DA+
Sbjct: 397 IAFSNPFLRHRARVRSMRMLVADDHEANRMVLQRLLEKAGHKVLCVNGAEQVLDAM---- 452
Query: 1073 GVERNTITSQTEILSFPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTAHA 1132
+ YD +++D MP M+G + K++R + + P+V L+A
Sbjct: 453 --------------AEEDYDAVIVDLHMPGMNGLDMLKQLRVMQASGMRYTPVVVLSADV 498
Query: 1133 MSCDEAKCLEVGMDAYLTKPIDFKLMESTILSL 1165
C + G A+L KP+ + T+ L
Sbjct: 499 TPEAIRACEQAGARAFLAKPVVAAKLLDTLADL 531
>LUXQ_VIBHA (P54302) Autoinducer 2 sensor kinase/phosphatase luxQ
(EC 2.7.3.-) (EC 3.1.3.-)
Length = 859
Score = 142 bits (357), Expect = 7e-33
Identities = 103/342 (30%), Positives = 171/342 (49%), Gaps = 43/342 (12%)
Query: 440 ILTNGVSKEMNLRAELISHLEARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLS 499
I+ G + AE S++ ARR+AE S+ ++ FLA MSHE+RTP+ ++G+ LL
Sbjct: 452 IIVQGQDITTLIEAEKQSNI-ARREAEKSAQARADFLAKMSHEIRTPINGILGVAQ-LLK 509
Query: 500 DDRLTNEQCATVTQIRKCSTALLHLLNNILDISKVESGKLVLEEAEFDFGRELEGLVDMF 559
D T EQ + + LL +LN+ILD SK+E GK +++ F F + L +++
Sbjct: 510 DSVDTQEQKNQIDVLCHSGEHLLAVLNDILDFSKIEQGKFNIQKHPFSFTDTMRTLENIY 569
Query: 560 SVQCINHNVEIILDLSDDMPKLVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPN 619
C N VE++++ D + D R+ QI NL++N++KFT G I L E
Sbjct: 570 RPICTNKGVELVIENELDPNVEIFTDQVRLNQILFNLVSNAVKFTPIGSIRLH--AELEQ 627
Query: 620 SYGDSENFTLEQKPFGCSRKTKMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDS 679
YG +EN +L E+ DTG GI+ K D
Sbjct: 628 FYG-AEN--------------------------------SVLVVELTDTGIGIESDKLDQ 654
Query: 680 VFESFEQADLSTTRLHGGTGLGLCIVRSLVNKMGGEIKIVKKEGPGTLMRLYM------Q 733
+FE F Q + +TTR +GG+GLGL IV++LV+ + G++++ +G GT + + +
Sbjct: 655 MFEPFVQEESTTTREYGGSGLGLTIVKNLVDMLEGDVQVRSSKGGGTTFVITLPVKDRER 714
Query: 734 LTAPVDATEQHCQVDFANNNLMVLLALHGNMSRLITSKWLQK 775
+ P++ +++ + +L VLL + + I + +K
Sbjct: 715 VLRPLEVSQRIKPEALFDESLKVLLVEDNHTNAFILQAFCKK 756
Score = 48.9 bits (115), Expect = 8e-05
Identities = 41/143 (28%), Positives = 59/143 (40%), Gaps = 23/143 (16%)
Query: 1027 EGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEIL 1086
E L++LL ED + +K V DG A+ E+L
Sbjct: 733 ESLKVLLVEDNHTNAFILQAFCKKYKMQVDWAKDGLDAM------------------ELL 774
Query: 1087 SFPSYDLILMDCQMPKMDGYEATKEIRKS-EIGTSFHIPIVALTAHAMSCDEAKCLEVGM 1145
S +YDLILMD Q+P + G E T EIR++ +GT PI A TA + G
Sbjct: 775 SDTTYDLILMDNQLPHLGGIETTHEIRQNLRLGT----PIYACTADTAKETSDAFMAAGA 830
Query: 1146 DAYLTKPIDFKLMESTILSLTKR 1168
+ + KPI + + +R
Sbjct: 831 NYVMLKPIKENALHEAFVDFKQR 853
>ETR1_PELHO (Q9XH58) Ethylene receptor 1 (EC 2.7.3.-) (PhETR1)
Length = 740
Score = 138 bits (348), Expect = 8e-32
Identities = 108/343 (31%), Positives = 164/343 (47%), Gaps = 46/343 (13%)
Query: 461 ARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTA 520
ARR+AE + ++ FLA M+HE+RTPM A+I L +L D LT+EQ V I K S
Sbjct: 333 ARREAETAIRARNDFLAVMNHEMRTPMHAIIALSSLLQETD-LTSEQRLMVETILKSSNL 391
Query: 521 LLHLLNNILDISKVESGKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPK 580
L L+N++LD+S++E G L L+ A F+ + ++ + I L++S D+P+
Sbjct: 392 LATLINDVLDLSRLEDGSLQLDIATFNLHAVFRQVFNLIKPIASVKKLFITLNVSPDLPE 451
Query: 581 LVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGD---SENFTLEQKPFGCS 637
V GD R+VQI N++ N++KF+ G I + + S D + F +
Sbjct: 452 YVIGDEKRLVQIMLNVVGNAVKFSKEGIISVTAFVAKSESVRDPRAPDFFPVSS------ 505
Query: 638 RKTKMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHGG 697
DN+ + +V D+G GI+P +F F Q+ T+ GG
Sbjct: 506 ------------------DNQFYMRVQVKDSGSGINPQDMPKLFTKFAQSQPVATKNSGG 547
Query: 698 TGLGLCIVRSLVNKMGGEIKIVKKEGP--GTLMRLYMQLTAPVDATE-----------QH 744
+GLGL I + VN M G I I EGP G + ++L P + E +
Sbjct: 548 SGLGLAISKRFVNLMDGHIWI-DSEGPSKGCTVTFVVKLGIPEGSNEPKLPLMPKVSANN 606
Query: 745 CQVDFANNNLMVLLALHGNMSRLITSKWLQKNG--VLTMEASE 785
Q DF L VLL +SR++T L G V ++ +SE
Sbjct: 607 SQTDFP--GLKVLLMDENGISRMVTKGLLMHLGCDVTSVSSSE 647
