

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149204.7 + phase: 0
(600 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
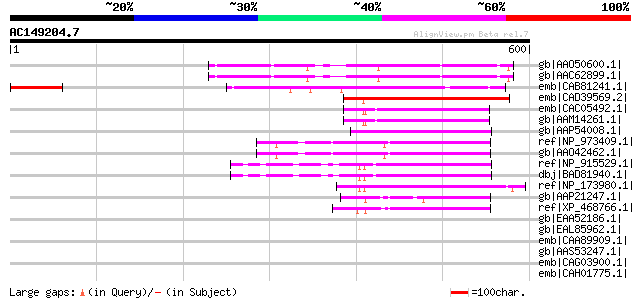
Score E
Sequences producing significant alignments: (bits) Value
gb|AAO50600.1| unknown protein [Arabidopsis thaliana] gi|2839318... 240 8e-62
gb|AAC62899.1| hypothetical protein [Arabidopsis thaliana] gi|25... 240 8e-62
emb|CAB81241.1| putative protein [Arabidopsis thaliana] gi|73210... 219 2e-55
emb|CAD39569.2| OSJNBa0019G23.13 [Oryza sativa (japonica cultiva... 187 6e-46
emb|CAC05492.1| putative protein [Arabidopsis thaliana] 145 4e-33
gb|AAM14261.1| putative DNA gyrase subunit B [Arabidopsis thalia... 145 4e-33
gb|AAP54008.1| putative polyprotein [Oryza sativa (japonica cult... 130 1e-28
ref|NP_973409.1| myb family transcription factor / ELM2 domain-c... 124 7e-27
gb|AAO42462.1| putative MYB family transcription factor [Arabido... 124 9e-27
ref|NP_915529.1| P0529E05.8 [Oryza sativa (japonica cultivar-gro... 119 2e-25
dbj|BAD81940.1| MYB transcription factor-like protein [Oryza sat... 119 2e-25
ref|NP_173980.1| expressed protein [Arabidopsis thaliana] gi|457... 112 3e-23
gb|AAP21247.1| At1g13880 [Arabidopsis thaliana] gi|8778387|gb|AA... 111 6e-23
ref|XP_468766.1| expressed protein [Oryza sativa (japonica culti... 109 3e-22
gb|EAA52186.1| hypothetical protein MG04878.4 [Magnaporthe grise... 48 0.001
gb|EAL85962.1| PHD transcription factor (Rum1), putative [Asperg... 45 0.007
emb|CAA89909.1| unknown [Saccharomyces cerevisiae] gi|6323830|re... 45 0.007
gb|AAS53247.1| AFL127Cp [Ashbya gossypii ATCC 10895] gi|45198394... 45 0.009
emb|CAG03900.1| unnamed protein product [Tetraodon nigroviridis] 45 0.009
emb|CAH01775.1| unnamed protein product [Kluyveromyces lactis NR... 43 0.025
>gb|AAO50600.1| unknown protein [Arabidopsis thaliana] gi|28393187|gb|AAO42024.1|
unknown protein [Arabidopsis thaliana]
gi|30690195|ref|NP_182128.2| ARID/BRIGHT DNA-binding
domain-containing protein / ELM2 domain-containing
protein [Arabidopsis thaliana]
Length = 562
Score = 240 bits (613), Expect = 8e-62
Identities = 145/366 (39%), Positives = 204/366 (55%), Gaps = 48/366 (13%)
Query: 231 DDVLMLDPSSVNRESFGRKRKRDDLMSEMQSWVIRTAKNPCDPVLGSMPEKSKWKSYGNQ 290
