

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149471.21 - phase: 0
(281 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
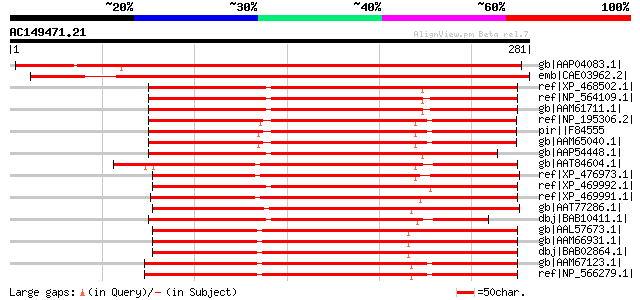
Score E
Sequences producing significant alignments: (bits) Value
gb|AAP04083.1| unknown protein [Arabidopsis thaliana] gi|2645045... 405 e-112
emb|CAE03962.2| OSJNBb0085H11.11 [Oryza sativa (japonica cultiva... 395 e-109
ref|XP_468502.1| putative prolyl 4-hydroxylase [Oryza sativa (ja... 224 2e-57
ref|NP_564109.1| oxidoreductase, 2OG-Fe(II) oxygenase family pro... 220 3e-56
gb|AAM61711.1| putative prolyl 4-hydroxylase, alpha subunit [Ara... 219 9e-56
ref|NP_195306.2| oxidoreductase, 2OG-Fe(II) oxygenase family pro... 214 2e-54
pir||F84555 similar to prolyl 4-hydroxylase alpha subunit [impor... 207 3e-52
gb|AAM65040.1| putative prolyl 4-hydroxylase, alpha subunit [Ara... 205 1e-51
gb|AAP54448.1| putative prolyl 4-hydroxylase, alpha subunit [Ory... 204 2e-51
gb|AAT84604.1| prolyl 4-hydroxylase [Dianthus caryophyllus] 197 3e-49
ref|XP_476973.1| prolyl 4-hydroxylase alpha-1 subunit precursor-... 194 3e-48
ref|XP_469992.1| putative oxidoreductase [Oryza sativa (japonica... 193 5e-48
ref|XP_469991.1| putative oxidoreductase [Oryza sativa (japonica... 188 1e-46
gb|AAT77286.1| putative prolyl 4-hydroxylase alpha subunit [Oryz... 184 3e-45
dbj|BAB10411.1| prolyl 4-hydroxylase, alpha subunit-like protein... 183 5e-45
gb|AAL57673.1| AT3g28480/MFJ20_16 [Arabidopsis thaliana] gi|2479... 183 5e-45
gb|AAM66931.1| prolyl 4-hydroxylase, putative [Arabidopsis thali... 182 9e-45
dbj|BAB02864.1| prolyl 4-hydroxylase alpha subunit-like protein ... 182 9e-45
gb|AAM67123.1| prolyl 4-hydroxylase alpha subunit-like protein [... 182 1e-44
ref|NP_566279.1| oxidoreductase, 2OG-Fe(II) oxygenase family pro... 182 1e-44
>gb|AAP04083.1| unknown protein [Arabidopsis thaliana] gi|26450452|dbj|BAC42340.1|
unknown protein [Arabidopsis thaliana]
gi|20197628|gb|AAM15158.1| hypothetical protein
[Arabidopsis thaliana] gi|3763917|gb|AAC64297.1|
hypothetical protein [Arabidopsis thaliana]
gi|25408834|pir||G84861 hypothetical protein At2g43080
[imported] - Arabidopsis thaliana
gi|15224220|ref|NP_181836.1| oxidoreductase, 2OG-Fe(II)
oxygenase family protein [Arabidopsis thaliana]
Length = 283
Score = 405 bits (1042), Expect = e-112
Identities = 195/278 (70%), Positives = 234/278 (84%), Gaps = 5/278 (1%)
Query: 4 PAMKIVFGLLTFVTIGMIIGALSQLAFIRRLELEEPFTTTTTTRSLLPRGYTYWNN---- 59
PAMKIVFGLLTFVT+GM+IG+L QLAFI RLE + T + R L + Y +
Sbjct: 3 PAMKIVFGLLTFVTVGMVIGSLLQLAFINRLE-DSYGTGFPSLRGLRGQNTRYLRDVSRW 61
Query: 60 NNDKEAQILRLGYVKPEVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGK 119
NDK+A++LR+G VKPEV+SWSPRII+LH+FLS EEC+YL+ +A PRL++STVVD TGK
Sbjct: 62 ANDKDAELLRIGNVKPEVVSWSPRIIVLHDFLSPEECEYLKAIARPRLQVSTVVDVKTGK 121
Query: 120 GIKSDVRTSSGMFLSHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHH 179
G+KSDVRTSSGMFL+H ER YP+I AIEKRI+V+SQ+P ENGEL+QVLRYE Q+Y+PHH