Score = 55.1 bits (131), Expect = 1e-06
Identities = 34/153 (22%), Positives = 67/153 (43%), Gaps = 19/153 (12%)
Query: 1019 AMNGKKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNT 1078
A N + GL++LL ++ + + V +L +G V +V ++ + ++
Sbjct: 604 ANNSQTDFPGLKVLLMDENGISRMVTKGLLMHLGCDVTSVSSSEECLRMVS--------- 654
Query: 1079 ITSQTEILSFPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTAHAMSCDEA 1138
+ ++ MD ++P +DG+E I + + IVALT++A +
Sbjct: 655 ----------QDHKVVFMDVRVPGLDGHELAVRIHEKFMKRHERPLIVALTSNADKVTKE 704
Query: 1139 KCLEVGMDAYLTKPIDFKLMESTILSLTKRETL 1171
CL VGM+ + KP+ M + + L + L
Sbjct: 705 NCLRVGMEGVILKPVSVDKMRNVLSKLLEHRIL 737
>TORS_ECOLI (P39453) Sensor protein torS (EC 2.7.3.-)
Length = 914
Score = 137 bits (344), Expect = 2e-31
Identities = 105/338 (31%), Positives = 164/338 (48%), Gaps = 56/338 (16%)
Query: 446 SKEMNLRAELISHLEARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTN 505
++ L+ +I H +AR +AE +S KS FLA MSHE+RTP+ ++G LL+D+ N
Sbjct: 418 ARTAELQELVIEHRQARAEAEKASQAKSAFLAAMSHEIRTPLYGILGTAQ-LLADNPALN 476
Query: 506 EQCATVTQIRKCSTALLHLLNNILDISKVESG--KLVLEEAEFDFGRELEGLVDMFSVQC 563
Q + I +LL +LN+ILD S +E+G + + + F+ LE + + S +
Sbjct: 477 AQRDDLRAITDSGESLLTILNDILDYSAIEAGGKNVSVSDEPFEPRPLLESTLQLMSGRV 536
Query: 564 INHNVEIILDLSDDMPKLVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGD 623
+ + ++DDMP + GD R+ Q+ NL++N+++FT G+IILR S D
Sbjct: 537 KGRPIRLATAIADDMPCALMGDPRRIRQVITNLLSNALRFTDEGYIILR-------SRTD 589
Query: 624 SENFTLEQKPFGCSRKTKMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFES 683
E + + EV+D+GCGIDP+K +F+
Sbjct: 590 GEQWLV----------------------------------EVEDSGCGIDPAKLAEIFQP 615
Query: 684 FEQADLSTTRLHGGTGLGLCIVRSLVNKMGGEIKIVKKEGPGTLMRLYMQL---TAPVDA 740
F Q + GGTGLGL I L MGGE+ G+ L + L TAPV
Sbjct: 616 FVQ----VSGKRGGTGLGLTISSRLAQAMGGELSATSTPEVGSCFCLRLPLRVATAPVPK 671
Query: 741 T-EQHCQVDFANNNLMVLLALHGNMSRLITSKWLQKNG 777
T Q ++D L +LL +++ IT + L+ +G
Sbjct: 672 TVNQAVRLD----GLRLLLIEDNPLTQRITIEMLKTSG 705
Score = 48.5 bits (114), Expect = 1e-04
Identities = 34/112 (30%), Positives = 53/112 (46%), Gaps = 24/112 (21%)
Query: 1018 KAMNGKKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDAL-NGMPGVER 1076
K +N L+GLR+LL ED + QR+ ML+ GA +VAVG+ QA++ L N P
Sbjct: 671 KTVNQAVRLDGLRLLLIEDNPLTQRITIEMLKTSGAQIVAVGNAAQALETLQNSEP---- 726
Query: 1077 NTITSQTEILSFPSYDLILMDCQMPKMDGYEATKEIRKSE-----IGTSFHI 1123
+ L+D +P +DG +++ + IG S H+
Sbjct: 727 --------------FAAALVDFDLPDIDGITLARQLAQQYPSLVLIGFSAHV 764
>ETR1_LYCES (Q41342) Ethylene receptor 1 (EC 2.7.3.-) (LeETR1)
Length = 754
Score = 135 bits (341), Expect = 5e-31
Identities = 108/333 (32%), Positives = 159/333 (47%), Gaps = 44/333 (13%)
Query: 461 ARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTA 520
ARR+AE + ++ FLA M+HE+RTPM A+I L +L D LT EQ V I K S
Sbjct: 347 ARREAEMAVRARNDFLAVMNHEMRTPMHAIIALSSLLQETD-LTPEQRLMVETILKSSNL 405
Query: 521 LLHLLNNILDISKVESGKLVLEEAEFDFG---RELEGLVD-MFSVQCINHNVEIILDLSD 576
L L+N++LD+S++E G L L+ F+ RE+ L+ + SV+ + + L LS
Sbjct: 406 LATLINDVLDLSRLEDGSLQLDIGTFNLHALFREVHSLIKPIASVK----KLFVTLSLSS 461
Query: 577 DMPKLVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGDSENFTLEQKPFGC 636
D+P+ V GD R++QI N++ N++KF+ G++ + + +S D P
Sbjct: 462 DLPEYVIGDEKRLMQILLNVVGNAVKFSKEGNVSISAFVAKSDSLRDPRAPEFFAVP--- 518
Query: 637 SRKTKMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHG 696
ENH L ++ DTG GI P ++F F Q+ T G
Sbjct: 519 --------SENH----------FYLRVQIKDTGIGITPQDIPNLFSKFTQSQALATTNSG 560
Query: 697 GTGLGLCIVRSLVNKMGGEIKIVKKE-GPGTLMRLYMQLTAPVDATE-----------QH 744
GTGLGL I + VN M G I I + G G+ ++L P A E H
Sbjct: 561 GTGLGLAICKRFVNLMEGHIWIESEGLGKGSTAIFIIKLGIPGRANESKLPFVTKLPANH 620
Query: 745 CQVDFANNNLMVLLALHGNMSRLITSKWLQKNG 777
