

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119412.1 - phase: 0
(411 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
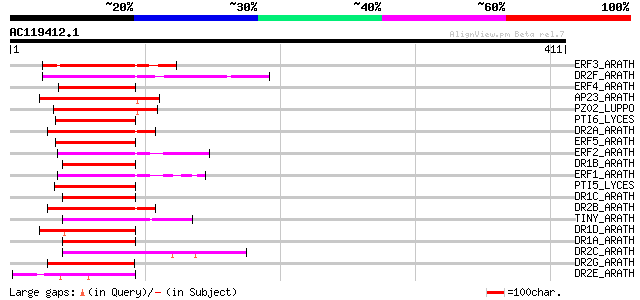
Score E
Sequences producing significant alignments: (bits) Value
ERF3_ARATH (O80339) Ethylene responsive element binding factor 3... 86 2e-16
DR2F_ARATH (Q9SVX5) Dehydration responsive element binding prote... 80 1e-14
ERF4_ARATH (O80340) Ethylene responsive element binding factor 4... 79 2e-14
AP23_ARATH (P42736) AP2 domain transcription factor RAP2.3 (Rela... 79 2e-14
PZ02_LUPPO (P16146) PPLZ02 protein 79 3e-14
PTI6_LYCES (O04682) Pathogenesis-related genes transcriptional a... 77 6e-14
DR2A_ARATH (O82132) Dehydration responsive element binding prote... 72 3e-12
ERF5_ARATH (O80341) Ethylene responsive element binding factor 5... 71 6e-12
ERF2_ARATH (O80338) Ethylene responsive element binding factor 2... 71 6e-12
DR1B_ARATH (P93835) Dehydration responsive element binding prote... 69 2e-11
ERF1_ARATH (O80337) Ethylene responsive element binding factor 1... 69 2e-11
PTI5_LYCES (O04681) Pathogenesis-related genes transcriptional a... 69 3e-11
DR1C_ARATH (Q9SYS6) Dehydration responsive element binding prote... 69 3e-11
DR2B_ARATH (O82133) Dehydration responsive element binding prote... 68 4e-11
TINY_ARATH (Q39127) Transcriptional factor TINY 67 6e-11
DR1D_ARATH (Q9FJ93) Dehydration responsive element binding prote... 67 6e-11
DR1A_ARATH (Q9M0L0) Dehydration responsive element binding prote... 67 6e-11
DR2C_ARATH (Q8LFR2) Dehydration responsive element binding prote... 67 8e-11
DR2G_ARATH (P61827) Putative dehydration responsive element bind... 65 3e-10
DR2E_ARATH (O80917) Dehydration responsive element binding prote... 65 3e-10
>ERF3_ARATH (O80339) Ethylene responsive element binding factor 3
(AtERF3)
Length = 225
Score = 85.5 bits (210), Expect = 2e-16
Identities = 50/99 (50%), Positives = 62/99 (62%), Gaps = 9/99 (9%)
Query: 25 AASLGGARRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLL 84
AA G + +R F GVR+RP GR+ AEI+D +K RVWLGTFD+AEEAARAYD AA L
Sbjct: 17 AAINGSVKEIR--FRGVRKRPWGRFAAEIRDPWKKARVWLGTFDSAEEAARAYDSAARNL 74
Query: 85 RGANTRTNFWPCSQSSTSPALSSKITNLLLQRLKERNMN 123
RG +TNF P SS P NL +++ +N N
Sbjct: 75 RGPKAKTNF-PIDSSSPPP------PNLRFNQIRNQNQN 106
>DR2F_ARATH (Q9SVX5) Dehydration responsive element binding protein
2F (DREB2F protein)
Length = 277
Score = 79.7 bits (195), Expect = 1e-14
Identities = 57/168 (33%), Positives = 81/168 (47%), Gaps = 9/168 (5%)
Query: 25 AASLGGARRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLL 84
A GG + ++ GVRQR G+WVAEI++ ++ R+WLG+F TAEEAA AYDEAA L
