

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119412.5 + phase: 0
(221 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
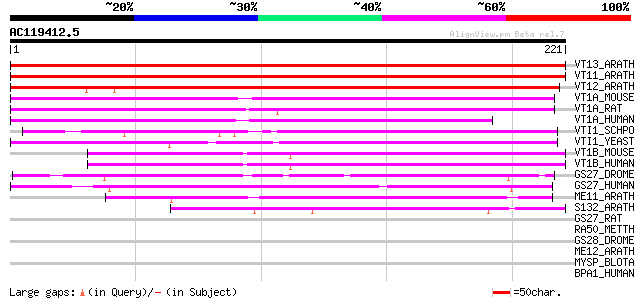
Score E
Sequences producing significant alignments: (bits) Value
VT13_ARATH (Q9LVP9) Vesicle transport v-SNARE 13 (AtVTI13) (Vesi... 339 3e-93
VT11_ARATH (Q9SEL6) Vesicle transport v-SNARE 11 (AtVTI11) (Vesi... 338 6e-93
VT12_ARATH (Q9SEL5) Vesicle transport v-SNARE 12 (AtVTI12) (Vesi... 286 2e-77
VT1A_MOUSE (O89116) Vesicle transport through interaction with t... 133 4e-31
VT1A_RAT (Q9JI51) Vesicle transport through interaction with t-S... 124 2e-28
VT1A_HUMAN (Q96AJ9) Vesicle transport through interaction with t... 120 3e-27
VTI1_SCHPO (P78768) Vesicle transport v-SNARE protein vti1 103 3e-22
VTI1_YEAST (Q04338) Vesicle transport v-SNARE protein VTI1 99 6e-21
VT1B_MOUSE (O88384) Vesicle transport through interaction with t... 94 3e-19
VT1B_HUMAN (Q9UEU0) Vesicle transport through interaction with t... 92 7e-19
GS27_DROME (Q9VRL2) Probable 27 kDa Golgi SNARE protein (Golgi S... 50 3e-06
GS27_HUMAN (O14653) 27 kDa Golgi SNARE protein (Golgi SNAP recep... 45 1e-04
ME11_ARATH (Q9SJL6) Membrin 11 (AtMEMB11) (Golgi SNAP receptor c... 44 4e-04
S132_ARATH (Q8VZU2) Syntaxin 132 (AtSYP132) 43 5e-04
GS27_RAT (O35165) 27 kDa Golgi SNARE protein (Golgi SNAP recepto... 42 0.001
RA50_METTH (O26640) DNA double-strand break repair rad50 ATPase 42 0.001
GS28_DROME (Q9VE50) Probable 28 kDa Golgi SNARE protein (Golgi S... 41 0.002
ME12_ARATH (Q9FK28) Membrin 12 (AtMEMB12) (Golgi SNAP receptor c... 40 0.004
MYSP_BLOTA (Q8MUF6) Paramyosin (Allergen Blo t 11) 39 0.007
BPA1_HUMAN (Q03001) Bullous pemphigoid antigen 1 isoforms 1/2/3/... 39 0.007
>VT13_ARATH (Q9LVP9) Vesicle transport v-SNARE 13 (AtVTI13) (Vesicle
transport v-SNARE protein VTI13) (Vesicle soluble NSF
attachment protein receptor 13)
Length = 221
Score = 339 bits (869), Expect = 3e-93
Identities = 166/221 (75%), Positives = 196/221 (88%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS FE YERQYCE+SANLSKKCT+A AL+GEQKKQ +SE+KSG++EAEAL++KMDLEAR
Sbjct: 1 MSQGFERYERQYCEISANLSKKCTSAIALDGEQKKQNLSEIKSGVEEAEALVKKMDLEAR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
++ PN+K LL KLREYKSDLNN K+EVK+I SGNLN +ARDELLE+GMADT+TASADQR
Sbjct: 61 NLPPNVKSSLLVKLREYKSDLNNFKTEVKRITSGNLNATARDELLEAGMADTLTASADQR 120
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
+RLM ST+ L +T +R+KDSRRT+LETEELGVSILQDLH QRQSLL AH TLHGVDDN+G
Sbjct: 121 SRLMMSTDHLGRTTDRIKDSRRTILETEELGVSILQDLHGQRQSLLRAHETLHGVDDNVG 180
Query: 181 KSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKLVK 221
KSKKI+T M+RRMN+NKW IG I+ VLV+AII ILYFKL +