+ +++L+ SV +E +++ + E W+ AK+PCDP LG +P++S+W SYG++
Sbjct: 227 EQIMILE--SVTQECSSPGKRKRECPLETLKWLSDVAKDPCDPSLGIVPDRSEWVSYGSE 284
Query: 291 EIWKKVLLFREAAFLKKDFGSDCEKLSWLAQRMHPSMYDVNLGVNYNLRQRI-------- 342
E WK++LLFR + + + S CEK Q+MHP +YD + G +YNLR+R+
Sbjct: 285 EPWKQLLLFRAS---RTNNDSACEKTWQKVQKMHPCLYDDSAGASYNLRERLSYEDYKRG 341
Query: 343 KRDNGVLVGKSASILQRTPDSEAKEKLLDSCVPESILDAPPTVNIPLGPNHQAEVPEWTG 402
K NG +G S D E + L +G QA+VPEWTG
Sbjct: 342 KTGNGSDIGSS--------DEEDRPCAL------------------VGSKFQAKVPEWTG 375
Query: 403 TTHKSDSKWLGTQIWPLQIVKSK---LLEGEPVGKGRQDSCRCQVQGSVECVRFHIAEKS 459
T +SDSKWLGT+IWPL ++K L+E + +GKGRQD C C GS+ECV+FHI K
Sbjct: 376 ITPESDSKWLGTRIWPLTKEQTKANLLIERDRIGKGRQDPCGCHNPGSIECVKFHITAKR 435
Query: 460 AKLKLEIGVAFYQWNLDKAGEEVRRCWTAEEEKKFKDVVKSNPASLDRCFWDDIFTTFPK 519
KLKLE+G AFY W D GE + WT E KK K ++ S+P SL F P
Sbjct: 436 DKLKLELGPAFYMWCFDVMGECTLQYWTDLELKKIKSLM-SSPPSLSPAFIHQAKMILPS 494
Query: 520 KSRESLVSYYFNVFLLQRRGYQNRHTPDNIDSDDEESEFTPLKGSFGHQTDKSNSF---T 576
KSR +VSY++NV LLQ R Q+R TP +IDSD ++ + + ++N+F
Sbjct: 495 KSRGKIVSYFYNVTLLQYRASQSRITPHDIDSDTDQIFLRATENE--NPAPEANTFQKPA 552
Query: 577 LLSPKK 582
LLSP K
Sbjct: 553 LLSPNK 558
>gb|AAC62899.1| hypothetical protein [Arabidopsis thaliana] gi|25333959|pir||A84898
hypothetical protein At2g46040 [imported] - Arabidopsis
thaliana
Length = 576
Score = 240 bits (613), Expect = 8e-62
Identities = 145/366 (39%), Positives = 204/366 (55%), Gaps = 48/366 (13%)
Query: 231 DDVLMLDPSSVNRESFGRKRKRDDLMSEMQSWVIRTAKNPCDPVLGSMPEKSKWKSYGNQ 290
+ +++L+ SV +E +++ + E W+ AK+PCDP LG +P++S+W SYG++
Sbjct: 241 EQIMILE--SVTQECSSPGKRKRECPLETLKWLSDVAKDPCDPSLGIVPDRSEWVSYGSE 298
Query: 291 EIWKKVLLFREAAFLKKDFGSDCEKLSWLAQRMHPSMYDVNLGVNYNLRQRI-------- 342
E WK++LLFR + + + S CEK Q+MHP +YD + G +YNLR+R+
Sbjct: 299 EPWKQLLLFRAS---RTNNDSACEKTWQKVQKMHPCLYDDSAGASYNLRERLSYEDYKRG 355
Query: 343 KRDNGVLVGKSASILQRTPDSEAKEKLLDSCVPESILDAPPTVNIPLGPNHQAEVPEWTG 402
K NG +G S D E + L +G QA+VPEWTG
Sbjct: 356 KTGNGSDIGSS--------DEEDRPCAL------------------VGSKFQAKVPEWTG 389
Query: 403 TTHKSDSKWLGTQIWPLQIVKSK---LLEGEPVGKGRQDSCRCQVQGSVECVRFHIAEKS 459
T +SDSKWLGT+IWPL ++K L+E + +GKGRQD C C GS+ECV+FHI K
Sbjct: 390 ITPESDSKWLGTRIWPLTKEQTKANLLIERDRIGKGRQDPCGCHNPGSIECVKFHITAKR 449