Sbjct: 122 GVKSDVRTSSGMFLTHVERSYPIIQAIEKRIAVFSQVPAENGELIQVLRYEPQQFYKPHH 181
Query: 180 DYFSDTFNLKRGGQRIATMLMYLGDNVEGGETHFPSAGSDECSCGGKLTKGLCVKPVKGN 239
DYF+DTFNLKRGGQR+ATMLMYL D+VEGGET+FP AG +C+CGGK+ KG+ VKP KG+
Sbjct: 182 DYFADTFNLKRGGQRVATMLMYLTDDVEGGETYFPLAGDGDCTCGGKIMKGISVKPTKGD 241
Query: 240 AVLFWSMGLDGQSDPDSVHGGCPVLAGEKWSATKWMRQ 277
AVLFWSMGLDGQSDP S+HGGC VL+GEKWSATKWMRQ
Sbjct: 242 AVLFWSMGLDGQSDPRSIHGGCEVLSGEKWSATKWMRQ 279
>emb|CAE03962.2| OSJNBb0085H11.11 [Oryza sativa (japonica cultivar-group)]
gi|50923279|ref|XP_472000.1| OSJNBb0085H11.11 [Oryza
sativa (japonica cultivar-group)]
Length = 267
Score = 395 bits (1015), Expect = e-109
Identities = 193/270 (71%), Positives = 219/270 (80%), Gaps = 16/270 (5%)
Query: 12 LLTFVTIGMIIGALSQLAFIRRLELEEPFTTTTTTRSLLPRGYTYWNNNNDKEAQILRLG 71
LLTFVT+GMI+G+L QLAF RR++ T + ND+EA LRLG
Sbjct: 14 LLTFVTLGMILGSLLQLAFFRRIDDHSNVT----------------HLENDQEAAFLRLG 57
Query: 72 YVKPEVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGM 131
VKPEV+SWSPRII+ HNFLS EECDYLR +A PRL+ISTVVD TGKG+KS+VRTSSGM
Sbjct: 58 LVKPEVISWSPRIIVFHNFLSSEECDYLRSIARPRLQISTVVDVATGKGVKSNVRTSSGM 117
Query: 132 FLSHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRG 191
F+S EERK P+I +IEKRISVYSQIP ENGEL+QVLRYE +QYYRPHHDYFSDTFN+KRG
Sbjct: 118 FVSSEERKLPVIQSIEKRISVYSQIPEENGELIQVLRYEPSQYYRPHHDYFSDTFNIKRG 177
Query: 192 GQRIATMLMYLGDNVEGGETHFPSAGSDECSCGGKLTKGLCVKPVKGNAVLFWSMGLDGQ 251
GQR+ATMLMYL D VEGGETHFP AG ECSCGGK+ KGLCVKP KG+AVLFWSMGLDG+
Sbjct: 178 GQRVATMLMYLTDGVEGGETHFPQAGDGECSCGGKMVKGLCVKPNKGDAVLFWSMGLDGE 237
Query: 252 SDPDSVHGGCPVLAGEKWSATKWMRQSVHV 281
+D +S+HGGCPVL GEKWSATKWMRQ V
Sbjct: 238 TDSNSIHGGCPVLEGEKWSATKWMRQKEFV 267
>ref|XP_468502.1| putative prolyl 4-hydroxylase [Oryza sativa (japonica
cultivar-group)] gi|48716447|dbj|BAD23054.1| putative
prolyl 4-hydroxylase [Oryza sativa (japonica
cultivar-group)]
Length = 310
Score = 224 bits (571), Expect = 2e-57
Identities = 118/206 (57%), Positives = 138/206 (66%), Gaps = 8/206 (3%)
Query: 76 EVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFLSH 135
EV+SW PR + HNFLS EECDYL G+A P + STVVD+ TGK S VRTSSGMFL
Sbjct: 100 EVISWEPRAFVYHNFLSKEECDYLIGLAKPHMVKSTVVDSTTGKSKDSRVRTSSGMFLQR 159
Query: 136 EERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQRI 195
K +I AIEKRI+ Y+ IP+E+GE +QVL YE Q Y PH DYF D +N K GGQR+
Sbjct: 160 GRDK--VIRAIEKRIADYTFIPMEHGEGLQVLHYEVGQKYEPHFDYFLDEYNTKNGGQRM 217
Query: 196 ATMLMYLGDNVEGGETHFPSAGSDECS------CGGKLTKGLCVKPVKGNAVLFWSMGLD 249
AT+LMYL D EGGET FP A + S KGL VKP G+A+LFWSM D
Sbjct: 218 ATLLMYLSDVEEGGETIFPDANVNSSSLPWYNELSECARKGLAVKPKMGDALLFWSMKPD 277
Query: 250 GQSDPDSVHGGCPVLAGEKWSATKWM 275
DP S+HGGCPV+ G KWS+TKWM
Sbjct: 278 ATLDPLSLHGGCPVIKGNKWSSTKWM 303
>ref|NP_564109.1| oxidoreductase, 2OG-Fe(II) oxygenase family protein [Arabidopsis
thaliana] gi|25513547|pir||D86336 F14O10.12 protein -
Arabidopsis thaliana gi|9558598|gb|AAF88161.1| Contains
similarity to a prolyl 4-hydroxylase alpha subunit
