

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149305.1 + phase: 0 /pseudo
(146 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
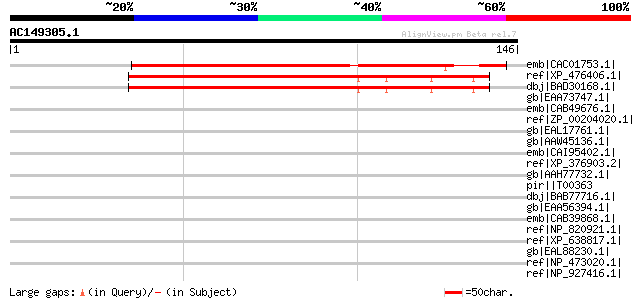
Sequences producing significant alignments: (bits) Value
emb|CAC01753.1| putative protein [Arabidopsis thaliana] gi|15242... 107 5e-23
ref|XP_476406.1| Nipped-B gene product-like protein [Oryza sativ... 96 1e-19
dbj|BAD30168.1| putative IDN3 protein isoform A [Oryza sativa (j... 96 1e-19
gb|EAA73747.1| hypothetical protein FG06256.1 [Gibberella zeae P... 35 0.38
emb|CAB49676.1| Hypothetical protein [Pyrococcus abyssi GE5] gi|... 35 0.49
ref|ZP_00204020.1| COG0055: F0F1-type ATP synthase, beta subunit... 34 0.64
gb|EAL17761.1| hypothetical protein CNBL2740 [Cryptococcus neofo... 34 0.84
gb|AAW45136.1| conserved hypothetical protein [Cryptococcus neof... 34 0.84
emb|CAI95402.1| KIAA0674 [Homo sapiens] gi|55859551|emb|CAI10964... 33 1.1
ref|XP_376903.2| PREDICTED: KIAA0674 [Homo sapiens] 33 1.1
gb|AAH77732.1| KIAA0674 protein [Homo sapiens] 33 1.1
pir||T00363 hypothetical protein KIAA0674 - human (fragment) gi|... 33 1.1
dbj|BAB77716.1| serine/threonine kinase [Nostoc sp. PCC 7120] gi... 33 1.1
gb|EAA56394.1| hypothetical protein MG06365.4 [Magnaporthe grise... 33 1.1
emb|CAB39868.1| putative small secreted hydrophilic protein [Str... 33 1.4
ref|NP_820921.1| ATP synthase F1, beta subunit [Coxiella burneti... 33 1.4
ref|XP_638817.1| hypothetical protein DDB0185762 [Dictyostelium ... 33 1.4
gb|EAL88230.1| hypothetical protein Afu1g05190 [Aspergillus fumi... 33 1.9
ref|NP_473020.1| hypothetical protein PFB0460c [Plasmodium falci... 33 1.9
ref|NP_927416.1| ATP synthase beta chain [Photorhabdus luminesce... 33 1.9
>emb|CAC01753.1| putative protein [Arabidopsis thaliana] gi|15242325|ref|NP_197058.1|
expressed protein [Arabidopsis thaliana]
gi|11358123|pir||T51532 hypothetical protein T20K14_150 -
Arabidopsis thaliana
Length = 1755
Score = 107 bits (268), Expect = 5e-23
Identities = 54/112 (48%), Positives = 78/112 (69%), Gaps = 13/112 (11%)
Query: 36 SEPPKPGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNALKEDTVDYSLYTANIKR 95
+EP KPGD SRQS+ F++ E++ LP++ Q+L+QRYQEFKNA++EDTVD+++Y+ N+KR
Sbjct: 1642 TEPLKPGDPLSRQSVAFDLSETRTDLPSTYQDLVQRYQEFKNAMREDTVDFTIYSTNVKR 1701
Query: 96 KRPTQTPRKVRKSGPPMVGGDNDEDDDED----WAGGSRNISFSGGRRSSLR 143
KRP TPRK +S V + D+DDD++ W GG GGR ++ R
Sbjct: 1702 KRP--TPRKTSRSAKKTVAYNEDDDDDDNDDRGWHGG-------GGRGAARR 1744