Query: 460 AKLKLEIGVAFYQWNLDKAGEEVRRCWTAEEEKKFKDVVKSNPASLDRCFWDDIFTTFPK 519
KLKLE+G AFY W D GE + WT E KK K ++ S+P SL F P
Sbjct: 450 DKLKLELGPAFYMWCFDVMGECTLQYWTDLELKKIKSLM-SSPPSLSPAFIHQAKMILPS 508
Query: 520 KSRESLVSYYFNVFLLQRRGYQNRHTPDNIDSDDEESEFTPLKGSFGHQTDKSNSF---T 576
KSR +VSY++NV LLQ R Q+R TP +IDSD ++ + + ++N+F
Sbjct: 509 KSRGKIVSYFYNVTLLQYRASQSRITPHDIDSDTDQIFLRATENE--NPAPEANTFQKPA 566
Query: 577 LLSPKK 582
LLSP K
Sbjct: 567 LLSPNK 572
>emb|CAB81241.1| putative protein [Arabidopsis thaliana] gi|7321037|emb|CAB82145.1|
putative protein [Arabidopsis thaliana]
gi|15233358|ref|NP_192879.1| ARID/BRIGHT DNA-binding
domain-containing protein / ELM2 domain-containing
protein / Myb-like DNA-binding domain-containing protein
[Arabidopsis thaliana] gi|7452448|pir||T10560
hypothetical protein F25E4.20 - Arabidopsis thaliana
Length = 573
Score = 219 bits (559), Expect = 2e-55
Identities = 133/346 (38%), Positives = 183/346 (52%), Gaps = 29/346 (8%)
Query: 251 KRDDLMSEMQSWVIRTAKNPCDPVLGSMPEKSKWKSYGNQEIWKKVLLFREAAFLKKDFG 310
KRDDL M W+ A +P DP +G +P SKWK Y + W +V + + +++D
Sbjct: 218 KRDDLPG-MLKWLALVATSPHDPAIGVIPHSSKWKQYNGNKCWLQVARAKNSLLVQRDNA 276
Query: 311 SDCEKLSWLAQRM---HPSMYDVNLGVNYNLRQRIKRDN-------GVLVGKSASILQRT 360
+ HPSMY+ + LR I+ N G S L ++
Sbjct: 277 ELRYRYHPFRGHQNIHHPSMYEDDRKSIGRLRYSIRPPNLSKHCSSSCCNGSSLVSLSKS 336
Query: 361 PDSEAKEKLLDSCVPESILDAP-----------PTVNIPLGPNHQAEVPEWTGTTHKSDS 409
++ ++ + + + P I +G HQA+V EWT + SDS
Sbjct: 337 RSTKCRKLTIIASERAGLTAGTSRARKRNKAEIPRRCIKVGHQHQAQVDEWTESGVDSDS 396
Query: 410 KWLGTQIWPLQIVKS--KLLEGEPVGKGRQDSCRCQVQGSVECVRFHIAEKSAKLKLEIG 467
KWLGT+IWP + ++ + L + VGKGR DSC C++ G VEC R HIAEK +LK E+G
Sbjct: 397 KWLGTRIWPPENSEALDQTLGNDLVGKGRPDSCSCELSGFVECTRLHIAEKRMELKRELG 456
Query: 468 VAFYQWNLDKAGEEVRRCWTAEEEKKFKDVVKSNPASLDRCFWDDIFTTFPKKSRESLVS 527
F+ W ++ GEEV WT EEEK+FKD++ ++P S FW + FPKK RE LVS
Sbjct: 457 DDFFHWRFNQMGEEVCLRWTEEEEKRFKDMIIADPQS----FWTNAAKNFPKKKREELVS 512
Query: 528 YYFNVFLLQRRGYQNRHTPDNIDSDDEESEFTPLKGSFGHQTDKSN 573
YYFNVFL+ RR YQNR TP +IDSDD E F + GSFG S+
Sbjct: 513 YYFNVFLINRRRYQNRVTPKSIDSDD-EGAFGSVGGSFGRDAVTSS 557
Score = 55.1 bits (131), Expect = 6e-06
Identities = 22/61 (36%), Positives = 42/61 (68%)
