

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119412.2 - phase: 0
(184 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
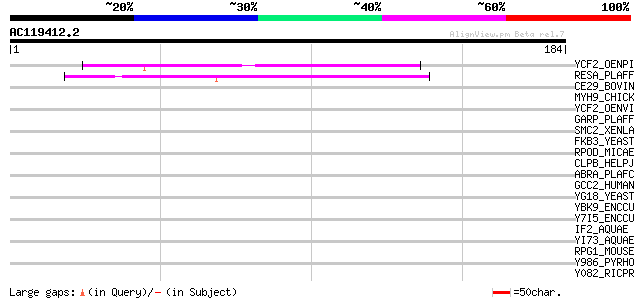
Score E
Sequences producing significant alignments: (bits) Value
YCF2_OENPI (P31568) Protein ycf2 (Fragment) 43 5e-04
RESA_PLAFF (P13830) Ring-infected erythrocyte surface antigen pr... 42 8e-04
CE29_BOVIN (Q9TU23) Centrosomal protein Cep290 41 0.001
MYH9_CHICK (P14105) Myosin heavy chain, nonmuscle (Cellular myos... 41 0.002
YCF2_OENVI (P31569) Protein ycf2 (Fragment) 40 0.003
GARP_PLAFF (P13816) Glutamic acid-rich protein precursor 40 0.003
SMC2_XENLA (P50533) Structural maintenance of chromosome 2 (Chro... 40 0.004
FKB3_YEAST (P38911) FK506-binding nuclear protein (EC 5.2.1.8) (... 40 0.004
RPOD_MICAE (P52322) RNA polymerase sigma factor rpoD1 39 0.005
CLPB_HELPJ (Q9ZMH1) Chaperone clpB 39 0.005
ABRA_PLAFC (P22620) 101 kDa malaria antigen (P101) (Acidic basic... 39 0.005
GCC2_HUMAN (Q8IWJ2) GRIP and coiled-coil domain-containing prote... 39 0.007
YG18_YEAST (P53207) Hypothetical 71.4 kDa protein in SEC9-MSB2 i... 39 0.009
YBK9_ENCCU (Q8STZ1) Hypothetical protein ECU11_2090 39 0.009
Y7I5_ENCCU (Q8ST44) Hypothetical protein ECU07_1850/ECU10_0050 39 0.009
IF2_AQUAE (O67825) Translation initiation factor IF-2 39 0.009
YI73_AQUAE (O67720) Hypothetical protein AQ_1873 38 0.011
RPG1_MOUSE (Q9EPQ2) X-linked retinitis pigmentosa GTPase regulat... 38 0.015
Y986_PYRHO (O58714) Hypothetical UPF0271 protein PH0986 37 0.020
Y082_RICPR (Q9ZE65) Hypothetical protein RP082 37 0.020
>YCF2_OENPI (P31568) Protein ycf2 (Fragment)
Length = 721
Score = 42.7 bits (99), Expect = 5e-04
Identities = 34/118 (28%), Positives = 54/118 (44%), Gaps = 10/118 (8%)
Query: 25 IMDIQANIQSTHGKQENNS------NTFEEVGKIVEEGVDDTEEDIILEECSTMKMVEEL 78
++D +A I + +E + T V + EEG D EED++LE +++EE
Sbjct: 483 LLDCEAEIPAEEIPEEEDELPEDALETEVAVWGVEEEGEADDEEDVLLEAQQEDELLEE- 541
Query: 79 EPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNKVEIDRI 136
EE +E +E E EE E+ E+E+ +EE EE + N E R+
Sbjct: 542 ---EDEELDEEEDELDEEEEEPKEEEDELHEEEEEEEEEEEEEEEDELQENDSEFFRV 596
Score = 41.2 bits (95), Expect = 0.001
Identities = 49/189 (25%), Positives = 80/189 (41%), Gaps = 29/189 (15%)
Query: 17 QMEMMQEQIMDIQANIQSTH-----GKQENNSNTFEEVGKIVEEGVDDTEEDIILEECST 71
++E +E++ + ++ T G +E T EEV +E V+ TEE++ E
Sbjct: 294 EVEGTEEEVEGTEEEVEGTEDEEVEGTEEEVEGTEEEVEGTEDEEVEGTEEEVEGTEEEV 353
Query: 72 MKMVEELEPLHPE-EFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNK 130
EE+E E E +E E E EE +E TE+E ++E +E K ++
Sbjct: 354 EGTEEEVEGTEEEVEGTEEEVEGTEEEVEGTEEEVEGTEEEVE-GTEDEEVEGTEKDSSQ 412
Query: 131 VEIDRIIDEICALFNKPK------LGRIWTPHQLYFKFME---------FLPTRRIEKDD 175
+ DR+ L +PK + R+ HQ Y +E F P + +E D
Sbjct: 413 FDNDRV-----TLLLRPKPRNPLDIQRLIYQHQKYESELEEDDDDDEDVFAPQKMLE--D 465
Query: 176 VLSVSFWPP 184
+ S W P
Sbjct: 466 LFSELVWSP 474
