

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149493.3 - phase: 0 /pseudo
(473 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
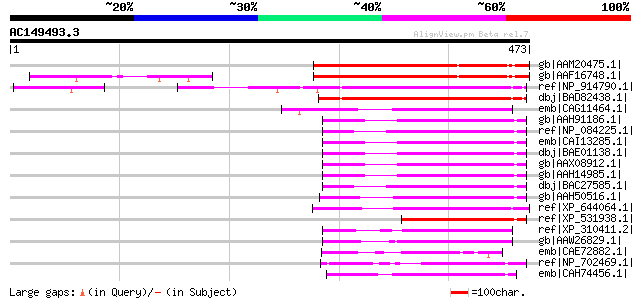
Score E
Sequences producing significant alignments: (bits) Value
gb|AAM20475.1| unknown protein [Arabidopsis thaliana] gi|2805953... 256 1e-66
gb|AAF16748.1| F3M18.1 [Arabidopsis thaliana] gi|25403350|pir||D... 256 1e-66
ref|NP_914790.1| P0470A12.23 [Oryza sativa (japonica cultivar-gr... 238 3e-61
dbj|BAD82438.1| small nuclear RNA activating complex, polypeptid... 236 1e-60
emb|CAG11464.1| unnamed protein product [Tetraodon nigroviridis] 130 1e-28
gb|AAH91186.1| Small nuclear RNA activating complex, polypeptide... 124 5e-27
ref|NP_084225.1| small nuclear RNA activating complex, polypepti... 124 8e-27
emb|CAI13285.1| small nuclear RNA activating complex, polypeptid... 122 3e-26
dbj|BAE01138.1| unnamed protein product [Macaca fascicularis] 122 3e-26
gb|AAX08912.1| small nuclear RNA activating complex, polypeptide... 122 3e-26
gb|AAH14985.1| small nuclear RNA activating complex, polypeptide... 122 3e-26
dbj|BAC27585.1| unnamed protein product [Mus musculus] 120 9e-26
gb|AAH50516.1| Similar to small nuclear RNA activating complex, ... 115 4e-24
ref|XP_644064.1| hypothetical protein DDB0203238 [Dictyostelium ... 113 1e-23
ref|XP_531938.1| PREDICTED: similar to small nuclear RNA activat... 106 1e-21
ref|XP_310411.2| ENSANGP00000017886 [Anopheles gambiae str. PEST... 79 3e-13
gb|AAW26829.1| unknown [Schistosoma japonicum] 77 1e-12
emb|CAE72882.1| Hypothetical protein CBG20195 [Caenorhabditis br... 67 1e-09
ref|NP_702469.1| hypothetical protein PF14_0580 [Plasmodium falc... 67 2e-09
emb|CAH74456.1| conserved hypothetical protein [Plasmodium chaba... 66 2e-09
>gb|AAM20475.1| unknown protein [Arabidopsis thaliana] gi|28059535|gb|AAO30067.1|
unknown protein [Arabidopsis thaliana]
gi|22329836|ref|NP_174178.2| snRNA activating complex
family protein [Arabidopsis thaliana]
Length = 375
Score = 256 bits (654), Expect = 1e-66
Identities = 122/196 (62%), Positives = 149/196 (75%), Gaps = 4/196 (2%)
Query: 278 QKKGQHDPSGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQK 337
QK G++DPSGYFLIEDVF+ DLR+PSA D + PILDWL NSK+EA KKWE ++ GELQ+K
Sbjct: 183 QKAGKYDPSGYFLIEDVFHNDLRNPSAKDYSYPILDWLWNSKDEALKKWECVLTGELQKK 242
Query: 338 QKAIVGEASVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHAD 397
QK ++GEA LPR+ + +M HFCD+ FR+GA Y+YCHQGDC HT+VIRDMR+ H +
Sbjct: 243 QKLVLGEAKSVDLPRYRTADMQSTHFCDIRFRVGASYVYCHQGDCKHTIVIRDMRMSHPE 302
Query: 398 DVQNWAVYPIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAED 457
DVQN A YPI+ F K R QKCGVCKI RA+KV VDDKW +N YFCD CF LLH +E+
Sbjct: 303 DVQNRAAYPIM-FWPKRRIQKCGVCKIKRASKVAVDDKWASENSSYFCDVCFELLH-SEE 360
Query: 458 GGSPMYTDFIEYDYNH 473
G P+ DF +DY H
Sbjct: 361 G--PLNCDFPVFDYVH 374
Score = 37.4 bits (85), Expect = 1.0
