

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149576.4 + phase: 0 /pseudo
(548 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
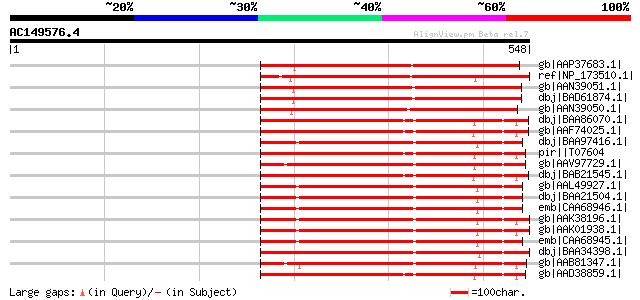
Score E
Sequences producing significant alignments: (bits) Value
gb|AAP37683.1| At1g76430 [Arabidopsis thaliana] gi|15223045|ref|... 353 8e-96
ref|NP_173510.1| phosphate transporter family protein [Arabidops... 349 1e-94
gb|AAN39051.1| putative phosphate transporter OsPT10 [Oryza sati... 300 1e-79
dbj|BAD61874.1| putative phosphate transporter [Oryza sativa (ja... 300 1e-79
gb|AAN39050.1| putative phosphate transporter OsPT9 [Oryza sativ... 295 3e-78
dbj|BAA86070.1| Phospate transpoter [Nicotiana tabacum] 278 2e-73
gb|AAF74025.1| inorganic phosphate transporter [Nicotiana tabacum] 276 9e-73
dbj|BAA97416.1| inorganic phosphate transporter [Arabidopsis tha... 276 2e-72
pir||T07604 phosphate transport protein PT1 - potato gi|1420871|... 275 2e-72
gb|AAV97729.1| phosphate transporter 3 [Lycopersicon esculentum] 275 3e-72
dbj|BAB21545.1| phosphate transporter [Nicotiana tabacum] 275 3e-72
gb|AAL49927.1| putative phosphate transporter protein [Arabidops... 275 4e-72
dbj|BAA21504.1| inorganic phosphate transporter [Arabidopsis tha... 275 4e-72
emb|CAA68946.1| phosphate transporter [Arabidopsis thaliana] 275 4e-72
gb|AAK38196.1| phosphate transporter 1 [Lupinus albus] 274 5e-72
gb|AAK01938.1| phosphate transporter 1 [Lupinus albus] 274 6e-72
emb|CAA68945.1| phosphate transporter [Arabidopsis thaliana] 273 8e-72
dbj|BAA34398.1| Phosphate Transporter 4 [Arabidopsis thaliana] g... 273 1e-71
gb|AAB81347.1| phosphate transporter [Medicago truncatula] gi|74... 273 1e-71
gb|AAD38859.1| inorganic phosphate transporter [Solanum tuberosum] 272 2e-71
>gb|AAP37683.1| At1g76430 [Arabidopsis thaliana] gi|15223045|ref|NP_177769.1|
phosphate transporter family protein [Arabidopsis
thaliana] gi|12323970|gb|AAG51941.1| putative phosphate
transporter; 18176-15417 [Arabidopsis thaliana]
gi|6554476|gb|AAF16658.1| putative phosphate
transporter; 23587-26346 [Arabidopsis thaliana]
gi|25309088|pir||A96792 probable phosphate transporter,
18176-15417 [imported] - Arabidopsis thaliana
Length = 532
Score = 353 bits (906), Expect = 8e-96
Identities = 175/277 (63%), Positives = 214/277 (77%), Gaps = 5/277 (1%)
Query: 266 YTALVEQNVLQAAKDMEKVLDVSM-SQITEEHPLP---PATNVAYPLLSREFLWRHGRDL 321
YTALVE NV+QAAKDM++V+ VSM SQITE+ P ++ +Y L SR FL HGRDL
Sbjct: 238 YTALVENNVVQAAKDMQRVMSVSMISQITEDSSSELEQPPSSSSYKLFSRRFLSLHGRDL 297
Query: 322 FACSANWFLLDIVFYSQVLFQSEIYKRYLNEKDDEDVYQEAFHLARIQAILAVCSTIPGY 381
FA SANWFL+D+VFY+ L S+I+ + +VY AF +A++ AI+A CSTIPGY
Sbjct: 298 FAASANWFLVDVVFYTSNLLLSQIFNFSNKPLNSTNVYDSAFEVAKLAAIVAACSTIPGY 357
Query: 382 FFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFA 441
+FTVYFID++GRVKIQMMGFF MAV + G PY +W+K E NKGFMV+YGL FFF+
Sbjct: 358 WFTVYFIDKIGRVKIQMMGFFLMAVVYLVAGIPYSWYWSKHEK-TNKGFMVLYGLIFFFS 416
Query: 442 NFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEKEEGYPKGIG 501
NFGPNTTTFI+PAELFPARFRSTCHGISGA GK GAI+G+VGFLWA+ +E+G+P
Sbjct: 417 NFGPNTTTFIIPAELFPARFRSTCHGISGAAGKFGAIVGTVGFLWATRHHEEDGFPDVKR 476
Query: 502 MKASLIILGGVCIVGMFVTYFFTKETMGRSLEENEDE 538
++ + +ILGGVCI GM VTY FT+ETMGRSLEENEDE
Sbjct: 477 VRIAFLILGGVCIAGMIVTYLFTRETMGRSLEENEDE 513
>ref|NP_173510.1| phosphate transporter family protein [Arabidopsis thaliana]
Length = 534
Score = 349 bits (895), Expect = 1e-94
Identities = 177/291 (60%), Positives = 219/291 (74%), Gaps = 11/291 (3%)
Query: 266 YTALVEQNVLQAAKDMEKVLDVSMSQITEE---HPLPPATNVAYPLLSREFLWRHGRDLF 322