protein from Gallus gallus gi|212530. [Arabidopsis
thaliana]
Length = 287
Score = 220 bits (561), Expect = 3e-56
Identities = 118/209 (56%), Positives = 138/209 (65%), Gaps = 14/209 (6%)
Query: 76 EVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFLSH 135
EVLSW PR + HNFLS EEC+YL +A P + STVVD+ TGK S VRTSSG FL
Sbjct: 77 EVLSWEPRAFVYHNFLSKEECEYLISLAKPHMVKSTVVDSETGKSKDSRVRTSSGTFLRR 136
Query: 136 EERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQRI 195
K +I IEKRI+ Y+ IP ++GE +QVL YE Q Y PH+DYF D FN K GGQR+
Sbjct: 137 GRDK--IIKTIEKRIADYTFIPADHGEGLQVLHYEAGQKYEPHYDYFVDEFNTKNGGQRM 194
Query: 196 ATMLMYLGDNVEGGETHFPSAGSDECS---------CGGKLTKGLCVKPVKGNAVLFWSM 246
ATMLMYL D EGGET FP+A + S CG KGL VKP G+A+LFWSM
Sbjct: 195 ATMLMYLSDVEEGGETVFPAANMNFSSVPWYNELSECG---KKGLSVKPRMGDALLFWSM 251
Query: 247 GLDGQSDPDSVHGGCPVLAGEKWSATKWM 275
D DP S+HGGCPV+ G KWS+TKWM
Sbjct: 252 RPDATLDPTSLHGGCPVIRGNKWSSTKWM 280
>gb|AAM61711.1| putative prolyl 4-hydroxylase, alpha subunit [Arabidopsis thaliana]
Length = 287
Score = 219 bits (557), Expect = 9e-56
Identities = 117/209 (55%), Positives = 138/209 (65%), Gaps = 14/209 (6%)
Query: 76 EVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFLSH 135
EVLSW PR + HNFLS EEC+YL +A P + STVVD+ TGK S VRTSSG FL
Sbjct: 77 EVLSWEPRAFVYHNFLSKEECEYLISLAKPHMVKSTVVDSETGKSKDSRVRTSSGTFLRR 136
Query: 136 EERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQRI 195
K +I IEKRI+ Y+ IP ++GE +QVL YE Q Y PH+DYF D FN K GGQR+
Sbjct: 137 GRDK--IIKTIEKRIADYTFIPADHGEGLQVLHYEAGQKYEPHYDYFVDEFNTKNGGQRM 194
Query: 196 ATMLMYLGDNVEGGETHFPSAGSDECS---------CGGKLTKGLCVKPVKGNAVLFWSM 246
ATMLMYL D EGGET FP+A + S CG KGL VKP G+A+LFWSM
Sbjct: 195 ATMLMYLSDVEEGGETVFPAANMNFSSVPWYNELSECG---KKGLSVKPRMGDALLFWSM 251
Query: 247 GLDGQSDPDSVHGGCPVLAGEKWSATKWM 275
D DP S+HGGCPV+ G KWS+TKW+
Sbjct: 252 RPDATLDPTSLHGGCPVIRGNKWSSTKWI 280
>ref|NP_195306.2| oxidoreductase, 2OG-Fe(II) oxygenase family protein [Arabidopsis
thaliana]
Length = 290
Score = 214 bits (545), Expect = 2e-54
Identities = 115/209 (55%), Positives = 140/209 (66%), Gaps = 16/209 (7%)
Query: 76 EVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFLS- 134
EV+SW PR + HNFL+ EEC++L +A P + S VVD TGK I S VRTSSG FL+
Sbjct: 81 EVISWEPRAFVYHNFLTNEECEHLISLAKPSMMKSKVVDVKTGKSIDSRVRTSSGTFLNR 140
Query: 135 -HEERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQ 193
H+E ++ IE RIS ++ IP ENGE +QVL YE Q Y PHHDYF D FN+++GGQ
Sbjct: 141 GHDE----IVEEIENRISDFTFIPPENGEGLQVLHYEVGQRYEPHHDYFFDEFNVRKGGQ 196
Query: 194 RIATMLMYLGDNVEGGETHFPSAGS--------DECSCGGKLTKGLCVKPVKGNAVLFWS 245
RIAT+LMYL D EGGET FP+A DE S GK +GL V P K +A+LFWS
Sbjct: 197 RIATVLMYLSDVDEGGETVFPAAKGNVSDVPWWDELSQCGK--EGLSVLPKKRDALLFWS 254
Query: 246 MGLDGQSDPDSVHGGCPVLAGEKWSATKW 274
M D DP S+HGGCPV+ G KWS+TKW
Sbjct: 255 MKPDASLDPSSLHGGCPVIKGNKWSSTKW 283
>pir||F84555 similar to prolyl 4-hydroxylase alpha subunit [imported] -
Arabidopsis thaliana gi|15227885|ref|NP_179363.1|
oxidoreductase, 2OG-Fe(II) oxygenase family protein