Q+ F L VL+ +SR++T L G
Sbjct: 621 TQMSF--QGLKVLVMDENGVSRMVTKGLLTHLG 651
Score = 48.1 bits (113), Expect = 1e-04
Identities = 37/148 (25%), Positives = 64/148 (43%), Gaps = 21/148 (14%)
Query: 1019 AMNGKKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNT 1078
A + + S +GL++L+ ++ V + V +L +G V VG + +
Sbjct: 618 ANHTQMSFQGLKVLVMDENGVSRMVTKGLLTHLGCDVTTVGSRDECL-----------RV 666
Query: 1079 ITSQTEILSFPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIP-IVALTAHAMSCDE 1137
+T + ++ ++MD M +D YE I + G P IVALT + +
Sbjct: 667 VTHEHKV--------VIMDVSMQGIDCYEVAVVIHE-RFGKRHGRPLIVALTGNTDRVTK 717
Query: 1138 AKCLEVGMDAYLTKPIDFKLMESTILSL 1165
C+ VGMD + KP+ M S + L
Sbjct: 718 ENCMRVGMDGVILKPVSVYKMRSVLSEL 745
>TORS_ECO57 (P58356) Sensor protein torS (EC 2.7.3.-)
Length = 914
Score = 134 bits (337), Expect = 1e-30
Identities = 102/338 (30%), Positives = 162/338 (47%), Gaps = 56/338 (16%)
Query: 446 SKEMNLRAELISHLEARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTN 505
++ L+ +I H +AR +AE +S KS FLA MSHE+RTP+ ++G LL+D+ N
Sbjct: 418 ARTAELQELVIEHRQARAEAEKASQAKSAFLAAMSHEIRTPLYGILGTAQ-LLADNPALN 476
Query: 506 EQCATVTQIRKCSTALLHLLNNILDISKVESG--KLVLEEAEFDFGRELEGLVDMFSVQC 563
Q + I +LL +LN+ILD S +E+G + + + F+ LE + + S +
Sbjct: 477 AQRDDLRAITDSGESLLTILNDILDYSAIEAGGKNVSVSDEPFEPRPLLESTLQLMSGRV 536
Query: 564 INHNVEIILDLSDDMPKLVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGD 623
+ + ++DD+P + GD R+ Q+ NL++N+++FT G I+LR S D
Sbjct: 537 KGRPIRLATAIADDVPTALMGDPRRIRQVITNLLSNALRFTDEGQIVLR-------SRTD 589
Query: 624 SENFTLEQKPFGCSRKTKMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFES 683
E + + EV+D+GCGIDP+K +F+
Sbjct: 590 GEQWLV----------------------------------EVEDSGCGIDPAKLAEIFQP 615
Query: 684 FEQADLSTTRLHGGTGLGLCIVRSLVNKMGGEIKIVKKEGPGTLMRLYMQL---TAPVDA 740
F + + GGTGLGL I L MGGE+ G+ L + L TAPV
Sbjct: 616 F----IQVSGKRGGTGLGLTISSRLAQAMGGELSATSTPEVGSCFCLRLPLRVATAPVPK 671
Query: 741 T-EQHCQVDFANNNLMVLLALHGNMSRLITSKWLQKNG 777
T Q ++D L +LL +++ IT + L +G
Sbjct: 672 TVNQAVRLD----GLRLLLIEDNPLTQRITVEMLNTSG 705
Score = 47.0 bits (110), Expect = 3e-04
Identities = 33/112 (29%), Positives = 52/112 (45%), Gaps = 24/112 (21%)
Query: 1018 KAMNGKKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDAL-NGMPGVER 1076
K +N L+GLR+LL ED + QR+ ML GA VVA+G+ QA++ L N P
Sbjct: 671 KTVNQAVRLDGLRLLLIEDNPLTQRITVEMLNTSGAQVVAIGNAAQALETLQNSEP---- 726
Query: 1077 NTITSQTEILSFPSYDLILMDCQMPKMDGYEATKEIRKSE-----IGTSFHI 1123
+ L+D +P ++G +++ + IG S H+
Sbjct: 727 --------------FAAALVDFDLPDVNGITLARQLARQYPSLVLIGFSAHV 764
>LUXQ_VIBCH (Q9KLK7) Autoinducer 2 sensor kinase/phosphatase luxQ
(EC 2.7.3.-) (EC 3.1.3.-)
Length = 857
Score = 134 bits (336), Expect = 2e-30
Identities = 112/359 (31%), Positives = 173/359 (47%), Gaps = 47/359 (13%)
Query: 427 SLCIIVIGCVCILILTNGVSKEMNLRAELISHLEARRKAEASSNYKSQFLANMSHELRTP 486
+L I++ I I+T G AE S ARR+AE S+ +++FLA MSHELRTP
Sbjct: 436 NLSPIMVEGQIISIITQGQDITTIAEAEKQSQA-ARREAEESARVRAEFLAKMSHELRTP 494
Query: 487 MAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTALLHLLNNILDISKVESGKLVLEEAEF 546
+ V+G+ LL L +EQ V + LL +LN+ILD S++E GK +++ EF
Sbjct: 495 LNGVLGVSQ-LLKRTPLNDEQREHVAVLCSSGEHLLAVLNDILDFSRLEQGKFRIQKNEF 553
Query: 547 DFGRELEGLVD-MFSVQCINHNVEIILDLSDDMPKLVRGDSARVVQIFANLINNSIKFTL 605
+EL +D ++ C +E++++ + +VR D R+ QI NL+NN+IKFT
Sbjct: 554 RL-KELVCAIDRIYRPLCNEKGLELVVNSNITTAAIVRSDQIRINQILFNLLNNAIKFTH 612
Query: 606 SGHIILRGWCENPNSYGDSENFTLEQKPFGCSRKTKMKQHENHSKKASNIDNKMILWFEV 665
G I + +++ E D L +V
Sbjct: 613 QGSI-----------------------------RVELQLIEG--------DPLAQLVIQV 635
Query: 666 DDTGCGIDPSKWDSVFESFEQADLSTTRLHGGTGLGLCIVRSLVNKMGGEIKIVKKEGPG 725
DTG GI +FE F QA+ +TTR +GG+GLGL IV SLV + G++ + + G G
Sbjct: 636 VDTGIGIREQDLTVIFEPFMQAESTTTREYGGSGLGLTIVHSLVEMLSGQLHVSSEYGIG 695
