

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149581.7 + phase: 0
(392 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
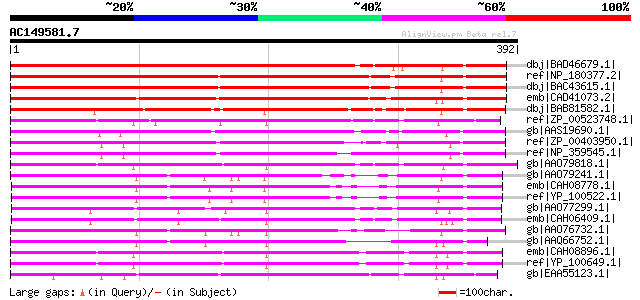
Score E
Sequences producing significant alignments: (bits) Value
dbj|BAD46679.1| unknown protein [Oryza sativa (japonica cultivar... 481 e-134
ref|NP_180377.2| glycosyl hydrolase family 29 / alpha-L-fucosida... 408 e-112
dbj|BAC43615.1| unknown protein [Arabidopsis thaliana] 405 e-112
emb|CAD41073.2| OSJNBa0084K11.7 [Oryza sativa (japonica cultivar... 393 e-108
dbj|BAB81582.1| conserved hypothetical protein [Clostridium perf... 328 1e-88
ref|ZP_00523748.1| Coagulation factor 5/8 type, C-terminal [Soli... 280 7e-74
gb|AAS19690.1| FucA [Streptococcus gordonii] 269 1e-70
ref|ZP_00403950.1| COG3669: Alpha-L-fucosidase [Streptococcus pn... 269 1e-70
ref|NP_359545.1| hypothetical protein spr1954 [Streptococcus pne... 267 4e-70
gb|AAO79818.1| conserved hypothetical protein [Bacteroides theta... 256 6e-67
gb|AAO79241.1| conserved hypothetical protein [Bacteroides theta... 254 3e-66
emb|CAH08778.1| putative exported fucosidase [Bacteroides fragil... 253 7e-66
ref|YP_100522.1| hypothetical protein BF3243 [Bacteroides fragil... 251 2e-65
gb|AAO77299.1| conserved hypothetical protein [Bacteroides theta... 251 4e-65
emb|CAH06409.1| conserved hypothetical exported protein [Bactero... 246 9e-64
gb|AAO76732.1| conserved hypothetical protein [Bacteroides theta... 244 4e-63
gb|AAQ66752.1| alpha-1,3/4-fucosidase, putative [Porphyromonas g... 243 6e-63
emb|CAH08896.1| putative lipoprotein [Bacteroides fragilis NCTC ... 243 6e-63
ref|YP_100649.1| hypothetical protein BF3371 [Bacteroides fragil... 243 6e-63
gb|EAA55123.1| hypothetical protein MG06780.4 [Magnaporthe grise... 229 1e-58
>dbj|BAD46679.1| unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 494
Score = 481 bits (1237), Expect = e-134
Identities = 226/390 (57%), Positives = 289/390 (73%), Gaps = 11/390 (2%)
Query: 1 MILTAKHHDGFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRH 60
++L AKHHDGFCLWPS +T HSV +S W+ G+GDVV+EF +AA +G+D+GIYLSPWDRH
Sbjct: 103 VVLVAKHHDGFCLWPSAHTAHSVRASPWRGGRGDVVREFADAARARGLDIGIYLSPWDRH 162
Query: 61 DSRYGHDLLYNEYYLAQLQELLKKYQDVREIWFDGAKDPRAQNVTYYFSDWFSMVKELQS 120
D RYG ++ YNEYYLAQL ELL Y V EIWFDGAK A N+TY+F +WF V++LQS
Sbjct: 163 DKRYGREVAYNEYYLAQLHELLTGYGSVSEIWFDGAKGKNATNMTYHFQEWFQTVRQLQS 222
Query: 121 SINIFSDAGPDVRWVGDETGTAGDTCWSTINRTSLSIGASNITQYLNTGDPKGTDWLPAE 180
SINIFSD GPD+RWVGDE G+AG TCWSTINR+ ++IG + I +YLNTGDP+G DW+P E
Sbjct: 223 SINIFSDDGPDLRWVGDENGSAGSTCWSTINRSKITIGEAGIEKYLNTGDPRGKDWVPPE 282
Query: 181 CDVSIRPGWFWHKSESPKKLSDLLDIYYKSVGRNCVLLLNVPPNTTGLISENDAHRLKEF 240
CDVSIRPGWFWHK+E+ K L +LL++YY SVGRNCVLLLN PPNTTGL+ D RL+EF
Sbjct: 283 CDVSIRPGWFWHKNETAKPLPELLEVYYNSVGRNCVLLLNAPPNTTGLVDAADIARLREF 342
Query: 241 RSAIDTIFHKNIAENRYVKVSSQRGGKEGGFGPENMLDSDHLWSYWTPREDDKEK--DHW 298
R+A+ IF ++A + SS+RGG+ F N+LD +YW P + E +W
Sbjct: 343 RTAVTAIFGTDLAAGSAARASSERGGR---FAAANVLDGRD-DTYWAPAAAEAEDGGGYW 398
Query: 299 IEIW--GNDGSLRFNVIRIQEAIGLGQRIERYEIYVD--GKSIIQGTTIGYKRLHRLDGD 354
IE+ + + +FNV+RIQE + +GQR+ER+E+YVD G ++ GTT+G+KRLHRL G
Sbjct: 399 IELRRPASAAARKFNVVRIQEHVAMGQRVERHEVYVDGGGAAVASGTTVGHKRLHRL-GA 457
Query: 355 VVHARVVRIRFIKARGVPLISSIGLHFDPF 384
V R VR+ RG PL+S++GLH DPF
Sbjct: 458 PVAGRTVRVWLASRRGPPLLSAVGLHLDPF 487
>ref|NP_180377.2| glycosyl hydrolase family 29 / alpha-L-fucosidase, putative
[Arabidopsis thaliana]
Length = 506
Score = 408 bits (1049), Expect = e-112
Identities = 202/390 (51%), Positives = 263/390 (66%), Gaps = 12/390 (3%)
Query: 1 MILTAKHHDGFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRH 60
+ILTAKHHDGFCLWPS+YT +SV SS+W+NG GDVV E +AA + GI +G+YLSPWDRH
Sbjct: 98 VILTAKHHDGFCLWPSEYTDYSVKSSQWRNGAGDVVAELASAAKEAGIGLGLYLSPWDRH 157
Query: 61 DSRYGHDLLYNEYYLAQLQELLKKYQDVREIWFDGAKDPRAQNVTYYFSDWFSMVKELQS 120
+ YG L YNE+YL+Q+ ELL KY +++E+W DGAK +++ Y+F WFS++ +LQ
Sbjct: 158 EQCYGKTLEYNEFYLSQMTELLTKYGEIKEVWLDGAKGDGEKDMEYFFDTWFSLIHQLQP 217
Query: 121 SINIFSDAGPDVRWVGDETGTAGDTCWSTINRTSLSIGASNITQYLNTGDPKGTDWLPAE 180
IFSDAGPDVRW+GDE G AG TCWS NRT+ IG + Y GD G DW+PAE
Sbjct: 218 KAVIFSDAGPDVRWIGDEAGLAGSTCWSLFNRTNAKIGDTE-PSYSQEGDGYGQDWVPAE 276
Query: 181 CDVSIRPGWFWHKSESPKKLSDLLDIYYKSVGRNCVLLLNVPPNTTGLISENDAHRLKEF 240
CDVSIRPGWFWH SESPK LLDIYY SVGRNC+ LLNVPPN++GLISE D L+EF
Sbjct: 277 CDVSIRPGWFWHASESPKPAVQLLDIYYNSVGRNCLFLLNVPPNSSGLISEQDIKVLEEF 336
