

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119414.6 - phase: 0
(347 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
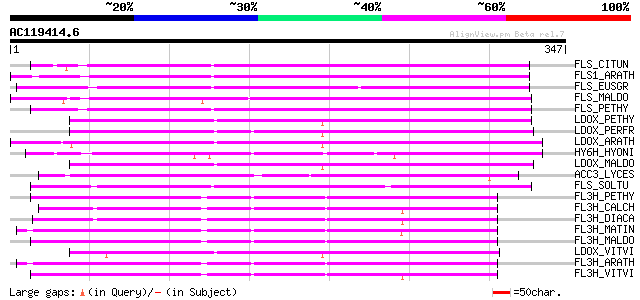
Sequences producing significant alignments: (bits) Value
FLS_CITUN (Q9ZWQ9) Flavonol synthase (EC 1.14.11.-) (FLS) (CitFLS) 214 2e-55
FLS1_ARATH (Q96330) Flavonol synthase 1 (EC 1.14.11.-) (FLS 1) 209 6e-54
FLS_EUSGR (Q9M547) Flavonol synthase (EC 1.14.11.-) (FLS) 205 1e-52
FLS_MALDO (Q9XHG2) Flavonol synthase (EC 1.14.11.-) (FLS) 203 5e-52
FLS_PETHY (Q07512) Flavonol synthase (EC 1.14.11.-) (FLS) 201 3e-51
LDOX_PETHY (P51092) Leucoanthocyanidin dioxygenase (EC 1.14.11.1... 199 7e-51
LDOX_PERFR (O04274) Leucoanthocyanidin dioxygenase (EC 1.14.11.1... 192 1e-48
LDOX_ARATH (Q96323) Leucoanthocyanidin dioxygenase (EC 1.14.11.1... 191 2e-48
HY6H_HYONI (P24397) Hyoscyamine 6-dioxygenase (EC 1.14.11.11) (H... 191 2e-48
LDOX_MALDO (P51091) Leucoanthocyanidin dioxygenase (EC 1.14.11.1... 191 2e-48
ACC3_LYCES (P10967) 1-aminocyclopropane-1-carboxylate oxidase ho... 191 2e-48
FLS_SOLTU (Q41452) Flavonol synthase (EC 1.14.11.-) (FLS) 190 4e-48
FL3H_PETHY (Q07353) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 184 3e-46
FL3H_CALCH (Q05963) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 182 1e-45
FL3H_DIACA (Q05964) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 181 2e-45
FL3H_MATIN (Q05965) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 179 7e-45
FL3H_MALDO (Q06942) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 177 3e-44
LDOX_VITVI (P51093) Leucoanthocyanidin dioxygenase (EC 1.14.11.1... 177 5e-44
FL3H_ARATH (Q9S818) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 177 5e-44
FL3H_VITVI (P41090) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 176 6e-44
>FLS_CITUN (Q9ZWQ9) Flavonol synthase (EC 1.14.11.-) (FLS) (CitFLS)
Length = 335
Score = 214 bits (546), Expect = 2e-55
Identities = 121/316 (38%), Positives = 181/316 (56%), Gaps = 12/316 (3%)
Query: 14 SLTPNFILPEHKRPHLSEVKY---LDSIPIIDLSYCDGNNPSSLEVIHKISKACEEFGFF 70
++ FI PE ++P + Y IP IDL ++P ++ I++A E+G F
Sbjct: 18 TIPAEFIRPEKEQP--ASTTYHGPAPEIPTIDL-----DDPVQDRLVRSIAEASREWGIF 70
Query: 71 QIVNHGVPDQVCTKMMKAITNFFELAPEEREHLSSTDNTKNVRLFNYYLQVDGGEKVKLW 130
Q+ NHG+P + K+ FFEL EE+E S + K+V+ + LQ + K K W
Sbjct: 71 QVTNHGIPSDLICKLQAVGKEFFELPQEEKEVYSRPADAKDVQGYGTKLQKEVEGK-KSW 129
Query: 131 SECFAHPWYPIDDIIQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLL 190
+ H +P I K YR EYAK + +V +L + +S+GLG+E L
Sbjct: 130 VDHLFHRVWPPSSINYRFWPKNPPSYRAVNEEYAKYMREVVDKLFTYLSLGLGVEGGVLK 189
Query: 191 KKLG-EQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQSEVSGLQVNKDGKWISI 249
+ G + + N+YPPCP P+L +G+ HTDL+ALTVL+ +EV GLQV KD +WI
Sbjct: 190 EAAGGDDIEYMLKINYYPPCPRPDLALGVVAHTDLSALTVLVPNEVPGLQVFKDDRWIDA 249
Query: 250 PCIPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNPETIIGPIHELI 309
IPNA VI++ DQIE+LSNG+YK+VLHR N + R+S +F P +T++GP+ +L+
Sbjct: 250 KYIPNALVIHIGDQIEILSNGKYKAVLHRTTVNKDKTRMSWPVFLEPPADTVVGPLPQLV 309
Query: 310 DEEHPPKYRNYHFSKF 325
D+E+PPKY+ F +
Sbjct: 310 DDENPPKYKAKKFKDY 325
>FLS1_ARATH (Q96330) Flavonol synthase 1 (EC 1.14.11.-) (FLS 1)
Length = 336
Score = 209 bits (533), Expect = 6e-54
Identities = 119/327 (36%), Positives = 184/327 (55%), Gaps = 12/327 (3%)
Query: 1 MESFQLANESSPLSLTPNFILPEHKRPHLSEVKY-LDSIPIIDLSYCDGNNPSSLEVIHK 59
+ S L E+ PL FI E ++P ++ + +IP++DLS +P V
Sbjct: 9 ISSSSLLTEAIPLE----FIRSEKEQPAITTFRGPTPAIPVVDLS-----DPDEESVRRA 59
Query: 60 ISKACEEFGFFQIVNHGVPDQVCTKMMKAITNFFELAPEEREHLSSTDNTKNVRLFNYYL 119
