

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149581.9 - phase: 0 /pseudo
(329 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
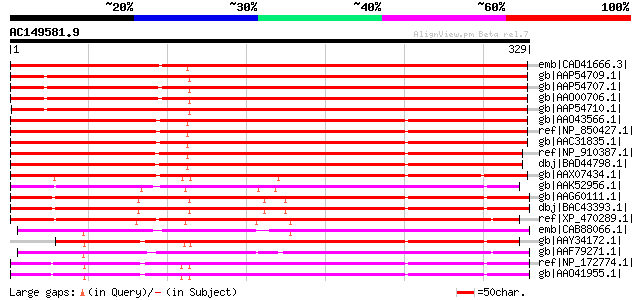
Score E
Sequences producing significant alignments: (bits) Value
emb|CAD41666.3| OSJNBa0019K04.13 [Oryza sativa (japonica cultiva... 372 e-102
gb|AAP54709.1| cytochrome P450-like protein [Oryza sativa (japon... 367 e-100
gb|AAP54707.1| cytochrome P450-like protein [Oryza sativa (japon... 367 e-100
gb|AAO00706.1| putative cytochrome P450-dependent fatty acid hyd... 367 e-100
gb|AAP54710.1| cytochrome P450-like protein [Oryza sativa (japon... 358 1e-97
gb|AAO43566.1| At2g45510 [Arabidopsis thaliana] gi|2979544|gb|AA... 328 1e-88
ref|NP_850427.1| cytochrome P450 family protein [Arabidopsis tha... 326 7e-88
gb|AAC31835.1| putative cytochrome P450 [Arabidopsis thaliana] g... 326 7e-88
ref|NP_910387.1| ESTs AU056036(S20239),C72753(E2173), AU056035(S... 324 2e-87
dbj|BAD44798.1| putative cytochrome P450 [Oryza sativa (japonica... 324 2e-87
gb|AAX07434.1| cytochrome P450 CYPD [Pinus taeda] 301 2e-80
gb|AAK52956.1| cytochrome P450-like protein [Zea mays] 261 2e-68
gb|AAG60111.1| cytochrome P450, putative [Arabidopsis thaliana] 259 1e-67
dbj|BAC43393.1| unknown protein [Arabidopsis thaliana] gi|306978... 259 1e-67
ref|XP_470289.1| putative plant cytochrome P-450 protein [Oryza ... 257 3e-67
emb|CAB88066.1| cytochrome P450-like protein [Arabidopsis thalia... 235 2e-60
gb|AAY34172.1| At1g13140 [Arabidopsis thaliana] 224 2e-57
gb|AAF79271.1| F12K21.15 [Arabidopsis thaliana] gi|15218671|ref|... 222 1e-56
ref|NP_172774.1| cytochrome P450, putative [Arabidopsis thaliana... 219 9e-56
gb|AAO41955.1| putative cytochrome P450 [Arabidopsis thaliana] 218 2e-55
>emb|CAD41666.3| OSJNBa0019K04.13 [Oryza sativa (japonica cultivar-group)]
gi|50928103|ref|XP_473579.1| OSJNBa0019K04.13 [Oryza
sativa (japonica cultivar-group)]
Length = 506
Score = 372 bits (956), Expect = e-102
Identities = 183/333 (54%), Positives = 252/333 (74%), Gaps = 6/333 (1%)
Query: 1 MKSTLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWYVNFLWKVQRFLNIGS 60
M++TLDS F+V GV L + G+SKEG F+ AF++AS ++F + + LWKV+RFLNI S
Sbjct: 175 MRATLDSFFRVGFGVNLGVLSGSSKEGLVFARAFDDASEQVLFRFFDLLWKVKRFLNISS 234
Query: 61 EAVLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLELNETD-----SKYL 115
EA +K+++R IN++VY+II KIEQ + Q + K DILSRFL E D +KY+
Sbjct: 235 EATMKQSIRTINDFVYSIIDRKIEQMSREQHEFAK-KEDILSRFLLEREKDPGCFDNKYI 293
Query: 116 KDVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATKVEDGSTTDELATRIT 175
+D+IL+F+IAG+DTT+ TLS F+Y +CK+ VQ+KIA+E+R+AT + + ++ +T
Sbjct: 294 RDIILNFVIAGRDTTAGTLSWFLYAVCKNQRVQDKIAREVRDATTGDRDVGVQDFSSFLT 353
Query: 176 EESMKKMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPYVMGRME 235
E+++ KMQYL AALTETLRL+P +P++ KYCFSDDTLPDG++V+KGD +++ PY MGRM+
Sbjct: 354 EDAINKMQYLHAALTETLRLYPGVPLDVKYCFSDDTLPDGHAVKKGDMVNYQPYPMGRMK 413
Query: 236 FLWGEDAEQFRPERWLDENGNFQRESPFKFTAFQAGPRICLGKEFVYRQMKIFSAVLLGS 295
FLWG++AE+F+PERWLD++G F ESPFKFTAFQAGPRICLGKEF YRQMKI SAVLL
Sbjct: 414 FLWGDNAEEFKPERWLDDSGMFVAESPFKFTAFQAGPRICLGKEFAYRQMKIVSAVLLYF 473
Query: 296 HNFKLADQNRLVKYRTTLTLQIDGGLHVNAFHR 328
F++ D + V YR LTL++DG ++ A R
Sbjct: 474 FRFEMWDDDATVGYRPMLTLKMDGPFYLRALAR 506
>gb|AAP54709.1| cytochrome P450-like protein [Oryza sativa (japonica
cultivar-group)] gi|37536240|ref|NP_922422.1| cytochrome
P450-like protein [Oryza sativa (japonica
cultivar-group)] gi|20146747|gb|AAM12483.1| cytochrome
P450-like protein [Oryza sativa (japonica
cultivar-group)]
Length = 515
Score = 367 bits (943), Expect = e-100
Identities = 173/333 (51%), Positives = 246/333 (72%), Gaps = 6/333 (1%)
Query: 1 MKSTLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWYVNFLWKVQRFLNIGS 60
M++T+DS+F + G +L+T+ G S EG +F+ AF++AS M Y+N WK+ R LN+G+
Sbjct: 184 MRATMDSIFTIAFGQDLNTLDG-SGEGRRFAAAFDDASEFTMLRYLNPFWKLSRLLNVGA 242
Query: 61 EAVLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLELNETDS-----KYL 115
EA+LK+ ++V++ +VY +I+ + ++ + + + + DIL+RF++ +DS KYL