YTALVE N++QAAKDM++V+ S S I++E P PP +Y L SR F HGRDLF
Sbjct: 230 YTALVENNIVQAAKDMQRVM--SRSHISDEATTDPPPPPPPPSYKLFSRCFFRLHGRDLF 287
Query: 323 ACSANWFLLDIVFYSQVLFQSEIYKRYLNEKDD-EDVYQEAFHLARIQAILAVCSTIPGY 381
A S NWFL+DIVFY+ L S I+ Y + E+VY AF +A + AI+A CSTIPGY
Sbjct: 288 AASFNWFLVDIVFYTSNLLLSHIFSHYSKKPSTAENVYDAAFEVAELGAIIAACSTIPGY 347
Query: 382 FFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFA 441
+FTVYFID++GRVKIQ+MGFFFMAV + G PY +W+K E H+NKGFMV+YGL FFF
Sbjct: 348 WFTVYFIDKIGRVKIQIMGFFFMAVIYLVAGIPYSWYWSKHE-HNNKGFMVLYGLVFFFC 406
Query: 442 NFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHK----EKEEGYP 497
NFGPNTTTFI+PAE FPARFRSTCHGISGA GK+GAI+G+VGFLWA+ K +K + YP
Sbjct: 407 NFGPNTTTFIIPAEHFPARFRSTCHGISGAAGKLGAIVGTVGFLWATKKMESDDKNQIYP 466
Query: 498 KGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENEDEQSHHAEEYND 548
+ M+ + +ILGGVCI G+ VTYFFTKETMGRSLEENE +Q ++AE ++
Sbjct: 467 EVNRMRIAFLILGGVCIAGILVTYFFTKETMGRSLEENEHDQDNNAESEDE 517
>gb|AAN39051.1| putative phosphate transporter OsPT10 [Oryza sativa (japonica
cultivar-group)]
Length = 549
Score = 300 bits (767), Expect = 1e-79
Identities = 153/284 (53%), Positives = 200/284 (69%), Gaps = 11/284 (3%)
Query: 266 YTALVEQNVLQAAKDMEKVL-DVSMSQITEEHPL------PPATNVAYPLLSREFLWRHG 318
YTALVE++V++A D+ +VL D+ + + EE PP +Y LLSR F+ +HG
Sbjct: 231 YTALVERDVVKATNDIGRVLADLDLGAVAEEEVAAALSRPPPPPRPSYGLLSRRFVRQHG 290
Query: 319 RDLFACSANWFLLDIVFYSQVLFQSEIYKRYLNEKDDEDVYQEAFHLARIQAILAVCSTI 378
RDLFAC+A WFLLDI +YS LFQS+IY+ + +QEAF++A+ QA++AV STI
Sbjct: 291 RDLFACAAAWFLLDIPYYSSTLFQSQIYRPLFPAPGLINAFQEAFNVAKFQAVIAVASTI 350
Query: 379 PGYFFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAF 438
PGYF V IDRVGR +QM GF MAV FAL PY +W H G++V+Y L F
Sbjct: 351 PGYFVAVLLIDRVGRRCLQMAGFLLMAVFLFALAGPYDGYWRDHGAH--AGYIVLYSLTF 408
Query: 439 FFANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKE--KEEGY 496
F AN GPNTTTFI+PAELFPARFRSTCHG+SGA GK+GA++GS+GFLWAS ++ G+
Sbjct: 409 FSANLGPNTTTFILPAELFPARFRSTCHGLSGAAGKLGALVGSIGFLWASQQKDGAAAGH 468
Query: 497 PKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENEDEQS 540
GIGM +L +LGG+C++G+ +TY FT ETM RSLEENE +++
Sbjct: 469 LPGIGMMYALFVLGGICLLGLALTYVFTPETMMRSLEENESDRA 512
>dbj|BAD61874.1| putative phosphate transporter [Oryza sativa (japonica
cultivar-group)] gi|50725719|dbj|BAD33230.1| putative
phosphate transporter [Oryza sativa (japonica
cultivar-group)]
Length = 552
Score = 300 bits (767), Expect = 1e-79
Identities = 153/284 (53%), Positives = 200/284 (69%), Gaps = 11/284 (3%)
Query: 266 YTALVEQNVLQAAKDMEKVL-DVSMSQITEEHPL------PPATNVAYPLLSREFLWRHG 318
YTALVE++V++A D+ +VL D+ + + EE PP +Y LLSR F+ +HG
Sbjct: 234 YTALVERDVVKATNDIGRVLADLDLGAVAEEEVAAALSRPPPPPRPSYGLLSRRFVRQHG 293
Query: 319 RDLFACSANWFLLDIVFYSQVLFQSEIYKRYLNEKDDEDVYQEAFHLARIQAILAVCSTI 378
RDLFAC+A WFLLDI +YS LFQS+IY+ + +QEAF++A+ QA++AV STI
Sbjct: 294 RDLFACAAAWFLLDIPYYSSTLFQSQIYRPLFPAPGLINAFQEAFNVAKFQAVIAVASTI 353
Query: 379 PGYFFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAF 438
PGYF V IDRVGR +QM GF MAV FAL PY +W H G++V+Y L F
Sbjct: 354 PGYFVAVLLIDRVGRRCLQMAGFLLMAVFLFALAGPYDGYWRDHGAH--AGYIVLYSLTF 411
Query: 439 FFANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKE--KEEGY 496
F AN GPNTTTFI+PAELFPARFRSTCHG+SGA GK+GA++GS+GFLWAS ++ G+
Sbjct: 412 FSANLGPNTTTFILPAELFPARFRSTCHGLSGAAGKLGALVGSIGFLWASQQKDGAAAGH 471
Query: 497 PKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENEDEQS 540
GIGM +L +LGG+C++G+ +TY FT ETM RSLEENE +++
Sbjct: 472 LPGIGMMYALFVLGGICLLGLALTYVFTPETMMRSLEENESDRA 515
>gb|AAN39050.1| putative phosphate transporter OsPT9 [Oryza sativa (japonica