+ KA EE+G FQ+VNHG+P ++ ++ FFEL E+E ++ +++K++ + L
Sbjct: 60 VVKASEEWGLFQVVNHGIPTELIRRLQDVGRKFFELPSSEKESVAKPEDSKDIEGYGTKL 119
Query: 120 QVDGGEKVKLWSECFAHPWYPIDDIIQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLIS 179
Q D K K W + H +P + K +YRE EYA V L LL ++S
Sbjct: 120 QKDPEGK-KAWVDHLFHRIWPPSCVNYRFWPKNPPEYREVNEEYAVHVKKLSETLLGILS 178
Query: 180 IGLGLEEDCLLKKLG-EQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQSEVSGL 238
GLGL+ D L + LG E + N+YPPCP P+L +G+ HTDL+ +T+L+ +EV GL
Sbjct: 179 DGLGLKRDALKEGLGGEMAEYMMKINYYPPCPRPDLALGVPAHTDLSGITLLVPNEVPGL 238
Query: 239 QVNKDGKWISIPCIPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNP 298
QV KD W IP+A ++++ DQI LSNGRYK+VLHR + + R+S +F P
Sbjct: 239 QVFKDDHWFDAEYIPSAVIVHIGDQILRLSNGRYKNVLHRTTVDKEKTRMSWPVFLEPPR 298
Query: 299 ETIIGPIHELIDEEHPPKYRNYHFSKF 325
E I+GP+ EL +++PPK++ + F +
Sbjct: 299 EKIVGPLPELTGDDNPPKFKPFAFKDY 325
>FLS_EUSGR (Q9M547) Flavonol synthase (EC 1.14.11.-) (FLS)
Length = 334
Score = 205 bits (522), Expect = 1e-52
Identities = 115/324 (35%), Positives = 190/324 (58%), Gaps = 10/324 (3%)
Query: 5 QLANESSPLSLTP-NFILPEHKRPHLSEVK-YLDSIPIIDLSYCDGNNPSSLEVIHKISK 62
++A+ S + P +I E+++P +S V + +P+IDLS D +++ +S+
Sbjct: 8 EIASLSKVIDTIPAEYIRSENEQPVISTVHGVVLEVPVIDLSDSDEK-----KIVGLVSE 62
Query: 63 ACEEFGFFQIVNHGVPDQVCTKMMKAITNFFELAPEEREHLSSTDNTKNVRLFNYYLQVD 122
A +E+G FQ+VNHG+P++V K+ + +FFEL EE+E ++ + ++++ + LQ +
Sbjct: 63 ASKEWGIFQVVNHGIPNEVIRKLQEVGKHFFELPQEEKELIAKPEGSQSIEGYGTRLQKE 122
Query: 123 GGEKVKLWSECFAHPWYPIDDIIQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLISIGL 182
K K W + H +P I K YREA EYAK + +V L +S+GL
Sbjct: 123 VDGK-KGWVDHLFHKIWPPSAINYQFWPKNPPAYREANEEYAKRLQLVVDNLFKYLSLGL 181
Query: 183 GLEEDCLLKKLG-EQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQSEVSGLQVN 241
LE + G + + N+YPPCP P+L +G+ HTD++A+TVL+ +EV GLQV
Sbjct: 182 DLEPNSFKDGAGGDDLVYLMKINYYPPCPRPDLALGV-AHTDMSAITVLVPNEVPGLQVY 240
Query: 242 KDGKWISIPCIPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNPETI 301
KDG W IPNA ++++ DQ+E++SNG+YKSV HR N + R+S +F P P+
Sbjct: 241 KDGHWYDCKYIPNALIVHIGDQVEIMSNGKYKSVYHRTTVNKEKTRMSWPVFLEPPPDHE 300
Query: 302 IGPIHELIDEEHPPKYRNYHFSKF 325
+GPI +L++EE+P K++ + +
Sbjct: 301 VGPIPKLVNEENPAKFKTKKYKDY 324
>FLS_MALDO (Q9XHG2) Flavonol synthase (EC 1.14.11.-) (FLS)
Length = 337
Score = 203 bits (517), Expect = 5e-52
Identities = 118/333 (35%), Positives = 193/333 (57%), Gaps = 14/333 (4%)
Query: 1 MESFQLANESSPLSLTPNFILPEHKRPHLSEV--KYLDSIPIIDLSYCDGNNPSSLEVIH 58
+ES + ES+ ++ FI E+++P ++ V K L+ +PIID S +P ++I
Sbjct: 3 VESVERERESNEGTIPAEFIRSENEQPGITTVHGKVLE-VPIIDFS-----DPDEEKLIV 56
Query: 59 KISKACEEFGFFQIVNHGVPDQVCTKMMKAITNFFELAPEEREHLSSTDNTKNVRLFNYY 118
+I++A +G +QIVNH +P +V +K+ FFEL EE+E + ++ ++ +
Sbjct: 57 QITEASSNWGMYQIVNHDIPSEVISKLQAVGKEFFELPQEEKEAYAKPPDSASIEGYGTK 116
Query: 119 L--QVDGGEKVKL-WSE-CFAHPWYPIDDIIQLLPEKIGTQYREAFTEYAKEVGSLVRRL 174
L ++ G+ K W + F W P Q P K YREA EYAK + ++V +L
Sbjct: 117 LFKEISEGDTTKKGWVDNLFNKIWPPSVVNYQFWP-KNPPSYREANEEYAKHLHNVVEKL 175
Query: 175 LSLISIGLGLEEDCLLKKLG-EQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQS 233
L+S+GLGLE L K G + + N+YPPCP P+L +G+ HTD++ +T+L+ +
Sbjct: 176 FRLLSLGLGLEGQELKKAAGGDNLEYLLKINYYPPCPRPDLALGVVAHTDMSTVTILVPN 235
Query: 234 EVSGLQVNKDGKWISIPCIPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMF 293
+V GLQ KDG+W + IPNA VI++ DQ+E++SNG+Y SVLHR N + RIS +F
Sbjct: 236 DVQGLQACKDGRWYDVKYIPNALVIHIGDQMEIMSNGKYTSVLHRTTVNKDKTRISWPVF 295
Query: 294 YGPNPETIIGPIHELIDEEHPPKYRNYHFSKFL 326
P + ++GP +L++ + PKY+ + ++
Sbjct: 296 LEPPADHVVGPHPQLVNAVNQPKYKTKKYGDYV 328
