

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119408.1 + phase: 0 /pseudo
(600 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
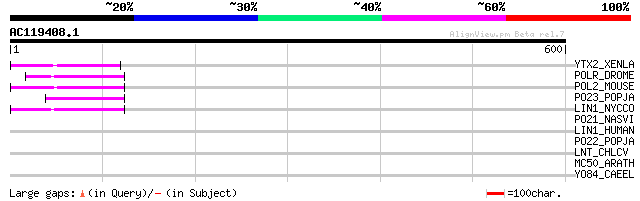
Score E
Sequences producing significant alignments: (bits) Value
YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein ... 56 2e-07
POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type... 50 2e-05
POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains... 49 4e-05
PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type... 47 1e-04
LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog 47 1e-04
PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type... 43 0.003
LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog 42 0.005
PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type... 39 0.039
LNT_CHLCV (Q824Q5) Apolipoprotein N-acyltransferase (EC 2.3.1.-)... 32 3.6
MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250... 32 4.8
YO84_CAEEL (P34620) Hypothetical protein ZK1236.4 in chromosome III 32 6.2
>YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein
(ORF 2)
Length = 1308
Score = 56.2 bits (134), Expect = 2e-07
Identities = 35/118 (29%), Positives = 56/118 (46%), Gaps = 3/118 (2%)
Query: 2 IANRIKYILPNITNDFQSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKA 61
I+ R+K +L + + QS VPGR I DN ++ + H+ R+T + + LD KA
Sbjct: 542 ISLRLKSVLAEVIHPDQSYTVPGRTIFDNVFLIRDLLHFARRTGLS---LAFLSLDQEKA 598
Query: 62 YDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDP 119
+D + ++ TL A F + + + IN T P + RG+RQG P
Sbjct: 599 FDRVDHQYLIGTLQAYSFGPQFVGYLKTMYASAECLVKINWSLTAPLAFGRGVRQGCP 656
>POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type II
retrotransposable element R2DM [Contains: Protease (EC
3.4.23.-); Reverse transcriptase (EC 2.7.7.49);
Endonuclease]
Length = 1057
Score = 50.1 bits (118), Expect = 2e-05
Identities = 36/108 (33%), Positives = 54/108 (49%), Gaps = 4/108 (3%)
Query: 18 QSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKAYDSLKWNFIEDTLTAM 77
Q F+P DN+ IV + K + Y+ LD++KA+DSL I DTL A
Sbjct: 446 QRGFLPTDGCADNATIVDLVLRHSHKHF--RSCYIA-NLDVSKAFDSLSHASIYDTLRAY 502
Query: 78 GFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDPF-PLIF 124
G P ++ + S+ +G +E F P RG++QGDP P++F
Sbjct: 503 GAPKGFVDYVQNTYEGGGTSLNGDGWSSEEFVPARGVKQGDPLSPILF 550
>POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains:
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1300
Score = 48.9 bits (115), Expect = 4e-05
Identities = 37/124 (29%), Positives = 55/124 (43%), Gaps = 3/124 (2%)
Query: 2 IANRIKYILPNITNDFQSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKA 61
+ANRI+ + I + Q F+PG N HYI K + N ++ I LD KA
Sbjct: 573 LANRIQEHIKAIIHPDQVGFIPGMQGWFNIRKSINVIHYINKLKDKN--HMIISLDAEKA 630
Query: 62 YDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDPF- 120
+D ++ F+ L G +N I +I +NG+ E + G RQG P
Sbjct: 631 FDKIQHPFMIKVLERSGIQGPYLNMIKAIYSKPVANIKVNGEKLEAIPLKSGTRQGCPLS 690
Query: 121 PLIF 124
P +F
Sbjct: 691 PYLF 694
>PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 606
Score = 47.4 bits (111), Expect = 1e-04
Identities = 27/87 (31%), Positives = 44/87 (50%), Gaps = 1/87 (1%)
Query: 39 HYIRKTRTNNNGYVGIKLDIAKAYDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSI 98
HYI+ R Y + LDI KA+D++ I + A G + + IM I +I
Sbjct: 11 HYIKLRRLKGKTYNVVSLDIRKAFDTVSHPAILRAMRAFGIDDGMQDFIMSTITDAYTNI 70
Query: 99 LINGQPTEPFSPQRGIRQGDPF-PLIF 124
++ G+ T + G++QGDP P++F
Sbjct: 71 VVGGRTTNKIYIRNGVKQGDPLSPVLF 97
>LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog
Length = 1260
Score = 47.4 bits (111), Expect = 1e-04
Identities = 34/124 (27%), Positives = 58/124 (46%), Gaps = 3/124 (2%)
Query: 2 IANRIKYILPNITNDFQSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKA 61
+ NRI+ + I + Q F+PG N +I K + N ++ + +D KA