Sbjct: 15 ARGKGGPQNALCQYRGVRQRTWGKWVAEIREPKKRARLWLGSFATAEEAAMAYDEAALKL 74
Query: 85 RGANTRTNFWPCSQSSTSPALSSKITNLLLQRLKERNMNNSCSSFSSSKYTSLLGSPEER 144
G + N P Q +T P+LS+ QR K S F S ++ P
Sbjct: 75 YGHDAYLNL-PHLQRNTRPSLSNS------QRFKWVPSRKFISMFPSCGMLNVNAQPSVH 127
Query: 145 REQQRANDHDRQEMQQQVDAYGEVSKDFSIDQFTDFLNDHEDYSTSNN 192
QQR + + + Q +Y S T FL++ ++N
Sbjct: 128 IIQQRLEELKKTGLLSQ--SYSSSSSSTESKTNTSFLDEKTSKGETDN 173
>ERF4_ARATH (O80340) Ethylene responsive element binding factor 4
(AtERF4)
Length = 222
Score = 79.3 bits (194), Expect = 2e-14
Identities = 39/57 (68%), Positives = 45/57 (78%)
Query: 37 RFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNF 93
R+ GVR+RP GR+ AEI+D +K RVWLGTFDTAEEAARAYD AA RGA +TNF
Sbjct: 24 RYRGVRKRPWGRYAAEIRDPGKKTRVWLGTFDTAEEAARAYDTAARDFRGAKAKTNF 80
>AP23_ARATH (P42736) AP2 domain transcription factor RAP2.3 (Related
to AP2 protein 3) (Cadmium-induced protein AS30)
Length = 248
Score = 79.0 bits (193), Expect = 2e-14
Identities = 45/94 (47%), Positives = 59/94 (61%), Gaps = 5/94 (5%)
Query: 23 KEAASLGGARRVRKR-FVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAA 81
KE A+ G RR RK + G+R+RP G+W AEI+D + +RVWLGTF+TAEEAA AYD AA
Sbjct: 63 KEQATEPGKRRKRKNVYRGIRKRPWGKWAAEIRDPRKGVRVWLGTFNTAEEAAMAYDVAA 122
Query: 82 CLLRGANTRTNF----WPCSQSSTSPALSSKITN 111
+RG + NF P + T P S + T+
Sbjct: 123 KQIRGDKAKLNFPDLHHPPPPNYTPPPSSPRSTD 156
>PZ02_LUPPO (P16146) PPLZ02 protein
Length = 164
Score = 78.6 bits (192), Expect = 3e-14
Identities = 42/80 (52%), Positives = 56/80 (69%), Gaps = 3/80 (3%)
Query: 33 RVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTN 92
R ++R+ G RQR G WV+EI+ +I K R+W GTF++AE+AARAYDEAA L+ G RTN
Sbjct: 3 RPQQRYRGFRQRHWGSWVSEIRHSILKTRIWQGTFESAEDAARAYDEAARLMCGTRARTN 62
Query: 93 F---WPCSQSSTSPALSSKI 109
F SQSS+S LS+ +
Sbjct: 63 FPYNPNASQSSSSKLLSATL 82
>PTI6_LYCES (O04682) Pathogenesis-related genes transcriptional
activator PTI6
Length = 248
Score = 77.4 bits (189), Expect = 6e-14
Identities = 37/59 (62%), Positives = 44/59 (73%)
Query: 35 RKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNF 93
RK+F GVRQRP GRW AEI+D + RVWLGT+DT EEAA YD+AA L+G + TNF
Sbjct: 95 RKKFRGVRQRPWGRWAAEIRDPTRGKRVWLGTYDTPEEAAVVYDKAAVKLKGPDAVTNF 153
>DR2A_ARATH (O82132) Dehydration responsive element binding protein
2A (DREB2A protein)
Length = 335
Score = 72.0 bits (175), Expect = 3e-12
Identities = 41/80 (51%), Positives = 51/80 (63%), Gaps = 1/80 (1%)
Query: 29 GGARRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGAN 88
GG R F GVRQR G+WVAEI++ + R+WLGTF TA+EAA AYDEAA + G
Sbjct: 70 GGPENSRCSFRGVRQRIWGKWVAEIREPNRGSRLWLGTFPTAQEAASAYDEAAKAMYGPL 129