Score = 37.4 bits (85), Expect = 0.020
Identities = 32/114 (28%), Positives = 50/114 (43%), Gaps = 7/114 (6%)
Query: 27 DIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKMVEELEPLHPEEF 86
+++ + G +E T EEV +E V+ TEE++ E EE+E E
Sbjct: 185 EVEGTEEEVEGTEEEVEGTEEEVEGTEDEEVEGTEEEVEGTEEEVEGTEEEVEGTEEE-- 242
Query: 87 PQERPYTEEAETVENEEV----MEVTEKEDRILIKEESMEEKGKKVNKVEIDRI 136
E EE E E+EEV EV E+ + EE +E ++V E + +
Sbjct: 243 -VEGTEDEEVEGTEDEEVEGTEEEVEGTEEEVEGTEEEVEGTEEEVEGTEDEEV 295
Score = 36.2 bits (82), Expect = 0.044
Identities = 35/136 (25%), Positives = 61/136 (44%), Gaps = 16/136 (11%)
Query: 4 MQDTNQRLKNLSFQMEMMQEQIMDIQANIQSTH-----GKQENNSNTFEEVGKIVEEGVD 58
++ T + ++ ++E +E++ + ++ T G +E T EEV +E V+
Sbjct: 259 VEGTEEEVEGTEEEVEGTEEEVEGTEEEVEGTEDEEVEGTEEEVEGTEEEVEGTEDEEVE 318
Query: 59 DTEEDIILEECSTMKMVEELEPLHPEEF--PQERPYTEEAETVENEEVMEVTEKEDRILI 116
TEE++ E EE+E EE +E E E EE +E TE+E +
Sbjct: 319 GTEEEVEGTE-------EEVEGTEDEEVEGTEEEVEGTEEEVEGTEEEVEGTEEE--VEG 369
Query: 117 KEESMEEKGKKVNKVE 132
EE +E ++V E
Sbjct: 370 TEEEVEGTEEEVEGTE 385
Score = 36.2 bits (82), Expect = 0.044
Identities = 31/115 (26%), Positives = 52/115 (44%), Gaps = 5/115 (4%)
Query: 27 DIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKMVEELEPLHPEEF 86
+++ + G +E T +E + EE V+ TEE++ E EE+E EE
Sbjct: 192 EVEGTEEEVEGTEEEVEGTEDEEVEGTEEEVEGTEEEVEGTEEEVEGTEEEVEGTEDEEV 251
Query: 87 PQERPYTEEAETVENEEVMEVTEKEDRILIKE-ESMEEKGKKVNKVEIDRIIDEI 140
TE+ E EE +E TE+E +E E EE+ + E++ +E+
Sbjct: 252 EG----TEDEEVEGTEEEVEGTEEEVEGTEEEVEGTEEEVEGTEDEEVEGTEEEV 302
Score = 33.5 bits (75), Expect = 0.28
Identities = 31/135 (22%), Positives = 61/135 (44%), Gaps = 13/135 (9%)
Query: 8 NQRLKNLSFQMEMMQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILE 67
++ ++ ++E +E++ + ++ T + E + EEV +E V+ TEE++
Sbjct: 212 DEEVEGTEEEVEGTEEEVEGTEEEVEGTEEEVEGTED--EEVEGTEDEEVEGTEEEVEGT 269
Query: 68 ECSTMKMVEELEPLHPEEFPQERPYTEEAETVENEEVMEVTEKE------DRILIKEESM 121
E EE+E +E TE+ E EE +E TE+E + + EE +
Sbjct: 270 EEEVEGTEEEVEGTE-----EEVEGTEDEEVEGTEEEVEGTEEEVEGTEDEEVEGTEEEV 324
Query: 122 EEKGKKVNKVEIDRI 136
E ++V E + +
Sbjct: 325 EGTEEEVEGTEDEEV 339
>RESA_PLAFF (P13830) Ring-infected erythrocyte surface antigen
precursor
Length = 1073
Score = 42.0 bits (97), Expect = 8e-04
Identities = 31/123 (25%), Positives = 61/123 (49%), Gaps = 4/123 (3%)
Query: 19 EMMQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILE--ECSTMKMVE 76
E ++E + +++ N++ +EN EEV + VEE V++ E+ + E E + + VE
Sbjct: 934 ENVEENVEEVEENVEEN--VEENVEENVEEVEENVEENVEENVEENVEENVEENVEENVE 991
Query: 77 ELEPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNKVEIDRI 136
E + EE+ +E E EN E E+ + EE++EE ++ + ++
Sbjct: 992 ENVEENVEEYDEENVEEVEENVEENVEENVEENVEENVEEVEENVEENVEENVEENVEEN 1051
Query: 137 IDE 139
++E
Sbjct: 1052 VEE 1054
Score = 39.3 bits (90), Expect = 0.005
Identities = 32/118 (27%), Positives = 54/118 (45%), Gaps = 7/118 (5%)