>FLS_PETHY (Q07512) Flavonol synthase (EC 1.14.11.-) (FLS)
Length = 348
Score = 201 bits (510), Expect = 3e-51
Identities = 107/316 (33%), Positives = 182/316 (56%), Gaps = 9/316 (2%)
Query: 14 SLTPNFILPEHKRPHLSEVK-YLDSIPIIDLSYCDGNNPSSLEVIHKISKACEEFGFFQI 72
++ +I E+++P + + + +P+IDL +P +++ I+ A +E+G FQ+
Sbjct: 30 TIPSEYIRSENEQPAATTLHGVVLQVPVIDL-----RDPDENKMVKLIADASKEWGIFQL 84
Query: 73 VNHGVPDQVCTKMMKAITNFFELAP-EEREHLSSTDNTKNVRLFNYYLQVDGGEKVKLWS 131
+NHG+PD+ + K FFE P EE+E ++ T + ++ + LQ + K K W
Sbjct: 85 INHGIPDEAIADLQKVGKEFFEHVPQEEKELIAKTPGSNDIEGYGTSLQKEVEGK-KGWV 143
Query: 132 ECFAHPWYPIDDIIQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLK 191
+ H +P + K YREA EY K + +V R+ +S+GLGLE +++
Sbjct: 144 DHLFHKIWPPSAVNYRYWPKNPPSYREANEEYGKRMREVVDRIFKSLSLGLGLEGHEMIE 203
Query: 192 KLG-EQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQSEVSGLQVNKDGKWISIP 250
G ++ + N+YPPCP P+L +G+ HTD++ +T+L+ +EV GLQV KDG W +
Sbjct: 204 AAGGDEIVYLLKINYYPPCPRPDLALGVVAHTDMSYITILVPNEVQGLQVFKDGHWYDVK 263
Query: 251 CIPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNPETIIGPIHELID 310
IPNA ++++ DQ+E+LSNG+YKSV HR N + R+S +F P E +GPI +L+
Sbjct: 264 YIPNALIVHIGDQVEILSNGKYKSVYHRTTVNKDKTRMSWPVFLEPPSEHEVGPIPKLLS 323
Query: 311 EEHPPKYRNYHFSKFL 326
E +PPK++ + ++
Sbjct: 324 EANPPKFKTKKYKDYV 339
>LDOX_PETHY (P51092) Leucoanthocyanidin dioxygenase (EC 1.14.11.19)
(LDOX) (Leucocyanidin oxygenase) (Leucoanthocyanidin
hydroxylase)
Length = 430
Score = 199 bits (507), Expect = 7e-51
Identities = 106/292 (36%), Positives = 172/292 (58%), Gaps = 4/292 (1%)
Query: 38 IPIIDLSYCDGNNPSSLEVIHKISKACEEFGFFQIVNHGVPDQVCTKMMKAITNFFELAP 97
+P IDL D + E H++ KA E+G +VNHG+ D++ ++ A FF+
Sbjct: 52 VPTIDLKEIDSEDKEIREKCHQLKKAAMEWGVMHLVNHGISDELINRVKVAGETFFDQPV 111
Query: 98 EEREHLSSTDNTKNVRLFNYYLQVDGGEKVKLWSECFAHPWYPIDDIIQLLPEKIGTQYR 157
EE+E ++ NV+ + L +++ W + F H +P D + K T Y
Sbjct: 112 EEKEKYANDQANGNVQGYGSKLANSACGQLE-WEDYFFHCAFPEDKRDLSIWPKNPTDYT 170
Query: 158 EAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLKKLG--EQPRQRAQSNFYPPCPDPELT 215
A +EYAK++ +L ++L+++SIGLGLEE L K++G E + + N+YP CP PEL
Sbjct: 171 PATSEYAKQIRALATKILTVLSIGLGLEEGRLEKEVGGMEDLLLQMKINYYPKCPQPELA 230
Query: 216 MGLNEHTDLNALTVLLQSEVSGLQVNKDGKWISIPCIPNAFVINLADQIEVLSNGRYKSV 275
+G+ HTD++ALT +L + V GLQ+ +G+W++ C+PN+ ++++ D IE+LSNG+YKS+
Sbjct: 231 LGVEAHTDVSALTFILHNMVPGLQLFYEGQWVTAKCVPNSIIMHIGDTIEILSNGKYKSI 290
Query: 276 LHRAVTNNVQPRISMAMFYGPNPETII-GPIHELIDEEHPPKYRNYHFSKFL 326
LHR V N + R S A+F P E II P+ E + E PP++ F++ +
Sbjct: 291 LHRGVVNKEKVRFSWAIFCEPPKEKIILKPLPETVTEAEPPRFPPRTFAQHM 342
>LDOX_PERFR (O04274) Leucoanthocyanidin dioxygenase (EC 1.14.11.19)
(LDOX) (Leucocyanidin oxygenase) (Leucoanthocyanidin
hydroxylase)
Length = 362
Score = 192 bits (487), Expect = 1e-48
Identities = 105/295 (35%), Positives = 175/295 (58%), Gaps = 7/295 (2%)
Query: 38 IPIIDLSYCDGNNPSSLEVIHK-ISKACEEFGFFQIVNHGVPDQVCTKMMKAITNFFELA 96
+P IDL D + + H+ + KA ++G ++NHG+P+++ ++ A FFEL
Sbjct: 53 LPTIDLEEMDSRDEEGRKKCHEELKKAATDWGVMHLINHGIPEELIDRVKAAGKEFFELP 112
Query: 97 PEEREHLSSTDNTKNVRLFNYYLQVDGGEKVKLWSECFAHPWYPIDDI-IQLLPEKIGTQ 155
EE+E ++ NV+ + L + +++ W + F H YP + + P K
Sbjct: 113 VEEKEAYANDQAAGNVQGYGSKLANNASGQLE-WEDYFFHCVYPEHKTDLSIWPTK-PPD 170
Query: 156 YREAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLKKLG--EQPRQRAQSNFYPPCPDPE 213
Y A +EYAK++ +L ++LS++SIGLGLE+ L K++G E + + NFYP CP PE
Sbjct: 171 YIPATSEYAKQLRALATKILSVLSIGLGLEKGRLEKEVGGAEDLIVQMKINFYPKCPQPE 230
Query: 214 LTMGLNEHTDLNALTVLLQSEVSGLQVNKDGKWISIPCIPNAFVINLADQIEVLSNGRYK 273
L +G HTD++ALT +L + V GLQ+ + KW++ C+PN+ ++++ D +E+LSNG+YK