Query: 89 TRTNFWPCSQSSTSPALSSK 108
R NF P S +S + SS+
Sbjct: 130 ARLNF-PRSDASEVTSTSSQ 148
>ERF5_ARATH (O80341) Ethylene responsive element binding factor 5
(AtERF5)
Length = 300
Score = 70.9 bits (172), Expect = 6e-12
Identities = 37/60 (61%), Positives = 45/60 (74%), Gaps = 1/60 (1%)
Query: 35 RKRFVGVRQRPSGRWVAEIKDTIQK-IRVWLGTFDTAEEAARAYDEAACLLRGANTRTNF 93
+K + GVRQRP G++ AEI+D ++ RVWLGTFDTA EAARAYDEAA LRG+ NF
Sbjct: 153 KKHYRGVRQRPWGKFAAEIRDPNKRGSRVWLGTFDTAIEAARAYDEAAFRLRGSKAILNF 212
>ERF2_ARATH (O80338) Ethylene responsive element binding factor 2
(AtERF2)
Length = 243
Score = 70.9 bits (172), Expect = 6e-12
Identities = 47/114 (41%), Positives = 65/114 (56%), Gaps = 11/114 (9%)
Query: 36 KRFVGVRQRPSGRWVAEIKDTIQK-IRVWLGTFDTAEEAARAYDEAACLLRGANTRTNFW 94
K + GVRQRP G++ AEI+D + RVWLGTF+TAE+AA AYD AA +RG+ NF
Sbjct: 115 KHYRGVRQRPWGKFAAEIRDPAKNGARVWLGTFETAEDAALAYDIAAFRMRGSRALLNF- 173
Query: 95 PCSQSSTSPALSSKITNLLLQRLKERNMNNSCSSFSSSKYTSLLGSPEERREQQ 148
P +S P R+ + ++S SS SSS +S G + RR+ +
Sbjct: 174 PLRVNSGEPD---------PVRITSKRSSSSSSSSSSSTSSSENGKLKRRRKAE 218
>DR1B_ARATH (P93835) Dehydration responsive element binding protein
1B (DREB1B protein) (C-repeat binding factor 1)
(C-repeat/dehydration responsive element binding factor
1) (CRT/DRE binding factor 1)
Length = 213
Score = 69.3 bits (168), Expect = 2e-11
Identities = 32/54 (59%), Positives = 41/54 (75%)
Query: 40 GVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNF 93
GVRQR SG+WV+E+++ +K R+WLGTF TAE AARA+D AA LRG + NF
Sbjct: 50 GVRQRNSGKWVSEVREPNKKTRIWLGTFQTAEMAARAHDVAALALRGRSACLNF 103
>ERF1_ARATH (O80337) Ethylene responsive element binding factor 1
(AtERF1) (EREBP-2 protein)
Length = 268
Score = 68.9 bits (167), Expect = 2e-11
Identities = 48/111 (43%), Positives = 63/111 (56%), Gaps = 19/111 (17%)
Query: 36 KRFVGVRQRPSGRWVAEIKDTIQK-IRVWLGTFDTAEEAARAYDEAACLLRGANTRTNFW 94
K + GVRQRP G++ AEI+D + RVWLGTF+TAE+AA AYD AA +RG+ NF
Sbjct: 146 KHYRGVRQRPWGKFAAEIRDPAKNGARVWLGTFETAEDAALAYDRAAFRMRGSRALLNF- 204
Query: 95 PCSQSSTSPALSSKITNLLLQRLKERNMNNSCSSFSSSKYTSLLGSPEERR 145
P +S P R+K + SSFSSS G+P++RR
Sbjct: 205 PLRVNSGEPD---------PVRIKSKR-----SSFSSSNEN---GAPKKRR 238
>PTI5_LYCES (O04681) Pathogenesis-related genes transcriptional
activator PTI5
Length = 161
Score = 68.6 bits (166), Expect = 3e-11
Identities = 34/61 (55%), Positives = 45/61 (73%), Gaps = 1/61 (1%)
Query: 34 VRKRFVGVRQRPSGRWVAEIKDTIQK-IRVWLGTFDTAEEAARAYDEAACLLRGANTRTN 92
+ K++ GVR+RP G++ AEI+D+ + RVWLGTF+TAEEAA AYD AA +RGA N
Sbjct: 55 IGKKYRGVRRRPWGKYAAEIRDSARHGARVWLGTFETAEEAALAYDRAAFRMRGAKALLN 114
Query: 93 F 93