cultivar-group)]
Length = 582
Score = 295 bits (754), Expect = 3e-78
Identities = 148/285 (51%), Positives = 199/285 (68%), Gaps = 16/285 (5%)
Query: 266 YTALVEQNVLQAAKDMEKVL-DVSMSQITEEH-----------PLPPATNVAYPLLSREF 313
YTALVE++V++A D+ +VL D+ ++ + EE PP +Y L SR F
Sbjct: 241 YTALVERDVVKATNDIGRVLADLDLAAVAEEEVAAAALSPPPVTTPPPPRPSYGLFSRRF 300
Query: 314 LWRHGRDLFACSANWFLLDIVFYSQVLFQSEIYKRYLNEKDDEDVYQEAFHLARIQAILA 373
+ +HGRDLFAC+A WFLLDI +YS LFQS+IY+ + + +QEAF++A+ QA++A
Sbjct: 301 VRQHGRDLFACAAAWFLLDIPYYSSTLFQSQIYRPWFPPAAKVNAFQEAFNVAKFQAVIA 360
Query: 374 VCSTIPGYFFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVI 433
V STIPGYF + I+R GR ++QM GF MAV FAL PY +W ++ G++V+
Sbjct: 361 VASTIPGYFAAMLLIERAGRRRLQMAGFLLMAVFLFALAGPYDGYWR--DHAKTAGYIVL 418
Query: 434 YGLAFFFANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKE-- 491
Y L FF AN GPNTTTFI+PAELFPARFRSTCHG+SGA GK+GA++GS+GFLWAS ++
Sbjct: 419 YSLTFFSANLGPNTTTFILPAELFPARFRSTCHGLSGAAGKLGALVGSIGFLWASQQKDG 478
Query: 492 KEEGYPKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENE 536
G+ GIGM +L +LGG+C++G+ +TY FT ETM RSLEENE
Sbjct: 479 AAAGHLPGIGMMYALFVLGGICLLGLALTYAFTPETMTRSLEENE 523
>dbj|BAA86070.1| Phospate transpoter [Nicotiana tabacum]
Length = 536
Score = 278 bits (712), Expect = 2e-73
Identities = 149/293 (50%), Positives = 196/293 (66%), Gaps = 15/293 (5%)
Query: 266 YTALVEQNVLQAAKDMEKVLDVSMSQITEE-HPLPPATNVAYPLLSREFLWRHGRDLFAC 324
YTALV +N+ QAA DM KVL V + + E+ + T + L S+EFL RHG L
Sbjct: 241 YTALVAKNLKQAANDMSKVLQVEIEEEQEKVENVSQNTGNEFGLFSKEFLRRHGLHLLGT 300
Query: 325 SANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIPGYF 382
++ WFLLDI FYSQ LFQ +I+ ++ + + +E + +AR Q ++A+CST+PGY+
Sbjct: 301 ASTWFLLDIAFYSQNLFQKDIFSAIGWIPPAETMNALEEVYRIARAQTLIALCSTVPGYW 360
Query: 383 FTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFAN 442
FTV+FIDR+GR IQ+MGFFFM V FAL PY+ HWT +N GF+++Y L FFFAN
Sbjct: 361 FTVFFIDRIGRFAIQLMGFFFMTVFMFALAIPYH-HWTLKDNRI--GFVIMYSLTFFFAN 417
Query: 443 FGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASH----KEKEEGYPK 498
FGPN TTF+VPAE+FPAR RSTCHGIS A GK GA+IG+ GFL+A+ K+ + GYP
Sbjct: 418 FGPNATTFVVPAEIFPARLRSTCHGISAAAGKAGAMIGAFGFLYAAQPTDPKKVDAGYPA 477
Query: 499 GIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLE----ENEDEQSHHAEEYN 547
GIG++ SLI+LG V +GM T F E+ G+SLE ENE E+ E N
Sbjct: 478 GIGVRNSLIVLGCVNFLGMVFT-FLVPESKGKSLEEMSRENEGEEESGTEMKN 529
>gb|AAF74025.1| inorganic phosphate transporter [Nicotiana tabacum]
Length = 537
Score = 276 bits (707), Expect = 9e-73
Identities = 148/293 (50%), Positives = 195/293 (66%), Gaps = 15/293 (5%)
Query: 266 YTALVEQNVLQAAKDMEKVLDVSMSQITEE-HPLPPATNVAYPLLSREFLWRHGRDLFAC 324
YTALV +N+ QAA DM KVL V + + E+ + T + L S+EFL RHG L
Sbjct: 242 YTALVAKNLKQAANDMSKVLQVDIEEEQEKVENVSQNTRNEFGLFSKEFLRRHGLHLLGT 301
Query: 325 SANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIPGYF 382
++ WFLLDI FYSQ LFQ +I+ ++ + +E + +AR Q ++A+CST+PGY+
Sbjct: 302 ASTWFLLDIAFYSQNLFQKDIFSAIGWIPPAQTMNALEEVYKIARAQTLIALCSTVPGYW 361
Query: 383 FTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFAN 442
FTV+FID++GR IQ+MGFFFM V FAL PY+ HWT +N GF+++Y L FFFAN
Sbjct: 362 FTVFFIDKIGRFAIQLMGFFFMTVFMFALAIPYH-HWTLKDNRI--GFVIMYSLTFFFAN 418
Query: 443 FGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWAS----HKEKEEGYPK 498
FGPN TTF+VPAE+FPAR RSTCHGIS A GK GA+IG+ GFL+A+ K+ + GYP
Sbjct: 419 FGPNATTFVVPAEIFPARLRSTCHGISAAAGKAGAMIGAFGFLYAAQPTDRKKADAGYPA 478
Query: 499 GIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLE----ENEDEQSHHAEEYN 547
GIG++ SLI+LG V +GM T F E+ G+SLE ENE E+ E N
