

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119417.4 - phase: 0 /pseudo
(459 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
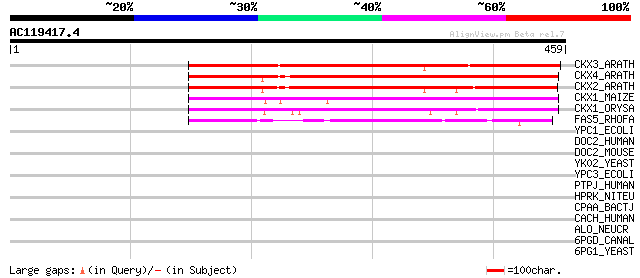
Score E
Sequences producing significant alignments: (bits) Value
CKX3_ARATH (Q9LTS3) Cytokinin dehydrogenase 3 precursor (EC 1.5.... 301 2e-81
CKX4_ARATH (Q9FUJ2) Cytokinin dehydrogenase 4 precursor (EC 1.5.... 273 5e-73
CKX2_ARATH (Q9FUJ3) Cytokinin dehydrogenase 2 precursor (EC 1.5.... 272 1e-72
CKX1_MAIZE (Q9T0N8) Cytokinin dehydrogenase 1 precursor (EC 1.5.... 242 2e-63
CKX1_ORYSA (Q9LDE6) Probable cytokinin dehydrogenase precursor (... 238 2e-62
FAS5_RHOFA (P46377) Hypothetical 47.9 kDa oxidoreductase in fasc... 130 9e-30
YPC1_ECOLI (P18351) Hypothetical 27.6 kDa protein (ORF 239) 32 2.6
DOC2_HUMAN (Q92608) Dedicator of cytokinesis protein 2 32 4.5
DOC2_MOUSE (Q8C3J5) Dedicator of cytokinesis protein 2 (Hch prot... 31 5.8
YK02_YEAST (P36118) Hypothetical 36.6 kDa protein in YPT52-DBP7 ... 31 7.6
YPC3_ECOLI (P18352) Hypothetical 6.6 kDa protein 30 9.9
PTPJ_HUMAN (Q12913) Protein-tyrosine phosphatase eta precursor (... 30 9.9
HPRK_NITEU (Q82Y30) HPr kinase/phosphorylase (EC 2.7.1.-) (EC 2.... 30 9.9
CPAA_BACTJ (O87906) Pesticidial crystal protein cry25Aa (Insecti... 30 9.9
CACH_HUMAN (Q8WYK0) Cytoplasmic acetyl-CoA hydrolase 1 (EC 3.1.2... 30 9.9
ALO_NEUCR (Q7SGY1) Putative D-arabinono-1,4-lactone oxidase (EC ... 30 9.9
6PGD_CANAL (O13287) 6-phosphogluconate dehydrogenase, decarboxyl... 30 9.9
6PG1_YEAST (P38720) 6-phosphogluconate dehydrogenase, decarboxyl... 30 9.9
>CKX3_ARATH (Q9LTS3) Cytokinin dehydrogenase 3 precursor (EC
1.5.99.12) (Cytokinin oxidase 3) (CKO 3)
Length = 523
Score = 301 bits (772), Expect = 2e-81
Identities = 144/310 (46%), Positives = 213/310 (68%), Gaps = 5/310 (1%)
Query: 149 SEEQNGELFYSVLGGLGQFGIITRARILLEPAPTMVKWIRVLYSDFTAFTKDQEKLIFAE 208
S++ N +LF++VLGGLGQFGIITRARI LE AP KW+R LY DF+ FT+DQE++I
Sbjct: 212 SKDMNSDLFFAVLGGLGQFGIITRARIKLEVAPKRAKWLRFLYIDFSEFTRDQERVISKT 271
Query: 209 KAFDYIEGFVIKNRTGLLNNWRLSFNP-QDPVQASKFKSDGRTLFCLELAKYFNMEETLE 267
D++EG ++ + G +NWR ++ P D ++ + R ++CLE+ KY++
Sbjct: 272 DGVDFLEGSIMVDH-GPPDNWRSTYYPPSDHLRIASMVKRHRVIYCLEVVKYYDETSQYT 330
Query: 268 VNQDIQKHLSHLNFIPSTLFQTEVTYVDFLDRVHISEVKLRSKGLWDVPHPWLNLFIPKS 327