Identities = 16/24 (66%), Positives = 20/24 (82%)
Query: 162 KVEEVVRIKQKQEEDKAQVRLHSF 185
KVE++ ++KQKQEEDKA V LH F
Sbjct: 70 KVEQLAKLKQKQEEDKAAVTLHCF 93
>gb|AAF16748.1| F3M18.1 [Arabidopsis thaliana] gi|25403350|pir||D86411 protein
F1K23.20 [imported] - Arabidopsis thaliana
gi|6691212|gb|AAF24550.1| F1K23.20 [Arabidopsis
thaliana]
Length = 482
Score = 256 bits (654), Expect = 1e-66
Identities = 122/196 (62%), Positives = 149/196 (75%), Gaps = 4/196 (2%)
Query: 278 QKKGQHDPSGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQK 337
QK G++DPSGYFLIEDVF+ DLR+PSA D + PILDWL NSK+EA KKWE ++ GELQ+K
Sbjct: 290 QKAGKYDPSGYFLIEDVFHNDLRNPSAKDYSYPILDWLWNSKDEALKKWECVLTGELQKK 349
Query: 338 QKAIVGEASVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHAD 397
QK ++GEA LPR+ + +M HFCD+ FR+GA Y+YCHQGDC HT+VIRDMR+ H +
Sbjct: 350 QKLVLGEAKSVDLPRYRTADMQSTHFCDIRFRVGASYVYCHQGDCKHTIVIRDMRMSHPE 409
Query: 398 DVQNWAVYPIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAED 457
DVQN A YPI+ F K R QKCGVCKI RA+KV VDDKW +N YFCD CF LLH +E+
Sbjct: 410 DVQNRAAYPIM-FWPKRRIQKCGVCKIKRASKVAVDDKWASENSSYFCDVCFELLH-SEE 467
Query: 458 GGSPMYTDFIEYDYNH 473
G P+ DF +DY H
Sbjct: 468 G--PLNCDFPVFDYVH 481
Score = 77.8 bits (190), Expect = 7e-13
Identities = 59/184 (32%), Positives = 88/184 (47%), Gaps = 42/184 (22%)
Query: 19 IPRGGPIYVSNMSGPIVRVPLFQDSILTQLHSLQSELPPDS------NHDISVDDLKVFT 72
+PRGGPIY+ NM G I P F+ S L L L++ L DS D+S+D LK++T
Sbjct: 22 VPRGGPIYLPNMVGNISTTPEFKSSFLNVLQDLETHLSLDSTSSSSQQFDVSIDSLKIYT 81
Query: 73 EDDLMDMALKQVFQGRDNNQDPPNAELVTSLPLFFFIL*YH*LIMMDVVVLNNID*ILNL 132
+++L +MA+K+ F +Q+ EL SL N+ N
Sbjct: 82 DEELTEMAMKEAFPEDYLSQE----ELEPSL---------------------NVSHHENP 116
Query: 133 VAG-------VRRSNSGESLV*QINPFLMLQSNCKE----KVEEVVRIKQKQEEDKAQVR 181
+AG V+ + + + M + +E KVE++ ++KQKQEEDKA V
Sbjct: 117 LAGRAKRKRTVKNTEVKKRTLKNTEVMKMTEKKTEEAYLVKVEQLAKLKQKQEEDKAAVT 176
Query: 182 LHSF 185
LH F
Sbjct: 177 LHCF 180
>ref|NP_914790.1| P0470A12.23 [Oryza sativa (japonica cultivar-group)]
Length = 545
Score = 238 bits (607), Expect = 3e-61
Identities = 138/348 (39%), Positives = 188/348 (53%), Gaps = 65/348 (18%)
Query: 154 MLQSNCKEKVEEVVRIKQKQEEDKAQVRLHSFQLVFLIFIHFIMCKITSI*SNSAIFL*S 213
+LQ + K ++ IK KQEEDK LHSF
Sbjct: 230 ILQGSYLTKAVKMAEIKAKQEEDKHAASLHSFS--------------------------- 262
Query: 214 *LQDQSICQQINQNSKDDVS*VHKFRQKG----QHTGSSGAHTCAGFRSCSLCRDFP*FS 269
S+ ++++ S + V R T G H + LC + +
Sbjct: 263 ---GDSVLAKVSKPSAEKVDVAKSLRYISTTWKNKTFKPGEHRPVVYPEVLLCVEV--YE 317
Query: 270 KRCQAIQCQK--------------------------KGQHDPSGYFLIEDVFYTDLRDPS 303
KR +++ Q+ QH SGYFLIED FY D R S
Sbjct: 318 KRYGSVKSQEFLVLGSQLLTDLRDNIYCFKDKLMNVAKQHVHSGYFLIEDTFYNDTRR-S 376
Query: 304 AIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGEASVSHLPRFASFEMHKIHF 363
+D ++PILDW++NS+ EA++KW+ I +G L+++QK ++ +VS++P F S +M K F