Query: 726 TL--MRLYMQLTAPVDATEQHCQV----DFANNNLMVLLALHGNMSRLITSKWLQKNGV 778
T ++L ++L DA +Q + L VLL + + I + +K G+
Sbjct: 696 TRFEIQLPIELVEKPDAPQQLLPAPDPQPLFDKTLRVLLVEDNHTNAFIAQAFCRKYGL 754
Score = 54.7 bits (130), Expect = 1e-06
Identities = 41/127 (32%), Positives = 54/127 (42%), Gaps = 25/127 (19%)
Query: 1029 LRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEILSF 1088
LR+LL ED +A K G V V DG QA+ E L
Sbjct: 730 LRVLLVEDNHTNAFIAQAFCRKYGLDVSWVTDGLQAI------------------EELKI 771
Query: 1089 PSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIV--ALTAHAMSCDEAKCLEVGMD 1146
YDL+LMD Q+P +DG E T+ I+K H+P+V A TA + G +
Sbjct: 772 HDYDLVLMDNQLPYLDGVETTRTIKK-----VLHLPVVVYACTADGLEETRQAFFHAGAE 826
Query: 1147 AYLTKPI 1153
L KP+
Sbjct: 827 YVLVKPL 833
>ETR1_BRAOL (O49230) Ethylene receptor (EC 2.7.3.-)
Length = 735
Score = 132 bits (332), Expect = 5e-30
Identities = 107/339 (31%), Positives = 162/339 (47%), Gaps = 44/339 (12%)
Query: 461 ARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTA 520
ARR+AE + ++ FLA M+HE+RTPM A+I L LL + LT EQ V + K S+
Sbjct: 333 ARREAETAIRARNDFLAVMNHEMRTPMHAIIALSS-LLQETELTPEQRLMVETVLKSSSL 391
Query: 521 LLHLLNNILDISKVESGKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPK 580
L L+N++LD+S++E G L LE F+ ++++ + + I L+L+ D+P+
Sbjct: 392 LATLMNDVLDLSRLEDGSLQLELGTFNLHTLFREVLNLIKPIAVVKKLPITLNLAPDLPE 451
Query: 581 LVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGDSENFTLEQKPFGCSRKT 640
V GD R++QI N++ N++KF+ G I + ++ + F + P G
Sbjct: 452 FVVGDEKRLMQIILNIVGNAVKFSKQGSISVTALVTKSDNRAPPDFFVV---PTG----- 503
Query: 641 KMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHGGTGL 700
+ L +V D G GI+P +F F Q TR GG+GL
Sbjct: 504 ----------------SHFYLRVKVKDLGAGINPQDIPKLFTKFAQTQSLATRSSGGSGL 547
Query: 701 GLCIVRSLVNKMGGEIKIVKKEGPG-----------TLMRLYMQLTAP-VDATEQHCQVD 748
GL I + VN M G I I + EG G + Q P V A QH V+
Sbjct: 548 GLAISKRFVNLMEGNIWI-ESEGVGKGCTAIFDVKLAISNESKQSGIPKVPANPQH--VN 604
Query: 749 FANNNLMVLLALHGNMSRLITSKWLQKNG--VLTMEASE 785
FA L VL+ +SR++T L G V T+ ++E
Sbjct: 605 FA--GLKVLVMDENGVSRMVTKGLLVHLGCEVTTVSSNE 641
Score = 41.2 bits (95), Expect = 0.017
Identities = 24/92 (26%), Positives = 42/92 (45%), Gaps = 2/92 (2%)
Query: 1073 GVERNTITSQTEILSFPSYD--LILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTA 1130
G E T++S E L S++ ++ MD P ++ Y+ I + +VALT
Sbjct: 631 GCEVTTVSSNEECLRVVSHEHRVVFMDVCTPGVENYQIALRIHEKFTKRHQRPLLVALTG 690
Query: 1131 HAMSCDEAKCLEVGMDAYLTKPIDFKLMESTI 1162
+ + KC+ G+D L KP+ M + +
Sbjct: 691 NTDKSTKEKCMSFGLDGVLLKPVSLDNMRNVL 722
>ETR1_PRUPE (Q9M7M1) Ethylene receptor (EC 2.7.3.-)
Length = 738
Score = 132 bits (331), Expect = 7e-30
Identities = 94/281 (33%), Positives = 136/281 (47%), Gaps = 34/281 (12%)
Query: 448 EMNLRAELISHLEARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQ 507
E N+ +L ARR+AE + ++ FLA M+HE+RTPM A+I L LL + LT EQ
Sbjct: 325 EQNIALDL-----ARREAETAIRARNDFLAVMNHEMRTPMHAIIALSS-LLQETELTPEQ 378
Query: 508 CATVTQIRKCSTALLHLLNNILDISKVESGKLVLEEAEFDFGRELEGLVDMFSVQCINHN 567
V I K S L L+N++LD+S++E G L LE A F+ + ++
Sbjct: 379 RLMVETILKSSHLLATLINDVLDLSRLEDGSLQLEIATFNLHSVFREVHNLIKPVASVKK 438
Query: 568 VEIILDLSDDMPKLVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGD---S 624
+ + L+L+ D+P GD R++QI N++ N++KF+ G I + + S D
Sbjct: 439 LSVSLNLAADLPVQAVGDEKRLMQIVLNVVGNAVKFSKEGSISITAFVAKSESLRDFRAP 498
Query: 625 ENFTLEQKPFGCSRKTKMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESF 684
E F + DN L +V D+G GI+P +F F
Sbjct: 499 EFFPAQS------------------------DNHFYLRVQVKDSGSGINPQDIPKLFTKF 534
Query: 685 EQADLSTTRLHGGTGLGLCIVRSLVNKMGGEIKIVKKEGPG 725
Q TR GG+GLGL I + VN M G I I + EGPG
Sbjct: 535 AQTQSLATRNSGGSGLGLAICKRFVNLMEGHIWI-ESEGPG 574
Score = 50.8 bits (120), Expect = 2e-05
