

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149305.4 - phase: 0 /partial
(542 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
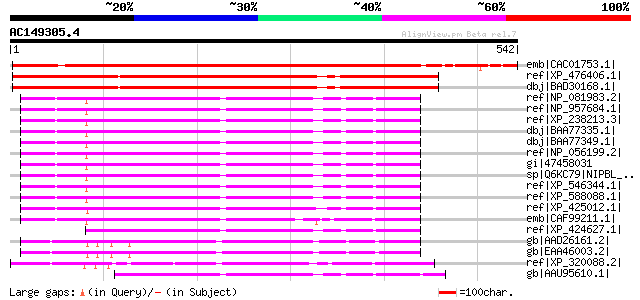
Score E
Sequences producing significant alignments: (bits) Value
emb|CAC01753.1| putative protein [Arabidopsis thaliana] gi|15242... 611 e-173
ref|XP_476406.1| Nipped-B gene product-like protein [Oryza sativ... 529 e-148
dbj|BAD30168.1| putative IDN3 protein isoform A [Oryza sativa (j... 529 e-148
ref|NP_081983.2| delangin isoform A [Mus musculus] gi|50400866|s... 164 5e-39
ref|NP_957684.1| delangin isoform B [Mus musculus] gi|48143965|e... 164 5e-39
ref|XP_238213.3| PREDICTED: similar to delangin [Rattus norvegicus] 164 5e-39
dbj|BAA77335.1| IDN3 [Homo sapiens] 164 7e-39
dbj|BAA77349.1| IDN3-B [Homo sapiens] 164 7e-39
ref|NP_056199.2| delangin isoform B [Homo sapiens] gi|48143963|e... 164 7e-39
gi|47458031 TPA: transcriptional regulator [Homo sapiens] 164 7e-39
sp|Q6KC79|NIPBL_HUMAN Nipped-B-like protein (Delangin) (SCC2 hom... 164 7e-39
ref|XP_546344.1| PREDICTED: similar to delangin isoform A [Canis... 164 9e-39
ref|XP_588088.1| PREDICTED: similar to delangin isoform A, parti... 164 9e-39
ref|XP_425012.1| PREDICTED: similar to transcriptional regulator... 163 1e-38
emb|CAF99211.1| unnamed protein product [Tetraodon nigroviridis] 146 1e-33
ref|XP_424627.1| PREDICTED: similar to transcriptional regulator... 146 2e-33
gb|AAD26161.2| adherin Nipped-B [Drosophila melanogaster] gi|504... 142 4e-32
gb|EAA46003.2| CG17704-PF.3 [Drosophila melanogaster] gi|5195100... 142 4e-32
ref|XP_320088.2| ENSANGP00000010656 [Anopheles gambiae str. PEST... 141 6e-32
gb|AAU95610.1| sister chromatid cohesion establishment factor [X... 131 5e-29
>emb|CAC01753.1| putative protein [Arabidopsis thaliana] gi|15242325|ref|NP_197058.1|
expressed protein [Arabidopsis thaliana]
gi|11358123|pir||T51532 hypothetical protein T20K14_150 -
Arabidopsis thaliana
Length = 1755
Score = 611 bits (1576), Expect = e-173
Identities = 324/545 (59%), Positives = 402/545 (73%), Gaps = 23/545 (4%)
Query: 4 QLLESIIFIIDSVLPLLRKLPPSIVEELEQDLKQMILRHSFLAVVHACIKCLCSMSELAG 63
QLLESIIFIIDSVLPL+RKLP S+ E+LEQDLK MI+RHSFL VVHAC+ S+LAG
Sbjct: 1067 QLLESIIFIIDSVLPLIRKLPLSVTEDLEQDLKHMIVRHSFLTVVHACV------SKLAG 1120
Query: 64 KGAAVIEHLIQVFFKCLDTEAVVNKQLVGRSLFCLGLLIRYGNCLLASSGNKLVDVKRSL 123
KG +++EHL+Q FFK L+ + N Q+ GRSLFCLGLLIR+GN L+++SG K ++ L
Sbjct: 1121 KGVSIVEHLLQFFFKRLEAQGSDNTQIAGRSLFCLGLLIRHGNSLISTSGGKNFNLSGCL 1180
Query: 124 NLFMKYLAGEDYALKARSLQALGYVLIARPEYMLENDIGKILEGTLSSIADDRLKIQALQ 183
NLF ++L ED ALK RSLQALG++LIARPEYMLE DIGKI+E TL+ A+ R+K+QALQ
Sbjct: 1181 NLFKRHLRTEDIALKVRSLQALGFILIARPEYMLEEDIGKIIETTLADEANGRMKMQALQ 1240
Query: 184 NMFEYLLDAESKMETEEVDGKVPGHSVRAGQSVPVAAGAGDTNICGGIIQLYWNNILGRC 243
NM+EYLLDAE ++ +E+ + G +VPVAAGAGDTNICGGI+QL+W+ ILGRC
Sbjct: 1241 NMYEYLLDAEKQLGSEKASDNTVNSVEQGGHNVPVAAGAGDTNICGGIVQLFWDKILGRC 1300
Query: 244 VDFNTQVRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSKMAHHLLMNMHEKYP 303
+DF+ Q+RQ++LKIVEVVLRQGLVHPITCVPYLIALETDP E+N K+AHHLLMNMHEKYP
Sbjct: 1301 LDFDDQIRQTSLKIVEVVLRQGLVHPITCVPYLIALETDPQEANQKLAHHLLMNMHEKYP 1360
Query: 304 AFFESCLGDGLQMSFMFMQSIF-VSPDENVNHKSQSKIAVSGKGKPEADSLAQSRVGVSR 362
AFFES LGDGLQMSF+FMQSI V+ + N + + + + GK + +L Q+R+GVSR
Sbjct: 1361 AFFESRLGDGLQMSFIFMQSISQVTSEPNQSLQQKGSTNMLGKNDHASSTLTQARLGVSR 1420
Query: 363 IYKLIRGNRISRNKFMSSIVRKFDNPKWNKFVIAFLTYCTEVLALLPFVAPDEPLYLIYT 422
IYKLIRGNR+SRNKFM+SIVRKFDNP WN VI+FL YCTE LALLPF +PDEPLYL+Y+
Sbjct: 1421 IYKLIRGNRVSRNKFMTSIVRKFDNPTWNGSVISFLKYCTETLALLPFTSPDEPLYLVYS 1480
Query: 423 INRVVQVRAGPLEANFKAWSSSLLQSEGQGTPHGNGMYQRATDETIHSTQGQSMDLNGPF 482
INRV+Q+RAG +E+N KA LL + T HGNG YQ+ + I MDLN
Sbjct: 1481 INRVMQIRAGAVESNLKA----LLHKDSAKTQHGNGAYQQ---DPIPGHMNM-MDLNTRI 1532
Query: 483 QQNVDVQPYVDDMTSVDLNG-----TNHQLPDYPLSHNGRLKVKPQAAGFADSFTFSKDD 537
Q+ T +DLNG + Q Y + HNG+ V + +D S DD
Sbjct: 1533 QEEPRHWNSYGHATLIDLNGSVYQDSRDQFTSYQV-HNGKADVHKMTS--SDPPELSTDD 1589
Query: 538 LEKVQ 542
L+K+Q
Sbjct: 1590 LQKIQ 1594
>ref|XP_476406.1| Nipped-B gene product-like protein [Oryza sativa (japonica
cultivar-group)]
Length = 1755
Score = 529 bits (1362), Expect = e-148
Identities = 278/458 (60%), Positives = 343/458 (74%), Gaps = 18/458 (3%)
Query: 4 QLLESIIFIIDSVLPLLRKLPPSIVEELEQDLKQMILRHSFLAVVHACIKCLCSMSELAG 63
QLLESIIF+ID+VLPL+ K P S+V ELEQDLKQMI+RHSFL VVHACIKCLC++S+ A
Sbjct: 1120 QLLESIIFVIDAVLPLIWKPPQSVVIELEQDLKQMIVRHSFLTVVHACIKCLCALSKAAD 1179
Query: 64 KGAAVIEHLIQVFFKCLDTEAVVNK--QLVGRSLFCLGLLIRYGNCLLASSGNKLVDVKR 121
+G ++E+L+ +F+K L N QL+GRSLFCLGLL+RYG+ L+A+S N+L D +
Sbjct: 1180 RGPRLLEYLVNIFYKHLSGSNSSNSDSQLLGRSLFCLGLLLRYGSQLMAASENQL-DFPK 1238