Sbjct: 377 TVDYSKPILDWIKNSRNEAEEKWDAITSGVLKKRQKDLLMGLNVSNVPDFKSAKMEKTRF 436
Query: 364 CDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAVYPIVTFQLKIRFQKCGVCK 423
DL FRLGAGYLYCHQG+C H +VIRDMRLIH +D QN A YP++TFQ++ R QKC VC+
Sbjct: 437 SDLNFRLGAGYLYCHQGNCKHMIVIRDMRLIHPEDTQNQAEYPLMTFQMQRRLQKCSVCQ 496
Query: 424 IFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGGSPMYTDFIEYDY 471
IF ATK+TVDDKWT +NPCYFCD+C+ LLH ED S +Y + YDY
Sbjct: 497 IFHATKMTVDDKWTLNNPCYFCDKCYYLLHYKED-NSLLYHHTV-YDY 542
Score = 55.8 bits (133), Expect = 3e-06
Identities = 30/86 (34%), Positives = 52/86 (59%), Gaps = 3/86 (3%)
Query: 4 DELELEFSIDDPSISIPRGGPIYVSNMSGPIVRVPLFQDSILTQLHSLQSEL--PPDS-N 60
+E E + + + RGGP++V M GP+ VP F S L +L SL+ EL P D +
Sbjct: 8 EEAESSAAGEQRRMPFARGGPVFVPFMVGPVSTVPEFMSSALHELQSLKDELGDPGDEFD 67
Query: 61 HDISVDDLKVFTEDDLMDMALKQVFQ 86
++ VD+L+V +E++L++ AL++ +
Sbjct: 68 EELCVDELRVLSEEELVERALREAME 93
>dbj|BAD82438.1| small nuclear RNA activating complex, polypeptide 3, 50kDa-like
[Oryza sativa (japonica cultivar-group)]
Length = 195
Score = 236 bits (602), Expect = 1e-60
Identities = 111/190 (58%), Positives = 144/190 (75%), Gaps = 3/190 (1%)
Query: 282 QHDPSGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAI 341
QH SGYFLIED FY D R S +D ++PILDW++NS+ EA++KW+ I +G L+++QK +
Sbjct: 6 QHVHSGYFLIEDTFYNDTRR-STVDYSKPILDWIKNSRNEAEEKWDAITSGVLKKRQKDL 64
Query: 342 VGEASVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQN 401
+ +VS++P F S +M K F DL FRLGAGYLYCHQG+C H +VIRDMRLIH +D QN
Sbjct: 65 LMGLNVSNVPDFKSAKMEKTRFSDLNFRLGAGYLYCHQGNCKHMIVIRDMRLIHPEDTQN 124
Query: 402 WAVYPIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGGSP 461
A YP++TFQ++ R QKC VC+IF ATK+TVDDKWT +NPCYFCD+C+ LLH ED S
Sbjct: 125 QAEYPLMTFQMQRRLQKCSVCQIFHATKMTVDDKWTLNNPCYFCDKCYYLLHYKED-NSL 183
Query: 462 MYTDFIEYDY 471
+Y + YDY
Sbjct: 184 LYHHTV-YDY 192
>emb|CAG11464.1| unnamed protein product [Tetraodon nigroviridis]
Length = 274
Score = 130 bits (326), Expect = 1e-28
Identities = 67/215 (31%), Positives = 101/215 (46%), Gaps = 34/215 (15%)
Query: 248 SGAHTCAGFRSCSLC----RDFP*FSKRCQAIQCQKKGQHDPSGYFLIEDVFYTDLRDPS 303
+G+HT A R C + FS A H S +F E VFY D+R P
Sbjct: 72 TGSHTLAELRDAICCVSDLQVCGEFSNNPDAAPAFISKDHFKSAFFYFEGVFYNDMRFPE 131
Query: 304 AIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGEASVSHLPRFASFEMHKIHF 363
+D++R ++W ++ + P F+ +M F
Sbjct: 132 CVDISRTTIEWAESH------------------------------NFPPFSQAKMEDTRF 161
Query: 364 CDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAVYPIVTFQLKIRFQKCGVCK 423
DL ++G YLYCHQGDC H ++I D+RL+H D + +YP++T + + QKC C
Sbjct: 162 VDLKVKVGFPYLYCHQGDCEHLVIITDIRLVHKSDCLDRKLYPLLTHKHRFSTQKCSACH 221
Query: 424 IFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDG 458
+F VT D++ P +PC FCD+CF +LH +G
Sbjct: 222 LFIGRWVTTGDRFAPSDPCLFCDKCFRMLHYDAEG 256
>gb|AAH91186.1| Small nuclear RNA activating complex, polypeptide 3 (predicted)