Sbjct: 243 EAMLKERIKVVDGFVYKLIRDRSDELSNTKAHDTDSRQDILTRFIQATTSDSGTVDYKYL 302
Query: 116 KDVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATKVEDGSTTDELATRIT 175
+D+IL+ +IAGKDTT+ +L+ F+Y +CKHP VQEKI E EAT + ++ DE + +T
Sbjct: 303 RDIILNIVIAGKDTTAGSLAWFLYMMCKHPEVQEKICHEAMEATNAGEAASIDEFSQSLT 362
Query: 176 EESMKKMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPYVMGRME 235
+E++ KM YL AALTETLRL+P +P+++K CFSDD LP+G++V KGD + + PY MGRME
Sbjct: 363 DEALNKMHYLHAALTETLRLYPAVPLDNKQCFSDDVLPNGFNVSKGDIVFYIPYAMGRME 422
Query: 236 FLWGEDAEQFRPERWLDENGNFQRESPFKFTAFQAGPRICLGKEFVYRQMKIFSAVLLGS 295
LWG+DAE FRPERWLDENG FQ+ESPFKFTAFQAGPRICLGK+F YRQMKIF+AVLL
Sbjct: 423 SLWGKDAESFRPERWLDENGVFQQESPFKFTAFQAGPRICLGKDFAYRQMKIFAAVLLRF 482
Query: 296 HNFKLADQNRLVKYRTTLTLQIDGGLHVNAFHR 328
KL D+ ++ YRT +TL +D GLH+ A R
Sbjct: 483 FVLKLRDEKEIISYRTMITLSVDQGLHLTAMAR 515
>gb|AAP54707.1| cytochrome P450-like protein [Oryza sativa (japonica
cultivar-group)] gi|37536236|ref|NP_922420.1| cytochrome
P450-like protein [Oryza sativa (japonica
cultivar-group)] gi|20146758|gb|AAM12494.1| cytochrome
P450-like protein [Oryza sativa (japonica
cultivar-group)]
Length = 495
Score = 367 bits (943), Expect = e-100
Identities = 181/332 (54%), Positives = 244/332 (72%), Gaps = 7/332 (2%)
Query: 1 MKSTLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWYVNFLWKVQRFLNIGS 60
+++T+DS+F + G +L+T+ G S EG++F+ AF++AS M Y++ LWK+ R LN+G
Sbjct: 167 LRATMDSIFTIAFGTDLNTLDG-SGEGSRFAAAFDDASEFTMLRYISPLWKLARLLNVGV 225
Query: 61 EAVLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLELNETDS----KYLK 116
EA+LK+ ++V++E+VY +I+++ ++ + + DILSRFL+ +DS KYL+
Sbjct: 226 EAMLKERIKVVDEFVYRLIRARSDELSNSHDSGS--RQDILSRFLQATTSDSVVDYKYLR 283
Query: 117 DVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATKVEDGSTTDELATRITE 176
D+IL+ +IAGKDTT+ L+ F+Y +CKHP VQEKI E AT D ++ DE +T+
Sbjct: 284 DIILNIVIAGKDTTAGALAWFLYMVCKHPEVQEKICHEAMVATSAGDTASVDEFLQSLTD 343
Query: 177 ESMKKMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPYVMGRMEF 236
+++ M YL AALTETLRL+P +PME+K CFSDD LP+G++V KGD + F PY MGRME
Sbjct: 344 QALNNMHYLHAALTETLRLYPSVPMENKQCFSDDVLPNGFNVSKGDIVFFIPYAMGRMES 403
Query: 237 LWGEDAEQFRPERWLDENGNFQRESPFKFTAFQAGPRICLGKEFVYRQMKIFSAVLLGSH 296
LWG+DAE FRPERWLDENG FQ+ESPFKFTAFQAGPRICLGKEF YRQMKIF+AVLL
Sbjct: 404 LWGKDAEYFRPERWLDENGVFQQESPFKFTAFQAGPRICLGKEFAYRQMKIFAAVLLRFF 463
Query: 297 NFKLADQNRLVKYRTTLTLQIDGGLHVNAFHR 328
KL D+ +V YRTTLTL ID GLH+ A R
Sbjct: 464 VLKLRDEKEIVSYRTTLTLAIDQGLHLTATAR 495
>gb|AAO00706.1| putative cytochrome P450-dependent fatty acid hydroxylase,
5'-partial [Oryza sativa (japonica cultivar-group)]
Length = 404
Score = 367 bits (943), Expect = e-100
Identities = 181/332 (54%), Positives = 244/332 (72%), Gaps = 7/332 (2%)
Query: 1 MKSTLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWYVNFLWKVQRFLNIGS 60
+++T+DS+F + G +L+T+ G S EG++F+ AF++AS M Y++ LWK+ R LN+G
Sbjct: 76 LRATMDSIFTIAFGTDLNTLDG-SGEGSRFAAAFDDASEFTMLRYISPLWKLARLLNVGV 134
Query: 61 EAVLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLELNETDS----KYLK 116
EA+LK+ ++V++E+VY +I+++ ++ + + DILSRFL+ +DS KYL+
Sbjct: 135 EAMLKERIKVVDEFVYRLIRARSDELSNSHDSGS--RQDILSRFLQATTSDSVVDYKYLR 192
Query: 117 DVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATKVEDGSTTDELATRITE 176
D+IL+ +IAGKDTT+ L+ F+Y +CKHP VQEKI E AT D ++ DE +T+
Sbjct: 193 DIILNIVIAGKDTTAGALAWFLYMVCKHPEVQEKICHEAMVATSAGDTASVDEFLQSLTD 252
Query: 177 ESMKKMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPYVMGRMEF 236
+++ M YL AALTETLRL+P +PME+K CFSDD LP+G++V KGD + F PY MGRME
Sbjct: 253 QALNNMHYLHAALTETLRLYPSVPMENKQCFSDDVLPNGFNVSKGDIVFFIPYAMGRMES 312
Query: 237 LWGEDAEQFRPERWLDENGNFQRESPFKFTAFQAGPRICLGKEFVYRQMKIFSAVLLGSH 296
LWG+DAE FRPERWLDENG FQ+ESPFKFTAFQAGPRICLGKEF YRQMKIF+AVLL
Sbjct: 313 LWGKDAEYFRPERWLDENGVFQQESPFKFTAFQAGPRICLGKEFAYRQMKIFAAVLLRFF 372
Query: 297 NFKLADQNRLVKYRTTLTLQIDGGLHVNAFHR 328
KL D+ +V YRTTLTL ID GLH+ A R
Sbjct: 373 VLKLRDEKEIVSYRTTLTLAIDQGLHLTATAR 404
>gb|AAP54710.1| cytochrome P450-like protein [Oryza sativa (japonica
cultivar-group)] gi|37536242|ref|NP_922423.1| cytochrome