VN+++++ LN + +++ +VTY+DFL+RV E+ L+SKG WDVPHPWLNLF+PK+
Sbjct: 331 VNEEMEELSDSLNHVRGFMYEKDVTYMDFLNRVRTGELNLKSKGQWDVPHPWLNLFVPKT 390
Query: 328 KIHNFAEVVFGNIV--KETSNGPVLIYPVHKSKWDKRTSVVIPDEDIFYLVGFLASSSGP 385
+I F + VF I+ ++GPVL+YP++++KW+ R S IP+ED+FY VGFL S+G
Sbjct: 391 QISKFDDGVFKGIILRNNITSGPVLVYPMNRNKWNDRMSAAIPEEDVFYAVGFL-RSAGF 449
Query: 386 DELEHILSQNKRILEYCERAHLGVKQYLPHYTTQEEWQTHYGHKWEIFKQRKSIYDPLAI 445
D E +N IL++CE A++GV QYLP++++QE W H+G +W IF +RK YDP I
Sbjct: 450 DNWEAFDQENMEILKFCEDANMGVIQYLPYHSSQEGWVRHFGPRWNIFVERKYKYDPKMI 509
Query: 446 LAPGQGIFSK 455
L+PGQ IF K
Sbjct: 510 LSPGQNIFQK 519
>CKX4_ARATH (Q9FUJ2) Cytokinin dehydrogenase 4 precursor (EC
1.5.99.12) (Cytokinin oxidase 4) (CKO 4)
Length = 524
Score = 273 bits (699), Expect = 5e-73
Identities = 137/311 (44%), Positives = 207/311 (66%), Gaps = 9/311 (2%)
Query: 149 SEEQNGELFYSVLGGLGQFGIITRARILLEPAPTMVKWIRVLYSDFTAFTKDQEKLIFA- 207
S + N ELFY VLGGLGQFGIITRARI L+ APT VKW R+LYSDF+AF +DQE+LI
Sbjct: 218 SPKLNPELFYGVLGGLGQFGIITRARIALDHAPTRVKWSRILYSDFSAFKRDQERLISMT 277
Query: 208 -EKAFDYIEGFVIKNRTGLLNNWRLSFNP-QDPVQASKFKSDGRTLFCLELAKYFNMEET 265
+ D++EG ++ + G ++ SF P D + + +D R ++ LE+AKY++
Sbjct: 278 NDLGVDFLEGQLMMSN-GFVDT---SFFPLSDQTRVASLVNDHRIIYVLEVAKYYDRTTL 333
Query: 266 LEVNQDIQKHLSHLNFIPSTLFQTEVTYVDFLDRVHISEVKLRSKGLWDVPHPWLNLFIP 325
++Q I L F P +F +V Y DFL+RV E KLRS GLW+VPHPWLN+F+P
Sbjct: 334 PIIDQVIDTLSRTLGFAPGFMFVQDVPYFDFLNRVRNEEDKLRSLGLWEVPHPWLNIFVP 393
Query: 326 KSKIHNFAE-VVFGNIVKETS-NGPVLIYPVHKSKWDKRTSVVIPDEDIFYLVGFLASSS 383
S+I +F + V+ G ++ +TS +G L YP +++KW+ R S + PDED+FY++G L S+
Sbjct: 394 GSRIQDFHDGVINGLLLNQTSTSGVTLFYPTNRNKWNNRMSTMTPDEDVFYVIGLLQSAG 453
Query: 384 GPDELEHILSQNKRILEYCERAHLGVKQYLPHYTTQEEWQTHYGHKWEIFKQRKSIYDPL 443
G + + + N +++++CE + + +K+YL HYT +E+W H+G KW+ F ++K ++DP
Sbjct: 454 GSQNWQELENLNDKVIQFCENSGIKIKEYLMHYTRKEDWVKHFGPKWDDFLRKKIMFDPK 513
Query: 444 AILAPGQGIFS 454
+L+PGQ IF+
Sbjct: 514 RLLSPGQDIFN 524
>CKX2_ARATH (Q9FUJ3) Cytokinin dehydrogenase 2 precursor (EC
1.5.99.12) (Cytokinin oxidase 2) (CKO 2)
Length = 501
Score = 272 bits (696), Expect = 1e-72
Identities = 136/311 (43%), Positives = 208/311 (66%), Gaps = 10/311 (3%)
Query: 149 SEEQNGELFYSVLGGLGQFGIITRARILLEPAPTMVKWIRVLYSDFTAFTKDQEKLIFA- 207