Sbjct: 181 KSKKILTTMTRRMNRNKWTIGAIITVLVLAIIFILYFKLTR 221
>VT11_ARATH (Q9SEL6) Vesicle transport v-SNARE 11 (AtVTI11) (Vesicle
transport v-SNARE protein VTI1a) (Vesicle soluble NSF
attachment protein receptor VTI1a) (AtVTI1a)
Length = 221
Score = 338 bits (867), Expect = 6e-93
Identities = 166/221 (75%), Positives = 195/221 (88%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS+VF+GYERQYCELSA+LSKKC++A +L+GEQKKQK+SE+KSG++ AE LIRKMDLEAR
Sbjct: 1 MSDVFDGYERQYCELSASLSKKCSSAISLDGEQKKQKLSEIKSGLENAEVLIRKMDLEAR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
++ PN+K LL KLRE+KSDLNN K+EVK+I SG LN +ARDELLE+GMADT TASADQR
Sbjct: 61 TLPPNLKSSLLVKLREFKSDLNNFKTEVKRITSGQLNAAARDELLEAGMADTKTASADQR 120
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
RLM STERL +T +RVKDSRRTM+ETEE+GVSILQDLH QRQSLL AH TLHGVDDNIG
Sbjct: 121 ARLMMSTERLGRTTDRVKDSRRTMMETEEIGVSILQDLHGQRQSLLRAHETLHGVDDNIG 180
Query: 181 KSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKLVK 221
KSKKI+T+M+RRMNKNKW IG I++ L+ AI ILYFKL K
Sbjct: 181 KSKKILTDMTRRMNKNKWTIGAIIIALIAAIFIILYFKLTK 221
>VT12_ARATH (Q9SEL5) Vesicle transport v-SNARE 12 (AtVTI12) (Vesicle
transport v-SNARE protein VTI1b) (Vesicle soluble NSF
attachment protein receptor VTI1b) (AtVTI1b)
Length = 223
Score = 286 bits (733), Expect = 2e-77
Identities = 144/221 (65%), Positives = 182/221 (82%), Gaps = 2/221 (0%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATAL-NGEQKKQKVSE-VKSGIDEAEALIRKMDLE 58
MS+VFEGYERQYCELS NLS+KC +A+ L NGE+KK K++E +KSGIDEA+ LIRKMDLE
Sbjct: 1 MSDVFEGYERQYCELSTNLSRKCHSASVLSNGEEKKGKIAEQIKSGIDEADVLIRKMDLE 60
Query: 59 ARSMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASAD 118
ARS+QP+ K V L+KLREYKSDLN LK E K++ S + PS+R+EL+ESGMAD SAD
Sbjct: 61 ARSLQPSAKAVCLSKLREYKSDLNQLKKEFKRVSSADAKPSSREELMESGMADLHAVSAD 120
Query: 119 QRTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDN 178
QR RL S ERL+++ +R+++SRR MLETEE+G+SI+QDL QRQ+LLHAHN LHGVDD
Sbjct: 121 QRGRLAMSVERLDQSSDRIRESRRLMLETEEVGISIVQDLSQQRQTLLHAHNKLHGVDDA 180
Query: 179 IGKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKL 219
I KSKK++T MSRRM +NKWII +++ LV+AII I+ +KL
Sbjct: 181 IDKSKKVLTAMSRRMTRNKWIITSVIVALVLAIILIISYKL 221
>VT1A_MOUSE (O89116) Vesicle transport through interaction with
t-SNAREs homolog 1A (Vesicle transport v-SNARE protein
Vti1-like 2) (Vti1-rp2)
Length = 217
Score = 133 bits (334), Expect = 4e-31
Identities = 67/217 (30%), Positives = 128/217 (58%), Gaps = 5/217 (2%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS+ FEGYE+ + L+A ++ K L ++KKQ V+ V+ ++EA L+ +MDLE R
Sbjct: 1 MSSDFEGYEQDFAVLTAEITSKIARVPRLPPDEKKQMVANVEKQLEEARELLEQMDLEVR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