P450-like protein [Oryza sativa (japonica
cultivar-group)] gi|20146744|gb|AAM12480.1| cytochrome
P450-like protein [Oryza sativa (japonica
cultivar-group)]
Length = 516
Score = 358 bits (919), Expect = 1e-97
Identities = 171/332 (51%), Positives = 240/332 (71%), Gaps = 6/332 (1%)
Query: 2 KSTLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWYVNFLWKVQRFLNIGSE 61
++T+DS+F + G +L+T+ G S EG F+ AF++A ++ Y+N WK+ R LN+G+E
Sbjct: 186 RATMDSIFTIAFGQDLNTLDG-SGEGRHFAKAFDDAGEYLLLRYLNPFWKLARLLNVGAE 244
Query: 62 AVLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLELNETDS-----KYLK 116
A LK+ ++V++E+VY +I+++ ++ + D+LSRF++ +DS KYL+
Sbjct: 245 ATLKERIKVVDEFVYKLIRARSDELSNTMAQDHRSRDDLLSRFIQATTSDSGTVDYKYLR 304
Query: 117 DVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATKVEDGSTTDELATRITE 176
D++L+ +IA KD+TS +L+ F+Y CK P VQEKI E+ E T D ++ DE T +T+
Sbjct: 305 DIVLNIVIAAKDSTSGSLAWFLYMACKRPEVQEKIFDEVMETTNAGDCASIDEFLTSLTD 364
Query: 177 ESMKKMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPYVMGRMEF 236
+++ KM YL AALTETLRL+P +P+E+K CFSDD LP+G+SV KGD + + PY MGRMEF
Sbjct: 365 QALNKMHYLHAALTETLRLYPSVPLENKQCFSDDVLPNGFSVSKGDGVFYMPYAMGRMEF 424
Query: 237 LWGEDAEQFRPERWLDENGNFQRESPFKFTAFQAGPRICLGKEFVYRQMKIFSAVLLGSH 296
LWG+DAE FRPERWLDE+G FQ+ESPFKFTAFQAGPRIC+GK+F YRQMKIF+AVL+ S
Sbjct: 425 LWGKDAEAFRPERWLDEHGVFQQESPFKFTAFQAGPRICIGKDFAYRQMKIFAAVLIRSF 484
Query: 297 NFKLADQNRLVKYRTTLTLQIDGGLHVNAFHR 328
FKL D+ V YRT +TL ID LH+ A R
Sbjct: 485 VFKLRDKKDNVSYRTAITLAIDQDLHLTATAR 516
>gb|AAO43566.1| At2g45510 [Arabidopsis thaliana] gi|2979544|gb|AAC06153.1| putative
cytochrome P450 [Arabidopsis thaliana]
gi|15225499|ref|NP_182075.1| cytochrome P450, putative
[Arabidopsis thaliana] gi|7430723|pir||T00864 cytochrome
P450 homolog F17K2.4 - Arabidopsis thaliana
Length = 511
Score = 328 bits (841), Expect = 1e-88
Identities = 171/334 (51%), Positives = 234/334 (69%), Gaps = 9/334 (2%)
Query: 1 MKSTLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWYVNFLWKVQRFLNIGS 60
M+ TLDS+FKV GVEL + G SKEG +F AF+E + +++ LWK++ F NIGS
Sbjct: 178 MRCTLDSIFKVGFGVELKCLDGFSKEGQEFMEAFDEGNVATSSRFIDPLWKLKWFFNIGS 237
Query: 61 EAVLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLELNETD-----SKYL 115
++ LKK++ I+++VY++I +K ++ K Q ++ DILSRFL +E D KYL
Sbjct: 238 QSKLKKSIATIDKFVYSLITTKRKELAKEQNTV--VREDILSRFLVESEKDPENMNDKYL 295
Query: 116 KDVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATKVEDGST-TDELATRI 174
+D+IL+F+IAGKDTT+ LS F+Y LCK+P VQEKI QEIR+ T + +T + I
Sbjct: 296 RDIILNFMIAGKDTTAALLSWFLYMLCKNPLVQEKIVQEIRDVTFSHEKTTDVNGFVESI 355
Query: 175 TEESMKKMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPYVMGRM 234
EE++ +M YL AAL+ETLRL+PP+P++ + +DD LPDG+ V KGD I + Y MGRM
Sbjct: 356 NEEALDEMHYLHAALSETLRLYPPVPVDMRCAENDDVLPDGHRVSKGDNIYYIAYAMGRM 415
Query: 235 EFLWGEDAEQFRPERWLDENGNFQRESPFKFTAFQAGPRICLGKEFVYRQMKIFSAVLLG 294
++WG+DAE+F+PERWL ++G FQ ESPFKF +F AGPRICLGK+F YRQMKI S LL
Sbjct: 416 TYIWGQDAEEFKPERWL-KDGLFQPESPFKFISFHAGPRICLGKDFAYRQMKIVSMALLH 474
Query: 295 SHNFKLADQNRLVKYRTTLTLQIDGGLHVNAFHR 328
FK+AD+N V Y+ LTL +DGGLH+ A R
Sbjct: 475 FFRFKMADENSKVYYKRMLTLHVDGGLHLCAIPR 508
>ref|NP_850427.1| cytochrome P450 family protein [Arabidopsis thaliana]
Length = 505
Score = 326 bits (835), Expect = 7e-88
Identities = 169/334 (50%), Positives = 233/334 (69%), Gaps = 9/334 (2%)
Query: 1 MKSTLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWYVNFLWKVQRFLNIGS 60
MK TLDS+FKV GVEL + G SKEG +F AF+E + + WK++ FLNIGS
Sbjct: 172 MKCTLDSIFKVGFGVELGCLDGFSKEGEEFMKAFDEGNGATSSRVTDPFWKLKCFLNIGS 231
Query: 61 EAVLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLELNETD-----SKYL 115
E+ LKK++ +I+++VY++I +K ++ K Q S ++ DILS+FL +E D KYL
Sbjct: 232 ESRLKKSIAIIDKFVYSLITTKRKELSKEQNTS--VREDILSKFLLESEKDPENMNDKYL 289
Query: 116 KDVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATKVEDGST-TDELATRI 174
+D+IL+ ++AGKDTT+ +LS F+Y LCK+P VQEKI QEIR+ T + +T + +
Sbjct: 290 RDIILNVMVAGKDTTAASLSWFLYMLCKNPLVQEKIVQEIRDVTSSHEKTTDVNGFIESV 349
Query: 175 TEESMKKMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPYVMGRM 234