[Arabidopsis thaliana]
Length = 291
Score = 207 bits (527), Expect = 3e-52
Identities = 111/209 (53%), Positives = 139/209 (66%), Gaps = 16/209 (7%)
Query: 76 EVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFL-- 133
EV+SW PR ++ HNFL+ EEC++L +A P + STVVD TG S VRTSSG FL
Sbjct: 81 EVISWEPRAVVYHNFLTNEECEHLISLAKPSMVKSTVVDEKTGGSKDSRVRTSSGTFLRR 140
Query: 134 SHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQ 193
H+E ++ IEKRIS ++ IP+ENGE +QVL Y+ Q Y PH+DYF D FN K GGQ
Sbjct: 141 GHDE----VVEVIEKRISDFTFIPVENGEGLQVLHYQVGQKYEPHYDYFLDEFNTKNGGQ 196
Query: 194 RIATMLMYLGDNVEGGETHFPSAGS--------DECSCGGKLTKGLCVKPVKGNAVLFWS 245
RIAT+LMYL D +GGET FP+A +E S GK +GL V P K +A+LFW+
Sbjct: 197 RIATVLMYLSDVDDGGETVFPAARGNISAVPWWNELSKCGK--EGLSVLPKKRDALLFWN 254
Query: 246 MGLDGQSDPDSVHGGCPVLAGEKWSATKW 274
M D DP S+HGGCPV+ G KWS+TKW
Sbjct: 255 MRPDASLDPSSLHGGCPVVKGNKWSSTKW 283
>gb|AAM65040.1| putative prolyl 4-hydroxylase, alpha subunit [Arabidopsis thaliana]
Length = 291
Score = 205 bits (521), Expect = 1e-51
Identities = 110/209 (52%), Positives = 138/209 (65%), Gaps = 16/209 (7%)
Query: 76 EVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFL-- 133
EV+SW PR ++ HNFL+ EEC++L +A P + STVVD TG S VRTSSG FL
Sbjct: 81 EVISWEPRAVVYHNFLTNEECEHLISLAKPSMVKSTVVDEKTGGSKDSRVRTSSGTFLRR 140
Query: 134 SHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQ 193
H+E ++ IEKRIS ++ IP+ENGE +QVL Y+ Q Y PH+DYF D FN K GGQ
Sbjct: 141 GHDE----VVEVIEKRISDFTFIPVENGEGLQVLHYQVGQKYEPHYDYFLDEFNTKNGGQ 196
Query: 194 RIATMLMYLGDNVEGGETHFPSAGS--------DECSCGGKLTKGLCVKPVKGNAVLFWS 245
RIAT+LMYL D +GGET FP+A +E S GK +GL V P +A+LFW+
Sbjct: 197 RIATVLMYLSDVDDGGETVFPAARGNISAVPWWNELSKCGK--EGLSVLPKXRDALLFWN 254
Query: 246 MGLDGQSDPDSVHGGCPVLAGEKWSATKW 274
M D DP S+HGGCPV+ G KWS+TKW
Sbjct: 255 MRPDASLDPSSLHGGCPVVKGNKWSSTKW 283
>gb|AAP54448.1| putative prolyl 4-hydroxylase, alpha subunit [Oryza sativa
(japonica cultivar-group)] gi|37535718|ref|NP_922161.1|
putative prolyl 4-hydroxylase, alpha subunit [Oryza
sativa (japonica cultivar-group)]
gi|18071415|gb|AAL58274.1| putative prolyl
4-hydroxylase, alpha subunit [Oryza sativa (japonica
cultivar-group)]
Length = 343
Score = 204 bits (520), Expect = 2e-51
Identities = 112/195 (57%), Positives = 129/195 (65%), Gaps = 8/195 (4%)
Query: 76 EVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFLSH 135
EVLSW PR L HNFLS EEC+YL +A P +K STVVDA+TG S VRTSSGMFL
Sbjct: 111 EVLSWEPRAFLYHNFLSKEECEYLISLAKPHMKKSTVVDASTGGSKDSRVRTSSGMFLGR 170
Query: 136 EERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQRI 195
+ K +I IEKRIS Y+ IP+ENGE +QVL YE Q Y PH DYF D FN K GGQRI
Sbjct: 171 GQDK--IIRTIEKRISDYTFIPVENGEGLQVLHYEVGQKYEPHFDYFHDEFNTKNGGQRI 228
Query: 196 ATMLMYLGDNVEGGETHFPSAGSDECS------CGGKLTKGLCVKPVKGNAVLFWSMGLD 249
AT+LMYL D EGGET FPS+ ++ S KGL VKP G+A+LFWSM D
Sbjct: 229 ATLLMYLSDVEEGGETIFPSSKANSSSSPFYNELSECAKKGLAVKPKMGDALLFWSMRPD 288
Query: 250 GQSDPDSVHGGCPVL 264
G D S+HG P+L
Sbjct: 289 GSLDATSLHGEIPIL 303