Identities = 34/122 (27%), Positives = 52/122 (41%), Gaps = 2/122 (1%)
Query: 1052 GATVVAVGDGQQAVDALNGMPGVERNTITSQTEILSFPSYD--LILMDCQMPKMDGYEAT 1109
G V+ + D L G + T++S E L S + ++ MD MP +DGYE
Sbjct: 613 GLKVLVMDDNGSVTKGLLVHLGCDVTTVSSIDEFLHVISQEHKVVFMDVCMPGIDGYELA 672
Query: 1110 KEIRKSEIGTSFHIPIVALTAHAMSCDEAKCLEVGMDAYLTKPIDFKLMESTILSLTKRE 1169
I + +VALT + + C+ VGMD + KP+ M S + L +
Sbjct: 673 VRIHEKFTKRHERPVLVALTGNIDKMTKENCMRVGMDGVILKPVSVDKMRSVLSELLEHR 732
Query: 1170 TL 1171
L
Sbjct: 733 VL 734
>ETR1_MALDO (O81122) Ethylene receptor (EC 2.7.3.-)
Length = 741
Score = 131 bits (330), Expect = 9e-30
Identities = 116/403 (28%), Positives = 173/403 (42%), Gaps = 59/403 (14%)
Query: 448 EMNLRAELISHLEARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQ 507
E N+ +L ARR+AE + ++ FLA M+HE+RTPM A+I L LL + LT EQ
Sbjct: 325 EQNIALDL-----ARREAETAIRARNDFLAVMNHEMRTPMHAIIALSS-LLQETELTAEQ 378
Query: 508 CATVTQIRKCSTALLHLLNNILDISKVESGKLVLEEAEFDFGRELEGLVDMFSVQCINHN 567
V I + S L L+N++LD+S++E G L LE A F+ + +M
Sbjct: 379 RLMVETILRSSNLLATLINDVLDLSRLEDGSLQLEIATFNLHSVFREVHNMIKPVASIKR 438
Query: 568 VEIILDLSDDMPKLVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGD--SE 625
+ + L+++ D+P GD R++Q N++ N++KF+ G I + + S D +
Sbjct: 439 LSVTLNIAADLPMYAIGDEKRLMQTILNVVGNAVKFSKEGSISITAFVAKSESLRDFRAP 498
Query: 626 NFTLEQKPFGCSRKTKMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESFE 685
+F Q DN L +V D+G GI+P +F F
Sbjct: 499 DFFPVQS-----------------------DNHFYLRVQVKDSGSGINPQDIPKLFTKFA 535
Query: 686 QADLSTTRLHGGTGLGLCIVRSLVNKMGGEIKIVKKE-GPGTLMRLYMQLTAPVDATE-- 742
Q TR GG+GLGL I + VN M G I I + G G ++L P + E
Sbjct: 536 QTQALATRNSGGSGLGLAICKRFVNLMEGHIWIESEGLGKGCTATFIVKLGFPERSNESK 595
Query: 743 ---------QHCQVDFANNNLMVLLALHGNMSRLITSKWLQKNGVLTMEASEWNGLTQIL 793
H Q +F L VL+ +SR +T L G S ++
Sbjct: 596 LPFAPKLQANHVQTNFP--GLKVLVMDDNGVSRSVTKGLLAHLGCDVTAVS-------LI 646
Query: 794 KELFEAKTSTHNNDF-DTHFPAPEG------LNSKFISIQELP 829
EL + H F D P +G ++ KF E P
Sbjct: 647 DELLHVISQEHKVVFMDVSMPGIDGYELAVRIHEKFTKRHERP 689
Score = 52.0 bits (123), Expect = 9e-06
Identities = 37/154 (24%), Positives = 64/154 (41%), Gaps = 19/154 (12%)
Query: 1018 KAMNGKKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERN 1077
+A + + + GL++L+ +D V + V +L +G V AV +
Sbjct: 603 QANHVQTNFPGLKVLVMDDNGVSRSVTKGLLAHLGCDVTAV------------------S 644
Query: 1078 TITSQTEILSFPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTAHAMSCDE 1137
I ++S + ++ MD MP +DGYE I + +VALT +
Sbjct: 645 LIDELLHVIS-QEHKVVFMDVSMPGIDGYELAVRIHEKFTKRHERPVLVALTGSIDKITK 703
Query: 1138 AKCLEVGMDAYLTKPIDFKLMESTILSLTKRETL 1171
C+ VG+D + KP+ M S + L + L
Sbjct: 704 ENCMRVGVDGVILKPVSVDKMRSVLSELLEHRVL 737
>ETR1_PASED (Q9ZWL6) Ethylene receptor (EC 2.7.3.-) (PE-ETR1)
Length = 738
Score = 130 bits (326), Expect = 3e-29
Identities = 106/339 (31%), Positives = 155/339 (45%), Gaps = 38/339 (11%)
Query: 461 ARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTA 520
ARR+AE + ++ FLA M+HE+RTPM AVI L LL + LT EQ V I K S
Sbjct: 333 ARREAETAIRARNDFLAVMNHEMRTPMHAVIALSS-LLQETELTPEQRLMVETILKSSNL 391
Query: 521 LLHLLNNILDISKVESGKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPK 580
L L+N++LD+SK+E G L L+ F+ ++++ + ++L+L+ D+P+
Sbjct: 392 LATLINDVLDLSKLEDGSLQLDSGTFNLHAVFREVLNLIKPIASVKKLLLLLNLAPDLPE 451
Query: 581 LVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGDSENFTLEQKPFGCSRKT 640
GD R++QI N++ N++KF+ G I + S D+ P
Sbjct: 452 YAVGDEKRLIQIILNIVGNAMKFSKEGSISITAIVAKLESLRDARVPDFFPTP------- 504
Query: 641 KMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHGGTGL 700
ENH L +V D+G GI+P +F F Q + R G+GL
Sbjct: 505 ----SENH----------FYLRVQVKDSGVGINPQDIPKLFIKFAQTQTTGARNSSGSGL 550
Query: 701 GLCIVRSLVNKMGGEIKIVKKE-GPGTLMRLYMQLTAPVDATE-----------QHCQVD 748