Sbjct: 546 LTNRIQQHIKKIIHHDQVGFIPGSQGWFNIRKSINVIQHINKLK--NKDHMILSIDAEKA 603
Query: 62 YDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDPF- 120
+D+++ F+ TL +G + I T +I++NG + F + G RQG P
Sbjct: 604 FDNIQHPFMIRTLKKIGIEGTFLKLIEAIYSKPTANIILNGVKLKSFPLRSGTRQGCPLS 663
Query: 121 PLIF 124
PL+F
Sbjct: 664 PLLF 667
>PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1025
Score = 42.7 bits (99), Expect = 0.003
Identities = 28/85 (32%), Positives = 40/85 (46%), Gaps = 1/85 (1%)
Query: 41 IRKTRTNNNGYVGIKLDIAKAYDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILI 100
I++ R G LD+ KA+DS++ I D L P+ + N IM R + +
Sbjct: 440 IKEARMKIKGLYIAILDVKKAFDSVEHRSILDALRRKKLPLEMRNYIMWVYRNSKTRLEV 499
Query: 101 NGQPTEPFSPQRGIRQGDPF-PLIF 124
P RG+RQGDP PL+F
Sbjct: 500 VKTKGRWIRPARGVRQGDPLSPLLF 524
>LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog
Length = 1259
Score = 42.0 bits (97), Expect = 0.005
Identities = 32/123 (26%), Positives = 56/123 (45%), Gaps = 3/123 (2%)
Query: 2 IANRIKYILPNITNDFQSTFVPGRLITDNSLIVFETFHYIRKTRTNNNGYVGIKLDIAKA 61
+AN+I+ + + + Q F+P N +I +T+ N ++ I +D KA
Sbjct: 546 LANQIQQHIKKLIHHDQVGFIPAMQGWFNIRKSINIIQHINRTKDTN--HMIISIDAEKA 603
Query: 62 YDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDPF- 120
+D ++ F+ L +G + I T +I++NGQ E + G RQG P
Sbjct: 604 FDKIQQPFMLKPLNKLGIDGTYLKIIRAIYDKPTANIILNGQKLEAPPLKTGTRQGCPLS 663
Query: 121 PLI 123
PL+
Sbjct: 664 PLL 666
>PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 711
Score = 38.9 bits (89), Expect = 0.039
Identities = 26/87 (29%), Positives = 40/87 (45%), Gaps = 2/87 (2%)
Query: 40 YIRKTRTNNNGYVGIKLDIAKAYDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSIL 99
YI R Y + LD+ KA+D++ + I L +G N I + T +I
Sbjct: 126 YISSRREQRKTYNVVSLDVRKAFDTVSHSSICRALQRLGIDEGTSNYITGSLSDSTTTIR 185
Query: 100 IN-GQPTEPFSPQRGIRQGDPF-PLIF 124
+ G T +RG++QGDP P +F
Sbjct: 186 VGPGSQTRKICIRRGVKQGDPLSPFLF 212
>LNT_CHLCV (Q824Q5) Apolipoprotein N-acyltransferase (EC 2.3.1.-)
(ALP N-acyltransferase)
Length = 541
Score = 32.3 bits (72), Expect = 3.6
Identities = 17/78 (21%), Positives = 37/78 (46%), Gaps = 6/78 (7%)
Query: 446 LTWTSLVG------ITVFSIHISCLLTKIK*NVSLLLIWPMMMRLCGILIKMANIRCTVV 499
L+W L +TV+ +H S +L+ + + ++W +++ L +L + +
Sbjct: 49 LSWRQLTSLLFLWSVTVYGVHFSWMLSDLYVGKFIYVVWGVLISLLALLFTAFSCLLFFI 108
Query: 500 IKLYRTEKLLLLPSLLIA 517
++ T+ L LP L +A
Sbjct: 109 VRKKHTKILWCLPGLWVA 126
>MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250
(ORF102)
Length = 122
Score = 32.0 bits (71), Expect = 4.8
Identities = 14/28 (50%), Positives = 19/28 (67%), Gaps = 1/28 (3%)
Query: 99 LINGQPTEPFSPQRGIRQGDPF-PLIFL 125
+ING P +P RG+RQGDP P +F+
Sbjct: 13 IINGAPQGLVTPSRGLRQGDPLSPYLFI 40
>YO84_CAEEL (P34620) Hypothetical protein ZK1236.4 in chromosome III
Length = 364
Score = 31.6 bits (70), Expect = 6.2
Identities = 19/74 (25%), Positives = 38/74 (50%), Gaps = 1/74 (1%)
Query: 56 LDIAKAYDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFSPQRGIR 115
LD +KA+D + + + D LT++ +LI + + + +F + + +EP G+
Sbjct: 23 LDFSKAFDKVSHDILLDKLTSIKINKHLIRWLDVFLTNRSFKVKVGNTLSEPKKTVCGVP 82
Query: 116 QGDPF-PLIFLYYV 128
QG P++F +V
Sbjct: 83 QGSVISPVLFGIFV 96
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.344 0.154 0.513
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 64,164,795
Number of Sequences: 164201
Number of extensions: 2497695
Number of successful extensions: 7946
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 7930
Number of HSP's gapped (non-prelim): 16
length of query: 600
length of database: 59,974,054
effective HSP length: 116
effective length of query: 484
effective length of database: 40,926,738
effective search space: 19808541192
effective search space used: 19808541192
T: 11
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.6 bits)
S2: 69 (31.2 bits)
Medicago: description of AC119408.1