>gb|AAT84604.1| prolyl 4-hydroxylase [Dianthus caryophyllus]
Length = 316
Score = 197 bits (501), Expect = 3e-49
Identities = 112/240 (46%), Positives = 147/240 (60%), Gaps = 28/240 (11%)
Query: 57 WNNNNDKEAQILRLGY--------VKPE---VLSWSPRIILLHNFLSYEECDYLRGVALP 105
W NN K +LRL V P LSW PR L FL++EECD+L +A
Sbjct: 26 WLNNVKKGKSVLRLKSENVPSSVGVDPSHVTQLSWKPRAFLYEGFLTHEECDHLIDMAKD 85
Query: 106 RLKISTVVDANTGKGIKSDVRTSSGMFLSHEERKYPMIHAIEKRISVYSQIPIENGELMQ 165
+L+ S V D +GK I S+VRTSSGMFL ++ + ++ AIE RI+ ++ +PIENGE MQ
Sbjct: 86 KLEKSMVADNESGKSIPSEVRTSSGMFL--QKAQDDVVAAIEARIAAWTFLPIENGEAMQ 143
Query: 166 VLRYEKNQYYRPHHDYFSDTFNLKRGGQRIATMLMYLGDNVEGGETHFPSAGS------- 218
+L YE+ Q Y PH DYF D N + GG RIAT+LMYL + EGGET FP+A +
Sbjct: 144 ILHYERGQKYEPHFDYFHDKVNQQLGGHRIATVLMYLSNVEEGGETVFPNAEAKLQLANN 203
Query: 219 ---DECSCGGKLTKGLCVKPVKGNAVLFWSMGLDGQSDPDSVHGGCPVLAGEKWSATKWM 275
+C+ G G VKP KG+A+LF+S+ D +D S+HG CPV+ GEKWSATKW+
Sbjct: 204 ESLSDCAKG-----GYSVKPKKGDALLFFSLHPDASTDSLSLHGSCPVIEGEKWSATKWI 258
>ref|XP_476973.1| prolyl 4-hydroxylase alpha-1 subunit precursor-like protein [Oryza
sativa (japonica cultivar-group)]
gi|34393269|dbj|BAC83179.1| prolyl 4-hydroxylase alpha-1
subunit precursor-like protein [Oryza sativa (japonica
cultivar-group)] gi|50509101|dbj|BAD30161.1| prolyl
4-hydroxylase alpha-1 subunit precursor-like protein
[Oryza sativa (japonica cultivar-group)]
Length = 313
Score = 194 bits (492), Expect = 3e-48
Identities = 101/209 (48%), Positives = 135/209 (64%), Gaps = 17/209 (8%)
Query: 78 LSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFLSHEE 137
+SW PR+ + FLS +ECD+L + +++ S V D +GK + S+VRTSSGMFL ++
Sbjct: 54 VSWRPRVFVYKGFLSDDECDHLVKLGKRKMQRSMVADNKSGKSVMSEVRTSSGMFL--DK 111
Query: 138 RKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQRIAT 197
R+ P++ IEKRI+ ++ +P EN E +Q+LRYE Q Y PH DYF D N GG R AT
Sbjct: 112 RQDPVVSRIEKRIAAWTFLPEENAENIQILRYEHGQKYEPHFDYFHDKVNQALGGHRYAT 171
Query: 198 MLMYLGDNVEGGETHFPSAGS----------DECSCGGKLTKGLCVKPVKGNAVLFWSMG 247
+LMYL +GGET FP+A EC+ KGL VKPVKG+ VLF+S+
Sbjct: 172 VLMYLSTVEKGGETVFPNAEGWENQPKDDTFSECA-----QKGLAVKPVKGDTVLFFSLH 226
Query: 248 LDGQSDPDSVHGGCPVLAGEKWSATKWMR 276
+DG DP S+HG CPV+ GEKWSA KW+R
Sbjct: 227 IDGVPDPLSLHGSCPVIEGEKWSAPKWIR 255
>ref|XP_469992.1| putative oxidoreductase [Oryza sativa (japonica cultivar-group)]
gi|29150365|gb|AAO72374.1| putative oxidoreductase
[Oryza sativa (japonica cultivar-group)]
Length = 299
Score = 193 bits (490), Expect = 5e-48
Identities = 100/203 (49%), Positives = 132/203 (64%), Gaps = 7/203 (3%)
Query: 78 LSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFLSHEE 137
LSW PR L FLS++ECD+L +A R++ S V D ++GK I S VRTSSG FLS E
Sbjct: 40 LSWRPRAFLYSGFLSHDECDHLVNLAKGRMEKSMVADNDSGKSIMSQVRTSSGTFLSKHE 99
Query: 138 RKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQRIAT 197
++ IEKR++ ++ +P EN E +Q+L YE Q Y H DYF D NLKRGG R+AT
Sbjct: 100 DD--IVSGIEKRVAAWTFLPEENAESIQILHYELGQKYDAHFDYFHDKNNLKRGGHRVAT 157