Sbjct: 231 LALGWEAHTDVSALTFILHNMVPGLQLFYEDKWVTAKCVPNSIIMHIGDTLEILSNGKYK 290
Query: 274 SVLHRAVTNNVQPRISMAMFYGPNPETII-GPIHELIDEEHPPKYRNYHFSKFLE 327
S+LHR + N + RIS A+F P E I+ P+ E + E PP++ F++ L+
Sbjct: 291 SILHRGLVNKEKVRISWAVFCEPPKEKIVLQPLPETVSEVEPPRFPPRTFAQHLK 345
>LDOX_ARATH (Q96323) Leucoanthocyanidin dioxygenase (EC 1.14.11.19)
(LDOX) (Leucocyanidin oxygenase) (Leucoanthocyanidin
hydroxylase) (Anthocyanidin synthase) (ANS)
Length = 356
Score = 191 bits (486), Expect = 2e-48
Identities = 110/345 (31%), Positives = 198/345 (56%), Gaps = 14/345 (4%)
Query: 1 MESFQLANESSPLSLTPNFILPEHKRPHLSEVKYLDS-------IPIIDLSYCDGNNPSS 53
+E + +S +S+ +I P+ + +++V +L+ +P IDL + ++
Sbjct: 4 VERVESLAKSGIISIPKEYIRPKEELESINDV-FLEEKKEDGPQVPTIDLKNIESDDEKI 62
Query: 54 LE-VIHKISKACEEFGFFQIVNHGVPDQVCTKMMKAITNFFELAPEEREHLSSTDNTKNV 112
E I ++ KA ++G ++NHG+P + ++ KA FF L+ EE+E ++ T +
Sbjct: 63 RENCIEELKKASLDWGVMHLINHGIPADLMERVKKAGEEFFSLSVEEKEKYANDQATGKI 122
Query: 113 RLFNYYLQVDGGEKVKLWSECFAHPWYPIDDIIQLLPEKIGTQYREAFTEYAKEVGSLVR 172
+ + L + +++ W + F H YP + + K + Y EA +EYAK + L
Sbjct: 123 QGYGSKLANNASGQLE-WEDYFFHLAYPEEKRDLSIWPKTPSDYIEATSEYAKCLRLLAT 181
Query: 173 RLLSLISIGLGLEEDCLLKKLG--EQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVL 230
++ +S+GLGLE D L K++G E+ + + N+YP CP PEL +G+ HTD++ALT +
Sbjct: 182 KVFKALSVGLGLEPDRLEKEVGGLEELLLQMKINYYPKCPQPELALGVEAHTDVSALTFI 241
Query: 231 LQSEVSGLQVNKDGKWISIPCIPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISM 290
L + V GLQ+ +GKW++ C+P++ V+++ D +E+LSNG+YKS+LHR + N + RIS
Sbjct: 242 LHNMVPGLQLFYEGKWVTAKCVPDSIVMHIGDTLEILSNGKYKSILHRGLVNKEKVRISW 301
Query: 291 AMFYGPNPETII-GPIHELIDEEHPPKYRNYHFSKFLE-EFFNQE 333
A+F P + I+ P+ E++ E P K+ F++ +E + F +E
Sbjct: 302 AVFCEPPKDKIVLKPLPEMVSVESPAKFPPRTFAQHIEHKLFGKE 346
>HY6H_HYONI (P24397) Hyoscyamine 6-dioxygenase (EC 1.14.11.11)
(Hyoscyamine 6-beta-hydroxylase)
Length = 344
Score = 191 bits (486), Expect = 2e-48
Identities = 114/333 (34%), Positives = 183/333 (54%), Gaps = 21/333 (6%)
Query: 11 SPLSLTPNFILPEHKRPHLSEVKYLDSIPIIDLSYCDGNNPSSLEVIHKISKACEEFGFF 70
S S++ +FI P KR +V + +PIIDL ++ +I+KAC++FG F
Sbjct: 9 STKSVSESFIAPLQKRAE-KDVPVGNDVPIIDLQQ------HHHLLVQQITKACQDFGLF 61
Query: 71 QIVNHGVPDQVCTKMMKAITNFFELAPEEREHLSSTDNTKNVRL---FNYYLQVDG---- 123
Q++NHG P+++ + M+ FF L EE+E L L V+G
Sbjct: 62 QVINHGFPEELMLETMEVCKEFFALPAEEKEKFKPKGEAAKFELPLEQKAKLYVEGEQLS 121
Query: 124 GEKVKLWSECFAHPWYPID-DIIQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLISIGL 182
E+ W + AH +P+D D++ PEK +YRE +Y+ EV L R+L I GL
Sbjct: 122 NEEFLYWKDTLAHGCHPLDQDLVNSWPEK-PAKYREVVAKYSVEVRKLTMRMLDYICEGL 180
Query: 183 GLEEDCLLKKLGEQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQSEVSGLQ--V 240
GL+ +L + Q +N+YPPCPDP T+G H D N +T LLQ ++ GLQ +
Sbjct: 181 GLKLGYFDNELSQI--QMMLTNYYPPCPDPSSTLGSGGHYDGNLIT-LLQQDLPGLQQLI 237
Query: 241 NKDGKWISIPCIPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNPET 300
KD WI++ IP AFV+NL ++V++N +++ +HR VT+ + R+S+A GP+
Sbjct: 238 VKDATWIAVQPIPTAFVVNLGLTLKVITNEKFEGSIHRVVTDPTRDRVSIATLIGPDYSC 297
Query: 301 IIGPIHELIDEEHPPKYRNYHFSKFLEEFFNQE 333
I P EL+++++PP Y+ Y +S+F + + + +
Sbjct: 298 TIEPAKELLNQDNPPLYKPYSYSEFADIYLSDK 330
>LDOX_MALDO (P51091) Leucoanthocyanidin dioxygenase (EC 1.14.11.19)
(LDOX) (Leucocyanidin oxygenase) (Leucoanthocyanidin
hydroxylase) (Anthocyanidin synthase)
Length = 357
Score = 191 bits (485), Expect = 2e-48
Identities = 102/294 (34%), Positives = 174/294 (58%), Gaps = 5/294 (1%)
Query: 38 IPIIDLSYCDGNNPS-SLEVIHKISKACEEFGFFQIVNHGVPDQVCTKMMKAITNFFELA 96
+P IDL + +N + K+ KA ++G +VNHG+ D++ K+ KA FF+L