GL I R VN M G I + + G G ++L P E H Q
Sbjct: 551 GLAICRRFVNLMDGHIWLESEGLGKGCTAIFIVKLGIPERLNESKPPFMSKVAVDHGQTT 610
Query: 749 FANNNLMVLLALHGNMSRLITSKWLQKNG--VLTMEASE 785
F L VLL +SR++T L G V T+ +SE
Sbjct: 611 FP--GLKVLLMDDNGVSRMVTKGLLLHLGCDVTTVGSSE 647
Score = 54.7 bits (130), Expect = 1e-06
Identities = 38/151 (25%), Positives = 66/151 (43%), Gaps = 21/151 (13%)
Query: 1021 NGKKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTIT 1080
+G+ + GL++LL +D V + V +L +G V VG ++ + +
Sbjct: 606 HGQTTFPGLKVLLMDDNGVSRMVTKGLLLHLGCDVTTVGSSEECI------------RVA 653
Query: 1081 SQTEILSFPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTAHAMSCDEAKC 1140
SQ + ++ MD MP +G+EA + + +VALTA + C
Sbjct: 654 SQ-------DHRVVFMDVGMP--EGFEAAVRLHEKFTKRHERPLVVALTASTDRMTKENC 704
Query: 1141 LEVGMDAYLTKPIDFKLMESTILSLTKRETL 1171
+ VGMD + KP+ M S + L + + L
Sbjct: 705 MRVGMDGAILKPVSVDKMRSVLSDLLEHKVL 735
>ETR2_LYCES (O49187) Ethylene receptor 2 (EC 2.7.3.-) (LeETR2)
Length = 736
Score = 128 bits (321), Expect = 1e-28
Identities = 101/327 (30%), Positives = 148/327 (44%), Gaps = 40/327 (12%)
Query: 461 ARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTA 520
ARR+AE + ++ FL M+HE+RTPM AV+ L LL + L EQ V I K S
Sbjct: 331 ARREAETAVRARNDFLGVMNHEMRTPMHAVVALSS-LLQESELIPEQRLMVETILKSSNL 389
Query: 521 LLHLLNNILDISKVESGKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPK 580
L L+N++LD+S++E G L L+ F+ ++++ + + L LS D P+
Sbjct: 390 LATLINDVLDLSRLEDGSLQLDVGTFNLHALFREVLNLIKPVAAVKKLFVTLSLSSDFPE 449
Query: 581 LVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGDSENFTLEQKPFGCSRKT 640
+ GD R++QI N++ N++KF+ G + + S D + +
Sbjct: 450 VAIGDEKRLMQILLNVVGNAVKFSKEGSVSVSAVNAKSESLIDPR-----------APEF 498
Query: 641 KMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHGGTGL 700
Q ENH L +V DTG GI+P + +F F Q T+ GTGL
Sbjct: 499 FPVQSENH----------FYLRVQVKDTGSGINPQDFPKLFCKFAQNQEPATKNSAGTGL 548
Query: 701 GLCIVRSLVNKMGGE--------------IKIVKKEGPGTLMRLYMQLTAPVDATEQHCQ 746
GL I + VN M G I IVK PG L + TA + A H Q
Sbjct: 549 GLAICKRFVNLMEGHIWIESEGVGKGSTAIFIVKLGIPGRLNESKLPFTAGLPA--NHMQ 606
Query: 747 VDFANNNLMVLLALHGNMSRLITSKWL 773
+ F L VL+ SR++T L
Sbjct: 607 MTF--QGLKVLVMDDNGFSRMVTKSLL 631
Score = 52.0 bits (123), Expect = 9e-06
Identities = 34/148 (22%), Positives = 63/148 (41%), Gaps = 21/148 (14%)
Query: 1025 SLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTE 1084
+ +GL++L+ +D + V +L +G V +G G + + L
Sbjct: 608 TFQGLKVLVMDDNGFSRMVTKSLLVHLGCDVTTIGSGDECLRILTR-------------- 653
Query: 1085 ILSFPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIP-IVALTAHAMSCDEAKCLEV 1143
+ +++MD + M+ Y+ + + + G P IVALT + + CL V
Sbjct: 654 -----EHKVLIMDASITGMNCYDVAVSVHE-KFGKRLERPLIVALTGNTDQVTKENCLRV 707
Query: 1144 GMDAYLTKPIDFKLMESTILSLTKRETL 1171
GMD + KP+ M S + L + T+
Sbjct: 708 GMDGVILKPVSIDKMRSVLSGLLEHGTV 735
>ETR1_ARATH (P49333) Ethylene receptor (EC 2.7.3.-)
Length = 738
Score = 126 bits (317), Expect = 3e-28
Identities = 85/258 (32%), Positives = 127/258 (48%), Gaps = 26/258 (10%)
Query: 461 ARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTA 520
ARR+AE + ++ FLA M+HE+RTPM A+I L LL + LT EQ V I K S
Sbjct: 333 ARREAETAIRARNDFLAVMNHEMRTPMHAIIALSS-LLQETELTPEQRLMVETILKSSNL 391
Query: 521 LLHLLNNILDISKVESGKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPK 580
L L+N++LD+S++E G L LE F+ ++++ + + I L+L+ D+P+
Sbjct: 392 LATLMNDVLDLSRLEDGSLQLELGTFNLHTLFREVLNLIKPIAVVKKLPITLNLAPDLPE 451
Query: 581 LVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGDSENFTLEQKPFGCSRKT 640
V GD R++QI N++ N++KF+ G I + + D+ P G
Sbjct: 452 FVVGDEKRLMQIILNIVGNAVKFSKQGSISVTALV----TKSDTRAADFFVVPTG----- 502
Query: 641 KMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHGGTGL 700