Query: 122 SLNLFMK-YLAGEDYALKARSLQALGYVLIARPEYMLENDIGKILEGTLSSIADDRLKIQ 180
++L K YL +D++LK R LQALGY+LIA+P++ML DI ++E +LSS+ D RLKIQ
Sbjct: 1239 IISLLKKEYLLKDDFSLKVRGLQALGYILIAKPDFMLRKDISTLIESSLSSVVDYRLKIQ 1298
Query: 181 ALQNMFEYLLDAESKMETEEVDGKVPGHSVRAGQSVPVAAGAGDTNICGGIIQLYWNNIL 240
LQN+FEYL DAES++ E GK ++ G VPVAAGAGDTNICGGIIQLYWN+IL
Sbjct: 1299 GLQNLFEYLRDAESQLNAEST-GKPTPNATNGGSEVPVAAGAGDTNICGGIIQLYWNSIL 1357
Query: 241 GRCVDFNTQVRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSKMAHHLLMNMHE 300
RC+D N QVRQ+ALKIVE+VLRQGLVHPITCVP+LIALETDPLE NSK+AHHLLMNM+E
Sbjct: 1358 ERCLDINDQVRQTALKIVEIVLRQGLVHPITCVPHLIALETDPLEGNSKLAHHLLMNMNE 1417
Query: 301 KYPAFFESCLGDGLQMSFMFMQSIFVSPDENVNHKSQSKIAVSGKGKPEADSLAQSRVGV 360
KYP+FFES LGDGLQMSF F +S + D +A + K P +A + G+
Sbjct: 1418 KYPSFFESRLGDGLQMSFRFFESTISNHD---------MVATNMKSNP----IAFVKPGI 1464
Query: 361 SRIYKLIRGNRISRNKFMSSIVRKFDNPKWNKFVIAFLTYCTEVLALLPFVAPDEPLYLI 420
SRIY+LIR NR SRNKF+ SIVRKF+ + I+FL YC EVLA LPF +PDEPLYLI
Sbjct: 1465 SRIYRLIRANRNSRNKFVHSIVRKFEGDNRSYPTISFLMYCAEVLASLPFTSPDEPLYLI 1524
Query: 421 YTINRVVQVRAGPLEANFKAWSSSLLQSEGQGTPHGNG 458
Y INRV+Q+RAG +EAN K W+S Q E G P G
Sbjct: 1525 YDINRVIQLRAGAVEANLKNWTSMYQQQEMVGMPRDTG 1562
>dbj|BAD30168.1| putative IDN3 protein isoform A [Oryza sativa (japonica
cultivar-group)]
Length = 764
Score = 529 bits (1362), Expect = e-148
Identities = 278/458 (60%), Positives = 343/458 (74%), Gaps = 18/458 (3%)
Query: 4 QLLESIIFIIDSVLPLLRKLPPSIVEELEQDLKQMILRHSFLAVVHACIKCLCSMSELAG 63
QLLESIIF+ID+VLPL+ K P S+V ELEQDLKQMI+RHSFL VVHACIKCLC++S+ A
Sbjct: 129 QLLESIIFVIDAVLPLIWKPPQSVVIELEQDLKQMIVRHSFLTVVHACIKCLCALSKAAD 188
Query: 64 KGAAVIEHLIQVFFKCLDTEAVVNK--QLVGRSLFCLGLLIRYGNCLLASSGNKLVDVKR 121
+G ++E+L+ +F+K L N QL+GRSLFCLGLL+RYG+ L+A+S N+L D +
Sbjct: 189 RGPRLLEYLVNIFYKHLSGSNSSNSDSQLLGRSLFCLGLLLRYGSQLMAASENQL-DFPK 247
Query: 122 SLNLFMK-YLAGEDYALKARSLQALGYVLIARPEYMLENDIGKILEGTLSSIADDRLKIQ 180
++L K YL +D++LK R LQALGY+LIA+P++ML DI ++E +LSS+ D RLKIQ
Sbjct: 248 IISLLKKEYLLKDDFSLKVRGLQALGYILIAKPDFMLRKDISTLIESSLSSVVDYRLKIQ 307
Query: 181 ALQNMFEYLLDAESKMETEEVDGKVPGHSVRAGQSVPVAAGAGDTNICGGIIQLYWNNIL 240
LQN+FEYL DAES++ E GK ++ G VPVAAGAGDTNICGGIIQLYWN+IL
Sbjct: 308 GLQNLFEYLRDAESQLNAEST-GKPTPNATNGGSEVPVAAGAGDTNICGGIIQLYWNSIL 366
Query: 241 GRCVDFNTQVRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSKMAHHLLMNMHE 300
RC+D N QVRQ+ALKIVE+VLRQGLVHPITCVP+LIALETDPLE NSK+AHHLLMNM+E
Sbjct: 367 ERCLDINDQVRQTALKIVEIVLRQGLVHPITCVPHLIALETDPLEGNSKLAHHLLMNMNE 426
Query: 301 KYPAFFESCLGDGLQMSFMFMQSIFVSPDENVNHKSQSKIAVSGKGKPEADSLAQSRVGV 360
KYP+FFES LGDGLQMSF F +S + D +A + K P +A + G+
Sbjct: 427 KYPSFFESRLGDGLQMSFRFFESTISNHD---------MVATNMKSNP----IAFVKPGI 473
Query: 361 SRIYKLIRGNRISRNKFMSSIVRKFDNPKWNKFVIAFLTYCTEVLALLPFVAPDEPLYLI 420
SRIY+LIR NR SRNKF+ SIVRKF+ + I+FL YC EVLA LPF +PDEPLYLI
Sbjct: 474 SRIYRLIRANRNSRNKFVHSIVRKFEGDNRSYPTISFLMYCAEVLASLPFTSPDEPLYLI 533
Query: 421 YTINRVVQVRAGPLEANFKAWSSSLLQSEGQGTPHGNG 458
Y INRV+Q+RAG +EAN K W+S Q E G P G
Sbjct: 534 YDINRVIQLRAGAVEANLKNWTSMYQQQEMVGMPRDTG 571
>ref|NP_081983.2| delangin isoform A [Mus musculus] gi|50400866|sp|Q6KCD5|NPBL_MOUSE
Nipped-B-like protein (Delangin homolog) (SCC2 homolog)
gi|48143960|emb|CAF25291.1| delangin [Mus musculus]
Length = 2798
Score = 164 bits (416), Expect = 5e-39
Identities = 119/443 (26%), Positives = 208/443 (46%), Gaps = 38/443 (8%)
Query: 12 IIDSVLPLLRKLPPSIVEELEQDLKQMILRHSFLAVVHACIKCLCSMSELAGKGAAVIEH 71
I++ V+PL+ + + +E+DL ++I+++ V H C+ CL ++ + +
Sbjct: 2044 ILELVVPLMEHPSETFLATIEEDLMKLIIKYGMTVVQH-CVSCLGAVVNKVTQNFKFVWA 2102
Query: 72 LIQVFFKCL------------DTEAVVNKQLVGRSLFCLGLLIRYGNCLLAS-SGNKLVD 118
++ + +T + NK + RSLF +G L R+ + L GN V+
Sbjct: 2103 CFNRYYGAISKLKSQHQEDPNNTSLLTNKPALLRSLFTVGALCRHFDFDLEDFKGNSKVN 2162
Query: 119 VK-RSLNLFMKYLAGEDYALKARSLQALGYVLIARPEYMLENDIGKILEGTLSSIADD-R 176
+K + L L M + D ++ +++ LG+ I P M E ++ + LS
Sbjct: 2163 IKDKVLELLMYFTKHSDEEVQTKAIIGLGFAFIQHPSLMFEQEVKNLYNSILSDKNSSVN 2222
Query: 177 LKIQALQNMFEYLLDAESKMETEEVDGKVPGHSVRAGQSVPVAAGAGDTNICGGIIQLYW 236
LKIQ L+N+ YL + +++M+ + D K + V++G + I+QLY
Sbjct: 2223 LKIQVLKNLQTYLQEEDTRMQQADRDWKKVAKQEDLKEMGDVSSGMSSS-----IMQLYL 2277
Query: 237 NNILGRCVDFNTQVRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSKMAHHLLM 296
+L + VR AL ++ + L QGL+HP+ CVPYLIA+ TDP + A L+
Sbjct: 2278 KQVLEAFFHTQSSVRHFALNVIALTLNQGLIHPVQCVPYLIAMGTDPEPAMRNKADQQLV 2337
Query: 297 NMHEKYPAFFESCLGDGLQMSFMFMQSIFVSPDENVNHKSQSKIAVSGKGKPEADSLAQS 356
+ +KY F G++MS+ Q+I + K V G + E+ S
Sbjct: 2338 EIDKKYAGFIHMKAVAGMKMSYQVQQAI----------NTCLKDPVRGFRQDESSSAL-- 2385
Query: 357 RVGVSRIYKLIRGNRISRNKFMSSIVRKFDNPKWNKFVIAFLTYCTEVLALLPFVAPDEP 416