Query: 27 DIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILE-----ECSTMKMVEELEPL 81
D + N++ H +EN EEV + VEE V++ E+ + E E + + VEE
Sbjct: 923 DAEENVE--HDAEENVEENVEEVEENVEENVEENVEENVEEVEENVEENVEENVEENVEE 980
Query: 82 HPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNKVEIDRIIDE 139
+ EE +E E VE + V E E+ + E E+ + N E++ ++E
Sbjct: 981 NVEENVEENVEENVEENVEEYDEENVEEVEENVEENVEENVEENVEENVEEVEENVEE 1038
Score = 28.9 bits (63), Expect = 7.0
Identities = 26/96 (27%), Positives = 47/96 (48%), Gaps = 5/96 (5%)
Query: 19 EMMQEQIMD-IQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTM-KMVE 76
E ++E + + ++ N++ +E + EEV + VEE V++ E+ + E + + VE
Sbjct: 980 ENVEENVEENVEENVEEN--VEEYDEENVEEVEENVEENVEENVEENVEENVEEVEENVE 1037
Query: 77 ELEPLHPEEFPQERPYTEEAETVENEEVMEVTEKED 112
E + EE +E E E + E V E E+ D
Sbjct: 1038 ENVEENVEENVEEN-VEENVEEYDEENVEEHNEEYD 1072
>CE29_BOVIN (Q9TU23) Centrosomal protein Cep290
Length = 1453
Score = 41.2 bits (95), Expect = 0.001
Identities = 36/135 (26%), Positives = 67/135 (48%), Gaps = 12/135 (8%)
Query: 1 MINMQDTNQRLKNLSFQMEMMQEQIMDIQANIQSTHGKQENNS--NTFEEVG----KIVE 54
M N Q ++ LKN + +ME+ + + D+ + ++ G Q+ S EE+ K+
Sbjct: 383 MKNSQQEHRSLKNKTLEMELKLKGLEDLISTLKDARGAQKVISWHTKIEELRLQELKLNR 442
Query: 55 EGVDDTEE-----DIILEECSTMKMVEELEPLHPEEFPQERPYTEEAETVENEEVMEVTE 109
E V D EE +II E +T+ +EE E + +F +ER + VE E ++V +
Sbjct: 443 ELVKDKEEIKYLNNIISEYENTISSLEE-EIVQQNKFHEERQMAWDQREVELERQLDVFD 501
Query: 110 KEDRILIKEESMEEK 124
++ +++E E+
Sbjct: 502 RQQSEILREAQKFEE 516
>MYH9_CHICK (P14105) Myosin heavy chain, nonmuscle (Cellular myosin
heavy chain) (NMMHC)
Length = 1959
Score = 40.8 bits (94), Expect = 0.002
Identities = 30/138 (21%), Positives = 68/138 (48%), Gaps = 12/138 (8%)
Query: 4 MQDTNQRLKNLSFQMEMMQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEG-VDDTEE 62
+++ +R ++L + + MQ+ I +++ ++ +E ++ ++ K+ E + EE
Sbjct: 924 VEEEEERCQHLQAEKKKMQQNIQELEEQLE-----EEESARQKLQLEKVTTEAKLKKLEE 978
Query: 63 DIILEECSTMKMVEELEPLHPEEFPQERPYTEEAET------VENEEVMEVTEKEDRILI 116
D+I+ E +K+ +E + L TEE E ++N+ +T+ E+R+
Sbjct: 979 DVIVLEDQNLKLAKEKKLLEDRMSEFTTNLTEEEEKSKSLAKLKNKHEAMITDLEERLRR 1038
Query: 117 KEESMEEKGKKVNKVEID 134
+E+ +E K K+E D
Sbjct: 1039 EEKQRQELEKTRRKLEGD 1056
>YCF2_OENVI (P31569) Protein ycf2 (Fragment)
Length = 630
Score = 40.0 bits (92), Expect = 0.003
Identities = 34/118 (28%), Positives = 54/118 (44%), Gaps = 12/118 (10%)
Query: 25 IMDIQANIQSTHGKQENNS------NTFEEVGKIVEEGVDDTEEDIILEECSTMKMVEEL 78
++D +A I + +E + T V + EEG D EED++LE +++EE
Sbjct: 394 LLDCEAEIPAEEIPEEEDELPEDALETEVAVWGVEEEGEADDEEDVLLEAQQEDELLEE- 452
Query: 79 EPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNKVEIDRI 136
EE +E +E E EE E+ E+E+ +EE EE + N E R+
Sbjct: 453 ---EDEELDEEEDELDEEEEEPKEEEDELHEEEEE--EEEEEEEEDELQENDSEFFRV 505
Score = 37.7 bits (86), Expect = 0.015
Identities = 49/215 (22%), Positives = 87/215 (39%), Gaps = 36/215 (16%)
