

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149293.2 - phase: 2
(212 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
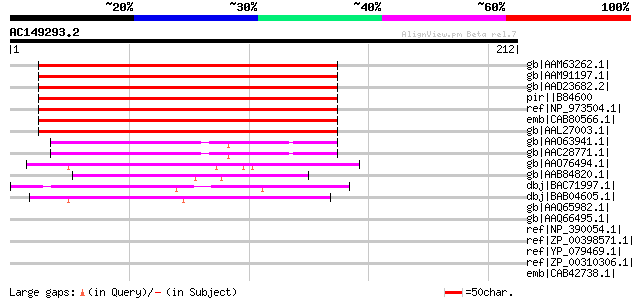
Score E
Sequences producing significant alignments: (bits) Value
gb|AAM63262.1| enhanced disease susceptibility 5 [Arabidopsis th... 204 1e-51
gb|AAM91197.1| unknown protein [Arabidopsis thaliana] gi|1706500... 204 1e-51
gb|AAD23682.2| expressed protein [Arabidopsis thaliana] gi|18399... 204 1e-51
pir||B84600 hypothetical protein At2g21340 [imported] - Arabidop... 204 1e-51
ref|NP_973504.1| enhanced disease susceptibility protein, putati... 204 1e-51
emb|CAB80566.1| putative protein [Arabidopsis thaliana] gi|45393... 179 6e-44
gb|AAL27003.1| enhanced disease susceptibility 5 [Arabidopsis th... 179 6e-44
gb|AAO63941.1| unknown protein [Arabidopsis thaliana] gi|2839322... 54 2e-06
gb|AAC28771.1| hypothetical protein [Arabidopsis thaliana] gi|74... 54 2e-06
gb|AAO76494.1| putative Na+-driven multidrug efflux pump [Bacter... 49 9e-05
gb|AAB84820.1| conserved protein [Methanothermobacter thermautot... 48 2e-04
dbj|BAC71997.1| putative DNA-damage-inducible protein F [Strepto... 47 3e-04
dbj|BAB04605.1| BH0886 [Bacillus halodurans C-125] gi|15613449|r... 47 4e-04
gb|AAQ65982.1| MATE efflux family protein [Porphyromonas gingiva... 45 0.002
gb|AAQ66495.1| MATE efflux family protein [Porphyromonas gingiva... 45 0.002
ref|NP_390054.1| hypothetical protein BSU21710 [Bacillus subtili... 44 0.003
ref|ZP_00398571.1| Multi antimicrobial extrusion protein MatE [A... 44 0.004
ref|YP_079469.1| putative Multi antimicrobial extrusion protein ... 44 0.004
ref|ZP_00310306.1| COG0534: Na+-driven multidrug efflux pump [Cy... 44 0.004
emb|CAB42738.1| putative membrane protein [Streptomyces coelicol... 43 0.005
>gb|AAM63262.1| enhanced disease susceptibility 5 [Arabidopsis thaliana]
Length = 555
Score = 204 bits (520), Expect = 1e-51
Identities = 99/125 (79%), Positives = 110/125 (87%)
Query: 13 LCGWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAA 72
L GWVAQSASLGMKDSWGPLKALA AS INGVGD+VLCT+LGYGIAGAAWATM SQVVAA
Sbjct: 248 LIGWVAQSASLGMKDSWGPLKALAVASAINGVGDVVLCTFLGYGIAGAAWATMVSQVVAA 307
Query: 73 YMMMRTLNMKGYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFATSMGTHT 132
YMMM LN KGY+AF+ +PS E +TI GLAAPVF+TMMSKV FY+LL+YFATSMGT+
Sbjct: 308 YMMMDALNKKGYSAFSFCVPSPSELLTIFGLAAPVFITMMSKVLFYTLLVYFATSMGTNI 367
Query: 133 MAAHQ 137
+AAHQ
Sbjct: 368 IAAHQ 372
>gb|AAM91197.1| unknown protein [Arabidopsis thaliana] gi|17065002|gb|AAL32655.1|
Unknown protein [Arabidopsis thaliana]
Length = 559
Score = 204 bits (520), Expect = 1e-51
Identities = 99/125 (79%), Positives = 110/125 (87%)