S +Y +IRGNR R F+ S++ FD+ K + L Y + LA P+ +EP
Sbjct: 2386 ---CSHLYSMIRGNRQHRRAFLISLLNLFDDTA--KTEVTMLLYIADNLACFPYQTQEEP 2440
Query: 417 LYLIYTINRVVQVRAGPLEANFK 439
L++++ I+ + V L +FK
Sbjct: 2441 LFIMHHIDITLSVSGSNLLQSFK 2463
>ref|NP_957684.1| delangin isoform B [Mus musculus] gi|48143965|emb|CAG26692.1|
delangin [Mus musculus]
Length = 2691
Score = 164 bits (416), Expect = 5e-39
Identities = 119/443 (26%), Positives = 208/443 (46%), Gaps = 38/443 (8%)
Query: 12 IIDSVLPLLRKLPPSIVEELEQDLKQMILRHSFLAVVHACIKCLCSMSELAGKGAAVIEH 71
I++ V+PL+ + + +E+DL ++I+++ V H C+ CL ++ + +
Sbjct: 2044 ILELVVPLMEHPSETFLATIEEDLMKLIIKYGMTVVQH-CVSCLGAVVNKVTQNFKFVWA 2102
Query: 72 LIQVFFKCL------------DTEAVVNKQLVGRSLFCLGLLIRYGNCLLAS-SGNKLVD 118
++ + +T + NK + RSLF +G L R+ + L GN V+
Sbjct: 2103 CFNRYYGAISKLKSQHQEDPNNTSLLTNKPALLRSLFTVGALCRHFDFDLEDFKGNSKVN 2162
Query: 119 VK-RSLNLFMKYLAGEDYALKARSLQALGYVLIARPEYMLENDIGKILEGTLSSIADD-R 176
+K + L L M + D ++ +++ LG+ I P M E ++ + LS
Sbjct: 2163 IKDKVLELLMYFTKHSDEEVQTKAIIGLGFAFIQHPSLMFEQEVKNLYNSILSDKNSSVN 2222
Query: 177 LKIQALQNMFEYLLDAESKMETEEVDGKVPGHSVRAGQSVPVAAGAGDTNICGGIIQLYW 236
LKIQ L+N+ YL + +++M+ + D K + V++G + I+QLY
Sbjct: 2223 LKIQVLKNLQTYLQEEDTRMQQADRDWKKVAKQEDLKEMGDVSSGMSSS-----IMQLYL 2277
Query: 237 NNILGRCVDFNTQVRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSKMAHHLLM 296
+L + VR AL ++ + L QGL+HP+ CVPYLIA+ TDP + A L+
Sbjct: 2278 KQVLEAFFHTQSSVRHFALNVIALTLNQGLIHPVQCVPYLIAMGTDPEPAMRNKADQQLV 2337
Query: 297 NMHEKYPAFFESCLGDGLQMSFMFMQSIFVSPDENVNHKSQSKIAVSGKGKPEADSLAQS 356
+ +KY F G++MS+ Q+I + K V G + E+ S
Sbjct: 2338 EIDKKYAGFIHMKAVAGMKMSYQVQQAI----------NTCLKDPVRGFRQDESSSAL-- 2385
Query: 357 RVGVSRIYKLIRGNRISRNKFMSSIVRKFDNPKWNKFVIAFLTYCTEVLALLPFVAPDEP 416
S +Y +IRGNR R F+ S++ FD+ K + L Y + LA P+ +EP
Sbjct: 2386 ---CSHLYSMIRGNRQHRRAFLISLLNLFDDTA--KTEVTMLLYIADNLACFPYQTQEEP 2440
Query: 417 LYLIYTINRVVQVRAGPLEANFK 439
L++++ I+ + V L +FK
Sbjct: 2441 LFIMHHIDITLSVSGSNLLQSFK 2463
>ref|XP_238213.3| PREDICTED: similar to delangin [Rattus norvegicus]
Length = 2798
Score = 164 bits (416), Expect = 5e-39
Identities = 119/443 (26%), Positives = 208/443 (46%), Gaps = 38/443 (8%)
Query: 12 IIDSVLPLLRKLPPSIVEELEQDLKQMILRHSFLAVVHACIKCLCSMSELAGKGAAVIEH 71
I++ V+PL+ + + +E+DL ++I+++ V H C+ CL ++ + +
Sbjct: 2044 ILELVVPLMEHPSETFLATIEEDLMKLIIKYGMTVVQH-CVSCLGAVVNKVTQNFKFVWA 2102
Query: 72 LIQVFFKCL------------DTEAVVNKQLVGRSLFCLGLLIRYGNCLLAS-SGNKLVD 118
++ + +T + NK + RSLF +G L R+ + L GN V+
Sbjct: 2103 CFNRYYGAISKLKSQHQEDPNNTSLLTNKPALLRSLFTVGALCRHFDFDLEDFKGNSKVN 2162
Query: 119 VK-RSLNLFMKYLAGEDYALKARSLQALGYVLIARPEYMLENDIGKILEGTLSSIADD-R 176
+K + L L M + D ++ +++ LG+ I P M E ++ + LS
Sbjct: 2163 IKDKVLELLMYFTKHSDEEVQTKAIIGLGFAFIQHPSLMFEQEVKNLYNSILSDKNSSVN 2222
Query: 177 LKIQALQNMFEYLLDAESKMETEEVDGKVPGHSVRAGQSVPVAAGAGDTNICGGIIQLYW 236
LKIQ L+N+ YL + +++M+ + D K + V++G + I+QLY
Sbjct: 2223 LKIQVLKNLQTYLQEEDTRMQQADRDWKKVAKQEDLKEMGDVSSGMSSS-----IMQLYL 2277
Query: 237 NNILGRCVDFNTQVRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSKMAHHLLM 296
+L + VR AL ++ + L QGL+HP+ CVPYLIA+ TDP + A L+
Sbjct: 2278 KQVLEAFFHTQSSVRHFALNVIALTLNQGLIHPVQCVPYLIAMGTDPEPAMRNKADQQLV 2337
Query: 297 NMHEKYPAFFESCLGDGLQMSFMFMQSIFVSPDENVNHKSQSKIAVSGKGKPEADSLAQS 356
+ +KY F G++MS+ Q+I + K V G + E+ S
Sbjct: 2338 EIDKKYAGFIHMKAVAGMKMSYQVQQAI----------NTCLKDPVRGFRQDESSSAL-- 2385
Query: 357 RVGVSRIYKLIRGNRISRNKFMSSIVRKFDNPKWNKFVIAFLTYCTEVLALLPFVAPDEP 416
S +Y +IRGNR R F+ S++ FD+ K + L Y + LA P+ +EP
Sbjct: 2386 ---CSHLYSMIRGNRQHRRAFLISLLNLFDDTA--KTEVTMLLYIADNLACFPYQTQEEP 2440
Query: 417 LYLIYTINRVVQVRAGPLEANFK 439
L++++ I+ + V L +FK
Sbjct: 2441 LFIMHHIDITLSVSGSNLLQSFK 2463
>dbj|BAA77335.1| IDN3 [Homo sapiens]
Length = 2265
Score = 164 bits (415), Expect = 7e-39
Identities = 119/443 (26%), Positives = 208/443 (46%), Gaps = 38/443 (8%)
Query: 12 IIDSVLPLLRKLPPSIVEELEQDLKQMILRHSFLAVVHACIKCLCSMSELAGKGAAVIEH 71
I++ V+PL+ + + +E+DL ++I+++ V H C+ CL ++ + +
Sbjct: 1511 ILELVVPLMEHPSETFLATIEEDLMKLIIKYGMTVVQH-CVSCLGAVVNKVTQNFKFVWA 1569
Query: 72 LIQVFFKCL------------DTEAVVNKQLVGRSLFCLGLLIRYGNCLLAS-SGNKLVD 118
++ + +T + NK + RSLF +G L R+ + L GN V+
Sbjct: 1570 CFNRYYGAISKLKSQHQEDPNNTSLLTNKPALLRSLFTVGALCRHFDFDLEDFKGNSKVN 1629
Query: 119 VK-RSLNLFMKYLAGEDYALKARSLQALGYVLIARPEYMLENDIGKILEGTLSSIADD-R 176
+K + L L M + D ++ +++ LG+ I P M E ++ + LS
Sbjct: 1630 IKDKVLELLMYFTKHSDEEVQTKAIIGLGFAFIQHPSLMFEQEVKNLYNNILSDKNSSVN 1689
Query: 177 LKIQALQNMFEYLLDAESKMETEEVDGKVPGHSVRAGQSVPVAAGAGDTNICGGIIQLYW 236
LKIQ L+N+ YL + +++M+ + D K + V++G + I+QLY
Sbjct: 1690 LKIQVLKNLQTYLQEEDTRMQQADRDWKKVAKQEDLKEMGDVSSGMSSS-----IMQLYL 1744
Query: 237 NNILGRCVDFNTQVRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSKMAHHLLM 296