+ P +G+ ++R YK ++ L+++ K+ + DE+ + D +S +QR
Sbjct: 61 EIPPQSRGMYSNRMRSYKQEMGKLETDFKRS-----RIAYSDEVRNELLGDAGNSSENQR 115
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
L+ +TERL ++ R++ + +ETE++G +L++L R+ + A + L D N+G
Sbjct: 116 AHLLDNTERLERSSRRLEAGYQIAVETEQIGQEMLENLSHDREKIQRARDRLRDADANLG 175
Query: 181 KSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYF 217
KS +I+T M RR+ +N+ ++ + +++V+AI+ + F
Sbjct: 176 KSSRILTGMLRRIIQNRILLVILGIIVVIAILTAIAF 212
>VT1A_RAT (Q9JI51) Vesicle transport through interaction with
t-SNAREs homolog 1A (Vesicle transport v-SNARE protein
Vti1-like 2) (Vti1-rp2)
Length = 224
Score = 124 bits (310), Expect = 2e-28
Identities = 65/220 (29%), Positives = 127/220 (57%), Gaps = 4/220 (1%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS FEGYE+ + L+A ++ K + L ++KKQ V+ V+ ++EA L+ +MDLE R
Sbjct: 1 MSADFEGYEQDFAVLTAEITSKISRVPRLPPDEKKQMVANVEKQLEEARELLEQMDLEVR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELL---ESGMADTMTASA 117
+ P +G+ ++R YK ++ L+++ K+ + R+ELL + + +
Sbjct: 61 EIPPQSRGMYSNRMRSYKQEMGKLETDFKRSRIA-YSDEVRNELLGDAGNSSENQLIKLR 119
Query: 118 DQRTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDD 177
++R L+ +TERL ++ R++ + +ETE++G +L++L R+ + A L D
Sbjct: 120 EERAHLLDNTERLERSSRRLEAGYQIAVETEQIGQEMLENLSHDRERIQRARERLRETDA 179
Query: 178 NIGKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYF 217
N+GKS +I+T M RR+ +N+ ++ + +++V+ I+ + F
Sbjct: 180 NLGKSSRILTGMLRRIIQNRILLVILGIIVVITILTAITF 219
>VT1A_HUMAN (Q96AJ9) Vesicle transport through interaction with
t-SNAREs homolog 1A (Vesicle transport v-SNARE protein
Vti1-like 2) (Vti1-rp2)
Length = 203
Score = 120 bits (301), Expect = 3e-27
Identities = 62/192 (32%), Positives = 111/192 (57%), Gaps = 5/192 (2%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS+ FEGYE+ + L+A ++ K L ++KKQ V+ V+ ++EA+ L+ +MDLE R
Sbjct: 1 MSSDFEGYEQDFAVLTAEITSKIARVPRLPPDEKKQMVANVEKQLEEAKELLEQMDLEVR 60
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
+ P +G+ ++R YK ++ L+++ K+ + DE+ + D +S +QR
Sbjct: 61 EIPPQSRGMYSNRMRSYKQEMGKLETDFKR-----SRIAYSDEVRNELLGDDGNSSENQR 115
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
L+ +TERL ++ R++ + +ETE++G +L++L R+ + A L D N+G
Sbjct: 116 AHLLDNTERLERSSRRLEAGYQIAVETEQIGQEMLENLSHDREKIQRARERLRETDANLG 175
Query: 181 KSKKIMTNMSRR 192
KS +I+T M RR
Sbjct: 176 KSSRILTGMLRR 187
>VTI1_SCHPO (P78768) Vesicle transport v-SNARE protein vti1
Length = 214
Score = 103 bits (257), Expect = 3e-22
Identities = 71/224 (31%), Positives = 122/224 (53%), Gaps = 24/224 (10%)
Query: 6 EGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSG---IDEAEALIRKMDLEARSM 62
E YE++Y L A++ +K LN K + S ++S ++E + +I +M++E +
Sbjct: 2 ETYEQEYRLLRADIEEK------LNDLSKSGENSVIQSCQRLLNEIDEVIGQMEIEITGI 55