Query: 1 MLGSEQTLDLYKLFMVVKDKGGYDVVCKNELWDLVGEEYGLGVKVGSSVKLVYSKYLSGL 60
++G + +DL+KLF++V+++ G+D V + LW++V E+ G + S+ L+Y KYL+ +
Sbjct: 50 VIGDGKNVDLFKLFVLVREREGFDTVSRKRLWEVVAEKLGFDCSLVPSLILIYLKYLNRM 109
Query: 61 E 61
E
Sbjct: 110 E 110
>emb|CAD39569.2| OSJNBa0019G23.13 [Oryza sativa (japonica cultivar-group)]
gi|50930121|ref|XP_474588.1| OSJNBa0019G23.13 [Oryza
sativa (japonica cultivar-group)]
Length = 538
Score = 187 bits (476), Expect = 6e-46
Identities = 100/200 (50%), Positives = 124/200 (62%), Gaps = 9/200 (4%)
Query: 387 IPLGPNHQAEVPEWTGTTHKS-----DSKWLGTQIWPLQIVKSKLLEG-EPVGKGRQDSC 440
IP+G QA+VP+WTG S KWLGT++WPL+ KL PVGKGR+ C
Sbjct: 327 IPVGSEFQAQVPQWTGELPVSYDNAETRKWLGTKVWPLENGNRKLSYFCNPVGKGREGVC 386
Query: 441 RCQVQGSVECVRFHIAEKSAKLKLEIGVAFYQWNLDKAGEEVRRCWTAEEEKKFKDVVKS 500
C + GSVECVRFH+AE+ +L+ E+ AFY W D+ GEE+ WT +EE FK V+
Sbjct: 387 GCNLPGSVECVRFHVAERRLQLRRELDSAFYAWGFDRMGEEIALSWTDKEEANFKACVQL 446
Query: 501 NPASLDRCFWDDIFTTFPKKSRESLVSYYFNVFLLQRRGYQNRHTPDNIDSDDE-ESEFT 559
N S R FW + F K R+ LVSYYFN FLL+RR YQNR TP+NIDSDDE E+EF
Sbjct: 447 NAPSSGRNFWKRLHMLFQSKGRKELVSYYFNCFLLRRRCYQNRMTPNNIDSDDEDETEFG 506
Query: 560 PLKGSFGHQTDK--SNSFTL 577
L GH K S+ +TL
Sbjct: 507 FLGNRLGHNATKYDSSKYTL 526
Score = 43.5 bits (101), Expect = 0.019
Identities = 23/76 (30%), Positives = 38/76 (49%), Gaps = 1/76 (1%)
Query: 2 LGSEQTLDLYKLFMVVKDKGGYDVVCKNE-LWDLVGEEYGLGVKVGSSVKLVYSKYLSGL 60
LG +DL +LF ++ GGY + +W E L + + VKL+Y KYL+ L
Sbjct: 63 LGDGSRVDLLRLFSAIRAAGGYAATSSSPAVWASAAESVCLDATLAAPVKLIYHKYLAAL 122
Query: 61 ETPLKNVVDGEFPKRD 76
+ ++ +V+ P D
Sbjct: 123 DRWIQRLVEAHGPFLD 138
>emb|CAC05492.1| putative protein [Arabidopsis thaliana]
Length = 351
Score = 145 bits (366), Expect = 4e-33
Identities = 76/179 (42%), Positives = 104/179 (57%), Gaps = 12/179 (6%)
Query: 387 IPLGPNHQAEVPEWTGTTHKS-------DS---KWLGTQIWPLQIVKSKLLEGEPVGKGR 436
IP+GP QAE+P W T K DS +WLGT +WP +K K + + VG+GR
Sbjct: 166 IPIGPRFQAEIPVWIAPTKKGKFYGSPGDSNTLRWLGTGVWPTYSLK-KTVHSKKVGEGR 224
Query: 437 QDSCRCQVQGSVECVRFHIAEKSAKLKLEIGVAFYQWNLDKAGEE-VRRCWTAEEEKKFK 495
DSC C S C++ H E L+ EI AF W D+ GEE V + WTA+EE++F+
Sbjct: 225 SDSCSCASPRSTNCIKRHKKEAQELLEKEINRAFSTWEFDQMGEEIVLKSWTAKEERRFE 284
Query: 496 DVVKSNPASLDRCFWDDIFTTFPKKSRESLVSYYFNVFLLQRRGYQNRHTPDNIDSDDE 554