>ref|XP_476406.1| Nipped-B gene product-like protein [Oryza sativa (japonica
cultivar-group)]
Length = 1755
Score = 96.3 bits (238), Expect = 1e-19
Identities = 51/114 (44%), Positives = 71/114 (61%), Gaps = 10/114 (8%)
Query: 35 VSEPPKPGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNALKEDTVDYSLYTANIK 94
+ +PPK G+ S+Q+IP NI + SLP+ PQ+ + YQ+FK L+EDTVDY +YT + +
Sbjct: 1634 LKDPPKSGETISKQNIPLNISNTNTSLPSCPQDAARVYQDFKTVLREDTVDYGMYTVSAQ 1693
Query: 95 RKRPT-QTPRKVRK-----SGPPMVGGDNDED-DDEDWAGGSRNI---SFSGGR 138
+KRPT ++ +VR+ G GG DED DDEDW GG + S GGR
Sbjct: 1694 KKRPTPRSSSRVRRPAAVTRGRGGGGGGGDEDTDDEDWTGGGARVLDFSAQGGR 1747
>dbj|BAD30168.1| putative IDN3 protein isoform A [Oryza sativa (japonica
cultivar-group)]
Length = 764
Score = 96.3 bits (238), Expect = 1e-19
Identities = 51/114 (44%), Positives = 71/114 (61%), Gaps = 10/114 (8%)
Query: 35 VSEPPKPGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNALKEDTVDYSLYTANIK 94
+ +PPK G+ S+Q+IP NI + SLP+ PQ+ + YQ+FK L+EDTVDY +YT + +
Sbjct: 643 LKDPPKSGETISKQNIPLNISNTNTSLPSCPQDAARVYQDFKTVLREDTVDYGMYTVSAQ 702
Query: 95 RKRPT-QTPRKVRK-----SGPPMVGGDNDED-DDEDWAGGSRNI---SFSGGR 138
+KRPT ++ +VR+ G GG DED DDEDW GG + S GGR
Sbjct: 703 KKRPTPRSSSRVRRPAAVTRGRGGGGGGGDEDTDDEDWTGGGARVLDFSAQGGR 756
>gb|EAA73747.1| hypothetical protein FG06256.1 [Gibberella zeae PH-1]
gi|46123757|ref|XP_386432.1| hypothetical protein
FG06256.1 [Gibberella zeae PH-1]
Length = 460
Score = 35.0 bits (79), Expect = 0.38
Identities = 22/73 (30%), Positives = 33/73 (45%), Gaps = 11/73 (15%)
Query: 82 DTVDYSLYTANIKRKRPTQTPRKVRKSGPPMVGGDNDED-----DDEDWAGGSR------ 130
D+ + T N KRK+ K P GD+D+D DD+D A G
Sbjct: 75 DSENEDFNTRNEKRKKEEADAAKSGSKAPGTTAGDDDDDDMFAVDDDDAANGDNNVPEGD 134
Query: 131 NISFSGGRRSSLR 143
N+ SGG+++ +R
Sbjct: 135 NVEESGGKKNKVR 147
>emb|CAB49676.1| Hypothetical protein [Pyrococcus abyssi GE5]
gi|14520970|ref|NP_126445.1| hypothetical protein
PAB1857 [Pyrococcus abyssi GE5] gi|7518261|pir||C75120
hypothetical protein PAB1857 - Pyrococcus abyssi (strain
Orsay)
Length = 602
Score = 34.7 bits (78), Expect = 0.49
Identities = 21/76 (27%), Positives = 36/76 (46%), Gaps = 9/76 (11%)
Query: 53 NIGESQFSLPTSPQELIQRYQEFKNALKEDTVDYSLYTANIKRKRPTQTPRKVRKSGPPM 112
N+ E F T EL++ +E N +E V+ + K P + PR + PP
Sbjct: 497 NLLEEYFEQGTFDVELLEELEEKDNLFQEVPVEAYI------SKEPPEVPRYIE---PPE 547
Query: 113 VGGDNDEDDDEDWAGG 128
V +N+E++ +W+ G
Sbjct: 548 VPDNNEEEEKGEWSSG 563
>ref|ZP_00204020.1| COG0055: F0F1-type ATP synthase, beta subunit [Psychrobacter sp.