Query: 13 LCGWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAA 72
L GWVAQSASLGMKDSWGPLKALA AS INGVGD+VLCT+LGYGIAGAAWATM SQVVAA
Sbjct: 252 LIGWVAQSASLGMKDSWGPLKALAVASAINGVGDVVLCTFLGYGIAGAAWATMVSQVVAA 311
Query: 73 YMMMRTLNMKGYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFATSMGTHT 132
YMMM LN KGY+AF+ +PS E +TI GLAAPVF+TMMSKV FY+LL+YFATSMGT+
Sbjct: 312 YMMMDALNKKGYSAFSFCVPSPSELLTIFGLAAPVFITMMSKVLFYTLLVYFATSMGTNI 371
Query: 133 MAAHQ 137
+AAHQ
Sbjct: 372 IAAHQ 376
>gb|AAD23682.2| expressed protein [Arabidopsis thaliana]
gi|18399664|ref|NP_565509.1| enhanced disease
susceptibility protein, putative / salicylic acid
induction deficient protein, putative [Arabidopsis
thaliana]
Length = 555
Score = 204 bits (520), Expect = 1e-51
Identities = 99/125 (79%), Positives = 110/125 (87%)
Query: 13 LCGWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAA 72
L GWVAQSASLGMKDSWGPLKALA AS INGVGD+VLCT+LGYGIAGAAWATM SQVVAA
Sbjct: 248 LIGWVAQSASLGMKDSWGPLKALAVASAINGVGDVVLCTFLGYGIAGAAWATMVSQVVAA 307
Query: 73 YMMMRTLNMKGYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFATSMGTHT 132
YMMM LN KGY+AF+ +PS E +TI GLAAPVF+TMMSKV FY+LL+YFATSMGT+
Sbjct: 308 YMMMDALNKKGYSAFSFCVPSPSELLTIFGLAAPVFITMMSKVLFYTLLVYFATSMGTNI 367
Query: 133 MAAHQ 137
+AAHQ
Sbjct: 368 IAAHQ 372
>pir||B84600 hypothetical protein At2g21340 [imported] - Arabidopsis thaliana
Length = 414
Score = 204 bits (520), Expect = 1e-51
Identities = 99/125 (79%), Positives = 110/125 (87%)
Query: 13 LCGWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAA 72
L GWVAQSASLGMKDSWGPLKALA AS INGVGD+VLCT+LGYGIAGAAWATM SQVVAA
Sbjct: 193 LIGWVAQSASLGMKDSWGPLKALAVASAINGVGDVVLCTFLGYGIAGAAWATMVSQVVAA 252
Query: 73 YMMMRTLNMKGYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFATSMGTHT 132
YMMM LN KGY+AF+ +PS E +TI GLAAPVF+TMMSKV FY+LL+YFATSMGT+
Sbjct: 253 YMMMDALNKKGYSAFSFCVPSPSELLTIFGLAAPVFITMMSKVLFYTLLVYFATSMGTNI 312
Query: 133 MAAHQ 137
+AAHQ
Sbjct: 313 IAAHQ 317
>ref|NP_973504.1| enhanced disease susceptibility protein, putative / salicylic acid
induction deficient protein, putative [Arabidopsis
thaliana]
Length = 552
Score = 204 bits (520), Expect = 1e-51
Identities = 99/125 (79%), Positives = 110/125 (87%)
Query: 13 LCGWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAA 72
L GWVAQSASLGMKDSWGPLKALA AS INGVGD+VLCT+LGYGIAGAAWATM SQVVAA
Sbjct: 245 LIGWVAQSASLGMKDSWGPLKALAVASAINGVGDVVLCTFLGYGIAGAAWATMVSQVVAA 304
Query: 73 YMMMRTLNMKGYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFATSMGTHT 132
YMMM LN KGY+AF+ +PS E +TI GLAAPVF+TMMSKV FY+LL+YFATSMGT+
Sbjct: 305 YMMMDALNKKGYSAFSFCVPSPSELLTIFGLAAPVFITMMSKVLFYTLLVYFATSMGTNI 364
Query: 133 MAAHQ 137
+AAHQ
Sbjct: 365 IAAHQ 369
>emb|CAB80566.1| putative protein [Arabidopsis thaliana] gi|4539322|emb|CAB38823.1|