+L + VR AL ++ + L QGL+HP+ CVPYLIA+ TDP + A L+
Sbjct: 1745 KQVLEAFFHTQSSVRHFALNVIALTLNQGLIHPVQCVPYLIAMGTDPEPAMRNKADQQLV 1804
Query: 297 NMHEKYPAFFESCLGDGLQMSFMFMQSIFVSPDENVNHKSQSKIAVSGKGKPEADSLAQS 356
+ +KY F G++MS+ Q+I + K V G + E+ S
Sbjct: 1805 EIDKKYAGFIHMKAVAGMKMSYQVQQAI----------NTCLKDPVRGFRQDESSSAL-- 1852
Query: 357 RVGVSRIYKLIRGNRISRNKFMSSIVRKFDNPKWNKFVIAFLTYCTEVLALLPFVAPDEP 416
S +Y +IRGNR R F+ S++ FD+ K + L Y + LA P+ +EP
Sbjct: 1853 ---CSHLYSMIRGNRQHRRAFLISLLNLFDDTA--KTDVTMLLYIADNLACFPYQTQEEP 1907
Query: 417 LYLIYTINRVVQVRAGPLEANFK 439
L++++ I+ + V L +FK
Sbjct: 1908 LFIMHHIDITLSVSGSNLLQSFK 1930
>dbj|BAA77349.1| IDN3-B [Homo sapiens]
Length = 2158
Score = 164 bits (415), Expect = 7e-39
Identities = 119/443 (26%), Positives = 208/443 (46%), Gaps = 38/443 (8%)
Query: 12 IIDSVLPLLRKLPPSIVEELEQDLKQMILRHSFLAVVHACIKCLCSMSELAGKGAAVIEH 71
I++ V+PL+ + + +E+DL ++I+++ V H C+ CL ++ + +
Sbjct: 1511 ILELVVPLMEHPSETFLATIEEDLMKLIIKYGMTVVQH-CVSCLGAVVNKVTQNFKFVWA 1569
Query: 72 LIQVFFKCL------------DTEAVVNKQLVGRSLFCLGLLIRYGNCLLAS-SGNKLVD 118
++ + +T + NK + RSLF +G L R+ + L GN V+
Sbjct: 1570 CFNRYYGAISKLKSQHQEDPNNTSLLTNKPALLRSLFTVGALCRHFDFDLEDFKGNSKVN 1629
Query: 119 VK-RSLNLFMKYLAGEDYALKARSLQALGYVLIARPEYMLENDIGKILEGTLSSIADD-R 176
+K + L L M + D ++ +++ LG+ I P M E ++ + LS
Sbjct: 1630 IKDKVLELLMYFTKHSDEEVQTKAIIGLGFAFIQHPSLMFEQEVKNLYNNILSDKNSSVN 1689
Query: 177 LKIQALQNMFEYLLDAESKMETEEVDGKVPGHSVRAGQSVPVAAGAGDTNICGGIIQLYW 236
LKIQ L+N+ YL + +++M+ + D K + V++G + I+QLY
Sbjct: 1690 LKIQVLKNLQTYLQEEDTRMQQADRDWKKVAKQEDLKEMGDVSSGMSSS-----IMQLYL 1744
Query: 237 NNILGRCVDFNTQVRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSKMAHHLLM 296
+L + VR AL ++ + L QGL+HP+ CVPYLIA+ TDP + A L+
Sbjct: 1745 KQVLEAFFHTQSSVRHFALNVIALTLNQGLIHPVQCVPYLIAMGTDPEPAMRNKADQQLV 1804
Query: 297 NMHEKYPAFFESCLGDGLQMSFMFMQSIFVSPDENVNHKSQSKIAVSGKGKPEADSLAQS 356
+ +KY F G++MS+ Q+I + K V G + E+ S
Sbjct: 1805 EIDKKYAGFIHMKAVAGMKMSYQVQQAI----------NTCLKDPVRGFRQDESSSAL-- 1852
Query: 357 RVGVSRIYKLIRGNRISRNKFMSSIVRKFDNPKWNKFVIAFLTYCTEVLALLPFVAPDEP 416
S +Y +IRGNR R F+ S++ FD+ K + L Y + LA P+ +EP
Sbjct: 1853 ---CSHLYSMIRGNRQHRRAFLISLLNLFDDTA--KTDVTMLLYIADNLACFPYQTQEEP 1907
Query: 417 LYLIYTINRVVQVRAGPLEANFK 439
L++++ I+ + V L +FK
Sbjct: 1908 LFIMHHIDITLSVSGSNLLQSFK 1930
>ref|NP_056199.2| delangin isoform B [Homo sapiens] gi|48143963|emb|CAG26691.1|
delangin [Homo sapiens]
Length = 2697
Score = 164 bits (415), Expect = 7e-39
Identities = 119/443 (26%), Positives = 208/443 (46%), Gaps = 38/443 (8%)
Query: 12 IIDSVLPLLRKLPPSIVEELEQDLKQMILRHSFLAVVHACIKCLCSMSELAGKGAAVIEH 71
I++ V+PL+ + + +E+DL ++I+++ V H C+ CL ++ + +
Sbjct: 2050 ILELVVPLMEHPSETFLATIEEDLMKLIIKYGMTVVQH-CVSCLGAVVNKVTQNFKFVWA 2108
Query: 72 LIQVFFKCL------------DTEAVVNKQLVGRSLFCLGLLIRYGNCLLAS-SGNKLVD 118
++ + +T + NK + RSLF +G L R+ + L GN V+
Sbjct: 2109 CFNRYYGAISKLKSQHQEDPNNTSLLTNKPALLRSLFTVGALCRHFDFDLEDFKGNSKVN 2168
Query: 119 VK-RSLNLFMKYLAGEDYALKARSLQALGYVLIARPEYMLENDIGKILEGTLSSIADD-R 176
+K + L L M + D ++ +++ LG+ I P M E ++ + LS
Sbjct: 2169 IKDKVLELLMYFTKHSDEEVQTKAIIGLGFAFIQHPSLMFEQEVKNLYNNILSDKNSSVN 2228
Query: 177 LKIQALQNMFEYLLDAESKMETEEVDGKVPGHSVRAGQSVPVAAGAGDTNICGGIIQLYW 236
LKIQ L+N+ YL + +++M+ + D K + V++G + I+QLY
Sbjct: 2229 LKIQVLKNLQTYLQEEDTRMQQADRDWKKVAKQEDLKEMGDVSSGMSSS-----IMQLYL 2283
Query: 237 NNILGRCVDFNTQVRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSKMAHHLLM 296
+L + VR AL ++ + L QGL+HP+ CVPYLIA+ TDP + A L+
Sbjct: 2284 KQVLEAFFHTQSSVRHFALNVIALTLNQGLIHPVQCVPYLIAMGTDPEPAMRNKADQQLV 2343
Query: 297 NMHEKYPAFFESCLGDGLQMSFMFMQSIFVSPDENVNHKSQSKIAVSGKGKPEADSLAQS 356
+ +KY F G++MS+ Q+I + K V G + E+ S
Sbjct: 2344 EIDKKYAGFIHMKAVAGMKMSYQVQQAI----------NTCLKDPVRGFRQDESSSAL-- 2391
Query: 357 RVGVSRIYKLIRGNRISRNKFMSSIVRKFDNPKWNKFVIAFLTYCTEVLALLPFVAPDEP 416
S +Y +IRGNR R F+ S++ FD+ K + L Y + LA P+ +EP
Sbjct: 2392 ---CSHLYSMIRGNRQHRRAFLISLLNLFDDTA--KTDVTMLLYIADNLACFPYQTQEEP 2446
Query: 417 LYLIYTINRVVQVRAGPLEANFK 439
L++++ I+ + V L +FK
Sbjct: 2447 LFIMHHIDITLSVSGSNLLQSFK 2469
>gi|47458031 TPA: transcriptional regulator [Homo sapiens]
Length = 2804
Score = 164 bits (415), Expect = 7e-39
Identities = 119/443 (26%), Positives = 208/443 (46%), Gaps = 38/443 (8%)
Query: 12 IIDSVLPLLRKLPPSIVEELEQDLKQMILRHSFLAVVHACIKCLCSMSELAGKGAAVIEH 71
I++ V+PL+ + + +E+DL ++I+++ V H C+ CL ++ + +
Sbjct: 2050 ILELVVPLMEHPSETFLATIEEDLMKLIIKYGMTVVQH-CVSCLGAVVNKVTQNFKFVWA 2108
Query: 72 LIQVFFKCL------------DTEAVVNKQLVGRSLFCLGLLIRYGNCLLAS-SGNKLVD 118
++ + +T + NK + RSLF +G L R+ + L GN V+
Sbjct: 2109 CFNRYYGAISKLKSQHQEDPNNTSLLTNKPALLRSLFTVGALCRHFDFDLEDFKGNSKVN 2168
Query: 119 VK-RSLNLFMKYLAGEDYALKARSLQALGYVLIARPEYMLENDIGKILEGTLSSIADD-R 176