Query: 198 MLMYLGDNVEGGETHFPSAGSDECSCGGK-----LTKGLCVKPVKGNAVLFWSMGLDGQS 252
+LMYL D +GGET FP+A + GL VKP KG+A+LF+S+ ++ +
Sbjct: 158 VLMYLTDVKKGGETVFPNAAGRHLQLKDETWSDCARSGLAVKPKKGDALLFFSLHVNATT 217
Query: 253 DPDSVHGGCPVLAGEKWSATKWM 275
DP S+HG CPV+ GEKWSATKW+
Sbjct: 218 DPASLHGSCPVIEGEKWSATKWI 240
>ref|XP_469991.1| putative oxidoreductase [Oryza sativa (japonica cultivar-group)]
gi|29150368|gb|AAO72377.1| putative oxidoreductase
[Oryza sativa (japonica cultivar-group)]
Length = 310
Score = 188 bits (478), Expect = 1e-46
Identities = 98/204 (48%), Positives = 132/204 (64%), Gaps = 7/204 (3%)
Query: 77 VLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFLSHE 136
++SW PRI FLS +ECD+L + +LK S V D +GK + S+VRTSSGMFL +
Sbjct: 50 IISWKPRIFFYKGFLSDDECDHLVKLGKEKLKRSMVADNESGKSVMSEVRTSSGMFL--D 107
Query: 137 ERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQRIA 196
+++ P++ IE+RI+ ++ +P EN E +Q+LRYE Q Y PH DYF D N +GG R A
Sbjct: 108 KQQDPVVSGIEERIAAWTLLPQENAENIQILRYENGQKYDPHFDYFQDKVNQLQGGHRYA 167
Query: 197 TMLMYLGDNVEGGETHFPSAGSDEC-----SCGGKLTKGLCVKPVKGNAVLFWSMGLDGQ 251
T+L YL +GGET FP+A E S KGL VK VKG++VLF+++ DG
Sbjct: 168 TVLTYLSTVEKGGETVFPNAEGWESQPKDDSFSDCAKKGLAVKAVKGDSVLFFNLQPDGT 227
Query: 252 SDPDSVHGGCPVLAGEKWSATKWM 275
DP S+HG CPV+ GEKWSA KW+
Sbjct: 228 PDPLSLHGSCPVIEGEKWSAPKWI 251
>gb|AAT77286.1| putative prolyl 4-hydroxylase alpha subunit [Oryza sativa (japonica
cultivar-group)]
Length = 319
Score = 184 bits (466), Expect = 3e-45
Identities = 96/207 (46%), Positives = 132/207 (63%), Gaps = 10/207 (4%)
Query: 78 LSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFLSHEE 137
+SW PR+ L +FLS +E ++L +A LK S V D +GK SD RTSSG F+ +
Sbjct: 61 ISWKPRVFLYQHFLSDDEANHLVSLARTELKRSAVADNLSGKSELSDARTSSGTFIRKSQ 120
Query: 138 RKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQRIAT 197
P++ IE++I+ ++ +P ENGE +QVLRY+ + Y H+DYFSD N RGG RIAT
Sbjct: 121 D--PIVAGIEEKIAAWTFLPKENGEDIQVLRYKHGEKYERHYDYFSDNVNTLRGGHRIAT 178
Query: 198 MLMYLGDNVEGGETHFPSA--------GSDECSCGGKLTKGLCVKPVKGNAVLFWSMGLD 249
+LMYL D EGGET FP A +++ + KG+ VKP KG+A+LF+++ D
Sbjct: 179 VLMYLTDVAEGGETVFPLAEEFTESGTNNEDSTLSECAKKGVAVKPRKGDALLFFNLSPD 238
Query: 250 GQSDPDSVHGGCPVLAGEKWSATKWMR 276
D S+H GCPV+ GEKWSATKW+R
Sbjct: 239 ASKDSLSLHAGCPVIKGEKWSATKWIR 265
>dbj|BAB10411.1| prolyl 4-hydroxylase, alpha subunit-like protein [Arabidopsis
thaliana] gi|15239240|ref|NP_201407.1| oxidoreductase,
2OG-Fe(II) oxygenase family protein [Arabidopsis
thaliana]
Length = 267
Score = 183 bits (464), Expect = 5e-45
Identities = 101/195 (51%), Positives = 123/195 (62%), Gaps = 18/195 (9%)
Query: 76 EVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFLSH 135
E++SW PR + HNFL+ EEC YL +A P ++ STVVD TGK S VRTSSG FL+
Sbjct: 79 EIISWEPRASVYHNFLTKEECKYLIELAKPHMEKSTVVDEKTGKSTDSRVRTSSGTFLAR 138
Query: 136 EERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQRI 195
K I IEKRIS ++ IP+E+GE +QVL YE Q Y PH+DYF D +N + GGQRI
Sbjct: 139 GRDK--TIREIEKRISDFTFIPVEHGEGLQVLHYEIGQKYEPHYDYFMDEYNTRNGGQRI 196