TEE++ +MQYL AAL+ET+RL+PP+P + +DD LPDG+ V KGD I + Y MGRM
Sbjct: 350 TEEALAQMQYLHAALSETMRLYPPVPEHMRCAENDDVLPDGHRVSKGDNIYYISYAMGRM 409
Query: 235 EFLWGEDAEQFRPERWLDENGNFQRESPFKFTAFQAGPRICLGKEFVYRQMKIFSAVLLG 294
++WG+DAE+F+PERWL ++G FQ ES FKF +F AGPRIC+GK+F YRQMKI S LL
Sbjct: 410 TYIWGQDAEEFKPERWL-KDGVFQPESQFKFISFHAGPRICIGKDFAYRQMKIVSMALLH 468
Query: 295 SHNFKLADQNRLVKYRTTLTLQIDGGLHVNAFHR 328
FK+AD+N V Y+ LTL +DGGLH+ A R
Sbjct: 469 FFRFKMADENSKVSYKKMLTLHVDGGLHLCAIPR 502
>gb|AAC31835.1| putative cytochrome P450 [Arabidopsis thaliana]
gi|7430726|pir||T00404 probable cytochrome P450
At2g44890 [imported] - Arabidopsis thaliana
Length = 490
Score = 326 bits (835), Expect = 7e-88
Identities = 169/334 (50%), Positives = 233/334 (69%), Gaps = 9/334 (2%)
Query: 1 MKSTLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWYVNFLWKVQRFLNIGS 60
MK TLDS+FKV GVEL + G SKEG +F AF+E + + WK++ FLNIGS
Sbjct: 157 MKCTLDSIFKVGFGVELGCLDGFSKEGEEFMKAFDEGNGATSSRVTDPFWKLKCFLNIGS 216
Query: 61 EAVLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLELNETD-----SKYL 115
E+ LKK++ +I+++VY++I +K ++ K Q S ++ DILS+FL +E D KYL
Sbjct: 217 ESRLKKSIAIIDKFVYSLITTKRKELSKEQNTS--VREDILSKFLLESEKDPENMNDKYL 274
Query: 116 KDVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATKVEDGST-TDELATRI 174
+D+IL+ ++AGKDTT+ +LS F+Y LCK+P VQEKI QEIR+ T + +T + +
Sbjct: 275 RDIILNVMVAGKDTTAASLSWFLYMLCKNPLVQEKIVQEIRDVTSSHEKTTDVNGFIESV 334
Query: 175 TEESMKKMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPYVMGRM 234
TEE++ +MQYL AAL+ET+RL+PP+P + +DD LPDG+ V KGD I + Y MGRM
Sbjct: 335 TEEALAQMQYLHAALSETMRLYPPVPEHMRCAENDDVLPDGHRVSKGDNIYYISYAMGRM 394
Query: 235 EFLWGEDAEQFRPERWLDENGNFQRESPFKFTAFQAGPRICLGKEFVYRQMKIFSAVLLG 294
++WG+DAE+F+PERWL ++G FQ ES FKF +F AGPRIC+GK+F YRQMKI S LL
Sbjct: 395 TYIWGQDAEEFKPERWL-KDGVFQPESQFKFISFHAGPRICIGKDFAYRQMKIVSMALLH 453
Query: 295 SHNFKLADQNRLVKYRTTLTLQIDGGLHVNAFHR 328
FK+AD+N V Y+ LTL +DGGLH+ A R
Sbjct: 454 FFRFKMADENSKVSYKKMLTLHVDGGLHLCAIPR 487
>ref|NP_910387.1| ESTs AU056036(S20239),C72753(E2173), AU056035(S20239) correspond to
a region of the predicted gene.~Similar to putative
cytochrome P-450 (AC003680) [Oryza sativa (japonica
cultivar-group)]
Length = 403
Score = 324 bits (831), Expect = 2e-87
Identities = 163/331 (49%), Positives = 236/331 (71%), Gaps = 9/331 (2%)
Query: 1 MKSTLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWYVNFLWKVQRFLNIGS 60
M++T+DS+FKV G EL+T+ G+ + G QFS AF+EA++ + + +V+ +WK++R+LNIGS
Sbjct: 69 MRATMDSIFKVGFGFELNTLSGSDESGIQFSKAFDEANSLVYYRFVDIMWKLKRYLNIGS 128
Query: 61 EAVLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLELNETD-----SKYL 115
EA LK+N+++I+ +V +I K EQ + + K DILSRF+ +E D +YL
Sbjct: 129 EAKLKRNIQIIDSFVMKLIHQKREQMKIAADY--KTKEDILSRFVLASEQDPGTMDDRYL 186
Query: 116 KDVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATK-VEDGSTTDELATRI 174
+D++L+F+IAGKDTT TL+ F Y LCK+P VQ+K+A EIRE + ++ +T + R+
Sbjct: 187 RDIVLNFLIAGKDTTGNTLTWFFYLLCKNPIVQDKVALEIREFVEWSKEDNTIESFTKRL 246
Query: 175 TEESMKKMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPYVMGRM 234
E ++ KM YL A ++ETLRL+P +P+++K DD LP+GY V KGD I++ Y MGRM
Sbjct: 247 DEGAISKMHYLQATISETLRLYPAVPVDAKMADEDDVLPNGYRVVKGDGINYMIYAMGRM 306
Query: 235 EFLWGEDAEQFRPERWLDENGNFQRESPFKFTAFQAGPRICLGKEFVYRQMKIFSAVLLG 294
+LWGEDA++FRPERWL NG +Q+ESPFKF +F AGPRICLGKEF +RQMKI +A L+
Sbjct: 307 TYLWGEDAQEFRPERWL-VNGVYQQESPFKFVSFNAGPRICLGKEFAHRQMKIMAATLIH 365
Query: 295 SHNFKLADQNRLVKYRTTLTLQIDGGLHVNA 325
F+L D+++ Y+T TL ID GLH+ A
Sbjct: 366 FFKFRLEDESKEPIYKTMFTLHIDNGLHLLA 396
>dbj|BAD44798.1| putative cytochrome P450 [Oryza sativa (japonica cultivar-group)]
Length = 525
Score = 324 bits (831), Expect = 2e-87
Identities = 163/331 (49%), Positives = 236/331 (71%), Gaps = 9/331 (2%)
Query: 1 MKSTLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWYVNFLWKVQRFLNIGS 60
M++T+DS+FKV G EL+T+ G+ + G QFS AF+EA++ + + +V+ +WK++R+LNIGS
Sbjct: 191 MRATMDSIFKVGFGFELNTLSGSDESGIQFSKAFDEANSLVYYRFVDIMWKLKRYLNIGS 250
Query: 61 EAVLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLELNETD-----SKYL 115