Query: 241 RSAIDTIFHKNIAENRYVKVSSQRGGKEGGFGPENMLDSDHLWSYWTPREDDKEKDHWIE 300
++IF N+A +V SS RG + FGP+N+L+ + L YW P E+ E W+
Sbjct: 337 SEMKNSIFSNNLARKAFVNSSSIRGDQSSQFGPKNVLE-EGLDKYWAPEENQNE---WVL 392
Query: 301 IWGNDGSLRFNVIRIQEAIGLGQRIERYEIYV------DGKSIIQGTTIGYKRLHRLDGD 354
+ FNV+ I+E I +GQRI + + + + ++ GTT+G KRL R +
Sbjct: 393 YLEFKDLVSFNVLEIREPIHMGQRIASFHLETRKTGSGEWERVVSGTTVGNKRLLRF-LN 451
Query: 355 VVHARVVRIRFIKARGVPLISSIGLHFDPF 384
VV +R +++ KAR PLIS +GL+ D F
Sbjct: 452 VVESRSLKLVVDKARTDPLISYLGLYMDKF 481
>dbj|BAC43615.1| unknown protein [Arabidopsis thaliana]
Length = 506
Score = 405 bits (1042), Expect = e-112
Identities = 201/390 (51%), Positives = 262/390 (66%), Gaps = 12/390 (3%)
Query: 1 MILTAKHHDGFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRH 60
+ILTAKHHDGFCLWPS+YT +SV SS+W+NG GDVV +AA + GI +G+YLSPWDRH
Sbjct: 98 VILTAKHHDGFCLWPSEYTDYSVKSSQWRNGAGDVVAGLASAAKEAGIGLGLYLSPWDRH 157
Query: 61 DSRYGHDLLYNEYYLAQLQELLKKYQDVREIWFDGAKDPRAQNVTYYFSDWFSMVKELQS 120
+ YG L YNE+YL+Q+ ELL KY +++E+W DGAK +++ Y+F WFS++ +LQ
Sbjct: 158 EQCYGKTLEYNEFYLSQMTELLTKYGEIKEVWLDGAKGDGEKDMEYFFDTWFSLIHQLQP 217
Query: 121 SINIFSDAGPDVRWVGDETGTAGDTCWSTINRTSLSIGASNITQYLNTGDPKGTDWLPAE 180
IFSDAGPDVRW+GDE G AG TCWS NRT+ IG + Y GD G DW+PAE
Sbjct: 218 KAVIFSDAGPDVRWIGDEAGLAGSTCWSLFNRTNAKIGDTE-PSYSQEGDGYGQDWVPAE 276
Query: 181 CDVSIRPGWFWHKSESPKKLSDLLDIYYKSVGRNCVLLLNVPPNTTGLISENDAHRLKEF 240
CDVSIRPGWFWH SESPK LLDIYY SVGRNC+ LLNVPPN++GLISE D L+EF
Sbjct: 277 CDVSIRPGWFWHASESPKPAVQLLDIYYNSVGRNCLFLLNVPPNSSGLISEQDIKVLEEF 336
Query: 241 RSAIDTIFHKNIAENRYVKVSSQRGGKEGGFGPENMLDSDHLWSYWTPREDDKEKDHWIE 300
++IF N+A +V SS RG + FGP+N+L+ + L YW P E+ E W+
Sbjct: 337 SEMKNSIFSNNLARKAFVNSSSIRGDQSSQFGPKNVLE-EGLDKYWAPEENQNE---WVL 392
Query: 301 IWGNDGSLRFNVIRIQEAIGLGQRIERYEIYV------DGKSIIQGTTIGYKRLHRLDGD 354
+ FNV+ I+E I +GQRI + + + + ++ GTT+G KRL R +
Sbjct: 393 YLEFKDLVSFNVLEIREPIHMGQRIASFHLETRKTGSGEWERVVSGTTVGNKRLLRF-LN 451
Query: 355 VVHARVVRIRFIKARGVPLISSIGLHFDPF 384
VV +R +++ KAR PLIS +GL+ D F
Sbjct: 452 VVESRSLKLVVDKARTDPLISYLGLYMDKF 481
>emb|CAD41073.2| OSJNBa0084K11.7 [Oryza sativa (japonica cultivar-group)]
gi|50927915|ref|XP_473485.1| OSJNBa0084K11.7 [Oryza
sativa (japonica cultivar-group)]
Length = 517
Score = 393 bits (1010), Expect = e-108
Identities = 200/389 (51%), Positives = 260/389 (66%), Gaps = 13/389 (3%)
Query: 1 MILTAKHHDGFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRH 60
++LTAKHHDGFCLWPS T +SV +S W+ G GDVV E AA +GI +G+YLSPWDRH
Sbjct: 100 VVLTAKHHDGFCLWPSALTNYSVAASPWKGGAGDVVGELAAAARAEGIGLGLYLSPWDRH 159
Query: 61 DSRYGHDLLYNEYYLAQLQELLKKYQDVREIWFDGAKDPRAQNVTYYFSDWFSMVKELQS 120
+ YG + YNE+YL Q+ ELL +Y DV E+W DGAK +++ Y F WF+++ +LQ
Sbjct: 160 EPVYGDTVAYNEHYLGQMTELLTRYGDVEEVWLDGAKG-EGKDMDYMFDAWFALIHQLQQ 218
Query: 121 SINIFSDAGPDVRWVGDETGTAGDTCWSTINRTSLSIGASNITQYLNTGDPKGTDWLPAE 180
+ IFSDAGPD RWVGDE G AG TCWS N+++++IG I +Y GDP G DW+PAE
Sbjct: 219 RVVIFSDAGPDTRWVGDEAGVAGYTCWSPFNKSTVTIGHI-IPEYSRCGDPFGQDWVPAE 277
Query: 181 CDVSIRPGWFWHKSESPKKLSDLLDIYYKSVGRNCVLLLNVPPNTTGLISENDAHRLKEF 240
CDVSIRPGWFWH SE PK + LLDIYYKSVGRNC+L+LNVPPN++GLIS D L+EF
Sbjct: 278 CDVSIRPGWFWHASEKPKNATTLLDIYYKSVGRNCLLILNVPPNSSGLISTEDMQVLQEF 337
Query: 241 RSAIDTIFHKNIAENRYVKVSSQRGG-KEGGFGPENMLDSDHLWSYWTPREDDKEKDHWI 299
TIF +N A N V S+ RGG F P N+L + ++SYW P E + W
Sbjct: 338 TEIRQTIFSQNFAANATVTASTVRGGLGNQQFAPSNVL-QESIYSYWAPEEG---QSSWE 393
Query: 300 EIWGNDGSLRFNVIRIQEAIGLGQRIERY--EIYVD--GKSIIQGTTIGYKRLHRLDGDV 355
++ S FNVI++QE I +GQR+ ++ EI VD ++I++GTTIGYKRL + V
Sbjct: 394 MLFDLGQSASFNVIQLQEPIQMGQRVIKFRVEILVDELWQTIVEGTTIGYKRLFQF--PV 451
Query: 356 VHARVVRIRFIKARGVPLISSIGLHFDPF 384
V + +++ AR PLIS G+ D F
Sbjct: 452 VEGQFLKLSIDGARADPLISFFGVFTDSF 480
>dbj|BAB81582.1| conserved hypothetical protein [Clostridium perfringens str. 13]
gi|18310858|ref|NP_562792.1| hypothetical protein
CPE1876 [Clostridium perfringens str. 13]
Length = 750
Score = 328 bits (842), Expect = 1e-88
Identities = 179/391 (45%), Positives = 242/391 (61%), Gaps = 18/391 (4%)
Query: 1 MILTAKHHDGFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRH 60
+ILTAKHHDGFCLW S YTKH V SS W+NGKGDVVKE A I G+YLSPWD++
Sbjct: 108 VILTAKHHDGFCLWDSAYTKHDVASSPWKNGKGDVVKEVSEACAKYNIKFGVYLSPWDQN 167
Query: 61 DSRY--GHDLLYNEYYLAQLQELLKKYQDVREIWFDGAKDPRAQNVTYYFSDWFSMVKEL 118
Y G+ YNE+Y+ QL+ELL Y + E+W DGAK + Y F +WF+++KEL
Sbjct: 168 SEHYGEGNGGDYNEFYMNQLRELLTNYGPIAEVWMDGAKGSNVKQ-EYNFEEWFALIKEL 226
Query: 119 QSSINIFSDAGPDVRWVGDETGTAGDTCWSTINRTSLSIGASNITQYLNTGDPKGTDWLP 178