Query: 196 ATMLMYLGDNVEGGETHFPSAGSD-----------ECSCGGKLTKGLCVKPVKGNAVLFW 244
AT+LMYL D EGGET FP+A + EC G GL VKP G+A+LFW
Sbjct: 197 ATVLMYLSDVEEGGETVFPAAKGNYSAVPWWNELSECGKG-----GLSVKPKMGDALLFW 251
Query: 245 SMGLDGQSDPDSVHG 259
SM D DP S+HG
Sbjct: 252 SMTPDATLDPSSLHG 266
>gb|AAL57673.1| AT3g28480/MFJ20_16 [Arabidopsis thaliana]
gi|24796986|gb|AAN64505.1| At3g28480/MFJ20_16
[Arabidopsis thaliana]
Length = 316
Score = 183 bits (464), Expect = 5e-45
Identities = 93/203 (45%), Positives = 137/203 (66%), Gaps = 7/203 (3%)
Query: 78 LSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFLSHEE 137
LSW+PR+ L FLS EECD+ +A +L+ S V D ++G+ ++S+VRTSSGMFLS +
Sbjct: 59 LSWTPRVFLYEGFLSDEECDHFIKLAKGKLEKSMVADNDSGESVESEVRTSSGMFLS--K 116
Query: 138 RKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQRIAT 197
R+ +++ +E +++ ++ +P ENGE MQ+L YE Q Y PH DYF D NL+ GG RIAT
Sbjct: 117 RQDDIVNNVEAKLAAWTFLPEENGESMQILHYENGQKYEPHFDYFHDQANLELGGHRIAT 176
Query: 198 MLMYLGDNVEGGETHFP-----SAGSDECSCGGKLTKGLCVKPVKGNAVLFWSMGLDGQS 252
+LMYL + +GGET FP + + S +G VKP KG+A+LF+++ + +
Sbjct: 177 VLMYLSNVEKGGETVFPMWKGKATQLKDDSWTECAKQGYAVKPRKGDALLFFNLHPNATT 236
Query: 253 DPDSVHGGCPVLAGEKWSATKWM 275
D +S+HG CPV+ GEKWSAT+W+
Sbjct: 237 DSNSLHGSCPVVEGEKWSATRWI 259
>gb|AAM66931.1| prolyl 4-hydroxylase, putative [Arabidopsis thaliana]
gi|18405808|ref|NP_566838.1| oxidoreductase, 2OG-Fe(II)
oxygenase family protein [Arabidopsis thaliana]
Length = 316
Score = 182 bits (462), Expect = 9e-45
Identities = 93/203 (45%), Positives = 136/203 (66%), Gaps = 7/203 (3%)
Query: 78 LSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFLSHEE 137
LSW+PR+ L FLS EECD+ +A +L+ S V D ++G+ ++S+VRTSSGMFLS +
Sbjct: 59 LSWTPRVFLYEGFLSDEECDHFIKLAKGKLEKSMVADNDSGESVESEVRTSSGMFLS--K 116
Query: 138 RKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQRIAT 197
R+ ++ +E +++ ++ +P ENGE MQ+L YE Q Y PH DYF D NL+ GG RIAT
Sbjct: 117 RQDDIVSNVEAKLAAWTFLPEENGESMQILHYENGQKYEPHFDYFHDQANLELGGHRIAT 176
Query: 198 MLMYLGDNVEGGETHFP-----SAGSDECSCGGKLTKGLCVKPVKGNAVLFWSMGLDGQS 252
+LMYL + +GGET FP + + S +G VKP KG+A+LF+++ + +
Sbjct: 177 VLMYLSNVEKGGETVFPMWKGKATQLKDDSWTECAKQGYAVKPRKGDALLFFNLHPNATT 236
Query: 253 DPDSVHGGCPVLAGEKWSATKWM 275
D +S+HG CPV+ GEKWSAT+W+
Sbjct: 237 DSNSLHGSCPVVEGEKWSATRWI 259
>dbj|BAB02864.1| prolyl 4-hydroxylase alpha subunit-like protein [Arabidopsis
thaliana]
Length = 332
Score = 182 bits (462), Expect = 9e-45
Identities = 93/203 (45%), Positives = 136/203 (66%), Gaps = 7/203 (3%)
Query: 78 LSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFLSHEE 137
LSW+PR+ L FLS EECD+ +A +L+ S V D ++G+ ++S+VRTSSGMFLS +
Sbjct: 75 LSWTPRVFLYEGFLSDEECDHFIKLAKGKLEKSMVADNDSGESVESEVRTSSGMFLS--K 132
Query: 138 RKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQRIAT 197
R+ ++ +E +++ ++ +P ENGE MQ+L YE Q Y PH DYF D NL+ GG RIAT
Sbjct: 133 RQDDIVSNVEAKLAAWTFLPEENGESMQILHYENGQKYEPHFDYFHDQANLELGGHRIAT 192
Query: 198 MLMYLGDNVEGGETHFP-----SAGSDECSCGGKLTKGLCVKPVKGNAVLFWSMGLDGQS 252