F
Sbjct: 115 F 115
>DR1C_ARATH (Q9SYS6) Dehydration responsive element binding protein
1C (DREB1C protein) (C-repeat binding factor 2)
(C-repeat/dehydration responsive element binding factor
2) (CRT/DRE binding factor 2)
Length = 216
Score = 68.6 bits (166), Expect = 3e-11
Identities = 32/54 (59%), Positives = 40/54 (73%)
Query: 40 GVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNF 93
GVRQR SG+WV E+++ +K R+WLGTF TAE AARA+D AA LRG + NF
Sbjct: 53 GVRQRNSGKWVCELREPNKKTRIWLGTFQTAEMAARAHDVAAIALRGRSACLNF 106
>DR2B_ARATH (O82133) Dehydration responsive element binding protein
2B (DREB2B protein)
Length = 330
Score = 68.2 bits (165), Expect = 4e-11
Identities = 40/80 (50%), Positives = 49/80 (61%), Gaps = 1/80 (1%)
Query: 29 GGARRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGAN 88
GG F GVRQR G+WVAEI++ R+WLGTF TAE+AA AYDEAA + G+
Sbjct: 69 GGPDNSHCSFRGVRQRIWGKWVAEIREPKIGTRLWLGTFPTAEKAASAYDEAATAMYGSL 128
Query: 89 TRTNFWPCSQSSTSPALSSK 108
R NF P S S + SS+
Sbjct: 129 ARLNF-PQSVGSEFTSTSSQ 147
>TINY_ARATH (Q39127) Transcriptional factor TINY
Length = 218
Score = 67.4 bits (163), Expect = 6e-11
Identities = 40/96 (41%), Positives = 58/96 (59%), Gaps = 1/96 (1%)
Query: 40 GVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNFWPCSQS 99
GVR+R G+WV+EI++ +K R+WLGTF + E AARA+D AA ++GA+ NF + S
Sbjct: 38 GVRKRNWGKWVSEIREPRKKSRIWLGTFPSPEMAARAHDVAALSIKGASAILNFPDLAGS 97
Query: 100 STSPALSSKITNLLLQRLKERNMNNSCSSFSSSKYT 135
P+ S ++ + LK +M S S SSS T
Sbjct: 98 FPRPS-SLSPRDIQVAALKAAHMETSQSFSSSSSLT 132
>DR1D_ARATH (Q9FJ93) Dehydration responsive element binding protein
1D (DREB1D protein) (C-repeat binding factor 4)
(C-repeat/dehydration responsive element binding factor
4) (CRT/DRE binding factor 4)
Length = 224
Score = 67.4 bits (163), Expect = 6e-11
Identities = 38/79 (48%), Positives = 48/79 (60%), Gaps = 8/79 (10%)
Query: 23 KEAASLGGARRVRKRFV--------GVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAA 74
K A+S R RK+F GVRQR SG+WV E+++ +K R+WLGTF T E AA
Sbjct: 31 KLASSCPKKRAGRKKFRETRHPIYRGVRQRNSGKWVCEVREPNKKSRIWLGTFPTVEMAA 90
Query: 75 RAYDEAACLLRGANTRTNF 93
RA+D AA LRG + NF
Sbjct: 91 RAHDVAALALRGRSACLNF 109
>DR1A_ARATH (Q9M0L0) Dehydration responsive element binding protein
1A (DREB1A protein) (C-repeat binding factor 3)
(C-repeat/dehydration responsive element binding factor
3) (CRT/DRE binding factor 3)
Length = 216
Score = 67.4 bits (163), Expect = 6e-11
Identities = 31/54 (57%), Positives = 40/54 (73%)
Query: 40 GVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNF 93
GVR+R SG+WV E+++ +K R+WLGTF TAE AARA+D AA LRG + NF