Q IFS GPD+RW+G+E G AG+ CWSTI+ + N T YLN G+ G DW+
Sbjct: 227 QPECLIFSPQGPDIRWIGNEKGYAGEPCWSTIDIEKMK-ERENPT-YLNNGEEGGPDWVV 284
Query: 179 AECDVSIRPGWFWHKSE--SPKKLSDLLDIYYKSVGRNCVLLLNVPPNTTGLISENDAHR 236
E DVSIRPGWF+H+S+ K L ++DIY+KS+GRN VLLLNVPPN G + END +R
Sbjct: 285 GESDVSIRPGWFYHESQDNEVKSLEKMMDIYFKSIGRNSVLLLNVPPNKEGKLHENDVNR 344
Query: 237 LKEFRSAIDTIFHKNIAENRYVKVSSQRGGKEGGFGPENMLDSDHLWSYWTPREDDKEKD 296
LKEF I +F+ ++A N+ V V ++ +G ++D D+ +YW P D+ K
Sbjct: 345 LKEFGETIKELFNDDLALNKEVIVDG-FANRDETYGANKIVDGDY-DTYWAP--DNSSKT 400
Query: 297 HWIEIWGNDGSLRFNVIRIQEAIGLGQRIERYEIYV----DGKSIIQGTTIGYKRLHRLD 352
IEI + S F+VI +QE I LGQR+ + + V + + +G TIGYKRL R+
Sbjct: 401 GTIEIDLGE-SKEFDVISLQEYIPLGQRVSSFNVEVLQGENWNKVYEGKTIGYKRLVRI- 458
Query: 353 GDVVHARVVRIRFIKARGVPLISSIGLHFDP 383
+RI + VPLI+++G++ P
Sbjct: 459 -APTKGEKIRINITGSLEVPLINNVGVYKQP 488
>ref|ZP_00523748.1| Coagulation factor 5/8 type, C-terminal [Solibacter usitatus
Ellin6076] gi|67862145|gb|EAM57168.1| Coagulation factor
5/8 type, C-terminal [Solibacter usitatus Ellin6076]
Length = 491
Score = 280 bits (715), Expect = 7e-74
Identities = 158/402 (39%), Positives = 227/402 (56%), Gaps = 31/402 (7%)
Query: 1 MILTAKHHDGFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRH 60
+ILT KHHDGFCLWP++ T HSV S+W+ G+GDVV+E AA + + G+YLSPWDR+
Sbjct: 91 VILTCKHHDGFCLWPTRTTDHSVAQSRWRGGRGDVVREISQAAARRKMKFGVYLSPWDRN 150
Query: 61 DSRYGHDLLYNEYYLAQLQELLKKYQDVREIWFD----------GAKDPRAQNVTYYFSD 110
++YG Y + Y AQL ELL Y + E+W D GAK+ R + Y+ D
Sbjct: 151 SAQYGTP-EYIKLYRAQLSELLTGYGPIFEVWHDGANGGDGYYGGAKEKRTIDKNTYY-D 208
Query: 111 W---FSMVKELQSSINIFSDAGPDVRWVGDETGTAGDTCWSTINRTSLSIGASN----IT 163
W + M++ +Q +FSD GPDVRWVG+E G AGD CW + GA++ T
Sbjct: 209 WPRTWEMIRAMQPEAVVFSDVGPDVRWVGNERGVAGDPCWVAFDPVGEDGGAASPGNVRT 268
Query: 164 QYLNTGDPKGTDWLPAECDVSIRPGWFWHKSESP--KKLSDLLDIYYKSVGRNCVLLLNV 221
+ G G+ WLPAECDVSIRPGWFWH++E+ KK S L+++YY+SVGR LLLNV
Sbjct: 269 KESGMGHRHGSKWLPAECDVSIRPGWFWHEAENARVKKASQLVNLYYESVGRGATLLLNV 328
Query: 222 PPNTTGLISENDAHRLKEFRSAIDTIFHKNIAENRYVKVSSQRGGKEGGFGPENMLDSDH 281
PPN GLIS D LK + F +N+A + R G + +GP +L+
Sbjct: 329 PPNRDGLISSEDVASLKGLGGYLSGTFAQNLAARAKTDATHVR-GNDKQYGPAQLLNGKP 387
Query: 282 LWSYWTPREDDKEKDHWIEIWGNDGSLRFNVIRIQEAIGLGQRIERYEI----YVDGKSI 337
++W + D ++ ++F VIR++EAI GQR++ + + + +
Sbjct: 388 -ETFWATDDGVTSADVTFDL---GRPVKFQVIRLREAIRFGQRVDAFTVERWQSDSWEQV 443
Query: 338 IQGTTIGYKRLHRLDGDVVHARVVRIRFIKARGVPLISSIGL 379
T+IG +RL RLD ++ R +R+R +A P +S +
Sbjct: 444 ASSTSIGPRRLIRLDAPIIATR-LRLRVTQASASPALSEFAV 484
>gb|AAS19690.1| FucA [Streptococcus gordonii]
Length = 576
Score = 269 bits (688), Expect = 1e-70
Identities = 155/395 (39%), Positives = 221/395 (55%), Gaps = 25/395 (6%)
Query: 1 MILTAKHHDGFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRH 60
+++ KHHDGF L+PS+Y+ ++V +S W++GKGD++ E +A+ +D+G+YLSPWD H
Sbjct: 81 VVVVVKHHDGFVLYPSRYSDYTVAASPWRDGKGDLLAEISRSASMYDMDLGLYLSPWDAH 140
Query: 61 DSRYGHDL--LYNEYYLAQLQELLKK-----YQDVREIWFDGAKDPRAQNVTYYFSDWFS 113
Y + YNEYY QL+E+L EIW DGA+ AQ +TY F WF
Sbjct: 141 SPLYHIETQEAYNEYYQNQLEEILSNPLYGNKGKFVEIWMDGARGEGAQKLTYDFGSWFE 200
Query: 114 MVKELQSSINIFSDAGPDVRWVGDETGTAGDTCWSTINRTSLSIGASNITQYLNTGDPKG 173
++ Q IFS ++RW+G+E G AGD W I LS + YL GDP+G
Sbjct: 201 PIRHYQKDALIFSTEATELRWIGNERGRAGDPLWQKIRPEKLS--ENTPATYLCHGDPQG 258
Query: 174 TDWLPAECDVSIRPGWFWHKSESPKKLSDLLDIYYKSVGRNCVLLLNVPPNTTGLISEND 233
T + E DVS+R GWF+H S+ PK L DLLDIY SVGR LLLNVPP GL++E D
Sbjct: 259 TQYSLGEADVSLRSGWFYHASQHPKSLPDLLDIYMDSVGRGTPLLLNVPPTKEGLLAEED 318
Query: 234 AHRLKEFRSAIDTIFHKNIAENRYVKVSSQRGGKEGGFGPENMLDSDHLWSYWTPREDDK 293
RL+EF I ++ N A V S+++ G P + L ++W P D
Sbjct: 319 VQRLQEFHQVISDLYTDNRAYQAKVSCSNEKEG-----FPSSHLTDGREDTFWAPASD-- 371
Query: 294 EKDHWIEIWGNDGSLR-FNVIRIQEAIGLGQRIERYEIYVDGKS----IIQGTTIGYKRL 348
E + +EI + G R FN++ I+E I GQRI + + V ++ G TIGYKRL
Sbjct: 372 ESAYVLEI--DLGQTRSFNLVEIREPITKGQRIAGFTLEVKQENGWIVFAHGQTIGYKRL 429
Query: 349 HRLDGDVVHARVVRIRFIKARGVPLISSIGLHFDP 383
L G +V AR +R+ + +PL++ + ++ P
Sbjct: 430 --LLGQMVEARYLRLILTDFQALPLLNKLAVYKTP 462
>ref|ZP_00403950.1| COG3669: Alpha-L-fucosidase [Streptococcus pneumoniae TIGR4]
gi|15901959|ref|NP_346563.1| hypothetical protein SP2146
[Streptococcus pneumoniae TIGR4]
gi|14973659|gb|AAK76203.1| conserved hypothetical
protein [Streptococcus pneumoniae TIGR4]
gi|25389606|pir||B95251 conserved hypothetical protein
SP2146 [imported] - Streptococcus pneumoniae (strain
TIGR4)
Length = 559