EA LK+N+++I+ +V +I K EQ + + K DILSRF+ +E D +YL
Sbjct: 251 EAKLKRNIQIIDSFVMKLIHQKREQMKIAADY--KTKEDILSRFVLASEQDPGTMDDRYL 308
Query: 116 KDVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATK-VEDGSTTDELATRI 174
+D++L+F+IAGKDTT TL+ F Y LCK+P VQ+K+A EIRE + ++ +T + R+
Sbjct: 309 RDIVLNFLIAGKDTTGNTLTWFFYLLCKNPIVQDKVALEIREFVEWSKEDNTIESFTKRL 368
Query: 175 TEESMKKMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPYVMGRM 234
E ++ KM YL A ++ETLRL+P +P+++K DD LP+GY V KGD I++ Y MGRM
Sbjct: 369 DEGAISKMHYLQATISETLRLYPAVPVDAKMADEDDVLPNGYRVVKGDGINYMIYAMGRM 428
Query: 235 EFLWGEDAEQFRPERWLDENGNFQRESPFKFTAFQAGPRICLGKEFVYRQMKIFSAVLLG 294
+LWGEDA++FRPERWL NG +Q+ESPFKF +F AGPRICLGKEF +RQMKI +A L+
Sbjct: 429 TYLWGEDAQEFRPERWL-VNGVYQQESPFKFVSFNAGPRICLGKEFAHRQMKIMAATLIH 487
Query: 295 SHNFKLADQNRLVKYRTTLTLQIDGGLHVNA 325
F+L D+++ Y+T TL ID GLH+ A
Sbjct: 488 FFKFRLEDESKEPIYKTMFTLHIDNGLHLLA 518
>gb|AAX07434.1| cytochrome P450 CYPD [Pinus taeda]
Length = 518
Score = 301 bits (770), Expect = 2e-80
Identities = 156/345 (45%), Positives = 241/345 (69%), Gaps = 21/345 (6%)
Query: 1 MKSTLDSVFKVILGVELDTMCGTSKEG---TQFSNAFNEASATIMFWY-VNFLWKVQRFL 56
M+S+LDS+ KV+ G++++++ + E F+ AF+ A+A + + V WKVQRF
Sbjct: 174 MRSSLDSICKVVFGIDINSLSSSKAESGPEASFAKAFDVANAMVFHRHMVGSFWKVQRFF 233
Query: 57 NIGSEAVLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLELN--ETDSK- 113
N+GSEA+L+ N++++++++Y +I + ++ +K + ++ DILSR++ ++ ETD K
Sbjct: 234 NVGSEAILRDNIKMVDDFLYKVIHFRRQEMFSAEKEN--VRPDILSRYIIISDKETDGKV 291
Query: 114 ---YLKDVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATKVEDGSTTDE- 169
YL+DVIL+F++A +DTT+I LS FIY LCKH HVQEK+ +EI +T V + + E
Sbjct: 292 SDKYLRDVILNFMVAARDTTAIALSWFIYMLCKHQHVQEKLLEEIISSTSVHEDQYSTEC 351
Query: 170 -----LATRITEESMKKMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYI 224
A +T+E++ KM YL A+L+ETLRL+P +P++ KY ++DTLPDG+ V+KGD +
Sbjct: 352 NDIASFAQSLTDEALGKMHYLHASLSETLRLYPALPVDGKYVVNEDTLPDGFKVKKGDSV 411
Query: 225 SFHPYVMGRMEFLWGEDAEQFRPERWLDENGNFQRESPFKFTAFQAGPRICLGKEFVYRQ 284
+F PY MGRM +LWG+DA++F+PERW+ ++G F +SPFKF AFQAGPR CLGK+F Y Q
Sbjct: 412 NFLPYAMGRMSYLWGDDAKEFKPERWI-QDGIFHPKSPFKFPAFQAGPRTCLGKDFAYLQ 470
Query: 285 MKIFSAVLLGSHNFKLADQNRLVKYRTTLTLQI-DGGLHVNAFHR 328
MKI +AVL+ F+ A + + V+YRT LTL + + GL+V R
Sbjct: 471 MKIVAAVLVRFFKFE-AVKTKEVRYRTMLTLHMNEDGLNVQVTPR 514
>gb|AAK52956.1| cytochrome P450-like protein [Zea mays]
Length = 543
Score = 261 bits (667), Expect = 2e-68
Identities = 146/357 (40%), Positives = 217/357 (59%), Gaps = 40/357 (11%)
Query: 1 MKSTLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWYVNFLWKVQRFLNIGS 60
M+ TLDS+ KV GVE+ T+ E + F+ AF+ A+ I +++ LW+++RF ++GS
Sbjct: 187 MRMTLDSICKVGFGVEIGTLSPDLPENS-FAQAFDAANIIITLRFIDPLWRIKRFFHVGS 245
Query: 61 EAVLKKNLRVINEYVYTIIKSK------IEQSQKPQKNSPELKGDILSRFLELNET---- 110
EA+L +++++++E+ Y++I+ + + S K +K +K DILSRF+EL E
Sbjct: 246 EALLAQSIKLVDEFTYSVIRRRKAEIVEVRASGKQEK----MKHDILSRFIELGEAGDDG 301
Query: 111 ----DSKYLKDVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIR--EATKVEDG 164
D K L+DV+L+F+IAG+DTT+ TLS F + HP V EK+ +E+ EA + +
Sbjct: 302 GGFGDDKSLRDVVLNFVIAGRDTTATTLSWFTHMAMSHPDVAEKLRRELCAFEAERAREE 361
Query: 165 STT------------------DELATRITEESMKKMQYLDAALTETLRLHPPIPMESKYC 206
T + A +T +S+ K+ YL A +TETLRL+P +P + K
Sbjct: 362 GVTLVLCGGADADDKAFAARVAQFAGLLTYDSLGKLVYLHACVTETLRLYPAVPQDPKGI 421
Query: 207 FSDDTLPDGYSVRKGDYISFHPYVMGRMEFLWGEDAEQFRPERWLDENGNFQRESPFKFT 266
DD LPDG VR G +++ PY MGRME+ WG DA FRPERW++E+G F+ SPFKFT
Sbjct: 422 LEDDVLPDGTKVRAGGMVTYVPYSMGRMEYNWGPDAASFRPERWINEDGAFRNASPFKFT 481
Query: 267 AFQAGPRICLGKEFVYRQMKIFSAVLLGSHNFKLADQNRLVKYRTTLTLQIDGGLHV 323
AFQAGPRICLGK+ Y QMK+ A+L ++F+L + V+YR L + GL V
Sbjct: 482 AFQAGPRICLGKDSAYLQMKMALAILFRFYSFRLL-EGHPVQYRMMTILSMAHGLKV 537
>gb|AAG60111.1| cytochrome P450, putative [Arabidopsis thaliana]
Length = 524
Score = 259 bits (661), Expect = 1e-67
Identities = 145/360 (40%), Positives = 219/360 (60%), Gaps = 34/360 (9%)
Query: 1 MKSTLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWYVNFLWKVQRFLNIGS 60
M+ TLDS+ KV GVE+ T+ E F+ AF+ A+ + +++ LWK+++FLNIGS