[Rattus norvegicus] gi|61557304|ref|NP_001013230.1|
small nuclear RNA activating complex, polypeptide 3
(predicted) [Rattus norvegicus]
Length = 407
Score = 124 bits (312), Expect = 5e-27
Identities = 65/186 (34%), Positives = 94/186 (49%), Gaps = 30/186 (16%)
Query: 286 SGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGEA 345
S +F E FY D R P DL+R I++W ++ K
Sbjct: 245 SAFFYFEGTFYNDKRYPECRDLSRTIIEWSESHDRGYGK--------------------- 283
Query: 346 SVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAVY 405
F + M F DL +LG YLYCHQGDC H +VI D+RL+H DD + +Y
Sbjct: 284 -------FQTARMEDFTFNDLNIKLGFPYLYCHQGDCEHVVVITDIRLVHHDDCLDRTLY 336
Query: 406 PIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGGSPMYTD 465
P++T + + +KC VCK++ A VT +D + P++PC+FCD CF +LH +G +
Sbjct: 337 PLLTKKHWLWTRKCFVCKMYTARWVTNNDTFAPEDPCFFCDVCFRMLHYDSEGNK--LGE 394
Query: 466 FIEYDY 471
F+ Y Y
Sbjct: 395 FLAYPY 400
>ref|NP_084225.1| small nuclear RNA activating complex, polypeptide 3 [Mus musculus]
gi|29145057|gb|AAH48713.1| Small nuclear RNA activating
complex, polypeptide 3 [Mus musculus]
gi|26340656|dbj|BAC33990.1| unnamed protein product [Mus
musculus] gi|26387042|dbj|BAB31890.2| unnamed protein
product [Mus musculus]
Length = 407
Score = 124 bits (310), Expect = 8e-27
Identities = 65/186 (34%), Positives = 94/186 (49%), Gaps = 30/186 (16%)
Query: 286 SGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGEA 345
S +F E FY D R P DL+R I++W E+
Sbjct: 245 SAFFYFEGTFYNDRRYPECRDLSRTIIEW----------------------------SES 276
Query: 346 SVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAVY 405
+F + M F DL +LG YLYCHQGDC H +VI D+RL+H DD + +Y
Sbjct: 277 HDRGYGKFQTARMEDFTFNDLHIKLGFPYLYCHQGDCEHVVVITDIRLVHHDDCLDRTLY 336
Query: 406 PIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGGSPMYTD 465
P++T + + +KC VCK++ A VT +D + P++PC+FCD CF +LH +G +
Sbjct: 337 PLLTKKHWLWTRKCFVCKMYTARWVTNNDTFAPEDPCFFCDVCFRMLHYDSEGNK--LGE 394
Query: 466 FIEYDY 471
F+ Y Y
Sbjct: 395 FLAYPY 400
>emb|CAI13285.1| small nuclear RNA activating complex, polypeptide 3, 50kDa [Homo
sapiens] gi|55633227|ref|XP_528547.1| PREDICTED: similar
to small nuclear RNA activating complex, polypeptide 3,
50kDa [Pan troglodytes] gi|4507105|ref|NP_003075.1|
small nuclear RNA activating complex, polypeptide 3,
50kDa [Homo sapiens] gi|4097682|gb|AAD09214.1| proximal
sequence element-binding transcription factor beta
subunit [Homo sapiens] gi|8134719|sp|Q92966|SNPC3_HUMAN
snRNA activating protein complex 50 kDa subunit (SNAPc
50 kDa subunit) (Small nuclear RNA activating complex
polypeptide 3) (Proximal sequence element-binding
transcription factor beta subunit) (PSE-binding factor
beta subunit) (PTF beta subunit)
gi|1619946|gb|AAC50948.1| snRNA activating protein
complex 50kD subunit [Homo sapiens]
Length = 411
Score = 122 bits (305), Expect = 3e-26