Score = 269 bits (687), Expect = 1e-70
Identities = 158/402 (39%), Positives = 222/402 (54%), Gaps = 40/402 (9%)
Query: 1 MILTAKHHDGFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRH 60
+IL KHHDGF L+P+ +T +SV S W+ GKGD++ E AAT+ +D+G+YLSPWD H
Sbjct: 71 LILVVKHHDGFVLYPTAHTDYSVKVSPWRRGKGDLLLEVSQAATEFDMDMGVYLSPWDAH 130
Query: 61 DSRYGHDLL--YNEYYLAQLQELLKKYQ-----DVREIWFDGAKDPRAQNVTYYFSDWFS 113
Y D YN YYLAQL+E+L E+W DGA+ AQ V Y F WF
Sbjct: 131 SPLYHVDREADYNAYYLAQLKEILSNPNYGNAGKFAEVWMDGARGEGAQKVNYEFEKWFE 190
Query: 114 MVKELQSSINIFSDAGPDVRWVGDETGTAGDTCWSTINRTSLSIGASNITQYLNTGDPKG 173
+++LQ IFS G +RW+G+E G AGD W +N L A YL GDP G
Sbjct: 191 TIRDLQGDCLIFSTEGTSIRWIGNERGYAGDPLWQKVNPDKLGTEAE--LNYLQHGDPSG 248
Query: 174 TDWLPAECDVSIRPGWFWHKSESPKKLSDLLDIYYKSVGRNCVLLLNVPPNTTGLISEND 233
T + E DVSIRPGWF+H+ + PK L +L++IY+ SVGR LLLN+PPN GL D
Sbjct: 249 TIFSIGEADVSIRPGWFYHEDQDPKSLEELVEIYFHSVGRGTPLLLNIPPNQAGLFDAKD 308
Query: 234 AHRLKEFRSAIDTIFHKNIAENRYVKVSSQRGGKEGGFGPENMLDSDHLWSYWTPREDDK 293
RL EF + + ++ +++A V GP L +D + T D
Sbjct: 309 IERLYEFATYRNELYKEDLALGAEVS------------GP--ALSADFACRHLT---DGL 351
Query: 294 EKDHW-------IEIWGNDGSLR-FNVIRIQEAIGLGQRIERYEIYVDGKSIIQ----GT 341
E W I++ + GS + F+VI ++E + LGQRI + + V+ + Q G
Sbjct: 352 ETSSWASDADLPIQLELDLGSPKTFDVIELREDLKLGQRIAAFHVQVEVDGVWQEFGSGH 411
Query: 342 TIGYKRLHRLDGDVVHARVVRIRFIKARGVPLISSIGLHFDP 383
T+GYKRL L G VV A+ +R+ +++ +PL++ I L+ P
Sbjct: 412 TVGYKRL--LRGAVVEAQKIRVVITESQALPLLTKISLYKTP 451
>ref|NP_359545.1| hypothetical protein spr1954 [Streptococcus pneumoniae R6]
gi|15459653|gb|AAL00756.1| Hypothetical protein
[Streptococcus pneumoniae R6] gi|25509375|pir||G98115
hypothetical protein spr1954 [imported] - Streptococcus
pneumoniae (strain R6)
Length = 583
Score = 267 bits (683), Expect = 4e-70
Identities = 153/395 (38%), Positives = 218/395 (54%), Gaps = 26/395 (6%)
Query: 1 MILTAKHHDGFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRH 60
+IL KHHDGF L+P+ +T +SV S W+ GKGD++ E AAT+ +D+G+YLSPWD H
Sbjct: 95 LILVVKHHDGFVLYPTAHTDYSVKVSPWRRGKGDLLLEVSQAATEFDMDMGVYLSPWDAH 154
Query: 61 DSRYGHDLL--YNEYYLAQLQELLKKYQ-----DVREIWFDGAKDPRAQNVTYYFSDWFS 113
Y D YN YYLAQL+E+L E+W DGA+ AQ V Y F WF
Sbjct: 155 SPLYHVDREADYNAYYLAQLKEILSNPNYGNAGKFAEVWMDGARGEGAQKVNYEFEKWFE 214
Query: 114 MVKELQSSINIFSDAGPDVRWVGDETGTAGDTCWSTINRTSLSIGASNITQYLNTGDPKG 173
+++LQ IFS G +RW+G+E G AGD W +N L A YL GDP G
Sbjct: 215 TIRDLQGDCLIFSTEGTSIRWIGNERGYAGDPLWQKVNLDKLGTEAE--LNYLQHGDPSG 272
Query: 174 TDWLPAECDVSIRPGWFWHKSESPKKLSDLLDIYYKSVGRNCVLLLNVPPNTTGLISEND 233
T + E DVSIRPGWF+H+ + PK L +L++IY+ SVGR LLLN+PPN GL D
Sbjct: 273 TIFSIGEADVSIRPGWFYHEDQDPKSLEELVEIYFHSVGRGTPLLLNIPPNQAGLFDAKD 332
Query: 234 AHRLKEFRSAIDTIFHKNIAENRYVKVSSQRGGKEGGFGPENMLDSDHLWSYWTPREDDK 293
R EF + + ++ +++A G + G D HL
Sbjct: 333 IERPYEFATYRNELYKEDLA----------LGAEVSGPALSADFDCRHLTDGLETSSWAS 382
Query: 294 EKDHWIEIWGNDGSLR-FNVIRIQEAIGLGQRIERYEIYVDGKSIIQ----GTTIGYKRL 348
+ D I++ + GS + F+VI ++E + LGQRI + + V+ + Q G T+G+KRL
Sbjct: 383 DADLPIQLELDLGSPKTFDVIELREDLKLGQRIAAFHVQVEVDGVWQEFGRGFTVGHKRL 442
Query: 349 HRLDGDVVHARVVRIRFIKARGVPLISSIGLHFDP 383
L G +V A+ VR+ +A+ +P+++ I L+ P
Sbjct: 443 --LRGPLVEAQKVRVMITEAQSIPVLTKISLYKTP 475
>gb|AAO79818.1| conserved hypothetical protein [Bacteroides thetaiotaomicron
VPI-5482] gi|29350121|ref|NP_813624.1| hypothetical
protein BT4713 [Bacteroides thetaiotaomicron VPI-5482]
Length = 580
Score = 256 bits (655), Expect = 6e-67
Identities = 156/413 (37%), Positives = 216/413 (51%), Gaps = 32/413 (7%)
Query: 1 MILTAKHHDGFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRH 60
+ILTAKHHDGFCLWP++ T++ + ++ +++GKGD+V+E +A GI +YLSPWDRH
Sbjct: 48 VILTAKHHDGFCLWPTQLTEYCIRNTPYKDGKGDIVRELSDACKKYGIKFAVYLSPWDRH 107
Query: 61 DSRYGHDLLYNEYYLAQLQELLKKYQDVREIWFD----------GAKDPRA-QNVTYY-F 108
+ YG Y +Y+ QL ELL Y DV EIWFD GAKD R TYY +
Sbjct: 108 QANYGTP-EYVDYFYKQLHELLTNYGDVFEIWFDGANGGDGWYGGAKDARTIDRKTYYDY 166
Query: 109 SDWFSMVKELQSSINIFSDAGPDVRWVGDETGTAGDTCWSTINRTSLSIGASNITQYLNT 168
+ M+ ELQ IFSD GP RWVG+E G AG T WS + + G + L
Sbjct: 167 PRAYKMIDELQPQAVIFSDGGPGCRWVGNENGFAGATNWSFLRAGEVYPGYPKYRE-LQY 225
Query: 169 GDPKGTDWLPAECDVSIRPGWFWHKSESP--KKLSDLLDIYYKSVGRNCVLLLNVPPNTT 226
G G W+ AECDVSIRPGWF+H E K + L D+YY+SVG N LLLN P +
Sbjct: 226 GHADGNQWVAAECDVSIRPGWFYHPEEDDKVKTVDQLTDLYYRSVGHNATLLLNFPVDRN 285
Query: 227 GLISENDAHRLKEFRSAIDTIFHKNIAENRYVKVSSQRGGKEGGFGPENMLDSDHLWSYW 286
GLI D+ F + N+ + V +RGG+ F + D + +YW
Sbjct: 286 GLIHPTDSLNAVSFHQRVQKELADNLLSSAKVSAFDERGGQ---FKVRAVTDGKY-DTYW 341