+LMYL + +GGET FP + + S +G VKP KG+A+LF+++ + +
Sbjct: 193 VLMYLSNVEKGGETVFPMWKGKATQLKDDSWTECAKQGYAVKPRKGDALLFFNLHPNATT 252
Query: 253 DPDSVHGGCPVLAGEKWSATKWM 275
D +S+HG CPV+ GEKWSAT+W+
Sbjct: 253 DSNSLHGSCPVVEGEKWSATRWI 275
>gb|AAM67123.1| prolyl 4-hydroxylase alpha subunit-like protein [Arabidopsis
thaliana]
Length = 297
Score = 182 bits (461), Expect = 1e-44
Identities = 100/212 (47%), Positives = 133/212 (62%), Gaps = 14/212 (6%)
Query: 74 KPEVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFL 133
K + +S PR + FL+ ECD+L +A L+ S V D + G+ SDVRTSSG F+
Sbjct: 35 KVKQVSSKPRAFVYEGFLTDLECDHLISLAKENLQRSAVADNDNGESQVSDVRTSSGTFI 94
Query: 134 SHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQ 193
S + K P++ IE ++S ++ +P ENGE +QVLRYE Q Y H DYF D N+ RGG
Sbjct: 95 S--KGKDPIVSGIEDKLSTWTFLPKENGEDLQVLRYEHGQKYDAHFDYFHDKVNIARGGH 152
Query: 194 RIATMLMYLGDNVEGGETHFPSA----------GSDECSCGGKLTKGLCVKPVKGNAVLF 243
RIAT+L+YL + +GGET FP A D+ S K KG+ VKP KGNA+LF
Sbjct: 153 RIATVLLYLSNVTKGGETVFPDAQEFSRRSLSENKDDLSDCAK--KGIAVKPKKGNALLF 210
Query: 244 WSMGLDGQSDPDSVHGGCPVLAGEKWSATKWM 275
+++ D DP S+HGGCPV+ GEKWSATKW+
Sbjct: 211 FNLQQDAIPDPFSLHGGCPVIEGEKWSATKWI 242
>ref|NP_566279.1| oxidoreductase, 2OG-Fe(II) oxygenase family protein [Arabidopsis
thaliana]
Length = 299
Score = 182 bits (461), Expect = 1e-44
Identities = 100/212 (47%), Positives = 133/212 (62%), Gaps = 14/212 (6%)
Query: 74 KPEVLSWSPRIILLHNFLSYEECDYLRGVALPRLKISTVVDANTGKGIKSDVRTSSGMFL 133
K + +S PR + FL+ ECD+L +A L+ S V D + G+ SDVRTSSG F+
Sbjct: 37 KVKQVSSKPRAFVYEGFLTDLECDHLISLAKENLQRSAVADNDNGESQVSDVRTSSGTFI 96
Query: 134 SHEERKYPMIHAIEKRISVYSQIPIENGELMQVLRYEKNQYYRPHHDYFSDTFNLKRGGQ 193
S + K P++ IE ++S ++ +P ENGE +QVLRYE Q Y H DYF D N+ RGG
Sbjct: 97 S--KGKDPIVSGIEDKLSTWTFLPKENGEDLQVLRYEHGQKYDAHFDYFHDKVNIARGGH 154
Query: 194 RIATMLMYLGDNVEGGETHFPSA----------GSDECSCGGKLTKGLCVKPVKGNAVLF 243
RIAT+L+YL + +GGET FP A D+ S K KG+ VKP KGNA+LF
Sbjct: 155 RIATVLLYLSNVTKGGETVFPDAQEFSRRSLSENKDDLSDCAK--KGIAVKPKKGNALLF 212
Query: 244 WSMGLDGQSDPDSVHGGCPVLAGEKWSATKWM 275
+++ D DP S+HGGCPV+ GEKWSATKW+
Sbjct: 213 FNLQQDAIPDPFSLHGGCPVIEGEKWSATKWI 244
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.320 0.138 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 519,685,288
Number of Sequences: 2540612
Number of extensions: 22842341
Number of successful extensions: 42996
Number of sequences better than 10.0: 239
Number of HSP's better than 10.0 without gapping: 201
Number of HSP's successfully gapped in prelim test: 38
Number of HSP's that attempted gapping in prelim test: 42290
Number of HSP's gapped (non-prelim): 252
length of query: 281
length of database: 863,360,394
effective HSP length: 126
effective length of query: 155
effective length of database: 543,243,282
effective search space: 84202708710
effective search space used: 84202708710
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 74 (33.1 bits)
Medicago: description of AC149471.21