+ L +V D+G GI+P +F F Q TR GG+GL
Sbjct: 503 ----------------SHFYLRVKVKDSGAGINPQDIPKIFTKFAQTQSLATRSSGGSGL 546
Query: 701 GLCIVRSLVNKMGGEIKI 718
GL I + VN M G I I
Sbjct: 547 GLAISKRFVNLMEGNIWI 564
Score = 43.1 bits (100), Expect = 0.004
Identities = 38/172 (22%), Positives = 70/172 (40%), Gaps = 26/172 (15%)
Query: 1005 SKGGETSRVSGSSK--AMNGKKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQ 1062
S+ S+ SG K A+ + GL++L+ ++ V + V +L +G V
Sbjct: 584 SERSNESKQSGIPKVPAIPRHSNFTGLKVLVMDENGVSRMVTKGLLVHLGCEVT------ 637
Query: 1063 QAVDALNGMPGVERNTITSQTEILSFPSYD--LILMDCQMPKMDGYEATKEIRKSEIGTS 1120
T++S E L S++ ++ MD MP ++ Y+ I +
Sbjct: 638 ---------------TVSSNEECLRVVSHEHKVVFMDVCMPGVENYQIALRIHEKFTKQR 682
Query: 1121 FHIPI-VALTAHAMSCDEAKCLEVGMDAYLTKPIDFKLMESTILSLTKRETL 1171
P+ VAL+ + + KC+ G+D L KP+ + + L + L
Sbjct: 683 HQRPLLVALSGNTDKSTKEKCMSFGLDGVLLKPVSLDNIRDVLSDLLEPRVL 734
>ETR1_TOBAC (O48929) Ethylene receptor (EC 2.7.3.-) (NT-ETR1)
Length = 738
Score = 122 bits (305), Expect = 7e-27
Identities = 102/347 (29%), Positives = 152/347 (43%), Gaps = 36/347 (10%)
Query: 461 ARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTA 520
ARR+AE + ++ FLA M+HE+RTPM A+I L LL + LT EQ V I K S
Sbjct: 331 ARREAEMAVRARNDFLAVMNHEMRTPMHAIIALSS-LLQETELTPEQRLMVETILKSSNL 389
Query: 521 LLHLLNNILDISKVESGKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPK 580
L L+N++LD+S++E G L L+ F+ ++++ N L P+
Sbjct: 390 LATLINDVLDLSRLEDGSLQLDVGTFNLHVLFRKVLNLIKPIASVKNCLSRLTCLQICPE 449
Query: 581 LVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGDSENFTLEQKPFGCSRKT 640
GD R++QI N++ N++KF+ G + + S D + +
Sbjct: 450 FAIGDEKRLMQILLNVVGNAVKFSKEGSVSISAVAAKSESLSDPR-----------APEF 498
Query: 641 KMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHGGTGL 700
Q ENH L +V DTG GI+P +F F Q T+ GGTGL
Sbjct: 499 FPVQSENH----------FYLRVQVKDTGSGINPQDIPKLFCKFAQNQALATKSSGGTGL 548
Query: 701 GLCIVRSLVNKMGGEIKIVKKE-GPGTLMRLYMQLTAPVDATE-----------QHCQVD 748
GL I + VN M G I I + G G+ ++L P + E H Q+
Sbjct: 549 GLAISKRFVNLMEGHIWIESEGLGKGSTAIFIVKLGIPGRSNEPKLPFMPRLPANHMQMT 608
Query: 749 FANNNLMVLLALHGNMSRLITSKWLQKNGVLTMEASEWNGLTQILKE 795
F L VL+ SR++T L G S + ++L +
Sbjct: 609 F--QGLKVLIMDDNGFSRMVTKGLLVHLGCDVTTVSSGDECLRVLTQ 653
Score = 52.4 bits (124), Expect = 7e-06
Identities = 35/142 (24%), Positives = 61/142 (42%), Gaps = 21/142 (14%)
Query: 1025 SLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTE 1084
+ +GL++L+ +D + V +L +G V V G + + L
Sbjct: 608 TFQGLKVLIMDDNGFSRMVTKGLLVHLGCDVTTVSSGDECLRVLT--------------- 652
Query: 1085 ILSFPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIP-IVALTAHAMSCDEAKCLEV 1143
+ ++ MD +P +D YE +I + + G + P IVALT + + C+ V
Sbjct: 653 ----QEHKVVFMDVSIPGIDCYEVAVQIHE-KFGKHHNRPLIVALTGNTDRVTKENCMRV 707
Query: 1144 GMDAYLTKPIDFKLMESTILSL 1165
GMD + KP+ M S + L
Sbjct: 708 GMDGVILKPVSVDKMRSVLSEL 729
>ARCB_ECOLI (P22763) Aerobic respiration control sensor protein arcB
(EC 2.7.3.-)
Length = 778
Score = 111 bits (277), Expect = 1e-23
Identities = 86/314 (27%), Positives = 142/314 (44%), Gaps = 43/314 (13%)
Query: 466 EASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTALLHLL 525
E +S K+ F++ +SHELRTP+ ++GL ILL D LT EQ + I + L ++
Sbjct: 277 ERASRDKTTFISTISHELRTPLNGIVGLSRILL-DTELTAEQEKYLKTIHVSAVTLGNIF 335
Query: 526 NNILDISKVESGKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPKLVRGD 585
N+I+D+ K+E K+ L+ DF L L ++ ++Q + L+ + +P V D
Sbjct: 336 NDIIDMDKMERRKVQLDNQPVDFTSFLADLENLSALQAQQKGLRFNLEPTLPLPHQVITD 395
Query: 586 SARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGDSENFTLEQKPFGCSRKTKMKQH 645
R+ QI NLI+N++KFT G + +R
Sbjct: 396 GTRLRQILWNLISNAVKFTQQGQVTVR--------------------------------- 422
Query: 646 ENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESFEQA-DLSTTRLHGGTGLGLCI 704