Query: 1 MINMQDTNQRLKNLSFQMEMMQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDT 60
++ T + ++ ++E +E++ + ++ T +E T +E + EE V+ T
Sbjct: 176 LVGSSPTEEEVEGTEEEVEGTEEEVEGTEEEVEGT---EEEVEGTEDEEVEGTEEEVEGT 232
Query: 61 EEDIILEECSTMKMVEELEPLHPE---------EFPQERPYTEEAETVENEEVMEVTEK- 110
EE++ E EE+E E E +E E E EE +E TE+
Sbjct: 233 EEEVEGTEEEVEGTEEEVEGTEEEVEGTEDEEVEGTEEEVEGTEEEVEGTEEEVEGTEEE 292
Query: 111 ----EDRILIKEESMEEKGKKVNKVEID--RIIDEICALFNKPK------LGRIWTPHQL 158
E+ + EE +E ++V E D + ++ L +PK + R+ HQ
Sbjct: 293 VEGTEEEVEGTEEEVEGTEEEVEGTEKDSSQFDNDRVTLLLRPKPRNPLDIQRLIYQHQK 352
Query: 159 YFKFME---------FLPTRRIEKDDVLSVSFWPP 184
Y +E F P + +E D+ S W P
Sbjct: 353 YESELEEDDDDDEDVFAPQKMLE--DLFSELVWSP 385
Score = 29.3 bits (64), Expect = 5.3
Identities = 32/129 (24%), Positives = 55/129 (41%), Gaps = 26/129 (20%)
Query: 29 QANIQSTHGKQENNSNTFEEVGKIV----------------EEGVDDTEEDIILEECSTM 72
+ N+ +++G EN+S+ + IV EE V+ TEE++ E
Sbjct: 142 EQNLITSYGLVENDSDLVHGLSDIVHGLLELEGALVGSSPTEEEVEGTEEEVEGTEEEVE 201
Query: 73 KMVEELEPLHPEEFPQERPYTEEAETVENEEVMEVTEK-----EDRILIKEESMEEKGKK 127
EE+E +E TE+ E EE +E TE+ E+ + EE +E ++
Sbjct: 202 GTEEEVEGTE-----EEVEGTEDEEVEGTEEEVEGTEEEVEGTEEEVEGTEEEVEGTEEE 256
Query: 128 VNKVEIDRI 136
V E + +
Sbjct: 257 VEGTEDEEV 265
>GARP_PLAFF (P13816) Glutamic acid-rich protein precursor
Length = 678
Score = 40.0 bits (92), Expect = 0.003
Identities = 37/130 (28%), Positives = 56/130 (42%), Gaps = 5/130 (3%)
Query: 10 RLKNLSFQMEMMQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILEEC 69
R N +M ++E + Q ++ K+E + EE K V+E ++ EED EE
Sbjct: 523 RRDNHKKKMAKIEEAELQKQKHVDKEEDKKEESKEVQEE-SKEVQEDEEEVEEDEEEEE- 580
Query: 70 STMKMVEELEPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVN 129
+ EE E EE +E EE E + E+ + E ED E+ EE + +
Sbjct: 581 ---EEEEEEEEEEEEEEEEEEEEEEEEEDEDEEDEDDAEEDEDDAEEDEDDAEEDDDEED 637
Query: 130 KVEIDRIIDE 139
E D DE
Sbjct: 638 DDEEDDDEDE 647
Score = 31.6 bits (70), Expect = 1.1
Identities = 26/126 (20%), Positives = 57/126 (44%), Gaps = 4/126 (3%)
Query: 5 QDTNQRLKNLSFQMEMMQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDI 64
+D + K + + + +QE +++ + + ++E EE + EE ++ E++
Sbjct: 549 EDKKEESKEVQEESKEVQEDEEEVEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEDE- 607
Query: 65 ILEECSTMKMVEELEPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEK 124
+E E+ + +E E EE + E+++ E ++ED +EE EE
Sbjct: 608 --DEEDEDDAEEDEDDAEEDEDDAEEDDDEEDDDEEDDDEDEDEDEEDEEE-EEEEEEES 664
Query: 125 GKKVNK 130
KK+ +
Sbjct: 665 EKKIKR 670
Score = 30.8 bits (68), Expect = 1.8
Identities = 24/110 (21%), Positives = 52/110 (46%)
Query: 21 MQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKMVEELEP 80
+ E D + N + + + + EE+ K +++ +E + E+ K ++ E
Sbjct: 228 VHETSNDTKDNDKENISEDKKEDHQQEEMLKTLDKKERKQKEKEMKEQEKIEKKKKKQEE 287
Query: 81 LHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNK 130