Sbjct: 168 MRMTLDSICKVGFGVEIGTLAPELPEN-HFAKAFDTANIIVTLRFIDPLWKMKKFLNIGS 226
Query: 61 EAVLKKNLRVINEYVYTIIK--------SKIEQSQKPQKNSPELKGDILSRFLELNETDS 112
EA+L K+++V+N++ Y++I+ ++I + N+ ++K DILSRF+E+++
Sbjct: 227 EALLGKSIKVVNDFTYSVIRRRKAELLEAQISPTNNNNNNNNKVKHDILSRFIEISDDPD 286
Query: 113 -----KYLKDVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATK------- 160
K L+D++L+F+IAG+DTT+ TL+ IY + + +V EK+ E++E K
Sbjct: 287 SKETEKSLRDIVLNFVIAGRDTTATTLTWAIYMIMMNENVAEKLYSELQELEKESAEATN 346
Query: 161 -------VEDGSTTDELATR----ITEESMKKMQYLDAALTETLRLHPPIPMESKYCFSD 209
ED ++ +E T + +S+ K+ YL A +TETLRL+P +P + K D
Sbjct: 347 TSLHQYDTEDFNSFNEKVTEFAGLLNYDSLGKLHYLHAVITETLRLYPAVPQDPKGVLED 406
Query: 210 DTLPDGYSVRKGDYISFHPYVMGRMEFLWGEDAEQFRPERWLDENGNFQRESPFKFTAFQ 269
D LP+G V+ G +++ PY MGRME+ WG DA F+PERWL ++G FQ SPFKFTAFQ
Sbjct: 407 DMLPNGTKVKAGGMVTYVPYSMGRMEYNWGSDAALFKPERWL-KDGVFQNASPFKFTAFQ 465
Query: 270 AGPRICLGKEFVYRQMKIFSAVLLGSHNFKLADQNRLVKYRTTLTLQIDGGLHVNAFHRN 329
AGPRICLGK+ Y QMK+ A+L + F L N VKYR L + GL V R+
Sbjct: 466 AGPRICLGKDSAYLQMKMAMAILCRFYKFHLV-PNHPVKYRMMTILSMAHGLKVTVSRRS 524
>dbj|BAC43393.1| unknown protein [Arabidopsis thaliana] gi|30697865|ref|NP_177109.2|
cytochrome P450 family protein [Arabidopsis thaliana]
Length = 478
Score = 259 bits (661), Expect = 1e-67
Identities = 145/360 (40%), Positives = 219/360 (60%), Gaps = 34/360 (9%)
Query: 1 MKSTLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWYVNFLWKVQRFLNIGS 60
M+ TLDS+ KV GVE+ T+ E F+ AF+ A+ + +++ LWK+++FLNIGS
Sbjct: 122 MRMTLDSICKVGFGVEIGTLAPELPEN-HFAKAFDTANIIVTLRFIDPLWKMKKFLNIGS 180
Query: 61 EAVLKKNLRVINEYVYTIIK--------SKIEQSQKPQKNSPELKGDILSRFLELNETDS 112
EA+L K+++V+N++ Y++I+ ++I + N+ ++K DILSRF+E+++
Sbjct: 181 EALLGKSIKVVNDFTYSVIRRRKAELLEAQISPTNNNNNNNNKVKHDILSRFIEISDDPD 240
Query: 113 -----KYLKDVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATK------- 160
K L+D++L+F+IAG+DTT+ TL+ IY + + +V EK+ E++E K
Sbjct: 241 SKETEKSLRDIVLNFVIAGRDTTATTLTWAIYMIMMNENVAEKLYSELQELEKESAEATN 300
Query: 161 -------VEDGSTTDELATR----ITEESMKKMQYLDAALTETLRLHPPIPMESKYCFSD 209
ED ++ +E T + +S+ K+ YL A +TETLRL+P +P + K D
Sbjct: 301 TSLHQYDTEDFNSFNEKVTEFAGLLNYDSLGKLHYLHAVITETLRLYPAVPQDPKGVLED 360
Query: 210 DTLPDGYSVRKGDYISFHPYVMGRMEFLWGEDAEQFRPERWLDENGNFQRESPFKFTAFQ 269
D LP+G V+ G +++ PY MGRME+ WG DA F+PERWL ++G FQ SPFKFTAFQ
Sbjct: 361 DMLPNGTKVKAGGMVTYVPYSMGRMEYNWGSDAALFKPERWL-KDGVFQNASPFKFTAFQ 419
Query: 270 AGPRICLGKEFVYRQMKIFSAVLLGSHNFKLADQNRLVKYRTTLTLQIDGGLHVNAFHRN 329
AGPRICLGK+ Y QMK+ A+L + F L N VKYR L + GL V R+
Sbjct: 420 AGPRICLGKDSAYLQMKMAMAILCRFYKFHLV-PNHPVKYRMMTILSMAHGLKVTVSRRS 478
>ref|XP_470289.1| putative plant cytochrome P-450 protein [Oryza sativa (japonica
cultivar-group)] gi|19071651|gb|AAL84318.1| putative
plant cytochrome P-450 protein [Oryza sativa (japonica
cultivar-group)]
Length = 544
Score = 257 bits (657), Expect = 3e-67
Identities = 144/353 (40%), Positives = 217/353 (60%), Gaps = 33/353 (9%)
Query: 1 MKSTLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWYVNFLWKVQRFLNIGS 60
M+ TLDS+ KV GVE+ T+ E + F+ AF+ A+ + +++ LW++++FL++GS
Sbjct: 189 MRMTLDSICKVGFGVEIGTLSPDLPENS-FAQAFDAANIIVTLRFIDPLWRLKKFLHVGS 247
Query: 61 EAVLKKNLRVINEYVYTII---KSKIEQSQKPQKNSPELKGDILSRFLELNET------- 110
EA+L++++++++++ Y++I K++I Q++ K ++K DILSRF+EL E
Sbjct: 248 EALLEQSMKLVDDFTYSVIRRRKAEILQARASGKQE-KIKHDILSRFIELGEAGGDEGGG 306
Query: 111 ---DSKYLKDVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEI---------REA 158
D K L+DV+L+F+IAG+DTT+ TLS F Y HP V +K+ +E+ E
Sbjct: 307 SFGDDKSLRDVVLNFVIAGRDTTATTLSWFTYMAMTHPAVADKLRRELAAFEDERAREEG 366
Query: 159 TKVEDGSTTDELATRITE-------ESMKKMQYLDAALTETLRLHPPIPMESKYCFSDDT 211
+ D + A R+ + +++ K+ YL A +TETLRL+P +P + K DD
Sbjct: 367 VALADAAGEASFAARVAQFASLLSYDAVGKLVYLHACVTETLRLYPAVPQDPKGIVEDDV 426
Query: 212 LPDGYSVRKGDYISFHPYVMGRMEFLWGEDAEQFRPERWLD-ENGNFQRESPFKFTAFQA 270
LPDG VR G +++ PY MGRME+ WG DA FRPERWL + G F+ SPFKFTAFQA
Sbjct: 427 LPDGTKVRAGGMVTYVPYSMGRMEYNWGPDAASFRPERWLSGDGGAFRNASPFKFTAFQA 486
Query: 271 GPRICLGKEFVYRQMKIFSAVLLGSHNFKLADQNRLVKYRTTLTLQIDGGLHV 323
GPRICLGK+ Y QMK+ A+L + F L + + VKYR L + GL V