S + N ELFY VLGGLGQFGIITRARI+L+ AP KW R+LYSDFT FTKDQE+LI
Sbjct: 195 SRQLNPELFYGVLGGLGQFGIITRARIVLDHAPKRAKWFRMLYSDFTTFTKDQERLISMA 254
Query: 208 -EKAFDYIEGFVIKNRTGLLNNWRLSFNPQDPVQASKFKSDGRTLFCLELAKYFNMEETL 266
+ DY+EG + + G+++ F P D + + ++ LE+AKY++
Sbjct: 255 NDIGVDYLEGQIFLS-NGVVDT--SFFPPSDQSKVADLVKQHGIIYVLEVAKYYDDPNLP 311
Query: 267 EVNQDIQKHLSHLNFIPSTLFQTEVTYVDFLDRVHISEVKLRSKGLWDVPHPWLNLFIPK 326
+++ I L+++P + +V Y DFL+RVH+ E KLRS GLW++PHPWLNL++PK
Sbjct: 312 IISKVIDTLTKTLSYLPGFISMHDVAYFDFLNRVHVEENKLRSLGLWELPHPWLNLYVPK 371
Query: 327 SKIHNFAEVVFGNIV--KETSNGPVLIYPVHKSKWDKRTSVVIP--DEDIFYLVGFLASS 382
S+I +F V +I+ +++++G L+YP +++KWD R S +IP DED+ Y++G L S+
Sbjct: 372 SRILDFHNGVVKDILLKQKSASGLALLYPTNRNKWDNRMSAMIPEIDEDVIYIIGLLQSA 431
Query: 383 SGPDELEHILSQNKRILEYCERAHLGVKQYLPHYTTQEEWQTHYGHKWEIFKQRKSIYDP 442
+ P +L + S N++I+ +C+ + + +KQYL HYT++E+W H+G KW+ F +RK ++DP
Sbjct: 432 T-PKDLPEVESVNEKIIRFCKDSGIKIKQYLMHYTSKEDWIEHFGSKWDDFSKRKDLFDP 490
Query: 443 LAILAPGQGIF 453
+L+PGQ IF
Sbjct: 491 KKLLSPGQDIF 501
>CKX1_MAIZE (Q9T0N8) Cytokinin dehydrogenase 1 precursor (EC
1.5.99.12) (Cytokinin oxidase 1) (CKO 1)
Length = 534
Score = 242 bits (617), Expect = 2e-63
Identities = 128/321 (39%), Positives = 188/321 (57%), Gaps = 15/321 (4%)
Query: 149 SEEQNGELFYSVLGGLGQFGIITRARILLEPAPTMVKWIRVLYSDFTAFTKDQEKLIFAE 208
S++ N +LF +VLGGLGQFG+ITRARI +EPAP +W+R++Y+DF AF+ DQE+L
Sbjct: 214 SKQLNADLFDAVLGGLGQFGVITRARIAVEPAPARARWVRLVYTDFAAFSADQERLTAPR 273
Query: 209 KA--------FDYIEGFVIKNR---TGLLNNWRLSFNPQDPVQASKFKSDGRTLFCLELA 257
Y+EG V N+ T L N + + A + + T++ +E
Sbjct: 274 PGGGGASFGPMSYVEGSVFVNQSLATDLANTGFFTDADVARIVALAGERNATTVYSIEAT 333
Query: 258 KYFN--MEETLEVNQDIQKHLSHLNFIPSTLFQTEVTYVDFLDRVHISEVKLRSKGLWDV 315
++ V+Q++ L L+++ FQ +V Y FLDRVH EV L GLW V
Sbjct: 334 LNYDNATAAAAAVDQELASVLGTLSYVEGFAFQRDVAYAAFLDRVHGEEVALNKLGLWRV 393
Query: 316 PHPWLNLFIPKSKIHNFAEVVFGNIVKETSN-GPVLIYPVHKSKWDKRTSVVIPDEDIFY 374
PHPWLN+F+P+S+I +F VF I++ T GP+++YP++KS WD S P ED+FY
Sbjct: 394 PHPWLNMFVPRSRIADFDRGVFKGILQGTDIVGPLIVYPLNKSMWDDGMSAATPSEDVFY 453
Query: 375 LVGFLASSSGPDELEHILSQNKRILEYCERAHLGVKQYLPHYTTQEEWQTHYG-HKWEIF 433
V L SS P++L + QN+RIL +C+ A + K YL +T + +W H+G KW F
Sbjct: 454 AVSLLFSSVAPNDLARLQEQNRRILRFCDLAGIQYKTYLARHTDRSDWVRHFGAAKWNRF 513
Query: 434 KQRKSIYDPLAILAPGQGIFS 454
+ K+ YDP +L+PGQ IF+