+VK NP S FW+ FP+KS++ L+SYY+NVFL++R +NIDSDD+
Sbjct: 285 ALVKKNPLSSSDGFWEFASNAFPQKSKKDLLSYYYNVFLIKRMRLLKSSAANNIDSDDD 343
>gb|AAM14261.1| putative DNA gyrase subunit B [Arabidopsis thaliana]
gi|18377676|gb|AAL66988.1| unknown protein [Arabidopsis
thaliana] gi|30680387|ref|NP_196031.2| DNA topoisomerase
II family protein [Arabidopsis thaliana]
gi|41619080|gb|AAS10019.1| MYB transcription factor
[Arabidopsis thaliana]
Length = 546
Score = 145 bits (366), Expect = 4e-33
Identities = 76/179 (42%), Positives = 104/179 (57%), Gaps = 12/179 (6%)
Query: 387 IPLGPNHQAEVPEWTGTTHKS-------DS---KWLGTQIWPLQIVKSKLLEGEPVGKGR 436
IP+GP QAE+P W T K DS +WLGT +WP +K K + + VG+GR
Sbjct: 361 IPIGPRFQAEIPVWIAPTKKGKFYGSPGDSNTLRWLGTGVWPTYSLK-KTVHSKKVGEGR 419
Query: 437 QDSCRCQVQGSVECVRFHIAEKSAKLKLEIGVAFYQWNLDKAGEE-VRRCWTAEEEKKFK 495
DSC C S C++ H E L+ EI AF W D+ GEE V + WTA+EE++F+
Sbjct: 420 SDSCSCASPRSTNCIKRHKKEAQELLEKEINRAFSTWEFDQMGEEIVLKSWTAKEERRFE 479
Query: 496 DVVKSNPASLDRCFWDDIFTTFPKKSRESLVSYYFNVFLLQRRGYQNRHTPDNIDSDDE 554
+VK NP S FW+ FP+KS++ L+SYY+NVFL++R +NIDSDD+
Sbjct: 480 ALVKKNPLSSSDGFWEFASNAFPQKSKKDLLSYYYNVFLIKRMRLLKSSAANNIDSDDD 538
>gb|AAP54008.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|37534838|ref|NP_921721.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 294
Score = 130 bits (327), Expect = 1e-28
Identities = 67/167 (40%), Positives = 95/167 (56%), Gaps = 4/167 (2%)
Query: 395 AEVPEWTG--TTHKSDSKWLGTQIWPLQIVKSKLLEGEPVGKGRQDSCRCQVQGSVECVR 452
A++PEWTG + D L P+ K+ + + +GKGR D C C+V GS CVR
Sbjct: 47 ADIPEWTGKPSLPYEDPDMLRFLGEPILTPKNNEVFDDTIGKGRPDKCNCEVPGSTSCVR 106
Query: 453 FHIAEKSAKLKLEIGVAFYQWNLDKAGEEVRRCWTAEEEKKFKDVVKSNPASLDRCFWDD 512
FH+AEK +LK E+G ++Y D+ G++ W +EEKKF+ +V+ N S FWD
Sbjct: 107 FHVAEKKTELKREMGSSYYAMKFDEIGQDAALTWQKDEEKKFETIVQQNLPSSKYNFWDK 166
Query: 513 IFTTFPKKSRESLVSYYFNVFLLQRRGYQNR--HTPDNIDSDDEESE 557
+ F K +LVSYY NVF +RR +QNR + +DSDD+ E
Sbjct: 167 LRAAFRYKGERALVSYYNNVFQPRRRAFQNRVAQHANGVDSDDDSIE 213
>ref|NP_973409.1| myb family transcription factor / ELM2 domain-containing protein
[Arabidopsis thaliana]
Length = 449
Score = 124 bits (312), Expect = 7e-27
Identities = 88/281 (31%), Positives = 136/281 (48%), Gaps = 20/281 (7%)