Query: 287 TPREDDKEKDHWIEIWGNDGSLRFNVIRIQEAIGLGQRIERYEI-YVDG------KSIIQ 339
+ D + N + IQE + LGQR++ + + Y +G K +
Sbjct: 342 ATNDGVTTADLTFTF---SQPTKMNRVMIQEYVPLGQRVKSFVVEYKEGDQWLSVKCNEE 398
Query: 340 GTTIGYKRLHRLDGDVVHARVVRIRFIKARGVPLISSIGLHFDPFWHSRFTAA 392
TT+GYKRL R +++ +RIRF AR I+ +G ++ P +T A
Sbjct: 399 TTTVGYKRLLRF--EMIETEELRIRFTDARACLCINEVGAYYAPDATENYTPA 449
>gb|AAO79241.1| conserved hypothetical protein [Bacteroides thetaiotaomicron
VPI-5482] gi|29349544|ref|NP_813047.1| hypothetical
protein BT4136 [Bacteroides thetaiotaomicron VPI-5482]
Length = 605
Score = 254 bits (649), Expect = 3e-66
Identities = 154/403 (38%), Positives = 217/403 (53%), Gaps = 57/403 (14%)
Query: 1 MILTAKHHDGFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRH 60
+ILTAKHHDGFCLWP+ TKHSV SS W+NG+GDVVKE A G+ G+YLSPWDR+
Sbjct: 111 VILTAKHHDGFCLWPTATTKHSVASSSWKNGQGDVVKELRKACKKYGMRFGLYLSPWDRN 170
Query: 61 DSRYGHDLLYNEYYLAQLQELLKKYQDVREIWFDGA--KDPRAQNVTYYFSDWFSMVKEL 118
YG YN++++ QL ELL Y +V E+WFDGA + P + Y + + + L
Sbjct: 171 AECYGDSPRYNKFFIRQLTELLTNYGEVHEVWFDGANGEGPNGKKQVYDWDTVYETIHRL 230
Query: 119 QSSINIFSDAGPDVRWVGDETGTAGDTCWST----------INRTSLSIGASNITQYLNT 168
Q + + G D+RWVG+E+G +T WST ++ + +G + + L +
Sbjct: 231 QPKA-VMAIMGDDIRWVGNESGLGRETEWSTTVLTPEIYARADKNNKKLGINGQSNDLGS 289
Query: 169 GD--PKGTD--WLPAECDVSIRPGWFWHKSE--SPKKLSDLLDIYYKSVGRNCVLLLNVP 222
K T+ W P+E DVSIRPGWF+HK E K L L DIY++SVG N VLLLN+P
Sbjct: 290 RKMLEKATELFWYPSEVDVSIRPGWFYHKEEDNKVKSLKHLADIYFQSVGYNSVLLLNIP 349
Query: 223 PNTTGLISENDAHRLKEFRSAIDTIFHKNIAENRYVKVSSQRGGKEGGFGPENMLDSDHL 282
P+ GLI E D RLKEF + + K + + YVK +G K
Sbjct: 350 PDHRGLIHEADVQRLKEFAA-----YRKQVFADNYVK----KGKK--------------- 385
Query: 283 WSYWTPREDDKEKDHWIEIWGNDGSLRFNVIRIQEAIGLGQRIERYEIYV----DGKSII 338
YW ++ +I+ NV+ +QE I GQRIE + + K +
Sbjct: 386 --YWNTASGNE------KIYQLKAGSEINVVMLQEDITKGQRIEAFTVEALTDNSWKEVA 437
Query: 339 QGTTIGYKRLHRLDGDVVHARVVRIRFIKARGVPLISSIGLHF 381
+GTT+GYKRL R + A +RI+ +++R IS + ++
Sbjct: 438 KGTTVGYKRLLRF--PTIKADQLRIKILESRLNANISQVAAYY 478
>emb|CAH08778.1| putative exported fucosidase [Bacteroides fragilis NCTC 9343]
gi|60682552|ref|YP_212696.1| putative exported
fucosidase [Bacteroides fragilis NCTC 9343]
Length = 605
Score = 253 bits (646), Expect = 7e-66
Identities = 160/402 (39%), Positives = 216/402 (52%), Gaps = 57/402 (14%)
Query: 2 ILTAKHHDGFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRHD 61
ILTAKHHDGFCLWP+K T HSV +S W++GKGDVV+E +A GI G+YLSPWDR+
Sbjct: 112 ILTAKHHDGFCLWPTKTTGHSVAASPWKDGKGDVVRELRDACDKYGIKFGVYLSPWDRNA 171
Query: 62 SRYGHDLLYNEYYLAQLQELLKKYQDVREIWFDGA--KDPRAQNVTYYFSDWFSMVKELQ 119
S YG YNE+++ QL ELL Y +V E+WFDGA + P + Y ++ S ++ LQ
Sbjct: 172 SCYGDSPKYNEFFIEQLTELLTNYGEVHEVWFDGANGEGPNGKKQEYDWTAILSTIRRLQ 231
Query: 120 SSINIFSDAGPDVRWVGDETGTAGDTCWSTINRT----------SLSIGASNITQYLNTG 169
+ + G DVRWVG+E G +T WS T + ++G ++ L
Sbjct: 232 PRA-VTAIMGDDVRWVGNERGLGRETEWSATVLTPGTYARCEEQNKALGVKATSKDLGGR 290
Query: 170 D----PKGTDWLPAECDVSIRPGWFWHKSE--SPKKLSDLLDIYYKSVGRNCVLLLNVPP 223
D K W P+E DVSIRPGWF+H+ E K L L DIY+KSVG N VLLLN+PP
Sbjct: 291 DMLVNAKELFWYPSEVDVSIRPGWFYHQQEDNQVKSLKHLTDIYFKSVGYNSVLLLNIPP 350
Query: 224 NTTGLISENDAHRLKEFRSAIDTIFHKNIAENRYVKVSSQRGGKEGGFGPENMLDSDHLW 283
+ G IS+ D +RLKEF IF A+NR +GG +
Sbjct: 351 DQRGRISDADVNRLKEFADYRKEIF----ADNRV------KGGLKA-------------- 386
Query: 284 SYWTPREDDKEKDHWIEIWGNDGSLRFNVIRIQEAIGLGQRIERYEI---YVDG-KSIIQ 339
WT R D ++ NV+ ++E I GQR+E + + DG K I +
Sbjct: 387 --WTARPGD------TRVYQLKPKSEINVVMLREDISKGQRMEAFTVEALTADGWKEIAK 438
Query: 340 GTTIGYKRLHRLDGDVVHARVVRIRFIKARGVPLISSIGLHF 381
GTT+GYKRL R+ V AR +R++ R IS + ++
Sbjct: 439 GTTVGYKRLIRI--PAVEARQLRVKVDACRLAANISEVAAYY 478
>ref|YP_100522.1| hypothetical protein BF3243 [Bacteroides fragilis YCH46]
gi|52217395|dbj|BAD49988.1| conserved hypothetical
protein [Bacteroides fragilis YCH46]
Length = 605
Score = 251 bits (642), Expect = 2e-65
Identities = 159/402 (39%), Positives = 215/402 (52%), Gaps = 57/402 (14%)
Query: 2 ILTAKHHDGFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRHD 61
ILTAKHHDGFCLWP+K T HSV +S W++GKGDVV+E +A GI G+YLSPWDR+
Sbjct: 112 ILTAKHHDGFCLWPTKTTGHSVAASPWKDGKGDVVRELRDACDKYGIKFGVYLSPWDRNA 171
Query: 62 SRYGHDLLYNEYYLAQLQELLKKYQDVREIWFDGA--KDPRAQNVTYYFSDWFSMVKELQ 119
S YG YNE+++ QL ELL Y +V E+WFDGA + P + Y ++ S ++ LQ
Sbjct: 172 SCYGDSPKYNEFFIEQLTELLTNYGEVHEVWFDGANGEGPNGKKQEYDWTAILSTIRRLQ 231
Query: 120 SSINIFSDAGPDVRWVGDETGTAGDTCWSTINRT----------SLSIGASNITQYLNTG 169