+K + L L M + D ++ +++ LG+ I P M E ++ + LS
Sbjct: 2169 IKDKVLELLMYFTKHSDEEVQTKAIIGLGFAFIQHPSLMFEQEVKNLYNNILSDKNSSVN 2228
Query: 177 LKIQALQNMFEYLLDAESKMETEEVDGKVPGHSVRAGQSVPVAAGAGDTNICGGIIQLYW 236
LKIQ L+N+ YL + +++M+ + D K + V++G + I+QLY
Sbjct: 2229 LKIQVLKNLQTYLQEEDTRMQQADRDWKKVAKQEDLKEMGDVSSGMSSS-----IMQLYL 2283
Query: 237 NNILGRCVDFNTQVRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSKMAHHLLM 296
+L + VR AL ++ + L QGL+HP+ CVPYLIA+ TDP + A L+
Sbjct: 2284 KQVLEAFFHTQSSVRHFALNVIALTLNQGLIHPVQCVPYLIAMGTDPEPAMRNKADQQLV 2343
Query: 297 NMHEKYPAFFESCLGDGLQMSFMFMQSIFVSPDENVNHKSQSKIAVSGKGKPEADSLAQS 356
+ +KY F G++MS+ Q+I + K V G + E+ S
Sbjct: 2344 EIDKKYAGFIHMKAVAGMKMSYQVQQAI----------NTCLKDPVRGFRQDESSSAL-- 2391
Query: 357 RVGVSRIYKLIRGNRISRNKFMSSIVRKFDNPKWNKFVIAFLTYCTEVLALLPFVAPDEP 416
S +Y +IRGNR R F+ S++ FD+ K + L Y + LA P+ +EP
Sbjct: 2392 ---CSHLYSMIRGNRQHRRAFLISLLNLFDDTA--KTDVTMLLYIADNLACFPYQTQEEP 2446
Query: 417 LYLIYTINRVVQVRAGPLEANFK 439
L++++ I+ + V L +FK
Sbjct: 2447 LFIMHHIDITLSVSGSNLLQSFK 2469
>sp|Q6KC79|NIPBL_HUMAN Nipped-B-like protein (Delangin) (SCC2 homolog)
gi|47578105|ref|NP_597677.2| delangin isoform A [Homo
sapiens] gi|48143958|emb|CAF25290.1| delangin [Homo
sapiens]
Length = 2804
Score = 164 bits (415), Expect = 7e-39
Identities = 119/443 (26%), Positives = 208/443 (46%), Gaps = 38/443 (8%)
Query: 12 IIDSVLPLLRKLPPSIVEELEQDLKQMILRHSFLAVVHACIKCLCSMSELAGKGAAVIEH 71
I++ V+PL+ + + +E+DL ++I+++ V H C+ CL ++ + +
Sbjct: 2050 ILELVVPLMEHPSETFLATIEEDLMKLIIKYGMTVVQH-CVSCLGAVVNKVTQNFKFVWA 2108
Query: 72 LIQVFFKCL------------DTEAVVNKQLVGRSLFCLGLLIRYGNCLLAS-SGNKLVD 118
++ + +T + NK + RSLF +G L R+ + L GN V+
Sbjct: 2109 CFNRYYGAISKLKSQHQEDPNNTSLLTNKPALLRSLFTVGALCRHFDFDLEDFKGNSKVN 2168
Query: 119 VK-RSLNLFMKYLAGEDYALKARSLQALGYVLIARPEYMLENDIGKILEGTLSSIADD-R 176
+K + L L M + D ++ +++ LG+ I P M E ++ + LS
Sbjct: 2169 IKDKVLELLMYFTKHSDEEVQTKAIIGLGFAFIQHPSLMFEQEVKNLYNNILSDKNSSVN 2228
Query: 177 LKIQALQNMFEYLLDAESKMETEEVDGKVPGHSVRAGQSVPVAAGAGDTNICGGIIQLYW 236
LKIQ L+N+ YL + +++M+ + D K + V++G + I+QLY
Sbjct: 2229 LKIQVLKNLQTYLQEEDTRMQQADRDWKKVAKQEDLKEMGDVSSGMSSS-----IMQLYL 2283
Query: 237 NNILGRCVDFNTQVRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSKMAHHLLM 296
+L + VR AL ++ + L QGL+HP+ CVPYLIA+ TDP + A L+
Sbjct: 2284 KQVLEAFFHTQSSVRHFALNVIALTLNQGLIHPVQCVPYLIAMGTDPEPAMRNKADQQLV 2343
Query: 297 NMHEKYPAFFESCLGDGLQMSFMFMQSIFVSPDENVNHKSQSKIAVSGKGKPEADSLAQS 356
+ +KY F G++MS+ Q+I + K V G + E+ S
Sbjct: 2344 EIDKKYAGFIHMKAVAGMKMSYQVQQAI----------NTCLKDPVRGFRQDESSSAL-- 2391
Query: 357 RVGVSRIYKLIRGNRISRNKFMSSIVRKFDNPKWNKFVIAFLTYCTEVLALLPFVAPDEP 416
S +Y +IRGNR R F+ S++ FD+ K + L Y + LA P+ +EP
Sbjct: 2392 ---CSHLYSMIRGNRQHRRAFLISLLNLFDDTA--KTDVTMLLYIADNLACFPYQTQEEP 2446
Query: 417 LYLIYTINRVVQVRAGPLEANFK 439
L++++ I+ + V L +FK
Sbjct: 2447 LFIMHHIDITLSVSGSNLLQSFK 2469
>ref|XP_546344.1| PREDICTED: similar to delangin isoform A [Canis familiaris]
Length = 2936
Score = 164 bits (414), Expect = 9e-39
Identities = 119/443 (26%), Positives = 208/443 (46%), Gaps = 38/443 (8%)
Query: 12 IIDSVLPLLRKLPPSIVEELEQDLKQMILRHSFLAVVHACIKCLCSMSELAGKGAAVIEH 71
I++ V+PL+ + + +E+DL ++I+++ V H C+ CL ++ + +
Sbjct: 2182 ILELVVPLMEHPSETFLATIEEDLMKLIIKYGMTVVQH-CVSCLGAVVNKVTQNFKFVWA 2240
Query: 72 LIQVFFKCL------------DTEAVVNKQLVGRSLFCLGLLIRYGNCLLAS-SGNKLVD 118
++ + +T + NK + RSLF +G L R+ + L GN V+
Sbjct: 2241 CFNRYYGAISKLKSQHQEDPNNTSLLTNKPALLRSLFTVGALCRHFDFDLEDFKGNSKVN 2300
Query: 119 VK-RSLNLFMKYLAGEDYALKARSLQALGYVLIARPEYMLENDIGKILEGTLSSIADD-R 176
+K + L L M + D ++ +++ LG+ I P M E ++ + LS
Sbjct: 2301 IKDKVLELLMYFTKHSDEEVQTKAIIGLGFAFIQHPSLMFEQEVKNLYNNILSDKNSSVN 2360
Query: 177 LKIQALQNMFEYLLDAESKMETEEVDGKVPGHSVRAGQSVPVAAGAGDTNICGGIIQLYW 236
LKIQ L+N+ YL + +++M+ + D K + V++G + I+QLY
Sbjct: 2361 LKIQVLKNLQTYLQEEDTRMQQADRDWKKVAKQEDLKEMGDVSSGMSSS-----IMQLYL 2415
Query: 237 NNILGRCVDFNTQVRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSKMAHHLLM 296
+L + VR AL ++ + L QGL+HP+ CVPYLIA+ TDP + A L+
Sbjct: 2416 KQVLEAFFHTQSSVRHFALNVIALTLNQGLIHPVQCVPYLIAMGTDPEPAMRNKADQQLV 2475
Query: 297 NMHEKYPAFFESCLGDGLQMSFMFMQSIFVSPDENVNHKSQSKIAVSGKGKPEADSLAQS 356
+ +KY F G++MS+ Q+I + K V G + E+ S
Sbjct: 2476 EIDKKYAGFIHMKAVAGMKMSYQVQQAI----------NTCLKDPVRGFRQDESSSAL-- 2523
Query: 357 RVGVSRIYKLIRGNRISRNKFMSSIVRKFDNPKWNKFVIAFLTYCTEVLALLPFVAPDEP 416
S +Y +IRGNR R F+ S++ FD+ K + L Y + LA P+ +EP
Sbjct: 2524 ---CSHLYSMIRGNRQHRRAFLISLLNLFDDTA--KTEVNMLLYIADNLACFPYQTQEEP 2578
Query: 417 LYLIYTINRVVQVRAGPLEANFK 439
L++++ I+ + V L +FK
Sbjct: 2579 LFIMHHIDITLSVSGSNLLQSFK 2601
>ref|XP_588088.1| PREDICTED: similar to delangin isoform A, partial [Bos taurus]
Length = 525
Score = 164 bits (414), Expect = 9e-39