putative protein [Arabidopsis thaliana]
gi|7485793|pir||T06063 hypothetical protein F19H22.130 -
Arabidopsis thaliana
Length = 484
Score = 179 bits (453), Expect = 6e-44
Identities = 86/125 (68%), Positives = 106/125 (84%)
Query: 13 LCGWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAA 72
L G VAQSASLGMK+SWGPLKALAAA++ING+GD +LC +LG GIAGAAWAT ASQ+V+A
Sbjct: 235 LVGLVAQSASLGMKNSWGPLKALAAATIINGLGDTILCLFLGQGIAGAAWATTASQIVSA 294
Query: 73 YMMMRTLNMKGYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFATSMGTHT 132
YMMM +LN +GYNA++ +IPS +E I LAAPVF+++ SK+AFYS +IY ATSMGTH
Sbjct: 295 YMMMDSLNKEGYNAYSFAIPSPQELWKISALAAPVFISIFSKIAFYSFIIYCATSMGTHV 354
Query: 133 MAAHQ 137
+AAHQ
Sbjct: 355 LAAHQ 359
>gb|AAL27003.1| enhanced disease susceptibility 5 [Arabidopsis thaliana]
gi|22329250|ref|NP_195614.2| enhanced disease
susceptibility 5 (EDS5) / salicylic acid induction
deficient 1 (SID1) [Arabidopsis thaliana]
gi|51970810|dbj|BAD44097.1| enhanced disease
susceptibility 5 (EDS5) [Arabidopsis thaliana]
gi|51970686|dbj|BAD44035.1| enhanced disease
susceptibility 5 (EDS5) [Arabidopsis thaliana]
gi|51970290|dbj|BAD43837.1| enhanced disease
susceptibility 5 (EDS5) [Arabidopsis thaliana]
gi|51969106|dbj|BAD43245.1| enhanced disease
susceptibility 5 (EDS5) [Arabidopsis thaliana]
gi|20137881|sp|Q945F0|EDS5_ARATH Enhanced disease
susceptibility 5 (Eds5) (Salicylic acid induction
deficient 1) (Sid1)
Length = 543
Score = 179 bits (453), Expect = 6e-44
Identities = 86/125 (68%), Positives = 106/125 (84%)
Query: 13 LCGWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAA 72
L G VAQSASLGMK+SWGPLKALAAA++ING+GD +LC +LG GIAGAAWAT ASQ+V+A
Sbjct: 235 LVGLVAQSASLGMKNSWGPLKALAAATIINGLGDTILCLFLGQGIAGAAWATTASQIVSA 294
Query: 73 YMMMRTLNMKGYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFATSMGTHT 132
YMMM +LN +GYNA++ +IPS +E I LAAPVF+++ SK+AFYS +IY ATSMGTH
Sbjct: 295 YMMMDSLNKEGYNAYSFAIPSPQELWKISALAAPVFISIFSKIAFYSFIIYCATSMGTHV 354
Query: 133 MAAHQ 137
+AAHQ
Sbjct: 355 LAAHQ 359
>gb|AAO63941.1| unknown protein [Arabidopsis thaliana] gi|28393220|gb|AAO42040.1|
unknown protein [Arabidopsis thaliana]
gi|30687474|ref|NP_181367.2| MATE efflux family protein
[Arabidopsis thaliana]
Length = 521
Score = 54.3 bits (129), Expect = 2e-06
Identities = 44/122 (36%), Positives = 62/122 (50%), Gaps = 6/122 (4%)
Query: 18 AQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAAYMMMR 77
AQ A G KD+ PL A+ A +V+N V D +L LG+GI+GAA AT+ S+ + A++++
Sbjct: 228 AQGAFRGFKDTTTPLYAVVAGNVLNAVLDPILIFVLGFGISGAAAATVISEYLIAFILLW 287
Query: 78 TLNMKGYNAFALS--IPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFATSMGTHTMAA 135
LN N LS I GR + + T+ V F +L A G MA