Sbjct: 51 VPTIDLKEIESDNEKVRAKCREKLKKAAVDWGVMHLVNHGISDELMDKVRKAGKAFFDLP 110
Query: 97 PEEREHLSSTDNTKNVRLFNYYLQVDGGEKVKLWSECFAHPWYPIDDIIQLLPEKIGTQY 156
E++E ++ + ++ + L + +++ W + F H YP D + + Y
Sbjct: 111 IEQKEKYANDQASGKIQGYGSKLANNASGQLE-WEDYFFHCVYPEDKRDLSIWPQTPADY 169
Query: 157 REAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLKKLG--EQPRQRAQSNFYPPCPDPEL 214
EA EYAK++ L ++L ++S+GLGL+E L K++G E+ + + N+YP CP PEL
Sbjct: 170 IEATAEYAKQLRELATKVLKVLSLGLGLDEGRLEKEVGGLEELLLQMKINYYPKCPQPEL 229
Query: 215 TMGLNEHTDLNALTVLLQSEVSGLQVNKDGKWISIPCIPNAFVINLADQIEVLSNGRYKS 274
+G+ HTD++ALT +L + V GLQ+ +GKW++ C+PN+ V+++ D +E+LSNG+YKS
Sbjct: 230 ALGVEAHTDVSALTFILHNMVPGLQLFYEGKWVTAKCVPNSIVMHIGDTLEILSNGKYKS 289
Query: 275 VLHRAVTNNVQPRISMAMFYGPNPETII-GPIHELIDEEHPPKYRNYHFSKFLE 327
+LHR + N + RIS A+F P E II P+ E + E+ P + F++ ++
Sbjct: 290 ILHRGMVNKEKVRISWAVFCEPPKEKIILKPLPETVSEDEPAMFPPRTFAEHIQ 343
>ACC3_LYCES (P10967) 1-aminocyclopropane-1-carboxylate oxidase
homolog (Protein E8)
Length = 363
Score = 191 bits (485), Expect = 2e-48
Identities = 107/303 (35%), Positives = 162/303 (53%), Gaps = 10/303 (3%)
Query: 19 FILPEHKRPHLSEVKYLDSIPIIDLSYCDGNNPSSLEVIHKISKACEEFGFFQIVNHGVP 78
F+LP R E ++ P+IDL D + E++ K+ A E++GFFQ+VNHG+P
Sbjct: 41 FVLPPKDRAKKCETHFV--FPVIDLQGIDEDPIKHKEIVDKVRDASEKWGFFQVVNHGIP 98
Query: 79 DQVCTKMMKAITNFFELAPEEREHLSSTDNTKNVRLFNYYLQVDGGEKVKLWSECFAHPW 138
V + ++ FFE E ++ + D K V + W +
Sbjct: 99 TSVLDRTLQGTRQFFEQDNEVKKQYYTRDTAKKVVYTSNLDLYKSSVPAASWRDTIFCYM 158
Query: 139 YPIDDIIQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLKKLGEQPR 198
P +Q P G E+ +++K+V L LL L+S GLGL+ LK +
Sbjct: 159 APNPPSLQEFPTPCG----ESLIDFSKDVKKLGFTLLELLSEGLGLDRS-YLKDYMDCFH 213
Query: 199 QRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQSEVSGLQVNKDGKWISIPCIPNAFVI 258
N+YPPCP PELTMG +HTD+ +T+LLQ ++ GLQV W+ +P P + V+
Sbjct: 214 LFCSCNYYPPCPQPELTMGTIQHTDIGFVTILLQDDMGGLQVLHQNHWVDVPPTPGSLVV 273
Query: 259 NLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNP---ETIIGPIHELIDEEHPP 315
N+ D +++LSN +Y SV HRA++NNV R+S+ F+G +P + GPI EL+ E++PP
Sbjct: 274 NIGDFLQLLSNDKYLSVEHRAISNNVGSRMSITCFFGESPYQSSKLYGPITELLSEDNPP 333
Query: 316 KYR 318
KYR
Sbjct: 334 KYR 336
>FLS_SOLTU (Q41452) Flavonol synthase (EC 1.14.11.-) (FLS)
Length = 349
Score = 190 bits (483), Expect = 4e-48
Identities = 107/316 (33%), Positives = 182/316 (56%), Gaps = 10/316 (3%)
Query: 14 SLTPNFILPEHKRPHLSEVK-YLDSIPIIDLSYCDGNNPSSLEVIHKISKACEEFGFFQI 72
++ +I E+++P + ++ + +P+ID+S D + +++ +I +A +E+G FQ+
Sbjct: 32 TIPSEYIRSENEQPAATTLQGVVLEVPVIDISNVDDDEE---KLVKEIVEASKEWGIFQV 88
Query: 73 VNHGVPDQVCTKMMKAITNFFELAP-EEREHLSSTDNTKNVRLFNYYLQVDGGEKVKLWS 131
+NHG+PD+V + K FFE P EE+E ++ +++ + LQ + K K W
Sbjct: 89 INHGIPDEVIENLQKVGKEFFEEVPQEEKELIAKKPGAQSLEGYGTSLQKEIEGK-KGWV 147
Query: 132 ECFAHPWYPIDDIIQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLK 191
+ H +P I K YREA EYAK + + + +S+GLGLE +++
Sbjct: 148 DHLFHKIWPPSAINYRYWPKNPPSYREANEEYAKWLRKVADGIFRSLSLGLGLEGHEMME 207
Query: 192 KLG-EQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQSEVSGLQVNKDGKWISIP 250
G E + N+YPPCP P+L +G+ HTD++ +T+L+ +EV QV KDG W +
Sbjct: 208 AAGSEDIVYMLKINYYPPCPRPDLALGVVAHTDMSYITLLVPNEV---QVFKDGHWYDVN 264
Query: 251 CIPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNPETIIGPIHELID 310
IPNA ++++ DQ+E+LSNG+YKSV HR N + R+S +F P+ E +GPI LI+
Sbjct: 265 YIPNAIIVHIGDQVEILSNGKYKSVYHRTTVNKYKTRMSWPVFLEPSSEHEVGPIPNLIN 324
Query: 311 EEHPPKYRNYHFSKFL 326
E +PPK++ + ++
Sbjct: 325 EANPPKFKTKKYKDYV 340
>FL3H_PETHY (Q07353) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
(Fragment)