Identities = 119/443 (26%), Positives = 208/443 (46%), Gaps = 38/443 (8%)
Query: 12 IIDSVLPLLRKLPPSIVEELEQDLKQMILRHSFLAVVHACIKCLCSMSELAGKGAAVIEH 71
I++ V+PL+ + + +E+DL ++I+++ V H C+ CL ++ + +
Sbjct: 14 ILELVVPLMEHPSETFLATIEEDLMKLIIKYGMTVVQH-CVSCLGAVVNKVTQNFKFVWA 72
Query: 72 LIQVFFKCL------------DTEAVVNKQLVGRSLFCLGLLIRYGNCLLAS-SGNKLVD 118
++ + +T + NK + RSLF +G L R+ + L GN V+
Sbjct: 73 CFNRYYGAISKLKSQHQEDPNNTSLLTNKPALLRSLFTVGALCRHFDFDLEDFKGNSKVN 132
Query: 119 VK-RSLNLFMKYLAGEDYALKARSLQALGYVLIARPEYMLENDIGKILEGTLSSIADD-R 176
+K + L L M + D ++ +++ LG+ I P M E ++ + LS
Sbjct: 133 IKDKVLELLMYFTKHSDEEVQTKAIIGLGFAFIQHPSLMFEQEVKNLYNNILSDKNSSVN 192
Query: 177 LKIQALQNMFEYLLDAESKMETEEVDGKVPGHSVRAGQSVPVAAGAGDTNICGGIIQLYW 236
LKIQ L+N+ YL + +++M+ + D K + V++G + I+QLY
Sbjct: 193 LKIQVLKNLQTYLQEEDTRMQQADRDWKKVAKQEDLKEMGDVSSGMSSS-----IMQLYL 247
Query: 237 NNILGRCVDFNTQVRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSKMAHHLLM 296
+L + VR AL ++ + L QGL+HP+ CVPYLIA+ TDP + A L+
Sbjct: 248 KQVLEAFFHTQSSVRHFALNVIALTLNQGLIHPVQCVPYLIAMGTDPEPAMRNKADQQLV 307
Query: 297 NMHEKYPAFFESCLGDGLQMSFMFMQSIFVSPDENVNHKSQSKIAVSGKGKPEADSLAQS 356
+ +KY F G++MS+ Q+I + K V G + E+ S
Sbjct: 308 EIDKKYAGFIHMKAVAGMKMSYQVQQAI----------NTCLKDPVRGFRQDESSSAL-- 355
Query: 357 RVGVSRIYKLIRGNRISRNKFMSSIVRKFDNPKWNKFVIAFLTYCTEVLALLPFVAPDEP 416
S +Y +IRGNR R F+ S++ FD+ K + L Y + LA P+ +EP
Sbjct: 356 ---CSHLYSMIRGNRQHRRAFLISLLNLFDDTA--KTEVNMLLYIADNLACFPYQTQEEP 410
Query: 417 LYLIYTINRVVQVRAGPLEANFK 439
L++++ I+ + V L +FK
Sbjct: 411 LFIMHHIDITLSVSGSNLLQSFK 433
>ref|XP_425012.1| PREDICTED: similar to transcriptional regulator [Gallus gallus]
Length = 2380
Score = 163 bits (412), Expect = 1e-38
Identities = 121/443 (27%), Positives = 204/443 (45%), Gaps = 38/443 (8%)
Query: 12 IIDSVLPLLRKLPPSIVEELEQDLKQMILRHSFLAVVHACIKCLCSMSELAGKGAAVIEH 71
I++ V+PL+ + + +E+DL ++I+++ V H C+ CL S+ + +
Sbjct: 1620 ILELVVPLMEHPSETFLATIEEDLMKLIIKYGMTVVQH-CVSCLGSVVNKVTQNYKFVWA 1678
Query: 72 LIQVFFKCLD------------TEAVVNKQLVGRSLFCLGLLIRYGNCLLAS-SGNKLVD 118
++ L T NK + RSLF +G L R+ + GN V+
Sbjct: 1679 CFNRYYGALSKLKSQHQEDPNSTILTANKPALLRSLFTVGALCRHFDFDHEDFKGNSKVN 1738
Query: 119 VK-RSLNLFMKYLAGEDYALKARSLQALGYVLIARPEYMLENDIGKILEGTLSSI-ADDR 176
+K + L L M + D ++ +++ LG+ I P M E ++ + LS
Sbjct: 1739 IKDKVLELLMYFTKHSDEEVQTKAIIGLGFAFIQHPSLMFEQEVKTLYNSILSDKNCSVN 1798
Query: 177 LKIQALQNMFEYLLDAESKMETEEVDGKVPGHSVRAGQSVPVAAGAGDTNICGGIIQLYW 236
LKIQ L+N+ YL + +++M+ + D K + +++G + I+QLY
Sbjct: 1799 LKIQVLKNLQTYLQEEDTRMQQADRDWKKVAKQEDLKEMGDISSGMSSS-----IMQLYL 1853
Query: 237 NNILGRCVDFNTQVRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSKMAHHLLM 296
+L + VR AL ++ + L QGL+HP+ CVPYLIA+ TDP S A L+
Sbjct: 1854 KQVLEAFFHTQSNVRHFALNVIALTLNQGLIHPVQCVPYLIAMGTDPEPSMRNKADQQLV 1913
Query: 297 NMHEKYPAFFESCLGDGLQMSFMFMQSIFVSPDENVNHKSQSKIAVSGKGKPEADSLAQS 356
+ +KY F G++MS+ Q+I P K V G E+ S
Sbjct: 1914 EIDKKYAGFIHMKAVAGMKMSYQVQQAINTCP----------KDPVRGFRHDESSSAL-- 1961
Query: 357 RVGVSRIYKLIRGNRISRNKFMSSIVRKFDNPKWNKFVIAFLTYCTEVLALLPFVAPDEP 416
S +Y +IRGNR R F+ S++ FD+ K + L Y + LA P+ +EP
Sbjct: 1962 ---CSHLYSMIRGNRQHRRAFLISLLNLFDDTA--KTEVNMLLYIADNLACFPYQTQEEP 2016
Query: 417 LYLIYTINRVVQVRAGPLEANFK 439
L++++ I+ + V L +FK
Sbjct: 2017 LFIMHHIDITLSVSGSNLLQSFK 2039
>emb|CAF99211.1| unnamed protein product [Tetraodon nigroviridis]
Length = 1794
Score = 146 bits (369), Expect = 1e-33
Identities = 114/446 (25%), Positives = 205/446 (45%), Gaps = 38/446 (8%)
Query: 12 IIDSVLPLLRKLPPSIVEELEQDLKQMILRHSFLAVVHACIKCLCSMSELAGKGAAVIEH 71
I++ V+PL+ + + +E+DL ++I+++ V H C+ CL ++ +
Sbjct: 1098 ILELVVPLMEHPSETFLTTIEEDLMKLIIKYGMTVVQH-CVSCLGAIINKVTHNYKFVWA 1156
Query: 72 LIQVFFKCL------------DTEAVVNKQLVGRSLFCLGLLIRYGNCLLAS--SGNKLV 117
++ L NK + RSLF +G L R+ + NK+V
Sbjct: 1157 CFNRYYGALAKLKTQHQEDPSSPTLASNKPTLLRSLFTVGALCRHFDFDQEDFKGANKIV 1216
Query: 118 DVKRSLNLFMKYLAGEDYALKARSLQALGYVLIARPEYMLENDIGKILEGTLSSIADD-R 176
+ L L + + ED ++ +++ LG+ I PE M D+ + LS
Sbjct: 1217 IKDKVLELLLYFTTHEDEEVQLKAIIGLGFQFIMHPELMFVQDVKVLYNSILSDENSSVS 1276
Query: 177 LKIQALQNMFEYLLDAESKMETEEVDGKVPGHSVRAGQSVPVAAGAGDTNICGGIIQLYW 236
LKIQ L+N+ YL + +S+M+ + + K + +++G + I+Q+Y
Sbjct: 1277 LKIQVLKNLQTYLQEEDSRMQEADREWKNKSKQEDLKEMGDISSGMSSS-----IMQIYL 1331
Query: 237 NNILGRCVDFNTQVRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSKMAHHLLM 296
+L + VR AL ++ + L QGL+HP+ CVPYLIA+ TDP + A L+
Sbjct: 1332 KQVLESFFHAQSTVRHFALSVITLTLSQGLIHPVQCVPYLIAMGTDPEPTMKNKADQQLV 1391
Query: 297 NMHEKYPAFFESCLGDGLQMSFMFMQSIFV---SPDENVNHKSQSKIAVSGKGKPEADSL 353
+ +KY F + +S F Q+ F +P ++ + SQ V+ P + L
Sbjct: 1392 EIDKKYSGF--------IHVSDCFFQNGFKTVRNPSFSI-YVSQLSACVA---LPPSYLL 1439