Sbjct: 288 KLN---ENVVLLSPQIKVGRANQYLKSGGLLIGRTVALLVPF-TLATSLAAQNGPTQMAG 343
Query: 136 HQ 137
HQ
Sbjct: 344 HQ 345
>gb|AAC28771.1| hypothetical protein [Arabidopsis thaliana] gi|7487160|pir||T02512
hypothetical protein At2g38330 [imported] - Arabidopsis
thaliana
Length = 539
Score = 54.3 bits (129), Expect = 2e-06
Identities = 44/122 (36%), Positives = 62/122 (50%), Gaps = 6/122 (4%)
Query: 18 AQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAAYMMMR 77
AQ A G KD+ PL A+ A +V+N V D +L LG+GI+GAA AT+ S+ + A++++
Sbjct: 228 AQGAFRGFKDTTTPLYAVVAGNVLNAVLDPILIFVLGFGISGAAAATVISEYLIAFILLW 287
Query: 78 TLNMKGYNAFALS--IPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFATSMGTHTMAA 135
LN N LS I GR + + T+ V F +L A G MA
Sbjct: 288 KLN---ENVVLLSPQIKVGRANQYLKSGGLLIGRTVALLVPF-TLATSLAAQNGPTQMAG 343
Query: 136 HQ 137
HQ
Sbjct: 344 HQ 345
>gb|AAO76494.1| putative Na+-driven multidrug efflux pump [Bacteroides
thetaiotaomicron VPI-5482] gi|29346797|ref|NP_810300.1|
putative Na+-driven multidrug efflux pump [Bacteroides
thetaiotaomicron VPI-5482]
Length = 454
Score = 48.9 bits (115), Expect = 9e-05
Identities = 45/170 (26%), Positives = 73/170 (42%), Gaps = 31/170 (18%)
Query: 8 AGLLYLCGWVAQSASL-GMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMA 66
AG+L++ G+ L G+ DS PL +A A VIN V D +L Y G GAA AT+
Sbjct: 141 AGILFIVGYNVVCGILRGLGDSKTPLYFVALACVINIVLDFILVGYFHLGATGAAVATIT 200
Query: 67 SQVVAAYMMMRTLNMKGYN-----------------AFALSIPSGRE---------FITI 100
+Q V+ + + L G++ L P + IT+
Sbjct: 201 AQGVSFMISLWFLYRHGFHFEFTRKDIRLNRNLAKKVMTLGAPIALQDALINISFLIITV 260
Query: 101 ----LGLAAPVFMTMMSKVAFYSLLIYFATSMGTHTMAAHQYGVNLSRKL 146
+G+ A + ++ K+ +++L A S TM A YG L +++
Sbjct: 261 IVNQMGVIASASLGVVEKIIVFAMLPPMAISSAVATMTAQNYGAGLIQRM 310
>gb|AAB84820.1| conserved protein [Methanothermobacter thermautotrophicus str.
Delta H] gi|15678342|ref|NP_275457.1| hypothetical
protein MTH314 [Methanothermobacter thermautotrophicus
str. Delta H] gi|7451039|pir||C69140 conserved
hypothetical protein MTH314 - Methanobacterium
thermoautotrophicum (strain Delta H)
Length = 452
Score = 48.1 bits (113), Expect = 2e-04
Identities = 32/116 (27%), Positives = 61/116 (52%), Gaps = 17/116 (14%)
Query: 27 DSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMASQVVAAYMMM------RTLN 80
D+ P+ A+ ++V+N V D +L G+GIAGAAWAT+ SQ++ + +++ RT
Sbjct: 162 DARRPMYAMGLSAVLNMVLDPILIYTAGWGIAGAAWATVISQLLVSLLIIYWFLAGRTFT 221
Query: 81 MKGYNAF-----------ALSIPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFA 125
G++ F +++IP+ EF+ + + A + + S ++ +Y A
Sbjct: 222 SIGWSHFRADRGVVWSILSVTIPASAEFLVMSMVTALLNWILTSVAGTSAVAVYSA 277
>dbj|BAC71997.1| putative DNA-damage-inducible protein F [Streptomyces avermitilis