Sbjct: 514 VEMKNKYDPKRLLSPGQDIFN 534
>CKX1_ORYSA (Q9LDE6) Probable cytokinin dehydrogenase precursor (EC
1.5.99.12) (Cytokinin oxidase) (CKO)
Length = 532
Score = 238 bits (607), Expect = 2e-62
Identities = 133/321 (41%), Positives = 192/321 (59%), Gaps = 16/321 (4%)
Query: 149 SEEQNGELFYSVLGGLGQFGIITRARILLEPAPTMVKWIRVLYSDFTAFTKDQEKLIFAE 208
S+ N +LF +VLGGLGQFG+ITRAR+ +EPAP +W+R++Y+DF AF+ DQE+L+ A
Sbjct: 213 SKAVNSDLFDAVLGGLGQFGVITRARVAVEPAPARARWVRLVYADFAAFSADQERLVAAR 272
Query: 209 K-----AFDYIEGFVIKNRTGLLNNWRLS---FNPQDP--VQASKFKSDGRTLFCLELAK 258
+ Y+EG V GL + S F+ D V A + ++ +E
Sbjct: 273 PDGSHGPWSYVEGAVYLAGRGLAVALKSSGGFFSDADAARVVALAAARNATAVYSIEATL 332
Query: 259 YFNMEET-LEVNQDIQKHLSHLNFIPSTLFQTEVTYVDFLDRVHISEVKLRSKGLWDVPH 317
+ T V+ + L L+F F +VTY +FLDRV+ E L GLW VPH
Sbjct: 333 NYAANATPSSVDAAVAAALGDLHFEEGFSFSRDVTYEEFLDRVYGEEEALEKAGLWRVPH 392
Query: 318 PWLNLFIPKSKIHNFAEVVFGNIVKETSN--GPVLIYPVHKSKWDKRTSVVIP--DEDIF 373
PWLNLF+P S+I +F VF I++ ++ GP++IYPV+KSKWD S V P +E++F
Sbjct: 393 PWLNLFVPGSRIADFDRGVFKGILQTATDIAGPLIIYPVNKSKWDAAMSAVTPEGEEEVF 452
Query: 374 YLVGFLASSSGPDELEHILSQNKRILEYCERAHLGVKQYLPHYTTQEEWQTHYGHKWEIF 433
Y+V L S+ D + + +QN+RIL +C+ A +G K YL HY ++ +W H+G KW+ F
Sbjct: 453 YVVSLLFSAVAND-VAALEAQNRRILRFCDLAGIGYKAYLAHYDSRGDWVRHFGAKWDRF 511
Query: 434 KQRKSIYDPLAILAPGQGIFS 454
QRK YDP +L+PGQ IF+
Sbjct: 512 VQRKDKYDPKKLLSPGQDIFN 532
>FAS5_RHOFA (P46377) Hypothetical 47.9 kDa oxidoreductase in
fasciation locus (ORF5)
Length = 438
Score = 130 bits (326), Expect = 9e-30
Identities = 91/304 (29%), Positives = 141/304 (45%), Gaps = 36/304 (11%)
Query: 149 SEEQNGELFYSVLGGLGQFGIITRARILLEPAPTMVKWIRVLYSDFTAFTKDQEKLIFAE 208
S N ELF +V GGLGQFG+I A I L A V+ ++ YS+ F DQ + + +
Sbjct: 162 SAVSNSELFDAVRGGLGQFGVIVNATIRLTAAHESVRQYKLQYSNLGVFLGDQLRAM-SN 220
Query: 209 KAFDYIEGFVIKNRTGLLNNWRLSFNPQDPVQASKFKSDGRTLFCLELAKYFNMEETLEV 268
+ FD+++G + + +DG + L+LAKYF T
Sbjct: 221 RLFDHVQGRI------------------------RVDADGHLRYRLDLAKYF----TPPR 252
Query: 269 NQDIQKHLSHLNFIPSTLFQTEVTYVDFLDRVHISEVKLRSKGLWDVPHPWLNLFIPKSK 328
D LS L + + ++V Y DF++R+ E+ LR G W PHPW +L IP K
Sbjct: 253 RPDDDALLSSLQYDSCAEYNSDVDYGDFINRMADQELDLRHTGEWFYPHPWASLLIPADK 312
Query: 329 IHNFAEVVFGNIVKETSN-GPVLIYPVHKSKWDKRTSVVIPDEDIFYLVGFLASSSGPDE 387
I F E ++ + N G +++YP+ + + IP D F+++ L ++S E