Query: 286 SYGNQEIWKKVLLFREAAFLKK-----DFGSDCEKLSWLAQRMHPSMYDVNLGVNYNLRQ 340
SY N + + + + A+ KK D CE +W H G+
Sbjct: 24 SYKNHNQFDEAIPYHHASMEKKTNVLEDLIGLCENPTWTNDANHVDKGFETTGL------ 77
Query: 341 RIKRDNGVLVGKSASILQRTPDSEAKEKLLDSCVPESILDAPPTVNIPLGPNHQAEVPEW 400
+ D+ V + + ++ S+ K ++ V +++ PP + +G NHQA++PE+
Sbjct: 78 -CQEDSQSGVTTQSDLSHQSSGSDFTWKPVED-VYTCLMNQPPRKQVLVGSNHQADIPEF 135
Query: 401 TGTTHKSDSKWLGTQIWPLQIVKSKLLEGEP-----VGKGRQDSCRCQVQGSVECVRFHI 455
S+ + ++++ ++ G+GR++ C C +GS+ CVR HI
Sbjct: 136 VKEEILDQSEARTKEDLEGKLMRKCVIPMSDSDLCGTGQGRKE-CLCLDKGSIRCVRRHI 194
Query: 456 AEKSAKLKLEIGVA-FYQWNLDKAGEEVRRCWTAEEEKKFKDVVKSNPASLDRCFWDDIF 514
E L IG F + L + GEEV WT EEE F VV SNP S R FW +
Sbjct: 195 IEARESLVETIGYERFMELGLCEMGEEVASLWTEEEEDLFHKVVYSNPFSAGRDFWKQLK 254
Query: 515 TTFPKKSRESLVSYYFNVFLLQRRGYQNRHTPDNIDSDDEE 555
TFP ++ + LVSYYFNVF+L+RRG QNR ++DSDD+E
Sbjct: 255 GTFPSRTMKELVSYYFNVFILRRRGIQNRFKALDVDSDDDE 295
>gb|AAO42462.1| putative MYB family transcription factor [Arabidopsis thaliana]
gi|27754610|gb|AAO22751.1| putative MYB family
transcription factor [Arabidopsis thaliana]
gi|4335752|gb|AAD17429.1| putative MYB family
transcription factor [Arabidopsis thaliana]
gi|25411010|pir||H84448 probable MYB family
transcription factor [imported] - Arabidopsis thaliana
gi|15227651|ref|NP_178446.1| myb family transcription
factor / ELM2 domain-containing protein [Arabidopsis
thaliana]
Length = 450
Score = 124 bits (311), Expect = 9e-27
Identities = 88/282 (31%), Positives = 136/282 (48%), Gaps = 21/282 (7%)
Query: 286 SYGNQEIWKKVLLFREAAFLKK------DFGSDCEKLSWLAQRMHPSMYDVNLGVNYNLR 339
SY N + + + + A+ KK D CE +W H G+
Sbjct: 24 SYKNHNQFDEAIPYHHASMEKKTNVLVEDLIGLCENPTWTNDANHVDKGFETTGL----- 78
Query: 340 QRIKRDNGVLVGKSASILQRTPDSEAKEKLLDSCVPESILDAPPTVNIPLGPNHQAEVPE 399
+ D+ V + + ++ S+ K ++ V +++ PP + +G NHQA++PE
Sbjct: 79 --CQEDSQSGVTTQSDLSHQSSGSDFTWKPVED-VYTCLMNQPPRKQVLVGSNHQADIPE 135
Query: 400 WTGTTHKSDSKWLGTQIWPLQIVKSKLLEGEP-----VGKGRQDSCRCQVQGSVECVRFH 454
+ S+ + ++++ ++ G+GR++ C C +GS+ CVR H
Sbjct: 136 FVKEEILDQSEARTKEDLEGKLMRKCVIPMSDSDLCGTGQGRKE-CLCLDKGSIRCVRRH 194
Query: 455 IAEKSAKLKLEIGVA-FYQWNLDKAGEEVRRCWTAEEEKKFKDVVKSNPASLDRCFWDDI 513
I E L IG F + L + GEEV WT EEE F VV SNP S R FW +