Query: 63 QPNIKGVLLAKLREYKSDLN----NLKSEV----KKIVSGNLNPSARDELLESGMADTMT 114
+ +G++ ++R Y+S L +LK E+ +K + GN RDE SG
Sbjct: 56 PTSERGLVNGRIRSYRSTLEEWRRHLKEEIGKSDRKALFGN-----RDET--SGDYIASD 108
Query: 115 ASADQRTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHG 174
DQRTRL+ T RL ++ +R+ +S+R ETE +G SIL+DLH QR L H+ L
Sbjct: 109 QDYDQRTRLLQGTNRLEQSSQRLLESQRIANETEGIGASILRDLHGQRNQLEHSLEMLGD 168
Query: 175 VDDNIGKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFK 218
++ +S + + M+RR+ N++ I+ +LV+ I+ +LY K
Sbjct: 169 TSGHLDRSLRTLKTMARRLAMNRFFTTAIIAILVILILLVLYSK 212
>VTI1_YEAST (Q04338) Vesicle transport v-SNARE protein VTI1
Length = 217
Score = 99.4 bits (246), Expect = 6e-21
Identities = 66/220 (30%), Positives = 113/220 (51%), Gaps = 7/220 (3%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MS++ YE + A + Q+ + V+ DE L+ +MD+E
Sbjct: 1 MSSLLISYESDFKTTLEQAKASLAEAPSQPLSQRNTTLKHVEQQQDELFDLLDQMDVEVN 60
Query: 61 SM--QPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASAD 118
+ + + AKLRE+K ++S++K+ + ++ RD L G + D
Sbjct: 61 NSIGDASERATYKAKLREWKK---TIQSDIKRPLQSLVDSGDRDRLF--GDLNASNIDDD 115
Query: 119 QRTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDN 178
QR +L+++ L K+G+R+KD+ R ETE +G I+ DL SQR++L +A TL D
Sbjct: 116 QRQQLLSNHAILQKSGDRLKDASRIANETEGIGSQIMMDLRSQRETLENARQTLFQADSY 175
Query: 179 IGKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFK 218
+ KS K + M+RR+ NK+I I+ VL++ I+ +L+ K
Sbjct: 176 VDKSIKTLKTMTRRLVANKFISYAIIAVLILLILLVLFSK 215
>VT1B_MOUSE (O88384) Vesicle transport through interaction with
t-SNAREs homolog 1B (Vesicle transport v-SNARE protein
Vti1-like 1) (Vti1-rp1)
Length = 232
Score = 93.6 bits (231), Expect = 3e-19
Identities = 53/193 (27%), Positives = 99/193 (50%), Gaps = 4/193 (2%)
Query: 32 EQKKQKVSEVKSGIDEAEALIRKMDLEARSMQPNIKGVLLAKLREYKSDLNNLKSEVKKI 91
E+KK+ V + EA + +M+ E R + +++KLR Y+ DL L EV+
Sbjct: 39 EEKKKLVRDFDENQQEANETLAEMEEELRYAPLTFRNPMMSKLRNYRKDLAKLHREVRST 98
Query: 92 VSGNLNPSARDELLESGMA---DTMTASADQRTRLMTSTERLNKTGERVKDSRRTMLETE 148
P R +L + + QR L+ TE LN+ + ++ S R ET+
Sbjct: 99 PL-TAAPGGRGDLKYGTYTLENEHLNRLQSQRALLLQGTESLNRATQSIERSHRIATETD 157
Query: 149 ELGVSILQDLHSQRQSLLHAHNTLHGVDDNIGKSKKIMTNMSRRMNKNKWIIGCIVLVLV 208
++G I+++L QR L + L ++N+ KS+KI+ +MSR++ NK ++ I+L+ +
Sbjct: 158 QIGTEIIEELGEQRDQLERTKSRLVNTNENLSKSRKILRSMSRKVITNKLLLSVIILLEL 217
Query: 209 VAIIAILYFKLVK 221
++ ++Y+K +
Sbjct: 218 AILVGLVYYKFFR 230
>VT1B_HUMAN (Q9UEU0) Vesicle transport through interaction with
t-SNAREs homolog 1B (Vesicle transport v-SNARE protein
Vti1-like 1) (Vti1-rp1)
Length = 232
Score = 92.4 bits (228), Expect = 7e-19
Identities = 53/193 (27%), Positives = 99/193 (50%), Gaps = 4/193 (2%)