Identities = 64/186 (34%), Positives = 93/186 (49%), Gaps = 30/186 (16%)
Query: 286 SGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGEA 345
S +F E FY D R P DL+R I++W ++ K
Sbjct: 249 SAFFYFEGTFYNDKRYPECRDLSRTIIEWSESHDRGYGK--------------------- 287
Query: 346 SVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAVY 405
F + M F DL +LG YLYCHQGDC H +VI D+RL+H DD + +Y
Sbjct: 288 -------FQTARMEDFTFNDLCIKLGFPYLYCHQGDCEHVIVITDIRLVHHDDCLDRTLY 340
Query: 406 PIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGGSPMYTD 465
P++ + + +KC VCK++ A VT +D + P++PC+FCD CF +LH +G +
Sbjct: 341 PLLIKKHWLWTRKCFVCKMYTARWVTNNDSFAPEDPCFFCDVCFRMLHYDSEGNK--LGE 398
Query: 466 FIEYDY 471
F+ Y Y
Sbjct: 399 FLAYPY 404
>dbj|BAE01138.1| unnamed protein product [Macaca fascicularis]
Length = 412
Score = 122 bits (305), Expect = 3e-26
Identities = 64/186 (34%), Positives = 93/186 (49%), Gaps = 30/186 (16%)
Query: 286 SGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGEA 345
S +F E FY D R P DL+R I++W ++ K
Sbjct: 250 SAFFYFEGTFYNDKRYPECRDLSRTIIEWSESHDRGYGK--------------------- 288
Query: 346 SVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAVY 405
F + M F DL +LG YLYCHQGDC H +VI D+RL+H DD + +Y
Sbjct: 289 -------FQTARMEDFTFNDLCIKLGFPYLYCHQGDCEHVIVITDIRLVHHDDCLDRTLY 341
Query: 406 PIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGGSPMYTD 465
P++ + + +KC VCK++ A VT +D + P++PC+FCD CF +LH +G +
Sbjct: 342 PLLIKKHWLWTRKCFVCKMYTARWVTNNDSFAPEDPCFFCDVCFRMLHYDSEGNK--LGE 399
Query: 466 FIEYDY 471
F+ Y Y
Sbjct: 400 FLAYPY 405
>gb|AAX08912.1| small nuclear RNA activating complex, polypeptide 3, 50kDa [Bos
taurus]
Length = 412
Score = 122 bits (305), Expect = 3e-26
Identities = 64/186 (34%), Positives = 94/186 (50%), Gaps = 30/186 (16%)
Query: 286 SGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGEA 345
S +F E FY D R P DL+R I++W ++ K
Sbjct: 250 SAFFYFEGTFYNDKRYPECRDLSRTIIEWSESHDRGYGK--------------------- 288
Query: 346 SVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAVY 405
F + +M F DL +LG YLYCHQGDC H +VI D+RL+H DD + +Y
Sbjct: 289 -------FQTAKMEDFTFNDLYIKLGFPYLYCHQGDCEHVVVITDIRLVHHDDCLDRTLY 341
Query: 406 PIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGGSPMYTD 465
P++ + + +KC VCK++ A VT +D + P++PC+FCD CF +LH +G +
Sbjct: 342 PLLIKKHWLWTRKCFVCKMYTARWVTNNDSFAPEDPCFFCDVCFRMLHYDSEGNK--LGE 399
Query: 466 FIEYDY 471
F+ Y Y
Sbjct: 400 FLAYPY 405
>gb|AAH14985.1| small nuclear RNA activating complex, polypeptide 3, 50kD [Homo
sapiens]
Length = 410
Score = 122 bits (305), Expect = 3e-26
Identities = 64/186 (34%), Positives = 93/186 (49%), Gaps = 30/186 (16%)
Query: 286 SGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGEA 345
S +F E FY D R P DL+R I++W ++ K
Sbjct: 248 SAFFYFEGTFYNDKRYPECRDLSRTIIEWSESHDRGYGK--------------------- 286
Query: 346 SVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAVY 405
F + M F DL +LG YLYCHQGDC H +VI D+RL+H DD + +Y