+ + G DVRWVG+E G +T WS T + ++G ++ L
Sbjct: 232 PRA-VTAIMGDDVRWVGNERGLGRETEWSATVLTPGTYARCEEQNKALGVKATSKDLGGR 290
Query: 170 D----PKGTDWLPAECDVSIRPGWFWHKSE--SPKKLSDLLDIYYKSVGRNCVLLLNVPP 223
D K W P+E DVSIRPGWF+H+ E K L L DIY+KSVG N VLLLN+PP
Sbjct: 291 DMLVNAKELFWYPSEVDVSIRPGWFYHQQEDNQVKSLKHLTDIYFKSVGYNSVLLLNIPP 350
Query: 224 NTTGLISENDAHRLKEFRSAIDTIFHKNIAENRYVKVSSQRGGKEGGFGPENMLDSDHLW 283
+ G IS+ D +RLKEF IF +NR +GG +
Sbjct: 351 DQRGRISDADVNRLKEFADYRKEIF----TDNRV------KGGLKA-------------- 386
Query: 284 SYWTPREDDKEKDHWIEIWGNDGSLRFNVIRIQEAIGLGQRIERYEI---YVDG-KSIIQ 339
WT R D ++ NV+ ++E I GQR+E + + DG K I +
Sbjct: 387 --WTARPGD------TRVYQLKPKSEINVVMLREDISKGQRMEAFTVEALTADGWKEIAK 438
Query: 340 GTTIGYKRLHRLDGDVVHARVVRIRFIKARGVPLISSIGLHF 381
GTT+GYKRL R+ V AR +R++ R IS + ++
Sbjct: 439 GTTVGYKRLIRI--PAVEARQLRVKVDACRLAANISEVAAYY 478
>gb|AAO77299.1| conserved hypothetical protein [Bacteroides thetaiotaomicron
VPI-5482] gi|29347602|ref|NP_811105.1| hypothetical
protein BT2192 [Bacteroides thetaiotaomicron VPI-5482]
Length = 484
Score = 251 bits (640), Expect = 4e-65
Identities = 159/398 (39%), Positives = 219/398 (54%), Gaps = 34/398 (8%)
Query: 2 ILTAKHHDGFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRHD 61
ILTAKH DGFCLWPSKYT +SV ++ W+NGKGDVV+EFV+A + G+ GIYL P DRH+
Sbjct: 94 ILTAKHADGFCLWPSKYTDYSVKNAAWKNGKGDVVREFVDACEEYGLKAGIYLGPHDRHE 153
Query: 62 --SRYGHDLLYNEYYLAQLQELLKKYQDVREIWFDGAKDPRAQNVTYYFSDWFSMVKELQ 119
S Y EYY QL EL+ Y + E W+DGA + T + W+ +V+E Q
Sbjct: 154 HLSPLYTTERYKEYYAHQLGELMSDYGKIWETWWDGA--GADELTTPVYRHWYKIVREKQ 211
Query: 120 SSINIFSDAG----PDVRWVGDETGTAGDTCWSTINRTSLSIGASNITQY---LNTGDPK 172
IF DVRW+G+E G AGD CW+T + S+ + QY LN G
Sbjct: 212 PDCVIFGTKNSYPFADVRWMGNEAGEAGDPCWATTD----SVAIRDEAQYYKGLNEGMLD 267
Query: 173 GTDWLPAECDVSIRPGWFWHKSESP--KKLSDLLDIYYKSVGRNCVLLLNVPPNTTGLIS 230
G ++PAE DVSIRP WF+H E K + +L DIY SVGRN VLLLN PP+ GLI
Sbjct: 268 GDAYIPAETDVSIRPSWFYHAEEDSRVKSVRELWDIYCTSVGRNSVLLLNFPPDRRGLIH 327
Query: 231 ENDAHRLKEFRSAIDTIFHKNIAENRYVKVSSQRGGKEGGFGPENMLDSDHLWSYWTPRE 290
D+ + ID F N+ VK ++ RG K + PE MLD++ +Y+ ++
Sbjct: 328 STDSLHAALLKQGIDETFSTNLLRGAKVKATNVRGAK---YSPEKMLDNEKN-TYFAGKD 383
Query: 291 DDKEKDHWIEIWGNDGSLRFNVIRIQEAIGLGQRIERY--EIYVDGKSII------QGTT 342
+ + D I+ ++ F+ + I+E I LG R ++ E VDGK+ I
Sbjct: 384 GEVKAD---IIFTLPKTIEFDCLMIEEVIELGHRTTKWSVEYTVDGKNWITIPEATDKQA 440
Query: 343 IGYKRLHRLDGDVVHARVVRIRFIKARGVPLISSIGLH 380
IG+K + RL V A+ VR+R + P I + G++
Sbjct: 441 IGHKWIVRL--APVKAKQVRLRIQDGKACPAIHTFGVY 476
>emb|CAH06409.1| conserved hypothetical exported protein [Bacteroides fragilis NCTC
9343] gi|60680223|ref|YP_210367.1| hypothetical protein
BF0664 [Bacteroides fragilis NCTC 9343]
gi|53712028|ref|YP_098020.1| hypothetical protein BF0735
[Bacteroides fragilis YCH46] gi|52214893|dbj|BAD47486.1|
conserved hypothetical protein [Bacteroides fragilis
YCH46]
Length = 483
Score = 246 bits (628), Expect = 9e-64
Identities = 156/395 (39%), Positives = 217/395 (54%), Gaps = 28/395 (7%)
Query: 2 ILTAKHHDGFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRHD 61
I+TAKH DGFCLWPSKYT + V +S W+NGKGDVV+EFV+A + GI GIYL P DRH+
Sbjct: 94 IITAKHADGFCLWPSKYTDYGVKNSAWKNGKGDVVREFVDACEEYGIKAGIYLGPHDRHE 153
Query: 62 --SRYGHDLLYNEYYLAQLQELLKKYQDVREIWFDGAKDPRAQNVTYYFSDWFSMVKELQ 119
S Y +YY QL+EL+ Y V E W+DGA + T ++ W+ +V+E Q
Sbjct: 154 HLSPLYTTEKYKQYYGHQLEELMGDYGKVWETWWDGA--GADELTTPVYTHWYKIVREKQ 211
Query: 120 SSINIFSDAG----PDVRWVGDETGTAGDTCWSTINRTSLSIGASNITQYLNTGDPKGTD 175
IF DVRW+G+E+G AGD CWST + + N + LN G G
Sbjct: 212 PDCVIFGTKNSYPFADVRWMGNESGKAGDPCWSTTDSVCVRDEWKNY-EGLNEGVKGGDA 270
Query: 176 WLPAECDVSIRPGWFWHKSESP--KKLSDLLDIYYKSVGRNCVLLLNVPPNTTGLISEND 233
++PAE DVSIRP WF+H E K + +L DIY SVG N VLLLN PP+ GLI D
Sbjct: 271 YIPAETDVSIRPSWFYHAEEDSRVKSVKELWDIYCTSVGHNSVLLLNFPPDRRGLIHPTD 330
Query: 234 AHRLKEFRSAIDTIFHKNIAENRYVKVSSQRGGKEGGFGPENMLDSDHLWSYWTPREDDK 293
+ + +D F N+ VK ++ RGGK F PE + D++ +Y+ R+ K
Sbjct: 331 SLHAALLKQGLDETFGNNLLAKAKVKATNGRGGK---FRPEFLTDNNK-ETYFAGRDGAK 386
Query: 294 EKDHWIEIWGNDGSLRFNVIRIQEAIGLGQRIERYEIYV--DGKS---IIQGT---TIGY 345
D ++ F+ + IQE I LG R ++ + DG++ I + T T+GY
Sbjct: 387 TSD---IVFTLPRQTEFDCLMIQEVIELGHRTTKWSVEYSNDGRNWTPIPEATDKQTVGY 443
Query: 346 KRLHRLDGDVVHARVVRIRFIKARGVPLISSIGLH 380
K + R + V A+ VR+R + P I + G++
Sbjct: 444 KWIVRF--EPVKAKQVRLRILDGFACPAIHTFGVY 476
>gb|AAO76732.1| conserved hypothetical protein [Bacteroides thetaiotaomicron