273-4]
Length = 477
Score = 34.3 bits (77), Expect = 0.64
Identities = 16/39 (41%), Positives = 24/39 (61%)
Query: 41 PGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNAL 79
P D SRQ P IGE +++ QE++QRY+E K+ +
Sbjct: 349 PLDSTSRQLDPLVIGEEHYNVARGVQEVLQRYKELKDII 387
>gb|EAL17761.1| hypothetical protein CNBL2740 [Cryptococcus neoformans var.
neoformans B-3501A]
Length = 796
Score = 33.9 bits (76), Expect = 0.84
Identities = 28/85 (32%), Positives = 36/85 (41%), Gaps = 17/85 (20%)
Query: 74 EFKNALKEDTVDYSLYTANIKRKRPTQTP----------RKVRKSGPPMVGGDNDEDDDE 123
+F A E V IK+KR Q+P V S P GD+D+DDDE
Sbjct: 575 DFDEADDEGVVRQEEEGRAIKKKRQRQSPPVWVTRTGATTGVGSSSSPFSIGDDDDDDDE 634
Query: 124 DWAG-------GSRNISFSGGRRSS 141
D G G R++SF+ SS
Sbjct: 635 DGEGVKNRETQGDRSMSFNHPDNSS 659
>gb|AAW45136.1| conserved hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21] gi|58270574|ref|XP_572443.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21]
Length = 796
Score = 33.9 bits (76), Expect = 0.84
Identities = 28/85 (32%), Positives = 36/85 (41%), Gaps = 17/85 (20%)
Query: 74 EFKNALKEDTVDYSLYTANIKRKRPTQTP----------RKVRKSGPPMVGGDNDEDDDE 123
+F A E V IK+KR Q+P V S P GD+D+DDDE
Sbjct: 575 DFDEADDEGVVRQEEEGRAIKKKRQRQSPPVWVTRTGATTGVGSSSSPFSIGDDDDDDDE 634
Query: 124 DWAG-------GSRNISFSGGRRSS 141
D G G R++SF+ SS
Sbjct: 635 DGEGVKNRETQGDRSMSFNHPDNSS 659
>emb|CAI95402.1| KIAA0674 [Homo sapiens] gi|55859551|emb|CAI10964.1| KIAA0674 [Homo
sapiens]
Length = 1244
Score = 33.5 bits (75), Expect = 1.1
Identities = 26/101 (25%), Positives = 40/101 (38%), Gaps = 10/101 (9%)
Query: 37 EPP--KPGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNALKEDTVDYSLYTANIK 94
+PP K DV S + ++ + L + +++L D D A +K
Sbjct: 1144 QPPQLKKDDVTSSTGPHKELSSTEAGSTVAGAALRPSHHSQRSSLSGDEEDELFKGATLK 1203
Query: 95 RKRPTQTPRKVRKSGPPMVG--------GDNDEDDDEDWAG 127
RP P + + M G GD+D+DDD DW G
Sbjct: 1204 ALRPKAQPEEEDEDEVSMKGRPPPTPLFGDDDDDDDIDWLG 1244
>ref|XP_376903.2| PREDICTED: KIAA0674 [Homo sapiens]
Length = 1337
Score = 33.5 bits (75), Expect = 1.1
Identities = 26/101 (25%), Positives = 40/101 (38%), Gaps = 10/101 (9%)
Query: 37 EPP--KPGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNALKEDTVDYSLYTANIK 94
+PP K DV S + ++ + L + +++L D D A +K
Sbjct: 1237 QPPQLKKDDVTSSTGPHKELSSTEAGSTVAGAALRPSHHSQRSSLSGDEEDELFKGATLK 1296
Query: 95 RKRPTQTPRKVRKSGPPMVG--------GDNDEDDDEDWAG 127
RP P + + M G GD+D+DDD DW G
Sbjct: 1297 ALRPKAQPEEEDEDEVSMKGRPPPTPLFGDDDDDDDIDWLG 1337
>gb|AAH77732.1| KIAA0674 protein [Homo sapiens]
Length = 662
Score = 33.5 bits (75), Expect = 1.1
Identities = 26/101 (25%), Positives = 40/101 (38%), Gaps = 10/101 (9%)
Query: 37 EPP--KPGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNALKEDTVDYSLYTANIK 94
+PP K DV S + ++ + L + +++L D D A +K
Sbjct: 562 QPPQLKKDDVTSSTGPHKELSSTEAGSTVAGAALRPSHHSQRSSLSGDEEDELFKGATLK 621
Query: 95 RKRPTQTPRKVRKSGPPMVG--------GDNDEDDDEDWAG 127
RP P + + M G GD+D+DDD DW G
Sbjct: 622 ALRPKAQPEEEDEDEVSMKGRPPPTPLFGDDDDDDDIDWLG 662
>pir||T00363 hypothetical protein KIAA0674 - human (fragment)
gi|3327162|dbj|BAA31649.1| KIAA0674 protein [Homo
sapiens]
Length = 1234
Score = 33.5 bits (75), Expect = 1.1
Identities = 26/101 (25%), Positives = 40/101 (38%), Gaps = 10/101 (9%)
Query: 37 EPP--KPGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNALKEDTVDYSLYTANIK 94
+PP K DV S + ++ + L + +++L D D A +K
Sbjct: 1134 QPPQLKKDDVTSSTGPHKELSSTEAGSTVAGAALRPSHHSQRSSLSGDEEDELFKGATLK 1193
Query: 95 RKRPTQTPRKVRKSGPPMVG--------GDNDEDDDEDWAG 127
RP P + + M G GD+D+DDD DW G
Sbjct: 1194 ALRPKAQPEEEDEDEVSMKGRPPPTPLFGDDDDDDDIDWLG 1234
>dbj|BAB77716.1| serine/threonine kinase [Nostoc sp. PCC 7120]
gi|25534252|pir||AH1830 serine/threonine kinase
[imported] - Nostoc sp. (strain PCC 7120)
gi|17227688|ref|NP_484236.1| serine/threonine kinase
[Nostoc sp. PCC 7120]
Length = 553
Score = 33.5 bits (75), Expect = 1.1
Identities = 14/43 (32%), Positives = 23/43 (52%)
Query: 62 PTSPQELIQRYQEFKNALKEDTVDYSLYTANIKRKRPTQTPRK 104
PT+ Q ++QR ++ K + D + YS +K P Q+P K
Sbjct: 275 PTNTQGILQRLRDIKQQVNRDEIPYSEQQTQLKSHLPLQSPTK 317
>gb|EAA56394.1| hypothetical protein MG06365.4 [Magnaporthe grisea 70-15]
gi|39976925|ref|XP_369850.1| hypothetical protein
MG06365.4 [Magnaporthe grisea 70-15]
Length = 423
Score = 33.5 bits (75), Expect = 1.1
Identities = 15/40 (37%), Positives = 23/40 (57%), Gaps = 2/40 (5%)
Query: 89 YTANIKRKRPTQTPRKVRKSGPPMVGGDNDEDDDEDWAGG 128
Y N+ +K +Q P ++ P+ G D D DDD++ AGG
Sbjct: 8 YGLNLSKKSGSQKPNPAKRR--PVFGDDGDSDDDDNRAGG 45
>emb|CAB39868.1| putative small secreted hydrophilic protein [Streptomyces
coelicolor A3(2)] gi|7481422|pir||T35523 probable small
secreted hydrophilic protein - Streptomyces coelicolor
gi|21220618|ref|NP_626397.1| putative small secreted
hydrophilic protein [Streptomyces coelicolor A3(2)]
Length = 107
Score = 33.1 bits (74), Expect = 1.4
Identities = 22/78 (28%), Positives = 32/78 (40%), Gaps = 8/78 (10%)
Query: 50 IPFNIGESQFSLPTSPQELIQRYQEFKNALKEDTVDYSLYTANIKRKRPTQTPRKVRKSG 109
IP I + F+L SPQE + ++ T P TP +
Sbjct: 16 IPLGIAATSFALTDSPQE--------PKVPAKVELESGSPTPAPTSSSPEPTPSDQVVTR 67
Query: 110 PPMVGGDNDEDDDEDWAG 127
PP+ G +D+DDD+D G
Sbjct: 68 PPVTDGPSDDDDDDDRGG 85
>ref|NP_820921.1| ATP synthase F1, beta subunit [Coxiella burnetii RSA 493]
gi|29542501|gb|AAO91435.1| ATP synthase F1, beta subunit
[Coxiella burnetii RSA 493]
Length = 461
Score = 33.1 bits (74), Expect = 1.4
Identities = 16/39 (41%), Positives = 22/39 (56%)
Query: 41 PGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNAL 79