++ +ER E+ E + E+ M+ +K ++ K+E E+K KK +K
Sbjct: 288 KEKKKQEKERKKQEKKERKQKEKEMKKQKKIEKERKKKEEKEKKKKKHDK 337
>SMC2_XENLA (P50533) Structural maintenance of chromosome 2
(Chromosome-associated protein E) (Chromosome assembly
protein XCAP-E)
Length = 1203
Score = 39.7 bits (91), Expect = 0.004
Identities = 31/141 (21%), Positives = 69/141 (47%), Gaps = 17/141 (12%)
Query: 1 MINMQDTNQRLKNLSFQMEMMQEQIMDIQANIQ-STHGKQENNSNTFEEVGKIVEEGVDD 59
++ +++T +R + L Q EM E+ +Q +Q S++ KQ+ ++ ++ +++
Sbjct: 703 LMTLKNTVERYRQLKQQWEMKSEEAELLQTKLQQSSYHKQQEELDSLKQT-------IEE 755
Query: 60 TEEDIILEECSTMKMVEELEPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEE 119
+EE T+K +E++ E+F + AE E+ E +K D K +
Sbjct: 756 SEE--------TLKNTKEVQKKAEEKFKVLEHKMKNAEAERERELKEAQQKLDTAKKKAD 807
Query: 120 SMEEKGKKVNKVEIDRIIDEI 140
+ +K K+ + E+D ++ E+
Sbjct: 808 ASNKKMKE-KQQEVDALVLEL 827
>FKB3_YEAST (P38911) FK506-binding nuclear protein (EC 5.2.1.8)
(Peptidyl-prolyl cis-trans isomerase) (PPIase) (Proline
rotamase) (Nucleolar proline isomerase) (FKBP-70)
Length = 411
Score = 39.7 bits (91), Expect = 0.004
Identities = 30/93 (32%), Positives = 48/93 (51%), Gaps = 6/93 (6%)
Query: 47 EEVGKIVEEGVDDTEEDIILEECSTMKMVEELEPLHPEEFPQERPYTEEAETVENEEVME 106
E +G +++ D+ EE++ +EE E+ E EE +E E+ E V+ E
Sbjct: 197 EIIGDDMDDLDDEEEEEVRIEEVQE----EDEEDNDGEEEQEEEEEEEQKEEVKPEPKKS 252
Query: 107 VTEKEDRILIKEESMEEKGKKVNKVEIDRIIDE 139
EK+ + KEE E+K KKV KVE + ++E
Sbjct: 253 KKEKKRKHEEKEE--EKKAKKVKKVEFKKDLEE 283
>RPOD_MICAE (P52322) RNA polymerase sigma factor rpoD1
Length = 416
Score = 39.3 bits (90), Expect = 0.005
Identities = 31/129 (24%), Positives = 63/129 (48%), Gaps = 13/129 (10%)
Query: 53 VEEGVDDTEEDIILEECSTM---KMVEELEPLHPEEFPQERPYTEEAETVENEEVMEVTE 109
++E DD + ++ LEE S K L + +++PYTE++ + +E+ +
Sbjct: 22 LDEDFDDEDIELELEETSDSNDGKKTRRLASARRRDAARKKPYTEDSIRIYLQEIGRI-- 79
Query: 110 KEDRILIKEESMEEKGKKVNKVEIDRIIDEICALFNKPKLGRIWTPHQLYFKFMEFLPTR 169
R+L EE +E K + ++++RI ++ C L++ + W F+ +E +
Sbjct: 80 ---RLLRAEEEIELARKIADLLKLERIREDFC-LYSDAE----WGKQVFLFERIEKIIVE 131
Query: 170 RIEKDDVLS 178
+ EK+ LS
Sbjct: 132 KSEKEPKLS 140
>CLPB_HELPJ (Q9ZMH1) Chaperone clpB
Length = 856
Score = 39.3 bits (90), Expect = 0.005
Identities = 37/143 (25%), Positives = 63/143 (43%), Gaps = 17/143 (11%)
Query: 14 LSFQMEMMQEQIMDIQANIQSTHGKQ-----ENNSNTFEEVGKIVEEGVDDTEEDIILE- 67
L QME ++ ++ +IQ ++ EN + + + +I++E D EE I LE
Sbjct: 398 LKMQMESEPAKLSSVKRSIQRLEMEKQALEMENKESNHKRMQEILKELSDLKEEKIQLEA 457
Query: 68 ----------ECSTMKMVEELEPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIK 117
E S +KM E E F + Y + AE +E ++ E +KE + K
Sbjct: 458 QFENEKEVFKEISRLKMEMEGLKKEAERFKRNGDYQQAAE-IEYSKIPEKEKKEKELQHK 516
Query: 118 EESMEEKGKKVNKVEIDRIIDEI 140
E+M++ G + + I EI
Sbjct: 517 WETMQQNGALLQNALTENNIAEI 539
>ABRA_PLAFC (P22620) 101 kDa malaria antigen (P101) (Acidic basic
repeat antigen)
Length = 743
Score = 39.3 bits (90), Expect = 0.005