Length = 369
Score = 184 bits (467), Expect = 3e-46
Identities = 110/295 (37%), Positives = 166/295 (55%), Gaps = 8/295 (2%)
Query: 14 SLTPNFILPEHKRPHLSEVKYLDSIPIIDLSYCDGNNPSSLEVIHKISKACEEFGFFQIV 73
+L +FI E +RP ++ ++ + IPII L D E+ KI KACE++G FQ+V
Sbjct: 17 TLQTSFIRDEDERPKVAYNQFSNEIPIISLEGIDDETGKRAEICDKIVKACEDWGVFQVV 76
Query: 74 NHGVPDQVCTKMMKAITNFFELAPEEREHLSSTDNTKNVRLFNYYLQVDGGEKVKLWSEC 133
+HGV +V ++M FF L PEE+ + K + + +LQ GE V+ W E
Sbjct: 77 DHGVDAEVISQMTTFAKEFFALPPEEKLRFDMSGGKKGGFIVSSHLQ---GEVVQDWREI 133
Query: 134 FAHPWYPI-DDIIQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLKK 192
+ YP P+K + +Y++++ L +LL ++S +GLE++ L K
Sbjct: 134 VTYFSYPTRARDYSRWPDK-PEGWIAVTQKYSEKLMELACKLLDVLSEAMGLEKEALTKA 192
Query: 193 LGEQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQSEVSGLQVNKD-GK-WISIP 250
+ Q+ NFYP CP+P+LT+GL HTD +T+LLQ +V GLQ KD GK WI++
Sbjct: 193 CVDMD-QKVVVNFYPKCPEPDLTLGLKRHTDPGTITLLLQDQVGGLQATKDNGKTWITVQ 251
Query: 251 CIPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNPETIIGPI 305
+ AFV+NL D LSNGR+K+ H+AV N+ R+S+A F P PE I+ P+
Sbjct: 252 PVEGAFVVNLGDHGHFLSNGRFKNADHQAVVNSNSSRLSIATFQNPAPEAIVYPL 306
>FL3H_CALCH (Q05963) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
Length = 356
Score = 182 bits (462), Expect = 1e-45
Identities = 106/290 (36%), Positives = 163/290 (55%), Gaps = 10/290 (3%)
Query: 19 FILPEHKRPHLSEVKYLDSIPIIDLSYCDGNNPSSLEVIHKISKACEEFGFFQIVNHGVP 78
F+ E +RP + + + IP+I L+ DG + E+ +I KACE++G FQ+V+HGV
Sbjct: 19 FVRDEDERPKVPYNTFSNEIPVISLAGIDGCRRA--EICDEIVKACEDWGIFQVVDHGVD 76
Query: 79 DQVCTKMMKAITNFFELAPEEREHLSSTDNTKNVRLFNYYLQVDGGEKVKLWSECFAHPW 138
++ + M +FF L +E+ T K + + +LQ GE V+ W E +
Sbjct: 77 TKLLSDMTGLARDFFHLPTQEKLRFDMTGGKKGGFIVSSHLQ---GEAVQDWREIVTYFS 133
Query: 139 YPIDDI-IQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLKKLGEQP 197
YPI P+K ++R EY+K + L +LL ++S +GLE++ L K +
Sbjct: 134 YPIKARDYSRWPDK-PNEWRAVTEEYSKVLMGLACKLLEVLSEAMGLEKEALTKACVDMD 192
Query: 198 RQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQSEVSGLQVNKDG--KWISIPCIPNA 255
Q+ N+YP CP P+LT+GL HTD +T+LLQ +V GLQ +DG WI++ + A
Sbjct: 193 -QKVVVNYYPKCPQPDLTLGLKRHTDPGTITLLLQDQVGGLQATRDGGESWITVKPVEGA 251
Query: 256 FVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNPETIIGPI 305
FV+NL D LSNGR+K+ H+AV N+ R+S+A F P PE I+ P+
Sbjct: 252 FVVNLGDHGHYLSNGRFKNADHQAVVNSSTSRLSIATFQNPAPEAIVYPL 301
>FL3H_DIACA (Q05964) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
Length = 365
Score = 181 bits (460), Expect = 2e-45
Identities = 103/293 (35%), Positives = 161/293 (54%), Gaps = 8/293 (2%)
Query: 15 LTPNFILPEHKRPHLSEVKYLDSIPIIDLSYCDGNNPSSLEVIHKISKACEEFGFFQIVN 74
L NF+ E +RP ++ ++ + IP+I L+ DG E+ KI +ACE++G FQ+V+
Sbjct: 18 LNSNFVRDEDERPKVAYNEFSNDIPVISLAGIDGEKRG--EICRKIVEACEDWGIFQVVD 75
Query: 75 HGVPDQVCTKMMKAITNFFELAPEEREHLSSTDNTKNVRLFNYYLQVDGGEKVKLWSECF 134
HGV D + M + FF L EE+ + K + + +LQ GE V+ W E
Sbjct: 76 HGVGDDLIADMTRLAREFFALPAEEKLRFDMSGGKKGGFIVSSHLQ---GEVVQDWREIV 132
Query: 135 AHPWYPIDDIIQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLKKLG 194
+ YP + + + EY+ ++ +L LL ++S +GLE + L K
Sbjct: 133 TYFSYPTNSRDYTRWPDKPEGWIKVTEEYSNKLMTLACTLLGVLSEAMGLELEALTKACV 192
Query: 195 EQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQSEVSGLQVNKDG--KWISIPCI 252
+ Q+ N+YP CP P+LT+GL HTD +T+LLQ +V GLQ +DG WI++ +
Sbjct: 193 DMD-QKIVVNYYPKCPQPDLTLGLKRHTDPGTITLLLQDQVGGLQATRDGGKTWITVQPV 251
Query: 253 PNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNPETIIGPI 305
P AFV+NL D LSNGR+K+ H+AV N+ R+S+A F P+P+ + P+
Sbjct: 252 PGAFVVNLGDHGHFLSNGRFKNADHQAVVNSECSRLSIATFQNPSPDATVYPL 304