P D SRQ P IGE + + QE +QRY+E K+ +
Sbjct: 338 PLDSTSRQLDPLIIGEEHYRVARGVQETLQRYEELKDII 376
>ref|XP_638817.1| hypothetical protein DDB0185762 [Dictyostelium discoideum]
gi|60467434|gb|EAL65457.1| hypothetical protein
DDB0185762 [Dictyostelium discoideum]
Length = 1051
Score = 33.1 bits (74), Expect = 1.4
Identities = 13/31 (41%), Positives = 21/31 (66%)
Query: 116 DNDEDDDEDWAGGSRNISFSGGRRSSLRNSR 146
D+D+DDD+D GG+RNIS + + +S+
Sbjct: 332 DDDDDDDDDGGGGTRNISSLDSNTTEINSSQ 362
>gb|EAL88230.1| hypothetical protein Afu1g05190 [Aspergillus fumigatus Af293]
Length = 435
Score = 32.7 bits (73), Expect = 1.9
Identities = 18/53 (33%), Positives = 27/53 (49%), Gaps = 1/53 (1%)
Query: 60 SLPTSPQELIQRYQEFKNA-LKEDTVDYSLYTANIKRKRPTQTPRKVRKSGPP 111
++P SP L R+ +++ L D + Y N + PTQ+PR R S PP
Sbjct: 3 AIPPSPLRLSHRHSHSQSSNLDFDPISPGSYAPNGFSQSPTQSPRTPRSSAPP 55
>ref|NP_473020.1| hypothetical protein PFB0460c [Plasmodium falciparum 3D7]
gi|3845190|gb|AAC71881.1| hypothetical protein PFB0460c
[Plasmodium falciparum 3D7] gi|7494294|pir||D71614
hypothetical protein PFB0460c - malaria parasite
(Plasmodium falciparum)
Length = 2573
Score = 32.7 bits (73), Expect = 1.9
Identities = 16/59 (27%), Positives = 32/59 (54%), Gaps = 1/59 (1%)
Query: 67 ELIQRYQEFKNALKEDTVDY-SLYTANIKRKRPTQTPRKVRKSGPPMVGGDNDEDDDED 124
+L + Y+ K L + + D+ +Y ++ K+K+ + V K G P +G ND++ + D
Sbjct: 2286 KLFKEYKNIKEILNDQSSDFLDMYKSDKKKKKKKKELDDVEKEGQPKMGVGNDDNINGD 2344
>ref|NP_927416.1| ATP synthase beta chain [Photorhabdus luminescens subsp. laumondii
TTO1] gi|36783495|emb|CAE12335.1| ATP synthase beta
chain [Photorhabdus luminescens subsp. laumondii TTO1]
Length = 460
Score = 32.7 bits (73), Expect = 1.9
Identities = 14/39 (35%), Positives = 23/39 (58%)
Query: 41 PGDVFSRQSIPFNIGESQFSLPTSPQELIQRYQEFKNAL 79
P D SRQ P +G+ +++ Q ++QRYQE K+ +
Sbjct: 337 PLDSTSRQLDPLVVGQEHYNVARGVQSILQRYQELKDII 375
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.329 0.143 0.449
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 264,141,888
Number of Sequences: 2540612
Number of extensions: 10993251
Number of successful extensions: 61069
Number of sequences better than 10.0: 122
Number of HSP's better than 10.0 without gapping: 58
Number of HSP's successfully gapped in prelim test: 67
Number of HSP's that attempted gapping in prelim test: 60911
Number of HSP's gapped (non-prelim): 161
length of query: 146
length of database: 863,360,394
effective HSP length: 122
effective length of query: 24
effective length of database: 553,405,730
effective search space: 13281737520
effective search space used: 13281737520
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 67 (30.4 bits)
Medicago: description of AC149305.1