Identities = 33/129 (25%), Positives = 62/129 (47%), Gaps = 14/129 (10%)
Query: 18 MEMMQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEGV-------DDTEEDIILEECS 70
+E + I I+ +Q+ + + N +N + + +I ++ + +TEE++ + +
Sbjct: 598 IEAINNDIQIIKQELQAIYNELMNYTNGNKNIQQIFQQNILENDVLNQETEEEMEKQVEA 657
Query: 71 TMKMVEE----LEPLHPEEFPQERPYTEEAETVENEEVMEVT---EKEDRILIKEESMEE 123
K +E L P + EE +E+ +E E E EE + EKE++ KEE EE
Sbjct: 658 ITKQIEAEVDALAPKNKEEEEKEKEKEKEKEEKEKEEKEKEEKEKEKEEKEKEKEEKEEE 717
Query: 124 KGKKVNKVE 132
K +K + E
Sbjct: 718 KKEKEEEQE 726
>GCC2_HUMAN (Q8IWJ2) GRIP and coiled-coil domain-containing protein
2 (Golgi coiled coil protein GCC185) (CTCL tumor antigen
se1-1) (CLL-associated antigen KW-11)
Length = 1583
Score = 38.9 bits (89), Expect = 0.007
Identities = 26/128 (20%), Positives = 65/128 (50%), Gaps = 3/128 (2%)
Query: 9 QRLKNLSFQMEMMQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEG-VDDTEEDIILE 67
+ L+ + + E +Q +++++ + T + +N EEV + + + + +E ++
Sbjct: 744 KELEEIQSEKEALQSDLLEMKNANEKTRLENQNLLIQVEEVSQTCSKSEIHNEKEKCFIK 803
Query: 68 ECSTMKMVEELEPLHPE--EFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKG 125
E +K + E + L E + ++ +V+N+ + V E E++I E+ +EK
Sbjct: 804 EHENLKPLLEQKELRDRRAELILLKDSLAKSPSVKNDPLSSVKELEEKIENLEKECKEKE 863
Query: 126 KKVNKVEI 133
+K+NK+++
Sbjct: 864 EKINKIKL 871
Score = 32.3 bits (72), Expect = 0.63
Identities = 35/150 (23%), Positives = 71/150 (47%), Gaps = 24/150 (16%)
Query: 8 NQRLKNLSFQMEMMQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILE 67
+Q+L+ L QM+++ E + A ++S + + S+ +++ + +E + +ED+IL+
Sbjct: 562 DQKLEKLMVQMKVLSEDKEVLSAEVKSLYEENNKLSSEKKQLSRDLEVFLSQ-KEDVILK 620
Query: 68 ECST------MKMVEELEPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRI-LIKEES 120
E T MVEE + L+ + +ENE+V ++ K +KE
Sbjct: 621 EHITQLEKKLQLMVEEQDNLN--------------KLLENEQVQKLFVKTQLYGFLKEMG 666
Query: 121 ME--EKGKKVNKVEIDRIIDEICALFNKPK 148
E E ++ + V + + + E A N+ K
Sbjct: 667 SEVSEDSEEKDVVNVLQAVGESLAKINEEK 696
>YG18_YEAST (P53207) Hypothetical 71.4 kDa protein in SEC9-MSB2
intergenic region
Length = 620
Score = 38.5 bits (88), Expect = 0.009
Identities = 30/122 (24%), Positives = 56/122 (45%), Gaps = 11/122 (9%)
Query: 6 DTNQRLKNLSFQMEMMQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEGV-DDTEEDI 64
+ +++K+ + + Q+ + I+ H + ++NT EE + +E DDTE+D+
Sbjct: 296 EKEKKMKSTYEESRRQRHQMQKVFDQIRKNHSGAKGSANTEEEDTNMEDEDEEDDTEDDL 355
Query: 65 ILEECSTMKMVEE--------LEPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILI 116
LE+ + +EE L LH P+ + + + EN E E EK + +
Sbjct: 356 ALEKRKEERDLEESNRRYEDMLHQLHSNTEPKIKSIRADIMSAENYE--EHLEKNRSLYL 413
Query: 117 KE 118
KE
Sbjct: 414 KE 415
>YBK9_ENCCU (Q8STZ1) Hypothetical protein ECU11_2090
Length = 634
Score = 38.5 bits (88), Expect = 0.009
Identities = 29/96 (30%), Positives = 45/96 (46%), Gaps = 10/96 (10%)