Sbjct: 287 -------FQTARMEDFTFNDLCIKLGFPYLYCHQGDCEHVIVITDIRLVHHDDCLDRTLY 339
Query: 406 PIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGGSPMYTD 465
P++ + + +KC VCK++ A VT +D + P++PC+FCD CF +LH +G +
Sbjct: 340 PLLIKKHWLWTRKCFVCKMYTARWVTNNDSFAPEDPCFFCDVCFRMLHYDSEGNK--LGE 397
Query: 466 FIEYDY 471
F+ Y Y
Sbjct: 398 FLAYPY 403
>dbj|BAC27585.1| unnamed protein product [Mus musculus]
Length = 407
Score = 120 bits (301), Expect = 9e-26
Identities = 64/186 (34%), Positives = 93/186 (49%), Gaps = 30/186 (16%)
Query: 286 SGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGEA 345
S +F E F D R P DL+R I++W E+
Sbjct: 245 SAFFYFEGTFCNDRRYPECRDLSRTIIEW----------------------------SES 276
Query: 346 SVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAVY 405
+F + M F DL +LG YLYCHQGDC H +VI D+RL+H DD + +Y
Sbjct: 277 HDRGYGKFQTARMEDFTFNDLHIKLGFPYLYCHQGDCEHVVVITDIRLVHHDDCLDRTLY 336
Query: 406 PIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGGSPMYTD 465
P++T + + +KC VCK++ A VT +D + P++PC+FCD CF +LH +G +
Sbjct: 337 PLLTKKHWLWTRKCFVCKMYTARWVTNNDTFAPEDPCFFCDVCFRMLHYDSEGNK--LGE 394
Query: 466 FIEYDY 471
F+ Y Y
Sbjct: 395 FLAYPY 400
>gb|AAH50516.1| Similar to small nuclear RNA activating complex, polypeptide 3,
50kDa [Danio rerio] gi|41055458|ref|NP_957409.1| similar
to small nuclear RNA activating complex, polypeptide 3,
50kDa [Danio rerio]
Length = 378
Score = 115 bits (287), Expect = 4e-24
Identities = 57/189 (30%), Positives = 89/189 (46%), Gaps = 32/189 (16%)
Query: 283 HDPSGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIV 342
H S +F FY D R P D+++ I +W ++
Sbjct: 215 HYKSAFFFFNGTFYNDTRFPECQDISKVIKEWTRSRD----------------------- 251
Query: 343 GEASVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNW 402
P F + M F DL ++G YLY HQGDC H +V+ D+RL+H DD +
Sbjct: 252 -------FPDFKTARMEDTSFNDLQMKVGFPYLYTHQGDCEHVVVLTDVRLVHQDDCLDI 304
Query: 403 AVYPIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGGSPM 462
+YP++T + ++ +KC VC ++ + +T +D P +PC FCD+CF + H + G
Sbjct: 305 KLYPLITHKHRVMTRKCSVCHLYISRWITTNDALAPMDPCLFCDQCFRMFHYDDKGNK-- 362
Query: 463 YTDFIEYDY 471
DF+ Y Y
Sbjct: 363 VGDFLAYAY 371
>ref|XP_644064.1| hypothetical protein DDB0203238 [Dictyostelium discoideum]
gi|60472283|gb|EAL70236.1| hypothetical protein
DDB0203238 [Dictyostelium discoideum]
Length = 1004
Score = 113 bits (283), Expect = 1e-23
Identities = 59/197 (29%), Positives = 95/197 (47%), Gaps = 29/197 (14%)
Query: 277 CQKKGQHDPSGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQ 336
C G + SG+F I +VFY D RD ++ L WL+ ++
Sbjct: 836 CILDGSNRKSGFFFINNVFYNDNRDQRNYQYSKNTLAWLKERGKD--------------- 880
Query: 337 KQKAIVGEASVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHA 396
+ F M + F DL +G YLYCHQG C H + +R+++
Sbjct: 881 -------------ISNFKEESMDDVTFNDLEISIGERYLYCHQGSCEHLVTFESLRMVNE 927