Query: 32 EQKKQKVSEVKSGIDEAEALIRKMDLEARSMQPNIKGVLLAKLREYKSDLNNLKSEVKKI 91
E+KK+ + + EA + +M+ E R + + +++KLR Y+ DL L EV+
Sbjct: 39 EEKKKLIRDFDEKQQEANETLAEMEEELRYAPLSFRNPMMSKLRNYRKDLAKLHREVRST 98
Query: 92 VSGNLNPSARDELLESGMA---DTMTASADQRTRLMTSTERLNKTGERVKDSRRTMLETE 148
P R ++ A + M QR L+ TE LN+ + ++ S R ET+
Sbjct: 99 PL-TATPGGRGDMKYGIYAVENEHMNRLQSQRAMLLQGTESLNRATQSIERSHRIATETD 157
Query: 149 ELGVSILQDLHSQRQSLLHAHNTLHGVDDNIGKSKKIMTNMSRRMNKNKWIIGCIVLVLV 208
++G I+++L QR L + L +N+ KS+KI+ +MSR++ NK ++ I+L+ +
Sbjct: 158 QIGSEIIEELGEQRDQLERTKSRLVNTSENLSKSRKILRSMSRKVTTNKLLLSIIILLEL 217
Query: 209 VAIIAILYFKLVK 221
+ ++Y+K +
Sbjct: 218 AILGGLVYYKFFR 230
>GS27_DROME (Q9VRL2) Probable 27 kDa Golgi SNARE protein (Golgi SNAP
receptor complex member 2)
Length = 216
Score = 50.4 bits (119), Expect = 3e-06
Identities = 53/219 (24%), Positives = 102/219 (46%), Gaps = 17/219 (7%)
Query: 2 SNVFEGYERQYCELSANLSKKCTAATALNGEQKKQ-KVSEVKSGIDEAEALIRKMDLEAR 60
+NV + ER + LS + +A +L+ E Q K+++ + D + L+ K+ R
Sbjct: 9 NNVVKDIERDFQRLS-----QLSAQESLDVENGIQLKITQANANCDRLDVLLYKVPPSQR 63
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQR 120
L LR ++ L + ++ + S R++LL T ++ +
Sbjct: 64 QSSKLRVDQLKYDLRHLQTSLQTARERRQRRMQ---EISEREQLLNHRF--TANSAQPEE 118
Query: 121 TRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIG 180
TRL E + T ++ ++ R + + G IL+ L SQR +L AH + + +G
Sbjct: 119 TRLQLDYELQHHT--QLGNAHRGVDDMIASGSGILESLISQRMTLGGAHKRIQAIGSTLG 176
Query: 181 KSKKIMTNMSRRMNKNK--WIIGCIVLVLVVAIIAILYF 217
S M + RR+ +++ +I G +V +L++A+ I+YF
Sbjct: 177 LSNHTMKLIERRLVEDRRIFIGGVVVTLLIIAL--IIYF 213
>GS27_HUMAN (O14653) 27 kDa Golgi SNARE protein (Golgi SNAP receptor
complex member 2) (Membrin)
Length = 212
Score = 45.1 bits (105), Expect = 1e-04
Identities = 42/221 (19%), Positives = 102/221 (46%), Gaps = 16/221 (7%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKV----SEVKSGIDEAEALIRKMD 56
M +F+ +Q E+ + + + TA KQ V +E+++ ID+ + + +++
Sbjct: 1 MDPLFQQTHKQVHEIQSCMGRLETA--------DKQSVHIVENEIQASIDQIFSRLERLE 52
Query: 57 LEARSMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTAS 116
+ + PN + ++ + K D+ +L++ ++ ++ E ++ T T +
Sbjct: 53 ILSSKEPPNKRQNARLRVDQLKYDVQHLQTALRNFQHRRHAREQQERQREELLSRTFTTN 112
Query: 117 ADQRTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVD 176
T M + + N + ++V + ++ G +IL L +QR +L + +
Sbjct: 113 DSDTTIPMDESLQFNSSLQKVHNGMDDLILD---GHNILDGLRTQRLTLKGTQKKILDIA 169
Query: 177 DNIGKSKKIMTNMSRRMNKNKW-IIGCIVLVLVVAIIAILY 216
+ +G S +M + +R ++K+ +IG ++L VV + + Y
Sbjct: 170 NMLGLSNTVMRLIEKRAFQDKYFMIGGMLLTCVVMFLVVQY 210
>ME11_ARATH (Q9SJL6) Membrin 11 (AtMEMB11) (Golgi SNAP receptor
complex member 2-1) (27 kDa Golgi SNARE protein)