Query: 34 STHGKQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTM--KMVEELEPLHPEEFPQERP 91
STH K + E V K+ E D + D++ CS K +EE+ + +ER
Sbjct: 282 STHKKYKCMDVVAELVRKVFVEDKDIEDSDVMSAVCSVRERKRLEEMREM------EERK 335
Query: 92 YTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKK 127
EE EE++ + E+E+R K E + +GKK
Sbjct: 336 RKEEERAKNEEELLRMVEREER--EKREESKGRGKK 369
>Y7I5_ENCCU (Q8ST44) Hypothetical protein ECU07_1850/ECU10_0050
Length = 634
Score = 38.5 bits (88), Expect = 0.009
Identities = 29/96 (30%), Positives = 45/96 (46%), Gaps = 10/96 (10%)
Query: 34 STHGKQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTM--KMVEELEPLHPEEFPQERP 91
STH K + E V K+ E D + D++ CS K +EE+ + +ER
Sbjct: 282 STHKKYKCMDVVAELVRKVFVEDKDIEDSDVMSAVCSVRERKRLEEMREM------EERK 335
Query: 92 YTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKK 127
EE EE++ + E+E+R K E + +GKK
Sbjct: 336 RKEEERAKNEEELLRMVEREER--EKREESKGRGKK 369
>IF2_AQUAE (O67825) Translation initiation factor IF-2
Length = 805
Score = 38.5 bits (88), Expect = 0.009
Identities = 41/134 (30%), Positives = 63/134 (46%), Gaps = 23/134 (17%)
Query: 19 EMMQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILEEC--------- 69
E++ EQ QA + K+E EEV IVEE V++ + ++I+EE
Sbjct: 67 EVVTEQA---QAPAEVEEKKEEEKK---EEV--IVEEVVEEKKPEVIVEEIEEKKEEEEK 118
Query: 70 ----STMKMVEEL--EPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEE 123
K VEEL E L +E +E+ E+ E V+EV ++E + KEE EE
Sbjct: 119 KEEEKPKKSVEELIKEILEKKEKEKEKKKVEKERKEEKVRVVEVKKEERKEEKKEEKKEE 178
Query: 124 KGKKVNKVEIDRII 137
+ K+ + +R I
Sbjct: 179 EKPKIKMSKKEREI 192
Score = 37.7 bits (86), Expect = 0.015
Identities = 33/125 (26%), Positives = 59/125 (46%), Gaps = 10/125 (8%)
Query: 19 EMMQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMK----M 74
E E I D I+ K+E + + ++ E+ ++ +E++I+EE K +
Sbjct: 47 EQALEMIYDA-FGIKEEEEKEEVVTEQAQAPAEVEEKKEEEKKEEVIVEEVVEEKKPEVI 105
Query: 75 VEELEPLHPEEFPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNKVEID 134
VEE+E EE +E +++ +E++E EKE KE+ EK +K KV +
Sbjct: 106 VEEIEEKKEEEEKKEEEKPKKSVEELIKEILEKKEKE-----KEKKKVEKERKEEKVRVV 160
Query: 135 RIIDE 139
+ E
Sbjct: 161 EVKKE 165
Score = 32.7 bits (73), Expect = 0.48
Identities = 32/116 (27%), Positives = 53/116 (45%), Gaps = 12/116 (10%)
Query: 33 QSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKMVEELEPLHPEEFPQERPY 92
+++HG E + I EE + +E+++ E+ VEE + EE +E
Sbjct: 40 KASHGLDEQALEMIYDAFGIKEE---EEKEEVVTEQAQAPAEVEEKK----EEEKKEEVI 92
Query: 93 TEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNKVEIDRIIDEICALFNKPK 148
EE + EV+ V E E++ +E+ EEK KK ++ +I EI K K
Sbjct: 93 VEEVVEEKKPEVI-VEEIEEKKEEEEKKEEEKPKK----SVEELIKEILEKKEKEK 143
>YI73_AQUAE (O67720) Hypothetical protein AQ_1873
Length = 407
Score = 38.1 bits (87), Expect = 0.011
Identities = 27/98 (27%), Positives = 44/98 (44%), Gaps = 1/98 (1%)
Query: 41 NNSNTFEEVG-KIVEEGVDDTEEDIILEECSTMKMVEELEPLHPEEFPQERPYTEEAETV 99
N S + G + + E V+ + + EE +M+ E E EE P+E P E