Query: 397 DDVQNWAVYPIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAE 456
D + YPI+T+Q K+R +KC VC I+ A VT+ D++ + P ++CDEC+ H ++
Sbjct: 928 MDDLEPSRYPIITYQQKVRRRKCLVCDIYAAKYVTLGDQFADETPFFYCDECYRTFHYSK 987
Query: 457 DGGSPMYTDFIEYDYNH 473
D G +Y+++ + Y H
Sbjct: 988 D-GHLLYSNYQVFPYYH 1003
>ref|XP_531938.1| PREDICTED: similar to small nuclear RNA activating complex,
polypeptide 3, 50kDa [Canis familiaris]
Length = 119
Score = 106 bits (265), Expect = 1e-21
Identities = 49/114 (42%), Positives = 70/114 (60%), Gaps = 2/114 (1%)
Query: 358 MHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAVYPIVTFQLKIRFQ 417
M F DL +LG YLYCHQGDC H +VI D+RL H DD + +YP++ + + +
Sbjct: 1 MEDFTFNDLYIKLGFPYLYCHQGDCEHVVVITDIRLAHRDDCLDRTLYPLLIKKHWLWTR 60
Query: 418 KCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGGSPMYTDFIEYDY 471
KC VCK++ A VT +D + P++PC+FCD CF +LH +G +F+ Y Y
Sbjct: 61 KCFVCKMYTARWVTNNDSFAPEDPCFFCDVCFRMLHYDSEGNK--LGEFLAYPY 112
>ref|XP_310411.2| ENSANGP00000017886 [Anopheles gambiae str. PEST]
gi|55243982|gb|EAA06033.2| ENSANGP00000017886 [Anopheles
gambiae str. PEST]
Length = 244
Score = 79.0 bits (193), Expect = 3e-13
Identities = 49/173 (28%), Positives = 71/173 (40%), Gaps = 29/173 (16%)
Query: 286 SGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGEA 345
SG+F + D FY D RD + D + I W +++++GE
Sbjct: 71 SGFFFVHDTFYNDFRDDANHDYSGVIRKWAD---------------------RQSLIGEL 109
Query: 346 SVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAVY 405
+ M F DL FRLG +Y HQG+C H VI D RL+ A D+ + Y
Sbjct: 110 KTAR--------MEDTRFGDLKFRLGYPQMYQHQGNCEHLFVISDCRLLAATDILTRSRY 161
Query: 406 PIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDG 458
P + R C +C +A + + +P Y C+ C H EDG
Sbjct: 162 PWLNSYGFSRDVPCNICGHCQAQYIVQNSTRHIFDPAYICENCLETYHYTEDG 214
>gb|AAW26829.1| unknown [Schistosoma japonicum]
Length = 386
Score = 77.0 bits (188), Expect = 1e-12
Identities = 48/174 (27%), Positives = 76/174 (43%), Gaps = 27/174 (15%)
Query: 286 SGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGEA 345
S YF IE FY DLR+ ++ L + ++ W ++ +E
Sbjct: 220 SSYFFIEGKFYDDLRNANSKSLGQEVIQWAKSKRE------------------------- 254
Query: 346 SVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAVY 405
VS P F S M I +L +G Y + HQG+C H ++ D+RL+ D Q+ + +
Sbjct: 255 LVSCGP-FTSSPMESITLENLAVCIGKPYFFVHQGNCEHMIIFSDIRLVDRDSCQSESSF 313
Query: 406 PIVTFQLKIRFQKCGVCKIFRATKVTVDDKW-TPDNPCYFCDECFSLLHLAEDG 458
P++T + R C C+ + + + P +PC CD C LL DG
Sbjct: 314 PMLTGRCSARILHCFACRRLACRWIVTECRTILPVDPCPICDVCIRLLLYTADG 367
>emb|CAE72882.1| Hypothetical protein CBG20195 [Caenorhabditis briggsae]
Length = 416
Score = 67.0 bits (162), Expect = 1e-09
Identities = 46/167 (27%), Positives = 71/167 (41%), Gaps = 35/167 (20%)
Query: 285 PSGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGE 344
PS + + + FY D +A D++ PI ++Q+ E+ A+
Sbjct: 243 PSSFIFVHNTFYVDSAPENAQDISFPIRKFMQDR--------------EIFDPVDAV--- 285