MA-4680] gi|29830828|ref|NP_825462.1| putative
DNA-damage-inducible protein F [Streptomyces avermitilis
MA-4680]
Length = 448
Score = 47.4 bits (111), Expect = 3e-04
Identities = 44/149 (29%), Positives = 67/149 (44%), Gaps = 17/149 (11%)
Query: 1 VDSRSIPAGLLYLCGWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGA 60
+ + IPA L+ L A G++D+ PL A V N V ++VL G GIAG+
Sbjct: 144 ISALGIPAMLVVLA---ATGVLRGLQDTKTPLYVAVAGFVANAVLNVVLVYGAGLGIAGS 200
Query: 61 AWATMASQ----VVAAYMMMRTLNMKGYNAFALSIPSGREFITILGLA---APVFMTMMS 113
AW T+ +Q V Y+++R A L P + I A AP+ + +S
Sbjct: 201 AWGTVIAQYGMAVAYLYVVVR-------GARKLGAPLRPDIAGIRACAQAGAPLLVRTLS 253
Query: 114 KVAFYSLLIYFATSMGTHTMAAHQYGVNL 142
A + A +G +AAHQ ++L
Sbjct: 254 LRAVLMIATAVAARLGDADIAAHQIILSL 282
>dbj|BAB04605.1| BH0886 [Bacillus halodurans C-125] gi|15613449|ref|NP_241752.1|
hypothetical protein BH0886 [Bacillus halodurans C-125]
gi|25329655|pir||F83760 hypothetical protein BH0886
[imported] - Bacillus halodurans (strain C-125)
Length = 454
Score = 47.0 bits (110), Expect = 4e-04
Identities = 35/128 (27%), Positives = 62/128 (48%), Gaps = 2/128 (1%)
Query: 9 GLLYLCGWVAQSASL-GMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMAS 67
G+L+L G+ L M DS P++ + A ++N + D ++ GI GAA+AT+ S
Sbjct: 144 GILFLFGYNFIGTVLRAMGDSRSPVRFIFIAVILNIILDPLMIAGFNLGIDGAAYATIVS 203
Query: 68 QVVA-AYMMMRTLNMKGYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFAT 126
Q A Y ++ ++ G +P R F + L P ++M++ A + ++ T
Sbjct: 204 QGTAFIYGLIYSVRKAGVPFQIPYVPEKRYFFVLFKLGLPAGLSMITISAGVTAILSVVT 263
Query: 127 SMGTHTMA 134
S G +A
Sbjct: 264 SFGEEAVA 271
>gb|AAQ65982.1| MATE efflux family protein [Porphyromonas gingivalis W83]
gi|34540604|ref|NP_905083.1| MATE efflux family protein
[Porphyromonas gingivalis W83]
Length = 452
Score = 44.7 bits (104), Expect = 0.002
Identities = 38/136 (27%), Positives = 65/136 (46%), Gaps = 10/136 (7%)
Query: 9 GLLYLCGWVAQSASLGMKD---SWGPLKALAAA----SVINGVGDIVLCTYLGYGIAGAA 61
G L++ V + ++ M + S G K A+A SVIN D + G+G+ GAA
Sbjct: 137 GRLFIVSCVIGTLNVSMSNIIVSQGASKTSASAMIIGSVINVALDPLFIYAFGWGVKGAA 196
Query: 62 WATMASQVVAAYMMMRTLNMKG--YNAFALSIPSGREFITILGLAAPVFMTMMSKVAFYS 119
WAT+ S+++ ++ MK +FAL +P+ R ++ I+ + + + + S
Sbjct: 197 WATILSRIITT-IIYTCFFMKAEVKVSFALFVPTIRMYLDIIKIGISMLCLQLLQTLSIS 255
Query: 120 LLIYFATSMGTHTMAA 135
LL A G +AA
Sbjct: 256 LLQRAAVKYGAEAVAA 271
>gb|AAQ66495.1| MATE efflux family protein [Porphyromonas gingivalis W83]
gi|34541117|ref|NP_905596.1| MATE efflux family protein
[Porphyromonas gingivalis W83]
Length = 439
Score = 44.7 bits (104), Expect = 0.002
Identities = 38/136 (27%), Positives = 65/136 (46%), Gaps = 10/136 (7%)
Query: 9 GLLYLCGWVAQSASLGMKD---SWGPLKALAAA----SVINGVGDIVLCTYLGYGIAGAA 61
G L++ V + ++ M + S G K A+A SVIN D + G+G+ GAA