Query: 354 AQSRVGVSRIYKLIRGNRISRNKFMSSIVRKFDNPKWNKFVIAFLTYCTEVLALLPFVAP 413
+ +Y ++RGNR R F+ S++ FD+ +K + L + + LA P+
Sbjct: 1440 LSGSDTFAHLYTMVRGNRQHRRAFLISLLNLFDDS--SKTEVHMLLFVADSLACFPYQTQ 1497
Query: 414 DEPLYLIYTINRVVQVRAGPLEANFK 439
DEPL++++ I+ + V L +FK
Sbjct: 1498 DEPLFIMHHIDITLSVSGSNLLQSFK 1523
>ref|XP_424627.1| PREDICTED: similar to transcriptional regulator, partial [Gallus
gallus]
Length = 725
Score = 146 bits (368), Expect = 2e-33
Identities = 105/361 (29%), Positives = 171/361 (47%), Gaps = 25/361 (6%)
Query: 82 TEAVVNKQLVGRSLFCLGLLIRYGNCLLAS-SGNKLVDVK-RSLNLFMKYLAGEDYALKA 139
T NK + RSLF +G L R+ + GN V++K + L L M + D ++
Sbjct: 48 TVLTANKPALLRSLFTVGALCRHFDFDQEDFKGNSKVNIKDKVLELLMYFTKHSDEEVQT 107
Query: 140 RSLQALGYVLIARPEYMLENDIGKILEGTLSSI-ADDRLKIQALQNMFEYLLDAESKMET 198
+++ LG+ I P M E ++ + LS LKIQ L+N+ YL + +++M+
Sbjct: 108 KAIIGLGFAFIQHPSLMFEQEVKSLYNSILSDKNCSVNLKIQVLKNLQTYLQEEDTRMQQ 167
Query: 199 EEVDGKVPGHSVRAGQSVPVAAGAGDTNICGGIIQLYWNNILGRCVDFNTQVRQSALKIV 258
+ D K + +++G + I+QLY +L + VR AL ++
Sbjct: 168 ADRDWKKVAKQEDLKEMGDISSGMSSS-----IMQLYLKQVLEAFFHTQSSVRHFALNVI 222
Query: 259 EVVLRQGLVHPITCVPYLIALETDPLESNSKMAHHLLMNMHEKYPAFFESCLGDGLQMSF 318
+ L QGL+HP+ CVPYLIA+ TDP S A L+ + +KY F G++MS+
Sbjct: 223 ALTLNQGLIHPVQCVPYLIAMGTDPEPSMRNKADQQLVEIDKKYAGFIHMKAVAGMKMSY 282
Query: 319 MFMQSIFVSPDENVNHKSQSKIAVSGKGKPEADSLAQSRVGVSRIYKLIRGNRISRNKFM 378
Q+I + +K V G E+ S S +Y +IRGNR R F+
Sbjct: 283 QVQQAI----------NTCTKDPVRGFRHDESSSAL-----CSHLYSMIRGNRQHRRAFL 327
Query: 379 SSIVRKFDNPKWNKFVIAFLTYCTEVLALLPFVAPDEPLYLIYTINRVVQVRAGPLEANF 438
S++ FD+ K + L Y + LA P+ +EPL++++ I+ + V L +F
Sbjct: 328 ISLLNLFDDTA--KTEVNMLLYIADNLACFPYQTQEEPLFIMHHIDITLSVSGSNLLQSF 385
Query: 439 K 439
K
Sbjct: 386 K 386
>gb|AAD26161.2| adherin Nipped-B [Drosophila melanogaster]
gi|50400943|sp|Q7PLI2|NIPB_DROME Nipped-B protein (SCC2
homolog)
Length = 2077
Score = 142 bits (357), Expect = 4e-32
Identities = 116/446 (26%), Positives = 216/446 (48%), Gaps = 36/446 (8%)
Query: 12 IIDSVLPLLRKLPPSIVEELEQDLKQMILRHSFLAVVHACIKCLCSMSELAGKGAAVIEH 71
I++ V+PL+ S + LE+ L +++ + A V +C+ CL ++ +I
Sbjct: 1414 ILEKVVPLVNNASESFLASLEEHLMLLVVSRN-QAEVTSCVSCLGALVNKITHNFKLIRD 1472
Query: 72 LIQVFFKCLD---TEAVVNKQLVG--------RSLFCLGLLIRYGNC---LLASSGNKLV 117
Q F++ LD ++ + V RSLF +G+L+RY + + N +
Sbjct: 1473 CFQKFYRVLDVSRSQVIQGNNSVDNIYTPSFRRSLFTIGILMRYFDFKSPIALGETNDGL 1532
Query: 118 DVKRSLNLF---MKYLAGEDYALKARSLQALGYVLIARPEYMLENDIGKILEGTLSSIAD 174
V ++F M + + ++ ++L +LG + Y+ +++ + LSSIA+
Sbjct: 1533 PVSICEDVFHCLMFFCRCTNQEIRKQALISLGSFCVLNDGYLTRSELKNLYCEILSSIAN 1592
Query: 175 DR-LKIQALQNMFEYLLDAESKMETEEVDGKVPGHSVRAGQSVPVAAGAGDTNICGGIIQ 233
D KI ++N++ YL ++E M +E + + + V++G + IIQ
Sbjct: 1593 DAGFKIICMRNIWIYLTESEMFMHNKEKEWEKQSKHEDLKEMNDVSSG-----MASRIIQ 1647
Query: 234 LYWNNILGRCVDFNTQVRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSKMAHH 293
LY IL ++ + VR A+K++++VLRQGLVHP+ VPYLI L TD ++ A
Sbjct: 1648 LYLEEILECFLNRDDTVRLWAVKVIQIVLRQGLVHPVRMVPYLICLSTDHRIESAHRADA 1707
Query: 294 LLMNMHEKYPAFFESCLGDGLQMSFMFMQSIFVSPDENVNHKSQSKIAVSGKGKPEADSL 353
LL ++ + Y F + GLQ+ F + + +N++ + +I + G D+
Sbjct: 1708 LLKDIDKTYSGFVNMKVQFGLQLCFKLQKIL------QINNRGKLEI-IRGYASRGPDNT 1760
Query: 354 AQSRVGVSRIYKLIRGNRISRNKFMSSIVRKFDNPKWNKFVIAFLTYCTEVLALLPFVAP 413
+ +Y L+R + R + ++ ++FD+ K + + + Y + LA P+V
Sbjct: 1761 TTALNDF--LYTLLRTTKPQRRALVQTVTKQFDDQKTS---LQQMLYIADNLAYFPYVVQ 1815
Query: 414 DEPLYLIYTINRVVQVRAGPLEANFK 439
DEPLYLI+ I+ ++ + L A FK
Sbjct: 1816 DEPLYLIHQIDLLISMAGTHLLATFK 1841
>gb|EAA46003.2| CG17704-PF.3 [Drosophila melanogaster] gi|51951003|gb|EAA46004.2|
CG17704-PE.3 [Drosophila melanogaster]
gi|62862616|ref|NP_001015455.1| CG17704-PF.3 [Drosophila
melanogaster] gi|62862614|ref|NP_001015454.1|
CG17704-PE.3 [Drosophila melanogaster]
Length = 2069
Score = 142 bits (357), Expect = 4e-32
Identities = 116/446 (26%), Positives = 216/446 (48%), Gaps = 36/446 (8%)
Query: 12 IIDSVLPLLRKLPPSIVEELEQDLKQMILRHSFLAVVHACIKCLCSMSELAGKGAAVIEH 71
I++ V+PL+ S + LE+ L +++ + A V +C+ CL ++ +I
Sbjct: 1406 ILEKVVPLVNNASESFLASLEEHLMLLVVSRN-QAEVTSCVSCLGALVNKITHNFKLIRD 1464
Query: 72 LIQVFFKCLD---TEAVVNKQLVG--------RSLFCLGLLIRYGNC---LLASSGNKLV 117
Q F++ LD ++ + V RSLF +G+L+RY + + N +
Sbjct: 1465 CFQKFYRVLDVSRSQVIQGNNSVDNIYTPSFRRSLFTIGILMRYFDFKSPIALGETNDGL 1524
Query: 118 DVKRSLNLF---MKYLAGEDYALKARSLQALGYVLIARPEYMLENDIGKILEGTLSSIAD 174
V ++F M + + ++ ++L +LG + Y+ +++ + LSSIA+
Sbjct: 1525 PVSICEDVFHCLMFFCRCTNQEIRKQALISLGSFCVLNDGYLTRSELKNLYCEILSSIAN 1584
Query: 175 DR-LKIQALQNMFEYLLDAESKMETEEVDGKVPGHSVRAGQSVPVAAGAGDTNICGGIIQ 233
D KI ++N++ YL ++E M +E + + + V++G + IIQ
Sbjct: 1585 DAGFKIICMRNIWIYLTESEMFMHNKEKEWEKQSKHEDLKEMNDVSSG-----MASRIIQ 1639