Sbjct: 487 GPRICLGKDSAYLQMKMALAILFRFYTFDLVEDHP-VKYRMMTILSMAHGLKV 538
>emb|CAB88066.1| cytochrome P450-like protein [Arabidopsis thaliana]
gi|15228981|ref|NP_191222.1| cytochrome P450, putative
[Arabidopsis thaliana] gi|11249683|pir||T49064
cytochrome P450-like protein - Arabidopsis thaliana
Length = 499
Score = 235 bits (599), Expect = 2e-60
Identities = 129/333 (38%), Positives = 202/333 (59%), Gaps = 22/333 (6%)
Query: 6 DSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWY---VNFLWKVQRFLNIGSEA 62
D++ K+ V+ + G F AF A+ I + +++ WK+++ LNIGSE
Sbjct: 180 DNICKLAFNVDSACLGDDGAAGVNFMQAFETAATIISQRFQSVISYSWKIKKKLNIGSER 239
Query: 63 VLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLELNETDS-KYLKDVILS 121
VL++++ +++++ I++++IEQ + K D+LSRF+ E +S + L+D+++S
Sbjct: 240 VLRESIMIVHKFADEIVRNRIEQGKVSDH-----KEDLLSRFISKEEMNSPEILRDIVIS 294
Query: 122 FIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATKVEDGSTTDELATRITE----E 177
FI+AG+DTTS LS F + L HP V++KI QE+ S + RI E E
Sbjct: 295 FILAGRDTTSSALSWFFWLLSMHPEVKDKILQELN--------SIRERTGKRIGEVYGFE 346
Query: 178 SMKKMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPYVMGRMEFL 237
+K M YL AA+TE+LRL+PP+P+++ C D+ LPDG + K IS++ Y MGRME +
Sbjct: 347 DLKLMNYLHAAITESLRLYPPVPVDTMSCAEDNVLPDGTFIGKDWGISYNAYAMGRMESI 406
Query: 238 WGEDAEQFRPERWLDE-NGNFQRESPFKFTAFQAGPRICLGKEFVYRQMKIFSAVLLGSH 296
WG+D ++F PERW+DE NG F+ E+P+KF AF AGPR+CLGKE Y QMK A +L
Sbjct: 407 WGKDCDRFDPERWIDETNGGFRGENPYKFPAFHAGPRMCLGKEMAYIQMKSIVAAVLERF 466
Query: 297 NFKLADQNRLVKYRTTLTLQIDGGLHVNAFHRN 329
++ + + ++TL+I GGL+V R+
Sbjct: 467 VVEVPGKKERPEILMSVTLRIRGGLNVRVQERS 499
>gb|AAY34172.1| At1g13140 [Arabidopsis thaliana]
Length = 534
Score = 224 bits (572), Expect = 2e-57
Identities = 122/305 (40%), Positives = 185/305 (60%), Gaps = 15/305 (4%)
Query: 30 FSNAFNEASATIMFWYV--NFLWKVQRFLNIGSEAVLKKNLRVINEYVYTIIKSKIEQSQ 87
F+ AF EA+ + MF ++ F+WK +F +IG E L+K + V++E+V ++ +I +
Sbjct: 219 FAQAFEEATESTMFRFMIPPFIWKPLKFFDIGYEKGLRKAVDVVHEFVDKMVVDRI--CK 276
Query: 88 KPQKNSPELKGDILSRFLELNE---TDSK------YLKDVILSFIIAGKDTTSITLS*FI 138
++ + + D+LSR +E+ TD K + + SFI+AG+DT+S+ L+ F
Sbjct: 277 LKEEGTLGNRSDVLSRIIEIESHKTTDEKDPSTIKFFRQFCTSFILAGRDTSSVALTWFF 336
Query: 139 YQLCKHPHVQEKIAQEIREATKVEDGSTTDELATRITEESMKKMQYLDAALTETLRLHPP 198
+ + KHP V+ KI +EI E + S T + + T + + M YL AAL+ET+RL+PP
Sbjct: 337 WVIQKHPEVENKIIREISEILRQRGDSPTSKNESLFTVKELNDMVYLQAALSETMRLYPP 396
Query: 199 IPMESKYCFSDDTLPDGYSVRKGDYISFHPYVMGRMEFLWGEDAEQFRPERWLDENGNFQ 258
IPME K DD PDG +RKG + F Y MGRME +WG+D E F+PERW+ ++GNF
Sbjct: 397 IPMEMKQAIEDDVFPDGTFIRKGSRVYFATYAMGRMESIWGKDCESFKPERWI-QSGNFV 455
Query: 259 RESPFKFTAFQAGPRICLGKEFVYRQMKIFSAVLLGSHNFKLADQNRLVKYRTTLTLQID 318
+ FK+ F AGPR+CLGK F Y QMK +A +L ++ K+A ++ +V R T TL +
Sbjct: 456 NDDQFKYVVFNAGPRLCLGKTFAYLQMKTIAASVLSRYSIKVA-KDHVVVPRVTTTLYMR 514
Query: 319 GGLHV 323
GL V
Sbjct: 515 HGLKV 519
>gb|AAF79271.1| F12K21.15 [Arabidopsis thaliana] gi|15218671|ref|NP_174713.1|
cytochrome P450 family protein [Arabidopsis thaliana]
Length = 498
Score = 222 bits (566), Expect = 1e-56
Identities = 125/329 (37%), Positives = 199/329 (59%), Gaps = 15/329 (4%)
Query: 6 DSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWY---VNFLWKVQRFLNIGSEA 62
D++ K+ V+ + G F AF A+ I + + W++++ LNIGSE
Sbjct: 180 DNICKLAFNVDCACLGHDGAVGVNFMRAFETAATIISQRFRSVASCAWRIKKKLNIGSER 239
Query: 63 VLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLELNETDS-KYLKDVILS 121
VL++++ ++++ I++++I+Q + S + K D+LSRF+ E +S + L+D+++S
Sbjct: 240 VLRESIATVHKFADEIVRNRIDQGR-----SSDHKEDLLSRFISKEEMNSPEILRDIVIS 294
Query: 122 FIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATKVEDGSTTDELATRITEESMKK 181
FI+AG+DTTS LS F + L HP V++KI QE+ + + G E+ E +K
Sbjct: 295 FILAGRDTTSSALSWFFWLLSMHPEVEDKILQELN-SIRARTGKRIGEV---YGFEHLKM 350
Query: 182 MQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPYVMGRMEFLWGED 241
M YL AA+TE+LRL+PP+P++ K C D+ LPDG V KG I+++ + MGRME +WG+D
Sbjct: 351 MNYLHAAITESLRLYPPVPVDIKSCAEDNVLPDGTFVGKGWAITYNIFAMGRMESIWGKD 410
Query: 242 AEQFRPERWLDE-NGNFQRESPFKFTAFQAGPRICLGKEFVYRQMKIFSAVLLGSHNFKL 300