VPI-5482] gi|29347035|ref|NP_810538.1| hypothetical
protein BT1625 [Bacteroides thetaiotaomicron VPI-5482]
Length = 605
Score = 244 bits (622), Expect = 4e-63
Identities = 152/406 (37%), Positives = 215/406 (52%), Gaps = 58/406 (14%)
Query: 1 MILTAKHHDGFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRH 60
++LTAKHHDGFCLWP+ TKHSV SS W+NG+GDVVKE NA + G+YLSPWDR+
Sbjct: 111 VLLTAKHHDGFCLWPTATTKHSVASSPWKNGQGDVVKELRNACDKYDMKFGVYLSPWDRN 170
Query: 61 DSRYGHDLLYNEYYLAQLQELLKKYQDVREIWFDGA--KDPRAQNVTYYFSDWFSMVKEL 118
YG YNE+++ QL ELL Y +V E+WFDGA + P + Y + ++ +++L
Sbjct: 171 AECYGDSPKYNEFFIRQLTELLTNYGEVHEVWFDGANGEGPNGKKQIYDWDAFYKTIQQL 230
Query: 119 QSSINIFSDAGPDVRWVGDETGTAGDTCWSTI----------NRTSLSIGASNITQYLNT 168
Q + + G DVRWVG+E G +T WS + +G + + L +
Sbjct: 231 QPKA-VMAIMGDDVRWVGNEKGLGRETEWSATVLTPGIYARSEENNKRLGVFSKAEDLGS 289
Query: 169 GD--PKGTD--WLPAECDVSIRPGWFWHKSESP--KKLSDLLDIYYKSVGRNCVLLLNVP 222
K T+ W P+E DVSIRPGWF+H E K L L DIY++SVG N VLLLN+P
Sbjct: 290 RAMLEKATELFWYPSEVDVSIRPGWFYHAEEDSKVKSLKHLADIYFQSVGYNSVLLLNIP 349
Query: 223 PNTTGLISENDAHRLKEFRSAIDTIFHKNIAENRYVKVSSQRGGKEGGFGPENMLDSDHL 282
P+ GLI+E D RL EF + + IF N E +G K+ E + S+ +
Sbjct: 350 PDRRGLINEADVQRLNEFAAYREKIFTNNRVE---------KGRKDW----EAVSGSETV 396
Query: 283 WSYWTPREDDKEKDHWIEIWGNDGSLRFNVIRIQEAIGLGQRIERYEIYV----DGKSII 338
+S E NV+ +QE I GQR+E + + + +
Sbjct: 397 YSLKPESE-------------------INVVMLQEDITKGQRVESFTVEALTEQGWQEVA 437
Query: 339 QGTTIGYKRLHRLDGDVVHARVVRIRFIKARGVPLISSIGLHF-DP 383
+GTT+GYKR+ R V A +R++ + R IS + ++ DP
Sbjct: 438 KGTTVGYKRMVRF--PAVKATQLRVKINECRLTAHISQVAAYYADP 481
>gb|AAQ66752.1| alpha-1,3/4-fucosidase, putative [Porphyromonas gingivalis W83]
gi|34541374|ref|NP_905853.1| alpha-1,3/4-fucosidase,
putative [Porphyromonas gingivalis W83]
Length = 606
Score = 243 bits (621), Expect = 6e-63
Identities = 151/393 (38%), Positives = 206/393 (51%), Gaps = 61/393 (15%)
Query: 1 MILTAKHHDGFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRH 60
+ILTAKHHDGFCLWP+ T+HSV SS W+ GKGDVVKE A + G+ GIYLSPWDR+
Sbjct: 113 VILTAKHHDGFCLWPTATTRHSVASSPWREGKGDVVKEVRAACEEYGMKFGIYLSPWDRN 172
Query: 61 DSRYGHDLLYNEYYLAQLQELLKKYQDVREIWFDGA--KDPRAQNVTYYFSDWFSMVKEL 118
YG YN ++++QL ELL Y +V E+WFDGA + P + Y + ++ ++ L
Sbjct: 173 AECYGDSHRYNRFFVSQLTELLTHYGEVHEVWFDGANGEGPNGKRQEYDWETFYDTIRRL 232
Query: 119 QSSINIFSDAGPDVRWVGDETGTAGDTCWSTI--------------NRTSLSIGASNITQ 164
Q + + G DVRWVG+E G T WS NR + + ++
Sbjct: 233 QPQA-VMAIMGDDVRWVGNERGLGRTTEWSATVLTPGIYSRSKTERNRLGIRENSPDLGS 291
Query: 165 YLNTGDPKGTDWLPAECDVSIRPGWFWHKSESP--KKLSDLLDIYYKSVGRNCVLLLNVP 222
+ W P+E DVSIRPGWF+H +E K L L+DIY++SVG N VLLLNVP
Sbjct: 292 REVLKEAGEIFWYPSEVDVSIRPGWFYHAAEDKKVKSLDKLVDIYFQSVGYNSVLLLNVP 351
Query: 223 PNTTGLISENDAHRLKEFRSAIDTIF-HKNIAEN-RYVKVSSQRGGKEGGFGPENMLDSD 280
P+ GLI E DA RLKE+ + F H + ++ RY V
Sbjct: 352 PDRRGLIHEADALRLKEWADYLGRAFAHDRVVDSARYAVV-------------------- 391
Query: 281 HLWSYWTPREDDKEKDHWIEIWGNDGSLRFNVIRIQEAIGLGQRIERY--EIYVDG--KS 336
+ H +E + + N++ +QE I GQR E + E +VDG +
Sbjct: 392 --------------QGHAVEEYALESKTHINILMLQEDITKGQRTEAFTVEAWVDGAWQE 437
Query: 337 IIQGTTIGYKRLHRLDGDVVHARVVRIRFIKAR 369
I +GTTIGYKRL R+ V +R+R + R
Sbjct: 438 IGRGTTIGYKRLLRI--PAVETDRIRVRIEQCR 468
>emb|CAH08896.1| putative lipoprotein [Bacteroides fragilis NCTC 9343]
gi|60682670|ref|YP_212814.1| putative lipoprotein
[Bacteroides fragilis NCTC 9343]
Length = 626
Score = 243 bits (621), Expect = 6e-63
Identities = 152/403 (37%), Positives = 211/403 (51%), Gaps = 34/403 (8%)
Query: 1 MILTAKHHDGFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRH 60
+ILTAKHHDGFCLWP++ T++ + ++ +++GKGD+V E A GI +YLSPWDRH
Sbjct: 93 VILTAKHHDGFCLWPTQLTEYCIRNTPYKDGKGDIVGELAAACKKYGIKFAVYLSPWDRH 152
Query: 61 DSRYGHDLLYNEYYLAQLQELLKKYQDVREIWFD----------GAKDPRA-QNVTYY-F 108
+ YG Y +Y+ QL EL+ Y +V E+WFD GAKD R TYY +
Sbjct: 153 QANYGTP-EYVDYFHKQLTELMTNYGEVFEVWFDGANGGDGWYGGAKDSRTIDRKTYYNY 211
Query: 109 SDWFSMVKELQSSINIFSDAGPDVRWVGDETGTAGDTCWSTINRTSLSIGASNITQYLNT 168
+ ++ +LQ +FSD GP RWVG+E G AG T WS + + G + L
Sbjct: 212 PRIYEILDKLQPQAIVFSDGGPGCRWVGNENGFAGATNWSFLRAGEVYPGYPKYRE-LQY 270
Query: 169 GDPKGTDWLPAECDVSIRPGWFWHKSESP--KKLSDLLDIYYKSVGRNCVLLLNVPPNTT 226
G G W+PAECDVSIRPGWF+H E K + L D+YY+SVG N LLLN P +
Sbjct: 271 GHADGNQWVPAECDVSIRPGWFYHPEEDDRVKTVEQLTDLYYRSVGHNATLLLNFPVDRD 330
Query: 227 GLISENDAHRLKEFRSAIDTIFHKNIAENRYVKVSSQRGGKEGGFGPENMLDSDHLW-SY 285
GLI D+ F + N+ K S +RGG+ +D W +Y
Sbjct: 331 GLIHPIDSANAVNFHKNVQKQLAHNLLAGIRPKASDERGGQFSA-----KAATDESWDTY 385