D +L FEV+D+G GI + D +F + Q D + GTG+GL +
Sbjct: 423 -------VRYDEGDMLHFEVEDSGIGIPQDELDKIFAMYYQVKDSHGGKPATGTGIGLAV 475
Query: 705 VRSLVNKMGGEIKIVKKEGPGTLMRLYMQLTAPVDATEQ-HCQVDFANNNLMVLLALHGN 763
R L MGG+I + ++G G+ L + + + + + D L VLL
Sbjct: 476 SRRLAKNMGGDITVTSEQGKGSTFTLTIHAPSVAEEVDDAFDEDDMPLPALNVLLVEDIE 535
Query: 764 MSRLITSKWLQKNG 777
++ ++ L+K G
Sbjct: 536 LNVIVARSVLEKLG 549
Score = 58.2 bits (139), Expect = 1e-07
Identities = 43/137 (31%), Positives = 67/137 (48%), Gaps = 20/137 (14%)
Query: 1026 LEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEI 1085
L L +LL ED + VA +LEK+G +V G+ A++ PG
Sbjct: 523 LPALNVLLVEDIELNVIVARSVLEKLGNSVDVAMTGKAALEMFK--PG------------ 568
Query: 1086 LSFPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTAHAMSCDEAKCLEVGM 1145
YDL+L+D Q+P M G + ++E+ K P+VALTA+ + D+ + L GM
Sbjct: 569 ----EYDLVLLDIQLPDMTGLDISRELTKRYPREDLP-PLVALTANVLK-DKQEYLNAGM 622
Query: 1146 DAYLTKPIDFKLMESTI 1162
D L+KP+ + + I
Sbjct: 623 DDVLSKPLSVPALTAMI 639
>ARCB_ECO57 (P58363) Aerobic respiration control sensor protein arcB
(EC 2.7.3.-)
Length = 778
Score = 111 bits (277), Expect = 1e-23
Identities = 86/314 (27%), Positives = 142/314 (44%), Gaps = 43/314 (13%)
Query: 466 EASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTALLHLL 525
E +S K+ F++ +SHELRTP+ ++GL ILL D LT EQ + I + L ++
Sbjct: 277 ERASRDKTTFISTISHELRTPLNGIVGLSRILL-DTELTAEQEKYLKTIHVSAVTLGNIF 335
Query: 526 NNILDISKVESGKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPKLVRGD 585
N+I+D+ K+E K+ L+ DF L L ++ ++Q + L+ + +P V D
Sbjct: 336 NDIIDMDKMERRKVQLDNQPVDFTSFLADLENLSALQAQQKGLRFNLEPTLPLPHQVITD 395
Query: 586 SARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGDSENFTLEQKPFGCSRKTKMKQH 645
R+ QI NLI+N++KFT G + +R
Sbjct: 396 GTRLRQILWNLISNAVKFTQQGQVTVR--------------------------------- 422
Query: 646 ENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESFEQA-DLSTTRLHGGTGLGLCI 704
D +L FEV+D+G GI + D +F + Q D + GTG+GL +
Sbjct: 423 -------VRYDEGDMLHFEVEDSGIGIPQDELDKIFAMYYQVKDSHGGKPATGTGIGLAV 475
Query: 705 VRSLVNKMGGEIKIVKKEGPGTLMRLYMQLTAPVDATEQ-HCQVDFANNNLMVLLALHGN 763
R L MGG+I + ++G G+ L + + + + + D L VLL
Sbjct: 476 SRRLAKNMGGDITVTSEQGKGSTFTLTIHAPSVAEEVDDAFDEDDMPLPALNVLLVEDIE 535
Query: 764 MSRLITSKWLQKNG 777
++ ++ L+K G
Sbjct: 536 LNVIVARSVLEKLG 549
Score = 58.2 bits (139), Expect = 1e-07
Identities = 43/137 (31%), Positives = 67/137 (48%), Gaps = 20/137 (14%)
Query: 1026 LEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEI 1085
L L +LL ED + VA +LEK+G +V G+ A++ PG
Sbjct: 523 LPALNVLLVEDIELNVIVARSVLEKLGNSVDVAMTGKAALEMFK--PG------------ 568
Query: 1086 LSFPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTAHAMSCDEAKCLEVGM 1145
YDL+L+D Q+P M G + ++E+ K P+VALTA+ + D+ + L GM
Sbjct: 569 ----EYDLVLLDIQLPDMTGLDISRELTKRYPREDLP-PLVALTANVLK-DKQEYLNAGM 622
Query: 1146 DAYLTKPIDFKLMESTI 1162
D L+KP+ + + I
Sbjct: 623 DDVLSKPLSVPALTAMI 639
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.133 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 130,886,427
Number of Sequences: 164201
Number of extensions: 5475047
Number of successful extensions: 17175
Number of sequences better than 10.0: 423
Number of HSP's better than 10.0 without gapping: 154
Number of HSP's successfully gapped in prelim test: 269
Number of HSP's that attempted gapping in prelim test: 16409
Number of HSP's gapped (non-prelim): 645
length of query: 1172
length of database: 59,974,054
effective HSP length: 121
effective length of query: 1051
effective length of database: 40,105,733
effective search space: 42151125383
effective search space used: 42151125383
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 72 (32.3 bits)
Medicago: description of AC121233.1