++F PERW+DE NG F+ E P KF AF AGPR+C+GK+ Y QMK A +L ++
Sbjct: 411 CDRFDPERWIDETNGCFRGEDPSKFPAFHAGPRMCVGKDMAYIQMKSIVAAVLERFVVEV 470
Query: 301 ADQNRLVKYRTTLTLQIDGGLHVNAFHRN 329
+ R + ++TL+I GGL R+
Sbjct: 471 PGKER-PEILLSMTLRIKGGLFARVQERS 498
>ref|NP_172774.1| cytochrome P450, putative [Arabidopsis thaliana]
gi|4850397|gb|AAD31067.1| Strong similarity to
gi|3313615 F21J9.9 from Arabidopsis thaliana and is a
member of the PF|00067 Cytochrome P450 family
gi|25282631|pir||G86265 F3F19.17 protein - Arabidopsis
thaliana
Length = 529
Score = 219 bits (558), Expect = 9e-56
Identities = 128/340 (37%), Positives = 196/340 (57%), Gaps = 16/340 (4%)
Query: 1 MKSTLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWYV--NFLWKVQRFLNI 58
++ T D + LG + +T+ + F+ AF EA+ + +F ++ F+WK RFL+
Sbjct: 184 LRLTFDIICLAGLGADPETLAVDLPQ-VPFAKAFEEATESTLFRFMIPPFIWKPMRFLDT 242
Query: 59 GSEAVLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLEL------NETDS 112
G E L+ + V++ +V +I +I + ++ + + + D+LSR +++ NE D
Sbjct: 243 GYEKGLRIAVGVVHGFVDKMIVDRI--CELKEEETLDNRSDVLSRIIQIESHKRENEIDP 300
Query: 113 ---KYLKDVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATKVEDGSTTDE 169
++ + SFI+AG+DT+S+ LS F + + KHP V+ KI EIRE + S T +
Sbjct: 301 STIRFFRQFCTSFILAGRDTSSVALSWFCWVIQKHPEVENKIICEIREILRQRGDSPTSK 360
Query: 170 LATRITEESMKKMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPY 229
+ T + + M YL AAL+ETLRL PPIPME K DD LPDG VRKG + F Y
Sbjct: 361 NESLFTVKELNNMVYLQAALSETLRLFPPIPMEMKQAIEDDVLPDGTFVRKGSRVYFSIY 420
Query: 230 VMGRMEFLWGEDAEQFRPERWLDENGNFQRESPFKFTAFQAGPRICLGKEFVYRQMKIFS 289
MGRME +WG+D E FRPERW+ + G F + FK+ F AGPR+C+GK F Y QMK+ +
Sbjct: 421 AMGRMESIWGKDCEIFRPERWI-QAGKFVSDDQFKYVVFNAGPRLCIGKTFAYLQMKMIA 479
Query: 290 AVLLGSHNFKLADQNRLVKYRTTLTLQIDGGLHVNAFHRN 329
A +L ++ K+ Q+ ++ R T L + GL V R+
Sbjct: 480 ASVLLRYSIKVV-QDHVIAPRVTTNLYMKYGLKVTITPRS 518
>gb|AAO41955.1| putative cytochrome P450 [Arabidopsis thaliana]
Length = 529
Score = 218 bits (555), Expect = 2e-55
Identities = 127/340 (37%), Positives = 196/340 (57%), Gaps = 16/340 (4%)
Query: 1 MKSTLDSVFKVILGVELDTMCGTSKEGTQFSNAFNEASATIMFWYV--NFLWKVQRFLNI 58
++ T D + LG + +T+ + F+ AF EA+ + +F ++ F+WK RFL+
Sbjct: 184 LRLTFDIICLAGLGADPETLAVDLPQ-VPFAKAFEEATESTLFRFMIPPFIWKPMRFLDT 242
Query: 59 GSEAVLKKNLRVINEYVYTIIKSKIEQSQKPQKNSPELKGDILSRFLEL------NETDS 112
G E L+ + V++ +V +I +I + ++ + + + D+LSR +++ NE D
Sbjct: 243 GYEKGLRIAVGVVHGFVDKMIVDRI--CELKEEETLDNRSDVLSRIIQIESHKRENEIDP 300
Query: 113 ---KYLKDVILSFIIAGKDTTSITLS*FIYQLCKHPHVQEKIAQEIREATKVEDGSTTDE 169
++ + SFI+AG+DT+S+ LS F + + KHP V+ KI EIRE + S T +
Sbjct: 301 STIRFFRQFCTSFILAGRDTSSVALSWFCWVIQKHPEVENKIICEIREILRQRGDSPTSK 360
Query: 170 LATRITEESMKKMQYLDAALTETLRLHPPIPMESKYCFSDDTLPDGYSVRKGDYISFHPY 229
+ T + + M YL AAL+ETLRL PPIPME K DD LPDG VRKG + F Y
Sbjct: 361 NESLFTVKELNNMVYLQAALSETLRLFPPIPMEMKQAIEDDVLPDGTFVRKGSRVYFSIY 420
Query: 230 VMGRMEFLWGEDAEQFRPERWLDENGNFQRESPFKFTAFQAGPRICLGKEFVYRQMKIFS 289
MGRME +WG+D E FRPERW+ + G F + FK+ F AGPR+C+GK F Y QM++ +
Sbjct: 421 AMGRMESIWGKDCEIFRPERWI-QAGKFVSDDQFKYVVFNAGPRLCIGKTFAYLQMRMIA 479
Query: 290 AVLLGSHNFKLADQNRLVKYRTTLTLQIDGGLHVNAFHRN 329
A +L ++ K+ Q+ ++ R T L + GL V R+
Sbjct: 480 ASVLLRYSIKVV-QDHVIAPRVTTNLYMKYGLKVTITPRS 518
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.322 0.137 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 526,056,980
Number of Sequences: 2540612
Number of extensions: 21221851
Number of successful extensions: 61883
Number of sequences better than 10.0: 4828
Number of HSP's better than 10.0 without gapping: 2427
Number of HSP's successfully gapped in prelim test: 2401
Number of HSP's that attempted gapping in prelim test: 51323
Number of HSP's gapped (non-prelim): 5045
length of query: 329
length of database: 863,360,394
effective HSP length: 128
effective length of query: 201
effective length of database: 538,162,058
effective search space: 108170573658
effective search space used: 108170573658
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 75 (33.5 bits)
Medicago: description of AC149581.9