Sbjct: 479 GIGVRNSLIVLGCVNFLGMVFT-FLVPESKGKSLEEMSRENEGEEESGTEMKN 530
>dbj|BAA97416.1| inorganic phosphate transporter [Arabidopsis thaliana]
gi|2780348|dbj|BAA24282.1| inorganic phosphate
transporter [Arabidopsis thaliana]
gi|15239860|ref|NP_199151.1| inorganic phosphate
transporter (PHT2) [Arabidopsis thaliana]
Length = 524
Score = 276 bits (705), Expect = 2e-72
Identities = 151/286 (52%), Positives = 189/286 (65%), Gaps = 16/286 (5%)
Query: 266 YTALVEQNVLQAAKDMEKVL--DVSMSQITEEHPLPPATNVAYPLLSREFLWRHGRDLFA 323
YTALV +N+ QA DM KVL D+ + + E+ P N Y L S+EFL RHG L
Sbjct: 242 YTALVAKNIKQATADMSKVLQTDIELEERVEDDVKDPRQN--YGLFSKEFLRRHGLHLLG 299
Query: 324 CSANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIPGY 381
++ WFLLDI FYSQ LFQ +I+ ++ + + E F +AR Q ++A+CST+PGY
Sbjct: 300 TTSTWFLLDIAFYSQNLFQKDIFSAIGWIPKAATMNATHEVFRIARAQTLIALCSTVPGY 359
Query: 382 FFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFA 441
+FTV FID +GR KIQ+ GFF M V FA+ FPY +HW K EN GF+V+Y L FFFA
Sbjct: 360 WFTVAFIDTIGRFKIQLNGFFMMTVFMFAIAFPY-NHWIKPENRI--GFVVMYSLTFFFA 416
Query: 442 NFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEK----EEGYP 497
NFGPN TTFIVPAE+FPAR RSTCHGIS A GK GAIIG+ GFL+A+ + + GYP
Sbjct: 417 NFGPNATTFIVPAEIFPARLRSTCHGISAAAGKAGAIIGAFGFLYAAQNQDKAKVDAGYP 476
Query: 498 KGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEE--NEDEQSH 541
GIG+K SLI+LG + +GM T F E G+SLEE E E SH
Sbjct: 477 PGIGVKNSLIVLGVLNFIGMLFT-FLVPEPKGKSLEELSGEAEVSH 521
>pir||T07604 phosphate transport protein PT1 - potato gi|1420871|emb|CAA67395.1|
inorganic phosphate transporter 1 [Solanum tuberosum]
Length = 540
Score = 275 bits (704), Expect = 2e-72
Identities = 150/291 (51%), Positives = 193/291 (65%), Gaps = 16/291 (5%)
Query: 266 YTALVEQNVLQAAKDMEKVLDVSMSQITEEHPLPPATNVA--YPLLSREFLWRHGRDLFA 323
YTALV +N+ QAA DM KVL V + E+ N A + L S+EFL RHG L
Sbjct: 241 YTALVAKNLKQAANDMSKVLQVEIEAEPEKVAAISVANGANEFGLFSKEFLRRHGLHLLG 300
Query: 324 CSANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIPGY 381
++ WFLLDI FYSQ LFQ +I+ ++ + +E + +AR Q ++A+CST+PGY
Sbjct: 301 TASTWFLLDIAFYSQNLFQKDIFSAIGWIPPAQTMNALEEVYKIARAQTLIALCSTVPGY 360
Query: 382 FFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFA 441
+FTV FIDR+GR IQ+MGFFFM V FAL PY+ HWT +N GF+V+Y L FFFA
Sbjct: 361 WFTVAFIDRIGRFAIQLMGFFFMTVFMFALALPYH-HWTLKDNRI--GFVVMYSLTFFFA 417
Query: 442 NFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASH----KEKEEGYP 497
NFGPN TTF+VPAE+FPAR RSTCHGIS A GK GA++G+ GFL+A+ K+ + GYP
Sbjct: 418 NFGPNATTFVVPAEIFPARLRSTCHGISAAAGKAGAMVGAFGFLYAAQPTDPKKTDAGYP 477
Query: 498 KGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLE----ENEDEQSHHAE 544
GIG++ SLI+LG V +GM T F E+ G+SLE ENE E+ AE
Sbjct: 478 AGIGVRNSLIVLGCVNFLGMLFT-FLVPESKGKSLEEMSRENEGEEETVAE 527
>gb|AAV97729.1| phosphate transporter 3 [Lycopersicon esculentum]
Length = 532
Score = 275 bits (703), Expect = 3e-72
Identities = 144/285 (50%), Positives = 189/285 (65%), Gaps = 12/285 (4%)
Query: 266 YTALVEQNVLQAAKDMEKVLDVSMSQITEEHPLPPATNVAYPLLSREFLWRHGRDLFACS 325
YTALV +N +AA DM KVL+V + E+ + ++ L ++EFL RHG L +
Sbjct: 242 YTALVAKNATKAASDMSKVLNVEIE--AEKEKIVETQGNSFGLFTKEFLHRHGLHLLGTT 299
Query: 326 ANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIPGYFF 383
+ WFLLDI FYSQ LFQ +I+ + ++ + + E F +A+ Q ++A+CST+PGY+F
Sbjct: 300 STWFLLDIAFYSQNLFQKDIFSKIGWIPHPETMNALDEVFKIAKAQTLIALCSTVPGYWF 359
Query: 384 TVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFANF 443
TV FIDR+GR IQ+MGFFFM V FAL PY +HWTK EN GF+++Y L FFFANF
Sbjct: 360 TVAFIDRMGRFAIQLMGFFFMTVFMFALAIPY-NHWTKKENRI--GFVIMYSLTFFFANF 416