Query: 286 WTPREDDKEKDHWIEIWGNDGSLRFNVIRIQEAIGLGQRIERY--EIYVDGKSI-----I 338
W + D + + + N + IQE I LGQR++ + E DGK +
Sbjct: 386 WATNDGVTAADIEFDFPKTE---KVNRMMIQEYIPLGQRVKSFIVEYDKDGKWLPVKLNE 442
Query: 339 QGTTIGYKRLHRLDGDVVHARVVRIRFIKARGVPLISSIGLHF 381
+ TT+GYKRL R + V +RIRF AR I++I ++
Sbjct: 443 ETTTVGYKRLLRF--ETVSTDKLRIRFTDARACLCINNIEAYY 483
>ref|YP_100649.1| hypothetical protein BF3371 [Bacteroides fragilis YCH46]
gi|52217522|dbj|BAD50115.1| conserved hypothetical
protein [Bacteroides fragilis YCH46]
Length = 626
Score = 243 bits (621), Expect = 6e-63
Identities = 152/403 (37%), Positives = 211/403 (51%), Gaps = 34/403 (8%)
Query: 1 MILTAKHHDGFCLWPSKYTKHSVISSKWQNGKGDVVKEFVNAATDKGIDVGIYLSPWDRH 60
+ILTAKHHDGFCLWP++ T++ + ++ +++GKGD+V E A GI +YLSPWDRH
Sbjct: 93 VILTAKHHDGFCLWPTQLTEYCIRNTPYKDGKGDIVGELAAACKKYGIKFAVYLSPWDRH 152
Query: 61 DSRYGHDLLYNEYYLAQLQELLKKYQDVREIWFD----------GAKDPRA-QNVTYY-F 108
+ YG Y +Y+ QL EL+ Y +V E+WFD GAKD R TYY +
Sbjct: 153 QANYGTP-EYVDYFHKQLTELMTNYGEVFEVWFDGANGGDGWYGGAKDSRTIDRKTYYNY 211
Query: 109 SDWFSMVKELQSSINIFSDAGPDVRWVGDETGTAGDTCWSTINRTSLSIGASNITQYLNT 168
+ ++ +LQ +FSD GP RWVG+E G AG T WS + + G + L
Sbjct: 212 PRIYEILDKLQPQAIVFSDGGPGCRWVGNENGFAGATNWSFLRAGEVYPGYPKYRE-LQY 270
Query: 169 GDPKGTDWLPAECDVSIRPGWFWHKSESP--KKLSDLLDIYYKSVGRNCVLLLNVPPNTT 226
G G W+PAECDVSIRPGWF+H E K + L D+YY+SVG N LLLN P +
Sbjct: 271 GHADGNQWVPAECDVSIRPGWFYHPEEDDRVKTVEQLTDLYYRSVGHNATLLLNFPVDRD 330
Query: 227 GLISENDAHRLKEFRSAIDTIFHKNIAENRYVKVSSQRGGKEGGFGPENMLDSDHLW-SY 285
GLI D+ F + N+ K S +RGG+ +D W +Y
Sbjct: 331 GLIHPIDSANAVNFHKNVQKQLAHNLLAGIRPKASDERGGQFSA-----KAATDESWDTY 385
Query: 286 WTPREDDKEKDHWIEIWGNDGSLRFNVIRIQEAIGLGQRIERY--EIYVDGKSI-----I 338
W + D + + + N + IQE I LGQR++ + E DGK +
Sbjct: 386 WATNDGVTAADIEFDFPKTE---KVNRMMIQEYIPLGQRVKSFIVEYDKDGKWLPVKLNE 442
Query: 339 QGTTIGYKRLHRLDGDVVHARVVRIRFIKARGVPLISSIGLHF 381
+ TT+GYKRL R + V +RIRF AR I++I ++
Sbjct: 443 ETTTVGYKRLLRF--ETVSTDKLRIRFTDARACLCINNIEAYY 483
>gb|EAA55123.1| hypothetical protein MG06780.4 [Magnaporthe grisea 70-15]
gi|39977791|ref|XP_370283.1| hypothetical protein
MG06780.4 [Magnaporthe grisea 70-15]
Length = 519
Score = 229 bits (584), Expect = 1e-58
Identities = 149/413 (36%), Positives = 208/413 (50%), Gaps = 44/413 (10%)
Query: 1 MILTAKHHDGFCLWPSKYTKHSVISSKWQNGK------GDVVKEFVNAATDKGIDVGIYL 54
MILTAKHHDG LW + T + + + KW + DVV+ +A G+ G+YL
Sbjct: 107 MILTAKHHDGMALWNTSSTTYKIANGKWAKDREAERLDADVVRMAATSAKKHGLKFGVYL 166
Query: 55 SPWDRH-DSRYGHDLL------------------YNEYYLAQLQELL-KKYQD-----VR 89
SPWD H D L YNE Y+ QL EL+ K +D +
Sbjct: 167 SPWDIHRDPAMPKPTLAGTIFDEAQIFGDGSPGDYNELYVQQLTELVDMKLEDGSNVELF 226
Query: 90 EIWFDGAKDPRAQNVTYYFSDWFSMVKELQSSINIFSDAGPDVRWVGDETGTAGDTCWST 149
E+W DGA A T+ +S + +++ Q ++ GPD RWVG+E G + T W T
Sbjct: 227 EVWLDGASG-SATVQTFDWSRYREVIRTHQPGAVMWGHQGPDARWVGNEDGYSVQTNWHT 285
Query: 150 INRTSLSIGASNITQYLNTGDPKGTDWLPAECDVSIRPGWFWHKSESPKKLSDLLDIYYK 209
I+RT + L TG G W PAE D +R GWFWH E PK DL+D+Y
Sbjct: 286 ISRTQDQERYGE--RELETGVRDGLYWTPAEADARMRAGWFWHAEEKPKTAKDLMDMYMG 343
Query: 210 SVGRNCVLLLNVPPNTTGLISENDAHRLKEFRSAIDTIFHKNIAE-NRYVKVSSQRGGKE 268
SVGR+ LLLNV P+ TG I + D L EF+ D F + + V SS R G
Sbjct: 344 SVGRSVNLLLNVGPDNTGRIPQVDVDALMEFKELRDGFFERKLLRPGLGVSASSVRAGDA 403
Query: 269 GGFGPENMLDSDHLWSYWTPREDDKEKDHWIEIWGNDGSLRFNVIRIQEAIGLGQRIERY 328
FGPEN+LD D +YWT +D +++ G++ + IQE I LGQR+ Y
Sbjct: 404 MQFGPENVLD-DRQDTYWTMEDDQTTGSLEVDV---GGTITIEAVAIQEHIALGQRVGGY 459
Query: 329 --EIYVDG--KSIIQGTTIGYKRLHRLDGDVVHARVVRIRFIKARGVPLISSI 377
+++ D K ++ GT++GY R+ RL+ + R +R+R +A VP+I I
Sbjct: 460 AFDVFSDDAWKEVVSGTSVGYGRIDRLNTTMTGTR-LRLRVTQANAVPMIQGI 511
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.319 0.138 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 739,812,355
Number of Sequences: 2540612
Number of extensions: 33050253
Number of successful extensions: 68298
Number of sequences better than 10.0: 124
Number of HSP's better than 10.0 without gapping: 60
Number of HSP's successfully gapped in prelim test: 64
Number of HSP's that attempted gapping in prelim test: 67984
Number of HSP's gapped (non-prelim): 193
length of query: 392
length of database: 863,360,394
effective HSP length: 130
effective length of query: 262
effective length of database: 533,080,834
effective search space: 139667178508
effective search space used: 139667178508
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 76 (33.9 bits)
Medicago: description of AC149581.7