Length = 225
Score = 43.5 bits (101), Expect = 4e-04
Identities = 43/179 (24%), Positives = 75/179 (41%), Gaps = 9/179 (5%)
Query: 39 SEVKSGIDEAEALIRKMDLEARSMQ-PNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLN 97
S VK I E +L MD RS+ + + + K + + L ++K +S N
Sbjct: 48 SSVKRDITEVRSLCSNMDTLWRSIPVKSQRDLWRRKTEQVGEEAEYLNLSLEKYMSRN-- 105
Query: 98 PSARDELLESGMADTMTASADQRTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQD 157
R L AD + ++ + ++ + + VK+S+R + E+ GV+IL
Sbjct: 106 --QRKMLEAKERADLLGRASGEGAHILQIFDEEAQAMSSVKNSKRMLEESFSSGVAILSK 163
Query: 158 LHSQRQSLLHAHNTLHGVDDNIGKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILY 216
QR L A V + +G S ++ + RR + W I ++A + ILY
Sbjct: 164 YAEQRDRLKSAQRKALDVLNTVGLSNSVLRLIERRNRVDTW----IKYAGMIATLVILY 218
>S132_ARATH (Q8VZU2) Syntaxin 132 (AtSYP132)
Length = 304
Score = 43.1 bits (100), Expect = 5e-04
Identities = 30/169 (17%), Positives = 83/169 (48%), Gaps = 14/169 (8%)
Query: 65 NIKGVLLAKLREYKSDLNNLKSEVKKIVSGNL--------NPSARDELLESGMADTMTAS 116
++K L K+ E++ N++ E + +V + + DEL+E+G ++ +
Sbjct: 131 SLKKKLKDKMAEFQVLRENIQQEYRDVVDRRVYTVTGERADEDTIDELIETGNSEQIFQK 190
Query: 117 ADQ---RTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLH 173
A Q R ++M + + + + V+D + +L+ +++ + + + +Q + L + + +
Sbjct: 191 AIQEQGRGQVMDTLAEIQERHDAVRDLEKKLLDLQQIFLDMAVLVDAQGEMLDNIESQVS 250
Query: 174 GVDDNIGKSKKIMTNM-SRRMNKNKWIIGCIVLVLVVAIIAILYFKLVK 221
D++ + S + N KW+ CI +++++ ++A++ ++K
Sbjct: 251 SAVDHVQSGNTALQRAKSLQKNSRKWM--CIAIIILLIVVAVIVVGVLK 297
>GS27_RAT (O35165) 27 kDa Golgi SNARE protein (Golgi SNAP receptor
complex member 2) (Membrin)
Length = 212
Score = 42.0 bits (97), Expect = 0.001
Identities = 39/221 (17%), Positives = 101/221 (45%), Gaps = 16/221 (7%)
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKV----SEVKSGIDEAEALIRKMD 56
M +++ +Q E+ +++ + TA KQ V +E+++ ID+ + + +++
Sbjct: 1 MEPLYQQTHKQVHEIQSHMGRLETA--------DKQSVHLVENEIQASIDQIFSHLERLE 52
Query: 57 LEARSMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTAS 116
+ + PN + ++ + K D+ +L++ ++ ++ + ++ T T +
Sbjct: 53 ILSSKEPPNRRQNAKLRVDQLKYDVQHLQTALRNFQHRRQAKEQQERQRDELLSRTFTTN 112
Query: 117 ADQRTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVD 176
T M + + N + + + ++ G SIL+ L +QR +L + +
Sbjct: 113 DYDTTIPMDESLQFNSSLQNIHHGMDDLIGG---GHSILEGLRAQRLTLKGTQKKILDIA 169
Query: 177 DNIGKSKKIMTNMSRRMNKNKW-IIGCIVLVLVVAIIAILY 216
+ +G S +M + +R ++K+ +IG ++L V + + Y
Sbjct: 170 NMLGLSNTVMRLIEKRALEDKYFMIGGMLLTCAVMFLVVQY 210
>RA50_METTH (O26640) DNA double-strand break repair rad50 ATPase
Length = 837
Score = 41.6 bits (96), Expect = 0.001
Identities = 34/147 (23%), Positives = 67/147 (45%), Gaps = 8/147 (5%)
Query: 46 DEAEALIRKMDLEARSMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELL 105
DE EA IRK+++E +Q + G+ K+ +Y+S + V +NP ++ L
Sbjct: 387 DEVEAGIRKLEVEVSDLQQELGGI-HGKIEDYES---IGRRGVCPTCDQEVNPDYINDKL 442
Query: 106 ESG---MADTMTASADQRTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQR 162
S + AD R +M E + + E + L E+L I ++ H +R
Sbjct: 443 SSARERKGSVESRIADLRNEIMNKNELMKRIAEYKRSREELRLLREDLEKRI-KECHRRR 501
Query: 163 QSLLHAHNTLHGVDDNIGKSKKIMTNM 189
+SL + + +++ +S+K++ +
Sbjct: 502 ESLDNIRERIRQIEEGKAESEKLIEEL 528
Score = 30.0 bits (66), Expect = 4.5
Identities = 31/132 (23%), Positives = 61/132 (45%), Gaps = 15/132 (11%)
Query: 29 LNGEQKKQKVSEVKSGIDEAEALIRKMDLEARSMQPNIKGVLLAKLREYKSDLNNLKSEV 88
+N + ++++E K +E L + DLE R + + + L +RE + K+E
Sbjct: 464 MNKNELMKRIAEYKRSREELRLL--REDLEKRIKECHRRRESLDNIRERIRQIEEGKAES 521
Query: 89 KKIVSGNLNPSARD------EL-----LESGMADTMTASADQRTRLMTSTERLN-KTGER 136
+K++ L + D EL +ESG + + A++ + +L T LN +T
Sbjct: 522 EKLIE-ELKDAESDFNKVEHELKSLREMESGYSSKLAAASGRMDQLRTRITELNPETSRE 580
Query: 137 VKDSRRTMLETE 148
+D+ + + E E
Sbjct: 581 YRDNLKKIDENE 592
>GS28_DROME (Q9VE50) Probable 28 kDa Golgi SNARE protein (Golgi SNAP
receptor complex member 1)
Length = 232
Score = 41.2 bits (95), Expect = 0.002
Identities = 36/177 (20%), Positives = 75/177 (42%), Gaps = 19/177 (10%)
Query: 46 DEAEALIRKMDLEARSMQ--PNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDE 103
+E E ++ K+ SM P + L+ ++ L + E KI + + R+E
Sbjct: 61 EEIEQMLEKLSSLNESMSDLPASGAAAMHTLQRHREILQGYRQEFNKICANHTMRIEREE 120
Query: 104 LLE-SGMADTMTASA----DQRTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDL 158
LL SG+A + + + ++R + + LN V D +ET + L
Sbjct: 121 LLRGSGLATSSGSPSISGLNRREMYLKESGHLNSASHLVNDQINIAIETRD-------HL 173
Query: 159 HSQRQSLLHAHNTLHGVDDNIGKSKKIMTNMSRRMNKNKWIIG-----CIVLVLVVA 210
H+QRQ+ + + + ++ ++ + ++ I+G C++L+L+ A
Sbjct: 174 HAQRQAFKRLQTRFNDISNRFPLISSLIQRINIKKRRDSLILGAVIGFCVILLLLYA 230
>ME12_ARATH (Q9FK28) Membrin 12 (AtMEMB12) (Golgi SNAP receptor
complex member 2-2)
Length = 219
Score = 40.0 bits (92), Expect = 0.004
Identities = 47/206 (22%), Positives = 84/206 (39%), Gaps = 17/206 (8%)
Query: 20 SKKCTAATALNGEQKKQK--------VSEVKSGIDEAEALIRKMDLEARSMQ-PNIKGVL 70
S K A NG +K ++ S VK I E ++L MD RS+ + + +
Sbjct: 15 SAKRILLRARNGIEKLERFDSDPTDLASSVKRDITEVQSLCSNMDGLWRSIPVKSQRDLW 74
Query: 71 LAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQRTRLMTSTERL 130
K + + L ++K + N R L AD + + + ++ +
Sbjct: 75 RRKSEQVGEEAEYLNQSLEKYMWRN----QRKMLEAKERADLLGRGSGEGAHILQIFDEE 130
Query: 131 NKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIGKSKKIMTNMS 190
+ VK+S+R + ++ + GV+IL QR L A V + +G S ++ +
Sbjct: 131 AQGMNSVKNSKRMLEDSFQSGVAILSKYAEQRDRLKSAQRKALDVLNTVGLSNSVLRLIE 190