Query: 444 GPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASH----KEKEEGYPKG 499
GPN TTF+VPAE+FPAR RSTCHGIS A GK GAI+G+ GFL+A+ + + GYP G
Sbjct: 417 GPNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQSTDPSKVDTGYPTG 476
Query: 500 IGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENEDEQSHHAE 544
IG+K +LI+LG V +GM T E+ GRSLEE E E
Sbjct: 477 IGVKNALIVLGCVNFLGMLFT-LLVPESKGRSLEEMSKENEGEEE 520
>dbj|BAB21545.1| phosphate transporter [Nicotiana tabacum]
Length = 537
Score = 275 bits (703), Expect = 3e-72
Identities = 147/293 (50%), Positives = 194/293 (66%), Gaps = 15/293 (5%)
Query: 266 YTALVEQNVLQAAKDMEKVLDVSMSQITEE-HPLPPATNVAYPLLSREFLWRHGRDLFAC 324
YTALV +N+ QAA DM KVL V + + E+ + T + L S+EFL RHG L
Sbjct: 242 YTALVAKNLKQAANDMSKVLQVDIEEEQEKVENVSQNTRNEFGLFSKEFLRRHGLHLLGT 301
Query: 325 SANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIPGYF 382
++ WFLLDI FYSQ LFQ +I+ ++ + +E + +AR Q ++A+CST+PGY+
Sbjct: 302 ASTWFLLDIAFYSQNLFQKDIFSAIGWIPPAQTMNALEEVYKIARAQTLIALCSTVPGYW 361
Query: 383 FTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFAN 442
FTV+FID++GR Q+MGFFFM V FAL PY+ HWT +N GF+++Y L FFFAN
Sbjct: 362 FTVFFIDKIGRFAFQLMGFFFMTVFMFALAIPYH-HWTLKDNRI--GFVIMYSLTFFFAN 418
Query: 443 FGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWAS----HKEKEEGYPK 498
FGPN TTF+VPAE+FPAR RSTCHGIS A GK GA+IG+ GFL+A+ K+ + GYP
Sbjct: 419 FGPNATTFVVPAEIFPARLRSTCHGISAAAGKAGAMIGAFGFLYAAQPTDRKKADAGYPA 478
Query: 499 GIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLE----ENEDEQSHHAEEYN 547
GIG++ SLI+LG V +GM T F E+ G+SLE ENE E+ E N
Sbjct: 479 GIGVRNSLIVLGCVNFLGMVFT-FLVPESKGKSLEEMSRENEGEEESGTEMKN 530
>gb|AAL49927.1| putative phosphate transporter protein [Arabidopsis thaliana]
Length = 524
Score = 275 bits (702), Expect = 4e-72
Identities = 150/286 (52%), Positives = 189/286 (65%), Gaps = 16/286 (5%)
Query: 266 YTALVEQNVLQAAKDMEKVL--DVSMSQITEEHPLPPATNVAYPLLSREFLWRHGRDLFA 323
YTALV +N+ QA DM KVL D+ + + E+ P N Y L S+EFL RHG L
Sbjct: 242 YTALVAKNIKQATADMSKVLQTDIELEERVEDDVKDPKQN--YGLFSKEFLRRHGLHLLG 299
Query: 324 CSANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIPGY 381
++ WFLLDI FYSQ LFQ +I+ ++ + + E F +AR Q ++A+CST+PGY
Sbjct: 300 TTSTWFLLDIAFYSQNLFQKDIFSAIGWIPKAATMNATHEVFRIARAQTLIALCSTVPGY 359
Query: 382 FFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFA 441
+FTV FID +GR KIQ+ GFF M V FA+ FPY +HW K EN GF+V+Y L FFFA
Sbjct: 360 WFTVAFIDTIGRFKIQLNGFFMMTVFMFAIAFPY-NHWIKPENRI--GFVVMYSLTFFFA 416
Query: 442 NFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEK----EEGYP 497
NFGPN TTFIVPAE+FPAR RSTCHGIS A GK GAI+G+ GFL+A+ + + GYP
Sbjct: 417 NFGPNATTFIVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQSQDKAKVDAGYP 476
Query: 498 KGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEE--NEDEQSH 541
GIG+K SLI+LG + +GM T F E G+SLEE E E SH
Sbjct: 477 PGIGVKNSLIMLGVLNFIGMLFT-FLVPEPKGKSLEELSGEAEVSH 521
>dbj|BAA21504.1| inorganic phosphate transporter [Arabidopsis thaliana]
gi|2258116|dbj|BAA21503.1| inorganic phosphate
transporter [Arabidopsis thaliana]
gi|8843888|dbj|BAA97414.1| phosphate transporter
[Arabidopsis thaliana] gi|1502428|gb|AAB17265.1|
phosphate transporter [Arabidopsis thaliana]
gi|15239848|ref|NP_199149.1| inorganic phosphate
transporter (PHT1) (PT1) [Arabidopsis thaliana]
Length = 524
Score = 275 bits (702), Expect = 4e-72
Identities = 150/286 (52%), Positives = 189/286 (65%), Gaps = 16/286 (5%)
Query: 266 YTALVEQNVLQAAKDMEKVL--DVSMSQITEEHPLPPATNVAYPLLSREFLWRHGRDLFA 323
YTALV +N+ QA DM KVL D+ + + E+ P N Y L S+EFL RHG L