Sbjct: 241 NKSTLNRDAGAEAIAEEVEKVAQKVKKEEIKSMETEEVKEEKVREEIPEEVPQEVEEIKE 300
Query: 100 ENEEVMEVTEKEDRILIKEESMEEKGKKVNKVEIDRII 137
E E+ E+ EK + I KE++ E K+ ++ I
Sbjct: 301 EKPEMSEIKEKTEAISTKEKTPEAVQVLAEKLSDEKFI 338
>RPG1_MOUSE (Q9EPQ2) X-linked retinitis pigmentosa GTPase
regulator-interacting protein 1 (RPGR-interacting protein
1)
Length = 1331
Score = 37.7 bits (86), Expect = 0.015
Identities = 34/130 (26%), Positives = 59/130 (45%), Gaps = 1/130 (0%)
Query: 26 MDIQANIQSTHGKQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMKMVEELEPLHPEE 85
+D +++ + G Q + + E K E G ++ +E+ + EE + EE++ EE
Sbjct: 887 LDWKSHYLAPEGFQMSEAEKPEGEEKEEEGGEEEVKEEEVEEEEEEEEEEEEVKEEKEEE 946
Query: 86 FPQERPYTEEAETVENEEVMEVTEKEDRILIKEESMEEKGKKVNKVEIDRIIDEICALFN 145
+ER EE E E EE E EKE+ +EE EE+ + V +E F+
Sbjct: 947 EEEEREEEEEKEE-EKEEEEEEDEKEEEEEEEEEEEEEEEDENKDVLEASFTEEWVPFFS 1005
Query: 146 KPKLGRIWTP 155
+ ++ P
Sbjct: 1006 QDQIASTEIP 1015
>Y986_PYRHO (O58714) Hypothetical UPF0271 protein PH0986
Length = 255
Score = 37.4 bits (85), Expect = 0.020
Identities = 31/118 (26%), Positives = 52/118 (43%), Gaps = 5/118 (4%)
Query: 36 HGKQENNSNTFEEVGKIVEEGVDDTEEDIILEECSTMK---MVEELEPLHPEEFPQERPY 92
HG N E++ + V EG+ D ++D+IL S + + EE+ E +R Y
Sbjct: 111 HGALYNAMVKEEDLARAVIEGILDFDKDLILVTLSNSRVADIAEEMGLKVAHEVFADRAY 170
Query: 93 TEEAETVENEEVMEVTEKEDRILIKEESMEEKG--KKVNKVEIDRIIDEICALFNKPK 148
+ V V E ++ I + SM + G + +N +D +D IC + PK
Sbjct: 171 NPDGTLVPRGRPGAVIEDKEEIAERVISMVKDGGIRAINGEWVDLKVDTICVHGDNPK 228
>Y082_RICPR (Q9ZE65) Hypothetical protein RP082
Length = 143
Score = 37.4 bits (85), Expect = 0.020
Identities = 37/122 (30%), Positives = 57/122 (46%), Gaps = 12/122 (9%)
Query: 5 QDTNQRLKNLSFQMEMMQEQIMDIQANIQSTHGKQENNSNTFEEVGKIVEEGV-DDTEED 63
+D NQ KNL E + + +Q I H + E NS+T + K + V ++ ED
Sbjct: 24 KDINQAKKNL----ENAEAKNEALQRQIILNHNQNEVNSHTTKNQEKFKTDNVTEEYLED 79
Query: 64 IIL---EECSTMKMVEELEPLHPEEF---PQERPYTEEAETVENEEVMEVTEKEDRILIK 117
+ L T + EE+ H EE QE TEE E V E+++ +TE+ + K
Sbjct: 80 MALMFKNSEDTAEQKEEVNCQHHEEQNRQKQEHINTEE-EAVHKEKIIHITEETETEAFK 138
Query: 118 EE 119
+E
Sbjct: 139 KE 140
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.313 0.132 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,794,209
Number of Sequences: 164201
Number of extensions: 977819
Number of successful extensions: 5531
Number of sequences better than 10.0: 401
Number of HSP's better than 10.0 without gapping: 82
Number of HSP's successfully gapped in prelim test: 323
Number of HSP's that attempted gapping in prelim test: 4772
Number of HSP's gapped (non-prelim): 756
length of query: 184
length of database: 59,974,054
effective HSP length: 104
effective length of query: 80
effective length of database: 42,897,150
effective search space: 3431772000
effective search space used: 3431772000
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC119412.2