Sbjct: 53 GVRRRNSGKWVCEVREPNKKTRIWLGTFQTAEMAARAHDVAALALRGRSACLNF 106
>DR2C_ARATH (Q8LFR2) Dehydration responsive element binding protein
2C (DREB2C protein)
Length = 341
Score = 67.0 bits (162), Expect = 8e-11
Identities = 49/148 (33%), Positives = 74/148 (49%), Gaps = 12/148 (8%)
Query: 40 GVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNFWPCSQS 99
GVRQR G+WVAEI++ R+WLGTF ++ EAA AYDEAA + G + R N +
Sbjct: 74 GVRQRRWGKWVAEIREPDGGARLWLGTFSSSYEAALAYDEAAKAIYGQSARLNLPEITNR 133
Query: 100 STSPALSSKITNLLLQRLKE-----RNMNNSCSSFSSSKYTS------LLGSPEERREQQ 148
S+S A ++ ++ + E R N+ S F K LL S + +E+
Sbjct: 134 SSSTAATATVSGSVTAFSDESEVCAREDTNASSGFGQVKLEDCSDEYVLLDSSQCIKEEL 193
Query: 149 RANDHDRQEMQQQVD-AYGEVSKDFSID 175
+ + R+E V G+ SK ++D
Sbjct: 194 KGKEEVREEHNLAVGFGIGQDSKRETLD 221
>DR2G_ARATH (P61827) Putative dehydration responsive element
binding protein 2G (DREB2G protein)
Length = 307
Score = 65.1 bits (157), Expect = 3e-10
Identities = 33/64 (51%), Positives = 41/64 (63%)
Query: 29 GGARRVRKRFVGVRQRPSGRWVAEIKDTIQKIRVWLGTFDTAEEAARAYDEAACLLRGAN 88
GG F GVRQR G+WVAEI++ + R+WLGTF+T+ EAA AYDEAA L G
Sbjct: 25 GGPENATCTFRGVRQRTWGKWVAEIREPNRGTRLWLGTFNTSVEAAMAYDEAAKKLYGHE 84
Query: 89 TRTN 92
+ N
Sbjct: 85 AKLN 88
>DR2E_ARATH (O80917) Dehydration responsive element binding protein
2E (DREB2E protein)
Length = 244
Score = 65.1 bits (157), Expect = 3e-10
Identities = 45/118 (38%), Positives = 58/118 (49%), Gaps = 32/118 (27%)
Query: 3 RKRKVSEAVEEGSMVWDDMMKEAASLGGARRVRK-------------------RFVGVRQ 43
R+R+V E VE W++ L ARRV+ RF GVRQ
Sbjct: 21 RRRRVVEPVEATLQRWEE-----EGLARARRVQAKGSKKGCMRGKGGPENPVCRFRGVRQ 75
Query: 44 RPSGRWVAEIKDTI--------QKIRVWLGTFDTAEEAARAYDEAACLLRGANTRTNF 93
R G+WVAEI++ + + R+WLGTF TA EAA AYD AA ++ G R NF
Sbjct: 76 RVWGKWVAEIREPVSHRGANSSRSKRLWLGTFATAAEAALAYDRAASVMYGPYARLNF 133
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.311 0.127 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 47,312,581
Number of Sequences: 164201
Number of extensions: 2004002
Number of successful extensions: 8641
Number of sequences better than 10.0: 199
Number of HSP's better than 10.0 without gapping: 44
Number of HSP's successfully gapped in prelim test: 156
Number of HSP's that attempted gapping in prelim test: 8107
Number of HSP's gapped (non-prelim): 518
length of query: 411
length of database: 59,974,054
effective HSP length: 113
effective length of query: 298
effective length of database: 41,419,341
effective search space: 12342963618
effective search space used: 12342963618
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 67 (30.4 bits)
Medicago: description of AC119412.1