Sbjct: 242 YTALVAKNIKQATADMSKVLQTDIELEERVEDDVKDPKQN--YGLFSKEFLRRHGLHLLG 299
Query: 324 CSANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIPGY 381
++ WFLLDI FYSQ LFQ +I+ ++ + + E F +AR Q ++A+CST+PGY
Sbjct: 300 TTSTWFLLDIAFYSQNLFQKDIFSAIGWIPKAATMNATHEVFRIARAQTLIALCSTVPGY 359
Query: 382 FFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFA 441
+FTV FID +GR KIQ+ GFF M V FA+ FPY +HW K EN GF+V+Y L FFFA
Sbjct: 360 WFTVAFIDTIGRFKIQLNGFFMMTVFMFAIAFPY-NHWIKPENRI--GFVVMYSLTFFFA 416
Query: 442 NFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEK----EEGYP 497
NFGPN TTFIVPAE+FPAR RSTCHGIS A GK GAI+G+ GFL+A+ + + GYP
Sbjct: 417 NFGPNATTFIVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQSQDKAKVDAGYP 476
Query: 498 KGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEE--NEDEQSH 541
GIG+K SLI+LG + +GM T F E G+SLEE E E SH
Sbjct: 477 PGIGVKNSLIMLGVLNFIGMLFT-FLVPEPKGKSLEELSGEAEVSH 521
>emb|CAA68946.1| phosphate transporter [Arabidopsis thaliana]
Length = 524
Score = 275 bits (702), Expect = 4e-72
Identities = 150/286 (52%), Positives = 189/286 (65%), Gaps = 16/286 (5%)
Query: 266 YTALVEQNVLQAAKDMEKVL--DVSMSQITEEHPLPPATNVAYPLLSREFLWRHGRDLFA 323
YTALV +N+ QA DM KVL D+ + + E+ P N Y L S+EFL RHG L
Sbjct: 242 YTALVAKNIKQATADMSKVLQTDIELEERVEDDVKDPKQN--YGLFSKEFLRRHGLHLLG 299
Query: 324 CSANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIPGY 381
++ WFLLDI FYSQ LFQ +I+ ++ + + E F +AR Q ++A+CST+PGY
Sbjct: 300 TTSTWFLLDIAFYSQNLFQKDIFSAIGWIPKAATMNATHEVFRIARAQTLIALCSTVPGY 359
Query: 382 FFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFA 441
+FTV FID +GR KIQ+ GFF M V FA+ FPY +HW K EN GF+V+Y L FFFA
Sbjct: 360 WFTVAFIDTIGRFKIQLNGFFMMTVFMFAIAFPY-NHWIKPENRI--GFVVMYSLTFFFA 416
Query: 442 NFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEK----EEGYP 497
NFGPN TTFIVPAE+FPAR RSTCHGIS A GK GAI+G+ GFL+A+ + + GYP
Sbjct: 417 NFGPNATTFIVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQSQDKPKVDAGYP 476
Query: 498 KGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEE--NEDEQSH 541
GIG+K SLI+LG + +GM T F E G+SLEE E E SH
Sbjct: 477 PGIGVKNSLIMLGVLNFIGMLFT-FLVPEPKGKSLEELSGEAEVSH 521
>gb|AAK38196.1| phosphate transporter 1 [Lupinus albus]
Length = 540
Score = 274 bits (701), Expect = 5e-72
Identities = 152/293 (51%), Positives = 192/293 (64%), Gaps = 16/293 (5%)
Query: 266 YTALVEQNVLQAAKDMEKVLDVSMSQITEEHPLPPATNVAYPLLSREFLWRHGRDLFACS 325
YTALV +N QAA DM KVL V + T + + + L S+EFL RHG L +
Sbjct: 242 YTALVAKNAQQAAADMSKVLQVEIQSETNKEEAQGKPS--FGLFSKEFLRRHGLHLLGTA 299
Query: 326 ANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIPGYFF 383
+ WFLLDI FYSQ LFQ +I+ ++ + E + +AR Q ++A+CST+PGY+F
Sbjct: 300 STWFLLDIAFYSQNLFQKDIFSAIGWIPPAKTMNALDEVYRIARAQTLIALCSTVPGYWF 359
Query: 384 TVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFANF 443
TV IDR+GR IQ+MGFFFM V FAL PY HWT +N GF+VIY L FFFANF
Sbjct: 360 TVALIDRIGRFAIQLMGFFFMTVFMFALAIPY-DHWTHKDNRI--GFVVIYSLTFFFANF 416
Query: 444 GPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLW-ASHKEK---EEGYPKG 499
GPN TTF+VPAE+FPARFRSTCHGIS A GK+GAI+G+ GFL+ A +K+K + GYP G
Sbjct: 417 GPNATTFVVPAEIFPARFRSTCHGISSASGKLGAIVGAFGFLYLAQNKDKSKTDAGYPAG 476
Query: 500 IGMKASLIILGGVCIVGMFVTYFFTKETMGRSLE----ENEDEQSHHAEEYND 548
IG+K SLI+LG V I+G T F E G+SLE ENE+E+ YN+
Sbjct: 477 IGVKNSLIVLGVVNILGFCFT-FLVPEPNGKSLEEMSGENEEEEPTKEGSYNN 528
>gb|AAK01938.1| phosphate transporter 1 [Lupinus albus]
Length = 543
Score = 274 bits (700), Expect = 6e-72
Identities = 152/293 (51%), Positives = 191/293 (64%), Gaps = 16/293 (5%)
Query: 266 YTALVEQNVLQAAKDMEKVLDVSMSQITEEHPLPPATNVAYPLLSREFLWRHGRDLFACS 325