Sbjct: 313 IEQFIETTSSSLTDDLGNSGLIMVYPIPTTP-ITAPFIPIPHCDTFFMLAVLRTASPGAE 371
Query: 388 LEHILSQNKRILEYCERAHLGVKQYLPHYTTQE--EWQTHYGHKWEIFKQRKSIYDPLAI 445
I S L Y + +G Y + +W TH+G +W+ + K +DP I
Sbjct: 372 ARMIASNR---LLYEQARDVGGVAYAVNAVPMSPGDWCTHFGSRWQAIARAKRRFDPYRI 428
Query: 446 LAPG 449
LAPG
Sbjct: 429 LAPG 432
>YPC1_ECOLI (P18351) Hypothetical 27.6 kDa protein (ORF 239)
Length = 239
Score = 32.3 bits (72), Expect = 2.6
Identities = 15/39 (38%), Positives = 23/39 (58%), Gaps = 2/39 (5%)
Query: 299 RVHISEVKLRSKGLWDVPHPWLN--LFIPKSKIHNFAEV 335
R++ S + +S GLW H WLN +P+S+ N AE+
Sbjct: 160 RLYESLAQFKSSGLWVTTHAWLNDRFLLPESQQKNLAEL 198
>DOC2_HUMAN (Q92608) Dedicator of cytokinesis protein 2
Length = 1830
Score = 31.6 bits (70), Expect = 4.5
Identities = 34/166 (20%), Positives = 70/166 (41%), Gaps = 22/166 (13%)
Query: 191 YSDFTAFTKDQEKLIFAEKAFDYIEGFVIKNRTGLLNNWRLSFNPQDPVQASKFKSDGRT 250
Y D ++ + E ++ KA +Y+ F++++RT L + + +F+ R
Sbjct: 720 YLDTSSRGEQCEPILRTLKALEYVFKFIVRSRT-------LFSQLYEGKEQMEFEESMRR 772
Query: 251 LFCLELAKYFNMEETLEVNQDIQKHLSHLNFIPSTLFQTEVTYVDFLDRVHISEVKLRSK 310
LF + N + I ++ L +IPS L E+ + + KL S+
Sbjct: 773 LF-----ESINNLMKSQYKTTILLQVAALKYIPSVLHDVEMVF----------DAKLLSQ 817
Query: 311 GLWDVPHPWLNLFIPKSKIHNFAEVVFGNIVKETSNGPVLIYPVHK 356
L++ + + K K+ + E+V N+ K+ +L+ + K
Sbjct: 818 LLYEFYTCIPPVKLQKQKVQSMNEIVQSNLFKKQECRDILLPVITK 863
>DOC2_MOUSE (Q8C3J5) Dedicator of cytokinesis protein 2 (Hch
protein)
Length = 1828
Score = 31.2 bits (69), Expect = 5.8
Identities = 34/166 (20%), Positives = 69/166 (41%), Gaps = 22/166 (13%)
Query: 191 YSDFTAFTKDQEKLIFAEKAFDYIEGFVIKNRTGLLNNWRLSFNPQDPVQASKFKSDGRT 250
Y D ++ + E ++ KA +Y+ F++++RT L + + +F+ R
Sbjct: 720 YLDTSSRGEQCEPILRTLKALEYVFKFIVRSRT-------LFSQLYEGKEQMEFEESMRR 772
Query: 251 LFCLELAKYFNMEETLEVNQDIQKHLSHLNFIPSTLFQTEVTYVDFLDRVHISEVKLRSK 310
LF + N + I ++ L +IPS L E + + KL S+
Sbjct: 773 LF-----ESINNLMKSQYKTTILLQVAALKYIPSVLHDVETVF----------DAKLLSQ 817
Query: 311 GLWDVPHPWLNLFIPKSKIHNFAEVVFGNIVKETSNGPVLIYPVHK 356
L++ + + K K+ + E+V N+ K+ +L+ + K
Sbjct: 818 LLYEFYTCIPPVKLQKQKVQSMNEIVQSNLFKKQECRDILLPVITK 863
>YK02_YEAST (P36118) Hypothetical 36.6 kDa protein in YPT52-DBP7
intergenic region
Length = 322
Score = 30.8 bits (68), Expect = 7.6
Identities = 16/42 (38%), Positives = 19/42 (45%)