Query: 345 ASVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAV 404
M + DL RLG Y++ H G+C H L+ D+RL+H D +
Sbjct: 286 ------------PMEGVRIIDLSLRLGQPYVFQHSGNCEHVLIFHDLRLLHESDPRGIDQ 333
Query: 405 YPIVTFQLKIRFQKCGVCKIFRATKVTVDDK--WTPDNPCYFCDECF 449
YP V ++ K +KC +CK V V D+ P+ YFC CF
Sbjct: 334 YPYVLYE-KGNERKCEICK---KGHVDVVDRHELLPNTYTYFCRSCF 376
>ref|NP_702469.1| hypothetical protein PF14_0580 [Plasmodium falciparum 3D7]
gi|23497653|gb|AAN37193.1| hypothetical protein
[Plasmodium falciparum 3D7]
Length = 635
Score = 66.6 bits (161), Expect = 2e-09
Identities = 52/189 (27%), Positives = 84/189 (43%), Gaps = 29/189 (15%)
Query: 284 DPSGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQK-KWEYIINGELQQKQKAIV 342
D S YF I+ + Y DLR +A+D + IL++ + K + K+ Y IN + KAI+
Sbjct: 471 DGSLYF-IDGILYPDLRSNNAVDYSTSILNFYKMKKMKTNFIKYPYKIN-----QDKAIL 524
Query: 343 GEASVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNW 402
+ + P F + HQG C H +V ++R + ++
Sbjct: 525 SQIEI---PLFKKC------------------CFLHQGTCEHRIVFNNIRQYNKLRDKHL 563
Query: 403 AVYPIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGGSPM 462
+ YP+ TF+ I + C C A K+ +D +NP Y C+ CF L L + P+
Sbjct: 564 SKYPLRTFKPNISNKYCISCHKNIAQKIVLDSYLLKENPSYMCNNCFDLF-LMDKNNQPI 622
Query: 463 YTDFIEYDY 471
T +DY
Sbjct: 623 DTSMKYFDY 631
>emb|CAH74456.1| conserved hypothetical protein [Plasmodium chabaudi]
Length = 610
Score = 66.2 bits (160), Expect = 2e-09
Identities = 45/174 (25%), Positives = 73/174 (41%), Gaps = 13/174 (7%)
Query: 289 FLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQKQKAIVGEASVS 348
+ I+ + Y D+RD A+D + I+++ +N K + N L
Sbjct: 437 YYIDGILYPDVRDKDALDYSTSIINFYKNRKNISSSTASSSSNSNLDD------------ 484
Query: 349 HLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHADDVQNWAVYPIV 408
L K D+ L + HQG+C H +V ++R + ++ + YPI
Sbjct: 485 FLKHPYKIYQDKAVLKDIKIPLYKKCCFLHQGNCEHRIVFSNVRQYNKFRDKDISQYPIR 544
Query: 409 TFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGGSPM 462
TF+ I + C C A K+ +D +NP Y CD CF L L + G P+
Sbjct: 545 TFKPNIATKYCICCHKNIAQKIIIDSYLFKENPSYVCDGCFELF-LLDSNGKPV 597
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.334 0.146 0.467
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 791,555,880
Number of Sequences: 2540612
Number of extensions: 32184645
Number of successful extensions: 108266
Number of sequences better than 10.0: 35
Number of HSP's better than 10.0 without gapping: 34
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 108184
Number of HSP's gapped (non-prelim): 56
length of query: 473
length of database: 863,360,394
effective HSP length: 132
effective length of query: 341
effective length of database: 527,999,610
effective search space: 180047867010
effective search space used: 180047867010
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 77 (34.3 bits)
Medicago: description of AC149493.3