YTALV +N QAA DM KVL V + T + + + L S+EFL RHG L +
Sbjct: 242 YTALVAKNAQQAAADMSKVLQVEIQSETNKEEAQGKPS--FGLFSKEFLRRHGLHLLGTA 299
Query: 326 ANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIPGYFF 383
WFLLDI FYSQ LFQ +I+ ++ + E + +AR Q ++A+CST+PGY+F
Sbjct: 300 GTWFLLDIAFYSQNLFQKDIFSAIGWIPPAKTMNALDEVYRIARAQTLIALCSTVPGYWF 359
Query: 384 TVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFANF 443
TV IDR+GR IQ+MGFFFM V FAL PY HWT +N GF+VIY L FFFANF
Sbjct: 360 TVALIDRIGRFAIQLMGFFFMTVFMFALAIPY-DHWTHKDNRI--GFVVIYSLTFFFANF 416
Query: 444 GPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLW-ASHKEK---EEGYPKG 499
GPN TTF+VPAE+FPARFRSTCHGIS A GK+GAI+G+ GFL+ A +K+K + GYP G
Sbjct: 417 GPNATTFVVPAEIFPARFRSTCHGISSASGKLGAIVGAFGFLYLAQNKDKSKTDAGYPAG 476
Query: 500 IGMKASLIILGGVCIVGMFVTYFFTKETMGRSLE----ENEDEQSHHAEEYND 548
IG+K SLI+LG V I+G T F E G+SLE ENE+E+ YN+
Sbjct: 477 IGVKNSLIVLGVVNILGFCFT-FLVPEPNGKSLEEMSGENEEEEPTKEGSYNN 528
>emb|CAA68945.1| phosphate transporter [Arabidopsis thaliana]
Length = 524
Score = 273 bits (699), Expect = 8e-72
Identities = 150/286 (52%), Positives = 188/286 (65%), Gaps = 16/286 (5%)
Query: 266 YTALVEQNVLQAAKDMEKVL--DVSMSQITEEHPLPPATNVAYPLLSREFLWRHGRDLFA 323
YTALV +N+ QA DM KVL D+ + + E+ P N Y L S+EFL RHG L
Sbjct: 242 YTALVAKNIKQATADMSKVLQTDIELEERVEDDVKDPRQN--YGLFSKEFLRRHGLHLLG 299
Query: 324 CSANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIPGY 381
++ WFLLDI FYSQ LFQ +I+ ++ + + E F +AR Q ++A+CST+PGY
Sbjct: 300 TTSTWFLLDIAFYSQNLFQKDIFSAIGWIPKAATMNATHEVFRIARAQTLIALCSTVPGY 359
Query: 382 FFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFA 441
+FTV FID +GR KIQ+ GFF M V FA+ FPY +HW K EN GF+V+Y L FFFA
Sbjct: 360 WFTVAFIDTIGRFKIQLNGFFMMTVFMFAIAFPY-NHWIKPENRI--GFVVMYSLTFFFA 416
Query: 442 NFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHKEK----EEGYP 497
N GPN TTFIVPAE+FPAR RSTCHGIS A GK GAIIG+ GFL+A+ + + GYP
Sbjct: 417 NLGPNATTFIVPAEIFPARLRSTCHGISAAAGKAGAIIGAFGFLYAAQNQDKAKVDAGYP 476
Query: 498 KGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEE--NEDEQSH 541
GIG+K SLI+LG + +GM T F E G+SLEE E E SH
Sbjct: 477 PGIGVKNSLIVLGVLNFIGMLFT-FLVPEPKGKSLEELSGEAEVSH 521
>dbj|BAA34398.1| Phosphate Transporter 4 [Arabidopsis thaliana]
gi|1502430|gb|AAB17266.1| phosphate transporter
[Arabidopsis thaliana] gi|3928081|gb|AAC79607.1|
phosphate transporter (AtPT2) [Arabidopsis thaliana]
gi|2564661|gb|AAB88291.1| phosphate transporter
[Arabidopsis thaliana] gi|15224985|ref|NP_181428.1|
phosphate transporter (PT2) [Arabidopsis thaliana]
gi|25309090|pir||C84811 phosphate transporter (AtPT2)
[imported] - Arabidopsis thaliana
Length = 534
Score = 273 bits (698), Expect = 1e-71
Identities = 146/290 (50%), Positives = 193/290 (66%), Gaps = 11/290 (3%)
Query: 266 YTALVEQNVLQAAKDMEKVLDVSMSQITEE-HPLPPATNVAYPLLSREFLWRHGRDLFAC 324
YTALV ++ QAA DM KVL V + ++ + + A+ L S+EF+ RHG L
Sbjct: 242 YTALVAKDAKQAASDMSKVLQVEIEPEQQKLEEISKEKSKAFGLFSKEFMSRHGLHLLGT 301
Query: 325 SANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIPGYF 382
++ WFLLDI FYSQ LFQ +I+ ++ + QE F +AR Q ++A+CST+PGY+
Sbjct: 302 TSTWFLLDIAFYSQNLFQKDIFSAIGWIPPAQSMNAIQEVFKIARAQTLIALCSTVPGYW 361
Query: 383 FTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFAN 442
FTV FID +GR IQMMGFFFM V FAL PY +HWT EN GF+++Y L FFFAN
Sbjct: 362 FTVAFIDVIGRFAIQMMGFFFMTVFMFALAIPY-NHWTHKENRI--GFVIMYSLTFFFAN 418
Query: 443 FGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLW-ASHKEKEE---GYPK 498
FGPN TTF+VPAE+FPARFRSTCHGIS A GK+GA++G+ GFL+ A + +K++ GYP
Sbjct: 419 FGPNATTFVVPAEIFPARFRSTCHGISAASGKLGAMVGAFGFLYLAQNPDKDKTDAGYPP 478