Query: 416 YTTQEEWQTHYGHKWEIFKQRKSIYDPLAILAPGQGIFSKSI 457
Y T EEW+T K I K + I PL +L P SI
Sbjct: 236 YRTNEEWETQLLSKGNINKSNEKIITPLPVLFPDDDESGNSI 277
>YPC3_ECOLI (P18352) Hypothetical 6.6 kDa protein
Length = 57
Score = 30.4 bits (67), Expect = 9.9
Identities = 14/35 (40%), Positives = 20/35 (57%), Gaps = 2/35 (5%)
Query: 303 SEVKLRSKGLWDVPHPWLN--LFIPKSKIHNFAEV 335
S + +S GLW H WLN +P+S+ N AE+
Sbjct: 3 SLAQFKSSGLWVTTHAWLNDRFLLPESQQKNLAEL 37
>PTPJ_HUMAN (Q12913) Protein-tyrosine phosphatase eta precursor (EC
3.1.3.48) (R-PTP-eta) (HPTP eta) (Protein-tyrosine
phosphatase receptor type J) (Density enhanced
phosphatase-1) (DEP-1) (CD148 antigen)
Length = 1337
Score = 30.4 bits (67), Expect = 9.9
Identities = 31/104 (29%), Positives = 46/104 (43%), Gaps = 18/104 (17%)
Query: 266 LEVNQDIQKHLSHLNFIPS---TLFQTEVTYVDFLDRVHISEVKLR------------SK 310
LEV+ + +HL S T ++TEVTY++F +IS + +
Sbjct: 752 LEVSSGAWNNATHLESCSSENGTEYRTEVTYLNFSTSYNISITTVSCGKMAAPTRNTCTT 811
Query: 311 GLWDVPHPWLNLFIPKSKIHNFAEVVFGNIVKETSNGPVLIYPV 354
G+ D P P + I S HN +V F E S+GP+ Y V
Sbjct: 812 GITDPPPPDGSPNI-TSVSHNSVKVKFSGF--EASHGPIKAYAV 852
>HPRK_NITEU (Q82Y30) HPr kinase/phosphorylase (EC 2.7.1.-) (EC
2.7.4.-) (HPrK/P) (HPr(Ser) kinase/phosphorylase)
Length = 323
Score = 30.4 bits (67), Expect = 9.9
Identities = 16/45 (35%), Positives = 24/45 (52%), Gaps = 4/45 (8%)
Query: 264 ETLEVNQDIQKHLSHLNFIP----STLFQTEVTYVDFLDRVHISE 304
E ++N Q+ + HLNF+ L QT V Y+D LD V + +
Sbjct: 33 ENHQINNSTQELIGHLNFVHPNWIQVLNQTSVNYLDQLDDVSLKK 77
>CPAA_BACTJ (O87906) Pesticidial crystal protein cry25Aa
(Insecticidal delta-endotoxin CryXXVA(a)) (Crystaline
entomocidal protoxin) (76 kDa crystal protein)
(Insecticidal protein Jeg74)
Length = 675
Score = 30.4 bits (67), Expect = 9.9
Identities = 16/48 (33%), Positives = 26/48 (53%)
Query: 175 ILLEPAPTMVKWIRVLYSDFTAFTKDQEKLIFAEKAFDYIEGFVIKNR 222
I L P PT + RV+Y+D +++ IF+ FDY+E + + R
Sbjct: 296 IKLYPIPTQTELSRVVYTDPVGCFGNRKSDIFSRLNFDYLENRLTRPR 343
>CACH_HUMAN (Q8WYK0) Cytoplasmic acetyl-CoA hydrolase 1 (EC 3.1.2.1)
(CACH-1) (hCACH-1)
Length = 555
Score = 30.4 bits (67), Expect = 9.9
Identities = 23/98 (23%), Positives = 44/98 (44%), Gaps = 4/98 (4%)
Query: 255 ELAKYFNMEETLEVNQDIQKHLSHLNFIPSTLFQTEVTYVDFLDRVHISEVKLRSKGLWD 314