Sbjct: 124 GRLFIVSCVIGTLNVSMSNIIVSQGASKTSASAMIIGSVINVALDPLFIYAFGWGVKGAA 183
Query: 62 WATMASQVVAAYMMMRTLNMKG--YNAFALSIPSGREFITILGLAAPVFMTMMSKVAFYS 119
WAT+ S+++ ++ MK +FAL +P+ R ++ I+ + + + + S
Sbjct: 184 WATILSRIITT-IIYTCFFMKAEVKVSFALFVPTIRMYLDIIKIGISMLCLQLLQTLSIS 242
Query: 120 LLIYFATSMGTHTMAA 135
LL A G +AA
Sbjct: 243 LLQRAAVKYGAEAVAA 258
>ref|NP_390054.1| hypothetical protein BSU21710 [Bacillus subtilis subsp. subtilis
str. 168] gi|2634591|emb|CAB14089.1| ypnP [Bacillus
subtilis subsp. subtilis str. 168]
gi|1256651|gb|AAA96645.1| 21% identity with the
NADH-dehydrogenase (ubiquinone) of Caenorhabditis
elegans; putative [Bacillus subtilis]
gi|7474682|pir||D69939 conserved hypothetical protein
ypnP - Bacillus subtilis gi|1730941|sp|P54181|YPNP_BACSU
Hypothetical protein ypnP
Length = 445
Score = 43.9 bits (102), Expect = 0.003
Identities = 38/128 (29%), Positives = 59/128 (45%), Gaps = 2/128 (1%)
Query: 9 GLLYLCGWVAQSASL-GMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMAS 67
G+L+L G+ S L + DS PL+ +A A V+N V + + GIAGAA++T+ S
Sbjct: 142 GILFLFGYNFISTVLRALGDSKTPLRFIAFAVVLNTVLAPLFISVFRMGIAGAAYSTILS 201
Query: 68 QVVA-AYMMMRTLNMKGYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFAT 126
Q +A Y + + K +P E IL L P + MM ++
Sbjct: 202 QGIAFLYGLFYVIKHKLVPFSIPRMPKWEESALILKLGIPAGLQMMVITGGMMAIMSVVN 261
Query: 127 SMGTHTMA 134
S G H ++
Sbjct: 262 SYGDHVVS 269
>ref|ZP_00398571.1| Multi antimicrobial extrusion protein MatE [Anaeromyxobacter
dehalogenans 2CP-C] gi|66779799|gb|EAL80875.1| Multi
antimicrobial extrusion protein MatE [Anaeromyxobacter
dehalogenans 2CP-C]
Length = 549
Score = 43.5 bits (101), Expect = 0.004
Identities = 38/137 (27%), Positives = 63/137 (45%), Gaps = 28/137 (20%)
Query: 5 SIPAGLLYLCGWVAQSASLGMKDSW-GPLKALAAASVINGVGDIVLCTYLGYGIAGAAWA 63
++PA LL + G ASL +W GPL+ + A G AGAA A
Sbjct: 259 NVPADLLLVFG----GASL---PAWTGPLRLVPAL-----------------GAAGAAIA 294
Query: 64 TMASQVVAAYMMMRT---LNMKGYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFYSL 120
T + A++++R +++ G L+ P T + + PV + M ++V F+SL
Sbjct: 295 TSLCIALQAFVLVRATRAVSLGGPRPGGLARPDRAALATAVRVGGPVGLHMFAEVGFFSL 354
Query: 121 LIYFATSMGTHTMAAHQ 137
+ + A +G + AHQ
Sbjct: 355 VSFLAGRLGAEALGAHQ 371
>ref|YP_079469.1| putative Multi antimicrobial extrusion protein [Bacillus
licheniformis ATCC 14580] gi|52003889|gb|AAU23831.1|
putative Multi antimicrobial extrusion protein [Bacillus
licheniformis ATCC 14580] gi|52786051|ref|YP_091880.1|
YpnP [Bacillus licheniformis ATCC 14580]
gi|52348553|gb|AAU41187.1| YpnP [Bacillus licheniformis
DSM 13]
Length = 443
Score = 43.5 bits (101), Expect = 0.004
Identities = 34/106 (32%), Positives = 55/106 (51%), Gaps = 2/106 (1%)