Query: 499 GIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLEENEDEQSHHAEEYND 548
GIG++ SLI+LG V +G+ T F E+ G+SLEE E + ND
Sbjct: 479 GIGVRNSLIVLGVVNFLGILFT-FLVPESKGKSLEEMSGENEDNENSNND 527
>gb|AAB81347.1| phosphate transporter [Medicago truncatula] gi|7446752|pir||T07894
probable inorganic phosphate transport protein PT2 -
barrel medic
Length = 533
Score = 273 bits (698), Expect = 1e-71
Identities = 151/293 (51%), Positives = 191/293 (64%), Gaps = 19/293 (6%)
Query: 266 YTALVEQNVLQAAKDMEKVLDVSMSQITEEHPLPPATNV---AYPLLSREFLWRHGRDLF 322
YTALV +N QAA DM KVL V + EE + T+ +Y L S++F RHG LF
Sbjct: 242 YTALVAKNAKQAAADMSKVLQVELE--VEEEKVEKMTSDKRNSYGLFSKQFAARHGLALF 299
Query: 323 ACSANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIPG 380
+ WFLLDI FYSQ LFQ +I+ ++ + + E + +AR Q ++A+CST+PG
Sbjct: 300 GTCSTWFLLDIAFYSQNLFQKDIFSAIGWIPPAKEMNAIHEVYKIARAQTLIALCSTVPG 359
Query: 381 YFFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFF 440
Y+FTV FID +GR IQMMGFFFM V FAL PY HW+K EN GF+VIY L FFF
Sbjct: 360 YWFTVAFIDHMGRFAIQMMGFFFMTVFMFALAIPY-DHWSKEENRI--GFVVIYSLTFFF 416
Query: 441 ANFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASHK----EKEEGY 496
ANFGPN TTF+VPAE+FPAR RSTCHGIS A GK GAI+G+ GFL+A+ + ++GY
Sbjct: 417 ANFGPNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQSKDPTKTDKGY 476
Query: 497 PKGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLE----ENEDEQSHHAEE 545
P GIG+K SLI+LG + VGM T E+ G+SLE ENE E + E+
Sbjct: 477 PTGIGIKNSLIMLGVINFVGMLCT-LLVPESKGKSLEELSGENEGEGAEATEQ 528
>gb|AAD38859.1| inorganic phosphate transporter [Solanum tuberosum]
Length = 540
Score = 272 bits (695), Expect = 2e-71
Identities = 149/291 (51%), Positives = 192/291 (65%), Gaps = 16/291 (5%)
Query: 266 YTALVEQNVLQAAKDMEKVLDVSMSQITEEHPLPPATNVA--YPLLSREFLWRHGRDLFA 323
YTALV +N+ QAA DM KVL V + E+ A + L S+EFL RHG L
Sbjct: 241 YTALVAKNLKQAANDMSKVLQVEIEAEPEKVGAISEAKGANEFGLFSKEFLRRHGLHLLG 300
Query: 324 CSANWFLLDIVFYSQVLFQSEIYKR--YLNEKDDEDVYQEAFHLARIQAILAVCSTIPGY 381
++ WFLLDI FYSQ LFQ +I+ ++ + +E + +AR Q ++A+CST+PGY
Sbjct: 301 TASTWFLLDIAFYSQNLFQKDIFSAIGWIPPAQTMNALEEVYKIARAQTLIALCSTVPGY 360
Query: 382 FFTVYFIDRVGRVKIQMMGFFFMAVSFFALGFPYYSHWTKGENHDNKGFMVIYGLAFFFA 441
+FTV FIDR+GR IQ+MGFFFM V FAL PY+ HWT +N GF+V+Y L FFFA
Sbjct: 361 WFTVAFIDRIGRFAIQLMGFFFMTVFMFALALPYH-HWTLKDNRI--GFVVMYSLTFFFA 417
Query: 442 NFGPNTTTFIVPAELFPARFRSTCHGISGAVGKVGAIIGSVGFLWASH----KEKEEGYP 497
NFGPN TTF+VPAE+FPAR RSTCHGIS A GK GA++G+ GFL+A+ K+ + GYP
Sbjct: 418 NFGPNATTFVVPAEIFPARLRSTCHGISAAAGKAGAMVGAFGFLYAAQPTDPKKTDAGYP 477
Query: 498 KGIGMKASLIILGGVCIVGMFVTYFFTKETMGRSLE----ENEDEQSHHAE 544
GIG++ SLI+LG V +GM T F E+ G+SLE ENE E+ AE
Sbjct: 478 PGIGVRNSLIVLGCVNFLGMLFT-FLVPESNGKSLEEMSRENEGEEETVAE 527
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.342 0.150 0.499
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 865,031,307
Number of Sequences: 2540612
Number of extensions: 34366524
Number of successful extensions: 115965
Number of sequences better than 10.0: 999
Number of HSP's better than 10.0 without gapping: 261
Number of HSP's successfully gapped in prelim test: 738
Number of HSP's that attempted gapping in prelim test: 114296
Number of HSP's gapped (non-prelim): 1245
length of query: 548
length of database: 863,360,394
effective HSP length: 133
effective length of query: 415
effective length of database: 525,458,998
effective search space: 218065484170
effective search space used: 218065484170
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (22.0 bits)
S2: 78 (34.7 bits)
Medicago: description of AC149576.4