++ + F+ + + +Q L+ + + S F T V +++H+ V L L +
Sbjct: 76 KVTRAFSTSMEISIKVMVQDMLTGIEKLVSVAFSTFVAKPVGKEKIHLKPVTL----LTE 131
Query: 315 VPHPWLNLFIPKSKIHNFAEVVFGNIVKETSNGPVLIY 352
H NL + K+ E F N++KE+S LI+
Sbjct: 132 QDHVEHNLAAERRKVRLQHEDTFNNLMKESSKFDDLIF 169
>ALO_NEUCR (Q7SGY1) Putative D-arabinono-1,4-lactone oxidase (EC
1.1.3.37) (ALO) (L-galactono-gamma-lactone oxidase)
Length = 556
Score = 30.4 bits (67), Expect = 9.9
Identities = 29/91 (31%), Positives = 41/91 (44%), Gaps = 7/91 (7%)
Query: 149 SEEQNGELFYSVLGGLGQFGIITRARILLEPAPTMVKWIRVLYSDFTAFTKDQEKLIFAE 208
S E +LF + L LG GIIT PA ++ W + + SD F K EK ++++
Sbjct: 186 SPEDKPDLFRAALISLGALGIITEVTFKAVPAFSLA-WSQAIDSDKRIFEK-WEKDLWSQ 243
Query: 209 KAFDYIEGFVIKNRTGLLNNWRLSFNPQDPV 239
F I F R + W + N DPV
Sbjct: 244 AEFVRIWWFPYMRRAAV---W--TANVVDPV 269
>6PGD_CANAL (O13287) 6-phosphogluconate dehydrogenase,
decarboxylating (EC 1.1.1.44)
Length = 517
Score = 30.4 bits (67), Expect = 9.9
Identities = 12/41 (29%), Positives = 22/41 (53%)
Query: 192 SDFTAFTKDQEKLIFAEKAFDYIEGFVIKNRTGLLNNWRLS 232
+D F D E+ ++A K Y +GF++ N+ W+L+
Sbjct: 341 TDKKQFIDDLEQALYASKIISYTQGFMLMNQAAKDYGWKLN 381
>6PG1_YEAST (P38720) 6-phosphogluconate dehydrogenase,
decarboxylating 1 (EC 1.1.1.44)
Length = 489
Score = 30.4 bits (67), Expect = 9.9
Identities = 19/77 (24%), Positives = 35/77 (44%), Gaps = 6/77 (7%)
Query: 193 DFTAFTKDQEKLIFAEKAFDYIEGFVIKNRTGLLNNWRLSFNPQDPVQASKFKSDG--RT 250
D F D E+ ++A K Y +GF++ W+L+ +P A ++ R+
Sbjct: 314 DREQFVDDLEQALYASKIISYAQGFMLIREAAATYGWKLN----NPAIALMWRGGCIIRS 369
Query: 251 LFCLELAKYFNMEETLE 267
+F ++ K + E LE
Sbjct: 370 VFLGQITKAYREEPDLE 386
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.336 0.148 0.478
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 51,950,215
Number of Sequences: 164201
Number of extensions: 2131083
Number of successful extensions: 7684
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 7652
Number of HSP's gapped (non-prelim): 22
length of query: 459
length of database: 59,974,054
effective HSP length: 114
effective length of query: 345
effective length of database: 41,255,140
effective search space: 14233023300
effective search space used: 14233023300
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 67 (30.4 bits)
Medicago: description of AC119417.4