Query: 9 GLLYLCGWVAQSASL-GMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGAAWATMAS 67
G+++L G+ S L + DS PL+ + A ++N D + + GIAGAA+AT+ S
Sbjct: 141 GIIFLFGYNFISTVLRAVGDSQTPLRFVLLAVILNLFMDPLFISVFNLGIAGAAYATVVS 200
Query: 68 QVVA-AYMMMRTLNMKGYNAFALSIPSGREFITILGLAAPVFMTMM 112
Q +A Y ++ T+ + ++PS E IL L P + MM
Sbjct: 201 QGIAFIYGVVYTVRKQLVPFSKPALPSLAETSVILKLGVPAGLQMM 246
>ref|ZP_00310306.1| COG0534: Na+-driven multidrug efflux pump [Cytophaga hutchinsonii]
Length = 454
Score = 43.5 bits (101), Expect = 0.004
Identities = 34/127 (26%), Positives = 60/127 (46%), Gaps = 13/127 (10%)
Query: 30 GPLKALAAASVINGVGDIVLCTYLGYG--------IAGAAWATMASQV-VAAYMMMRTLN 80
G K S++ V +IVL L YG + G A+AT+ +++ +A M++ L
Sbjct: 153 GNTKQAMVVSILGNVVNIVLIVILVYGWYGLPKLGLNGIAYATLIARIFMAVTMVLFFLF 212
Query: 81 MKGY----NAFALSIPSGREFITILGLAAPVFMTMMSKVAFYSLLIYFATSMGTHTMAAH 136
K Y +A+ + F+ + + PV + M + +SL F + GT ++AH
Sbjct: 213 TKNYAHYRSAYKRTKLEWHYFVDVFNKSYPVGLQMSFESGAFSLAAIFVGTFGTAQISAH 272
Query: 137 QYGVNLS 143
Q +NL+
Sbjct: 273 QIALNLA 279
>emb|CAB42738.1| putative membrane protein [Streptomyces coelicolor A3(2)]
gi|7480041|pir||T36597 hypothetical protein SCH24.32c -
Streptomyces coelicolor gi|21222317|ref|NP_628096.1|
putative membrane protein [Streptomyces coelicolor
A3(2)]
Length = 448
Score = 43.1 bits (100), Expect = 0.005
Identities = 37/144 (25%), Positives = 65/144 (44%), Gaps = 7/144 (4%)
Query: 1 VDSRSIPAGLLYLCGWVAQSASLGMKDSWGPLKALAAASVINGVGDIVLCTYLGYGIAGA 60
+ S IPA L+ L + G++++ PL A + N V +++L G GIAG+
Sbjct: 144 ISSLGIPAMLVVLA---STGVLRGLQNTRTPLYVAVAGFIANAVLNVLLVYGAGLGIAGS 200
Query: 61 AWATMASQ--VVAAYMMMRTLNMKGYNAFALSIPSGREFITILGLAAPVFMTMMSKVAFY 118
AW T+ +Q + A Y+ + + + A G G+ P+ + +S A
Sbjct: 201 AWGTVIAQCGMAAVYLWVVVRGARQHGASLRPDLVGIRASAQAGM--PLLVRTLSLRAIL 258
Query: 119 SLLIYFATSMGTHTMAAHQYGVNL 142
+ A +G +AAHQ ++L
Sbjct: 259 MIATAVAARLGDADIAAHQIVLSL 282
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.338 0.146 0.479
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 320,755,870
Number of Sequences: 2540612
Number of extensions: 11250517
Number of successful extensions: 47721
Number of sequences better than 10.0: 170
Number of HSP's better than 10.0 without gapping: 69
Number of HSP's successfully gapped in prelim test: 101
Number of HSP's that attempted gapping in prelim test: 47618
Number of HSP's gapped (non-prelim): 186
length of query: 212
length of database: 863,360,394
effective HSP length: 122
effective length of query: 90
effective length of database: 553,405,730
effective search space: 49806515700
effective search space used: 49806515700
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.8 bits)
S2: 72 (32.3 bits)
Medicago: description of AC149293.2