>FL3H_MATIN (Q05965) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
(Fragment)
Length = 357
Score = 179 bits (455), Expect = 7e-45
Identities = 105/304 (34%), Positives = 167/304 (54%), Gaps = 11/304 (3%)
Query: 5 QLANESSPLSLTPNFILPEHKRPHLSEVKYLDSIPIIDLSYCDGNNPSSLEVIHKISKAC 64
+LA ES L F+ E +RP ++ ++ D IP+I L+ D + E+ +I +AC
Sbjct: 7 ELAGESK---LNSKFVRDEDERPKVAYNEFSDEIPVISLAGIDDVDGKRGEICREIVEAC 63
Query: 65 EEFGFFQIVNHGVPDQVCTKMMKAITNFFELAPEEREHLSSTDNTKNVRLFNYYLQVDGG 124
E +G FQ+V+HGV + M + +FF L PEE+ + K + + +LQ G
Sbjct: 64 ENWGIFQVVDHGVDTSLVADMTRLARDFFALPPEEKLRFDMSGGKKGGFIVSSHLQ---G 120
Query: 125 EKVKLWSECFAHPWYPI-DDIIQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLISIGLG 183
E V+ W E + YP+ + P+K + + EY++++ L +LL ++S +G
Sbjct: 121 EAVQDWREIVTYFSYPVRNRDYSRWPDK-PQGWAKVTEEYSEKLMGLACKLLEVLSEAMG 179
Query: 184 LEEDCLLKKLGEQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQSEVSGLQVNKD 243
LE++ L + Q+ N+YP CP P+LT+GL HTD +T+LLQ +V GLQ +D
Sbjct: 180 LEKESLTNACVDMD-QKIVVNYYPKCPQPDLTLGLKRHTDPGTITLLLQDQVGGLQATRD 238
Query: 244 --GKWISIPCIPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNPETI 301
WI++ + AFV+NL D LSNGR+K+ H+AV N+ R+S+A F P PE
Sbjct: 239 DGNTWITVQPVEGAFVVNLGDHGHFLSNGRFKNADHQAVVNSNSSRLSIATFQNPAPEAT 298
Query: 302 IGPI 305
+ P+
Sbjct: 299 VYPL 302
>FL3H_MALDO (Q06942) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
Length = 364
Score = 177 bits (449), Expect = 3e-44
Identities = 108/295 (36%), Positives = 163/295 (54%), Gaps = 8/295 (2%)
Query: 14 SLTPNFILPEHKRPHLSEVKYLDSIPIIDLSYCDGNNPSSLEVIHKISKACEEFGFFQIV 73
+L F+ E +RP ++ + + IPII L+ D E+ KI ACE++G FQIV
Sbjct: 15 TLQQKFVRDEDERPKVAYNDFSNEIPIISLAGIDEVEGRRGEICKKIVAACEDWGIFQIV 74
Query: 74 NHGVPDQVCTKMMKAITNFFELAPEEREHLSSTDNTKNVRLFNYYLQVDGGEKVKLWSEC 133
+HGV ++ ++M FF L EE+ + K + + +LQ GE V+ W E
Sbjct: 75 DHGVDAELISEMTGLAREFFALPSEEKLRFDMSGGKKGGFIVSSHLQ---GEAVQDWREI 131
Query: 134 FAHPWYPI-DDIIQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLKK 192
+ YPI P+K +RE +Y+ E+ L +LL ++S +GL+ + L K
Sbjct: 132 VTYFSYPIRHRDYSRWPDK-PEAWREVTKKYSDELMGLACKLLGVLSEAMGLDTEALTKA 190
Query: 193 LGEQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQSEVSGLQVNK-DGK-WISIP 250
+ Q+ NFYP CP P+LT+GL HTD +T+LLQ +V GLQ + DGK WI++
Sbjct: 191 CVDMD-QKVVVNFYPKCPQPDLTLGLKRHTDPGTITLLLQDQVGGLQATRDDGKTWITVQ 249
Query: 251 CIPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNPETIIGPI 305
+ AFV+NL D +LSNGR+K+ H+AV N+ R+S+A F P E I+ P+
Sbjct: 250 PVEGAFVVNLGDHGHLLSNGRFKNADHQAVVNSNSSRLSIATFQNPAQEAIVYPL 304
>LDOX_VITVI (P51093) Leucoanthocyanidin dioxygenase (EC 1.14.11.19)
(LDOX) (Leucocyanidin oxygenase) (Leucoanthocyanidin
hydroxylase)
Length = 362
Score = 177 bits (448), Expect = 5e-44
Identities = 97/276 (35%), Positives = 158/276 (57%), Gaps = 8/276 (2%)
Query: 38 IPIIDLSYCDGNNPSSLEVIHK-----ISKACEEFGFFQIVNHGVPDQVCTKMMKAITNF 92
+P IDL + + I + + KA E+G +VNHG+ D + ++ A F
Sbjct: 49 VPTIDLKDIESEDEVVRREIRERCREELKKAAMEWGVMHLVNHGISDDLINRVKVAGETF 108
Query: 93 FELAPEEREHLSSTDNTKNVRLFNYYLQVDGGEKVKLWSECFAHPWYPIDDIIQLLPEKI 152
F L EE+E ++ + + + L + +++ W + F H +P D + K
Sbjct: 109 FNLPMEEKEKYANDQASGKIAGYGSKLANNASGQLE-WEDYFFHLIFPEDKRDMTIWPKT 167
Query: 153 GTQYREAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLKKLG--EQPRQRAQSNFYPPCP 210
+ Y A EY+ ++ SL ++LS++S+GLGLEE L K++G E+ + + N+YP CP
Sbjct: 168 PSDYVPATCEYSVKLRSLATKILSVLSLGLGLEEGRLEKEVGGMEELLLQKKINYYPKCP 227
Query: 211 DPELTMGLNEHTDLNALTVLLQSEVSGLQVNKDGKWISIPCIPNAFVINLADQIEVLSNG 270
PEL +G+ HTD++ALT +L + V GLQ+ +GKW++ C+PN+ ++++ D IE+LSNG