Query: 234 LYWNNILGRCVDFNTQVRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSKMAHH 293
LY IL ++ + VR A+K++++VLRQGLVHP+ VPYLI L TD ++ A
Sbjct: 1640 LYLEEILECFLNRDDTVRLWAVKVIQIVLRQGLVHPVRMVPYLICLSTDHRIESAHRADA 1699
Query: 294 LLMNMHEKYPAFFESCLGDGLQMSFMFMQSIFVSPDENVNHKSQSKIAVSGKGKPEADSL 353
LL ++ + Y F + GLQ+ F + + +N++ + +I + G D+
Sbjct: 1700 LLKDIDKTYSGFVNMKVQFGLQLCFKLQKIL------QINNRGKLEI-IRGYASRGPDNT 1752
Query: 354 AQSRVGVSRIYKLIRGNRISRNKFMSSIVRKFDNPKWNKFVIAFLTYCTEVLALLPFVAP 413
+ +Y L+R + R + ++ ++FD+ K + + + Y + LA P+V
Sbjct: 1753 TTALNDF--LYTLLRTTKPQRRALVQTVTKQFDDQKTS---LQQMLYIADNLAYFPYVVQ 1807
Query: 414 DEPLYLIYTINRVVQVRAGPLEANFK 439
DEPLYLI+ I+ ++ + L A FK
Sbjct: 1808 DEPLYLIHQIDLLISMAGTHLLATFK 1833
>ref|XP_320088.2| ENSANGP00000010656 [Anopheles gambiae str. PEST]
gi|55235638|gb|EAA15100.2| ENSANGP00000010656 [Anopheles
gambiae str. PEST]
Length = 1518
Score = 141 bits (355), Expect = 6e-32
Identities = 121/478 (25%), Positives = 223/478 (46%), Gaps = 51/478 (10%)
Query: 1 MVTQLLESIIFIIDSVLPLLRKLPPSIVEELEQDLKQMILRHSFLAVVHACIKCLCSMSE 60
++++ + SI I++ V+PL+ + +LE L +I+ S +V +C+ CL ++
Sbjct: 938 IISKFISSIAEILEQVVPLMDHPSEVFLADLESHLMMLIVTQS-RTIVLSCVSCLSTVVN 996
Query: 61 LAGKGAAVIEHLI-QVFFK---CLDTEAVVNKQL---------VGRSLFCLGLLIRY--- 104
K +I ++++K C+ + V + + RS+F +GL++RY
Sbjct: 997 KITKNYKLIRDCFSKLYYKGLVCIKDKLVSDPSIPIEQYFRPQFRRSIFTVGLIMRYFDF 1056
Query: 105 ------GNCLLASSGNKLVDVKRSLNLFMKYLAGEDYALKARSLQALGYVLIARPEYMLE 158
G+ L A N DV +L F L+ + + +L ++G + EY+++
Sbjct: 1057 QQPEVYGSTLPA---NICEDVFATLAFF---LSCDHSEICKEALTSMGNFCVKNYEYLMK 1110
Query: 159 NDIGKILEGTLSS--IADDRLKIQALQNMFEYLLDAESKMETEEVDGKVPGHSVRAGQSV 216
++ L+ + D +KI L+N+ YL + E++M ++ + + +
Sbjct: 1111 VELRDYYNYLLTQDKVLTD-MKITVLKNILMYLTEEENQMVRKDKEWSKQSKTEDLKEMG 1169
Query: 217 PVAAGAGDTNICGGIIQLYWNNILGRCVDFNTQVRQSALKIVEVVLRQGLVHPITCVPYL 276
V++G + +IQ+Y IL + + VR A++++EVVLRQGLVHP+ VPYL
Sbjct: 1170 DVSSG-----MASRVIQIYLKEILRSFLHRDYGVRSWAMRVIEVVLRQGLVHPVQIVPYL 1224
Query: 277 IALETDPLESNSKMAHHLLMNMHEKYPAFFESCLGDGLQMSFMFMQSIFVSPDENVNHKS 336
I L TDP + + A L + ++YP F G+Q+S+ E + +
Sbjct: 1225 ICLSTDPEKEVAHSADRHLQEIDKQYPGFVNMKSNAGMQLSYEL--------QELLQRRD 1276
Query: 337 QSKIAVSGKGKPEADSLAQSRVGVSRIYKLIRGNRISRNKFMSSIVRKFDNPKWNKFVIA 396
+S + +G D +Y L+RG + R + SI ++FD+ K +
Sbjct: 1277 ESSLV---RGYRIKDPQEPPSAMNGFLYTLLRGTKPQRRALIHSITKQFDD---GKISLR 1330
Query: 397 FLTYCTEVLALLPFVAPDEPLYLIYTINRVVQVRAGPLEANFKAWSSSLLQSEGQGTP 454
+ Y + LA P+V DEPL++I+ I+ ++ V L A F+ L +EG P
Sbjct: 1331 QMLYLADNLAYFPYVVQDEPLFIIHHIDVLISVTGTNLLATFREGLKPLPGTEGNADP 1388
>gb|AAU95610.1| sister chromatid cohesion establishment factor [Xenopus laevis]
Length = 537
Score = 131 bits (330), Expect = 5e-29
Identities = 96/356 (26%), Positives = 167/356 (45%), Gaps = 26/356 (7%)
Query: 113 GNKLVDVK-RSLNLFMKYLAGEDYALKARSLQALGYVLIARPEYMLENDIGKILEGTLSS 171
G+ V++K + L L + + D ++ +++ LG+ I P M E ++ + LS
Sbjct: 18 GSSKVNIKDKVLELLLYFTKHSDEEVQTKAIIGLGFSFIQHPVLMFEVEVKNLYNIILSD 77
Query: 172 I-ADDRLKIQALQNMFEYLLDAESKMETEEVDGKVPGHSVRAGQSVPVAAGAGDTNICGG 230
LKIQ L+N+ YL + +++M+ + D K + +++G +
Sbjct: 78 KNCSVNLKIQVLKNLQTYLQEEDTRMQQADRDWKKMSKQEDLKEMGDISSGMSSS----- 132
Query: 231 IIQLYWNNILGRCVDFNTQVRQSALKIVEVVLRQGLVHPITCVPYLIALETDPLESNSKM 290
I+QLY +L + VR AL ++ + L QGL+HP+ CVPYLIA+ TDP S
Sbjct: 133 IMQLYLKQVLESFFSTQSSVRHFALNVIALTLNQGLIHPVQCVPYLIAMGTDPEPSMRNK 192
Query: 291 AHHLLMNMHEKYPAFFESCLGDGLQMSFMFMQSIFVSPDENVNHKSQSKIAVSGKGKPEA 350
+ L+ + +KY F G++MS+ Q+I S + V G + E+
Sbjct: 193 SDQQLIEIDKKYTGFIHMKAVAGIKMSYQVQQAI----------NSNANQIVRGFRQDES 242
Query: 351 DSLAQSRVGVSRIYKLIRGNRISRNKFMSSIVRKFDNPKWNKFVIAFLTYCTEVLALLPF 410
+S S +Y +IR NR R F+ S++ FD+ K + L Y + LA P+
Sbjct: 243 NSAL-----CSHLYSMIRSNRQHRRAFLISLLNLFDDAA--KTEVNMLLYIADNLACFPY 295
Query: 411 VAPDEPLYLIYTINRVVQVRAGPLEANFKAWSSSLLQSEGQGTPHGNGMYQRATDE 466
+ +EPL++++ I+ + V L +FK S ++ + P N +DE
Sbjct: 296 QSQEEPLFIMHHIDITLSVSGSNLLQSFK--ESMVMAKQKVKAPSSNEDSSSESDE 349
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.321 0.137 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 890,332,382
Number of Sequences: 2540612
Number of extensions: 37129810
Number of successful extensions: 81121
Number of sequences better than 10.0: 52
Number of HSP's better than 10.0 without gapping: 43
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 80935
Number of HSP's gapped (non-prelim): 80
length of query: 542
length of database: 863,360,394
effective HSP length: 133
effective length of query: 409
effective length of database: 525,458,998
effective search space: 214912730182
effective search space used: 214912730182
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 78 (34.7 bits)
Medicago: description of AC149305.4