Query: 191 RRMNKNKWIIGCIVLVLVVAIIAILY 216
RR + W I ++A + ILY
Sbjct: 191 RRNRVDTW----IKYAGMIATLVILY 212
>MYSP_BLOTA (Q8MUF6) Paramyosin (Allergen Blo t 11)
Length = 875
Score = 39.3 bits (90), Expect = 0.007
Identities = 39/184 (21%), Positives = 84/184 (45%), Gaps = 8/184 (4%)
Query: 14 ELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEARSMQPNIKGVLLAK 73
E L ++ T A K + +E+++ +DE E L RKM + ++ LL K
Sbjct: 276 EARLELERQLTKANGDAASWKSKYEAELQAHVDEVEELRRKMAQKISEYGEQLE-ALLNK 334
Query: 74 LREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQRTRLMTSTERLNKT 133
+ L+SEV+ ++ +A + LE ++ + D +++L + L +T
Sbjct: 335 CSALEKQKARLQSEVEVLIMDLEKATAHAQALEKRVSQLEKINLDLKSKLEEVSMLLEQT 394
Query: 134 GERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIGKSKKIMTNMSRRM 193
KD R + + ++L + L Q+++L + L D++ ++K + + RR+
Sbjct: 395 ---QKDLRVKIADLQKLQHE-YEKLRDQKEALARENKKL---ADDLAEAKSQLNDAHRRI 447
Query: 194 NKNK 197
++ +
Sbjct: 448 HEQE 451
>BPA1_HUMAN (Q03001) Bullous pemphigoid antigen 1 isoforms 1/2/3/4/5/8
(230 kDa bullous pemphigoid antigen) (BPA)
(Hemidesmosomal plaque protein) (Dystonia musculorum
protein) (Dystonin) (Fragment)
Length = 3214
Score = 39.3 bits (90), Expect = 0.007
Identities = 41/172 (23%), Positives = 70/172 (39%), Gaps = 12/172 (6%)
Query: 23 CTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEARSMQPNIKGVLLAKLREYKSDLN 82
C A T L E KQK E+K +DE A RK + + R + + + L K +
Sbjct: 1911 CRAVTGLQQEHDKQKAEELKQQVDELTAANRKAEQDMRELTYELNALQLEKTSSEE---- 1966
Query: 83 NLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQRTRLMTSTERLNKTGERVKDSRR 142
K+ + K N + R LE D Q+ R + +LN+T + +++
Sbjct: 1967 --KARLLKDKLDETNNTLRCLKLELERKDQAEKGYSQQLREL--GRQLNQTTGKAEEA-- 2020
Query: 143 TMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIGKSKKIMTNMSRRMN 194
M E +L I ++ + +SL H L D I ++ + + +N
Sbjct: 2021 -MQEASDL-KKIKRNYQLELESLNHEKGKLQREVDRITRAHAVAEKNIQHLN 2070
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.314 0.129 0.342
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,675,792
Number of Sequences: 164201
Number of extensions: 786509
Number of successful extensions: 3409
Number of sequences better than 10.0: 234
Number of HSP's better than 10.0 without gapping: 24
Number of HSP's successfully gapped in prelim test: 210
Number of HSP's that attempted gapping in prelim test: 3306
Number of HSP's gapped (non-prelim): 309
length of query: 221
length of database: 59,974,054
effective HSP length: 106
effective length of query: 115
effective length of database: 42,568,748
effective search space: 4895406020
effective search space used: 4895406020
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 63 (28.9 bits)
Medicago: description of AC119412.5