Sbjct: 228 QPELALGVEAHTDVSALTFILHNMVPGLQLFYEGKWVTAKCVPNSIIMHIGDTIEILSNG 287
Query: 271 RYKSVLHRAVTNNVQPRISMAMFYGPNPETIIGPIH 306
+YKS+LHR + N + RIS A+F P E II H
Sbjct: 288 KYKSILHRGLVNKEKVRISWAVFCEPPKEKIILKAH 323
>FL3H_ARATH (Q9S818) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Naringenin 3-dioxygenase) (Flavanone
3-hydroxylase) (FH3) (TRANSPARENT TESTA 6 protein)
Length = 358
Score = 177 bits (448), Expect = 5e-44
Identities = 106/304 (34%), Positives = 168/304 (54%), Gaps = 11/304 (3%)
Query: 5 QLANESSPLSLTPNFILPEHKRPHLSEVKYLDSIPIIDLSYCDGNNPSSLEVIHKISKAC 64
+LA ES L F+ E +RP ++ + D IP+I L+ D + E+ +I +AC
Sbjct: 8 ELAGESK---LNSKFVRDEDERPKVAYNVFSDEIPVISLAGIDDVDGKRGEICRQIVEAC 64
Query: 65 EEFGFFQIVNHGVPDQVCTKMMKAITNFFELAPEEREHLSSTDNTKNVRLFNYYLQVDGG 124
E +G FQ+V+HGV + M + +FF L PE++ + K + + +LQ G
Sbjct: 65 ENWGIFQVVDHGVDTNLVADMTRLARDFFALPPEDKLRFDMSGGKKGGFIVSSHLQ---G 121
Query: 125 EKVKLWSECFAHPWYPI-DDIIQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLISIGLG 183
E V+ W E + YP+ + P+K + + EY++ + SL +LL ++S +G
Sbjct: 122 EAVQDWREIVTYFSYPVRNRDYSRWPDK-PEGWVKVTEEYSERLMSLACKLLEVLSEAMG 180
Query: 184 LEEDCLLKKLGEQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQSEVSGLQVNKD 243
LE++ L + Q+ N+YP CP P+LT+GL HTD +T+LLQ +V GLQ +D
Sbjct: 181 LEKESLTNACVDMD-QKIVVNYYPKCPQPDLTLGLKRHTDPGTITLLLQDQVGGLQATRD 239
Query: 244 -GK-WISIPCIPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNPETI 301
GK WI++ + AFV+NL D LSNGR+K+ H+AV N+ R+S+A F P P+
Sbjct: 240 NGKTWITVQPVEGAFVVNLGDHGHFLSNGRFKNADHQAVVNSNSSRLSIATFQNPAPDAT 299
Query: 302 IGPI 305
+ P+
Sbjct: 300 VYPL 303
>FL3H_VITVI (P41090) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
Length = 364
Score = 176 bits (447), Expect = 6e-44
Identities = 103/296 (34%), Positives = 163/296 (54%), Gaps = 9/296 (3%)
Query: 14 SLTPNFILPEHKRPHLSEVKYLDSIPIIDLSYCDGNNPSSL-EVIHKISKACEEFGFFQI 72
+L +F+ E +RP ++ + + IP+I L+ + + E+ KI +ACE++G FQ+
Sbjct: 14 TLQSSFVRDEDERPKVAYNDFSNEIPVISLTKESMKLAAVVDEICRKIVEACEDWGIFQV 73
Query: 73 VNHGVPDQVCTKMMKAITNFFELAPEEREHLSSTDNTKNVRLFNYYLQVDGGEKVKLWSE 132
VNHGV + ++M + FF L PEE + K + + +LQ GE V+ W E
Sbjct: 74 VNHGVDSNLISEMTRLAREFFALPPEENVRFDMSGGKKGGFIVSSHLQ---GEAVQDWRE 130
Query: 133 CFAHPWYPIDDI-IQLLPEKIGTQYREAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLK 191
+ YP+ P+K +R EY++++ L +LL ++S + L++D L
Sbjct: 131 IVTYFSYPLRTRDYSRWPDK-PEGWRSVTQEYSEKLMGLACKLLEVLSEAMDLDKDALTN 189
Query: 192 KLGEQPRQRAQSNFYPPCPDPELTMGLNEHTDLNALTVLLQSEVSGLQVNKDG--KWISI 249
+ Q+ NFYP CP P+LT+GL HTD +T+LLQ +V GLQ +DG WI++
Sbjct: 190 ACVDMD-QKVVVNFYPQCPQPDLTLGLKRHTDPGTITLLLQDQVGGLQATRDGGKTWITV 248
Query: 250 PCIPNAFVINLADQIEVLSNGRYKSVLHRAVTNNVQPRISMAMFYGPNPETIIGPI 305
+ AFV+NL D LSNGR+K+ H+AV N+ R+S+A F P PE + P+
Sbjct: 249 QPVEGAFVVNLGDHGHYLSNGRFKNADHQAVVNSNHSRLSIATFQNPAPEATVYPL 304
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.138 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 44,134,653
Number of Sequences: 164201
Number of extensions: 1913776
Number of successful extensions: 4598
Number of sequences better than 10.0: 82
Number of HSP's better than 10.0 without gapping: 67
Number of HSP's successfully gapped in prelim test: 15
Number of HSP's that attempted gapping in prelim test: 4366
Number of HSP's gapped (non-prelim): 86
length of query: 347
length of database: 59,974,054
effective HSP length: 111
effective length of query: 236
effective length of database: 41,747,743
effective search space: 9852467348
effective search space used: 9852467348
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 66 (30.0 bits)
Medicago: description of AC119414.6