Sbjct: 195 IIEARESLVETIGYERFMELGLCEMGEEVASLWTEEEEDLFHKVVYSNPFSAGRDFWKQL 254
Query: 514 FTTFPKKSRESLVSYYFNVFLLQRRGYQNRHTPDNIDSDDEE 555
TFP ++ + LVSYYFNVF+L+RRG QNR ++DSDD+E
Sbjct: 255 KGTFPSRTMKELVSYYFNVFILRRRGIQNRFKALDVDSDDDE 296
>ref|NP_915529.1| P0529E05.8 [Oryza sativa (japonica cultivar-group)]
Length = 523
Score = 119 bits (299), Expect = 2e-25
Identities = 96/314 (30%), Positives = 149/314 (46%), Gaps = 43/314 (13%)
Query: 256 MSEMQSWVIRTAKNPCDPVLGSMPEKSKWKSYGNQEIWKKVLLFREAAFLKKDFGSDCEK 315
+ E+ W A NP + + P ++ K+ Q VL R A +L+ + ++ ++
Sbjct: 94 LGELLRWTREVAANP----VAARPVPAEVKARKRQ-----VLALRRARYLRMEDVANADE 144
Query: 316 L-SWLAQRMHPSMYDVNLGVNYNLRQRIKRDNGVLVGKSASILQRTPDSEAKEKLLDSCV 374
L S+ +R + S N+ RQ+ K + KS + +R KL+ S +
Sbjct: 145 LPSFFKKRKYRSHN------NHAERQKGKGSMNMPTRKSERLAKRM-------KLMTSVL 191
Query: 375 PESILDAPPTVNIPLGPNHQAEVPEWTG------TTHKSD---SKWLGTQIWPLQIVKSK 425
I +G + QAEVP+WT +K+D SK LGT+IWP + +
Sbjct: 192 ------LTQRKKIGVGEHFQAEVPDWTEPPSDELARYKNDPNISKMLGTRIWPPE---GQ 242
Query: 426 LLEGEP--VGKGRQDSCRCQVQGSVECVRFHIAEKSAKLKLEIGVAFYQWNLDKAGEEVR 483
+L+ + G+GR +SC C GS C + H +L+ E+G AF +W D GEEV
Sbjct: 243 VLQTDKKIAGQGRMESCNCSYPGSFFCRQHHTDAARDQLRCELGRAFTEWRFDSMGEEVS 302
Query: 484 RCWTAEEEKKFKDVVKSNPASLDRCFWDDIFTTFPKKSRESLVSYYFNVFLLQRRGYQNR 543
+ WT EE+ KF + + P + FW K+R LV YY NVFL++R Q R
Sbjct: 303 KMWTREEQLKFNALERLVPVMDHKTFWAVASKHLASKTRIELVRYYLNVFLMRRVLSQCR 362
Query: 544 HTPDNIDSDDEESE 557
IDSD++E+E
Sbjct: 363 LNLLEIDSDEDETE 376
>dbj|BAD81940.1| MYB transcription factor-like protein [Oryza sativa (japonica
cultivar-group)]
Length = 403
Score = 119 bits (299), Expect = 2e-25
Identities = 96/314 (30%), Positives = 149/314 (46%), Gaps = 43/314 (13%)
Query: 256 MSEMQSWVIRTAKNPCDPVLGSMPEKSKWKSYGNQEIWKKVLLFREAAFLKKDFGSDCEK 315
+ E+ W A NP + + P ++ K+ Q VL R A +L+ + ++ ++
Sbjct: 94 LGELLRWTREVAANP----VAARPVPAEVKARKRQ-----VLALRRARYLRMEDVANADE 144
Query: 316 L-SWLAQRMHPSMYDVNLGVNYNLRQRIKRDNGVLVGKSASILQRTPDSEAKEKLLDSCV 374
L S+ +R + S N+ RQ+ K + KS + +R KL+ S +
Sbjct: 145 LPSFFKKRKYRSHN------NHAERQKGKGSMNMPTRKSERLAKRM-------KLMTSVL 191
Query: 375 PESILDAPPTVNIPLGPNHQAEVPEWTG------TTHKSD---SKWLGTQIWPLQIVKSK 425
I +G + QAEVP+WT +K+D SK LGT+IWP + +
Sbjct: 192 ------LTQRKKIGVGEHFQAEVPDWTEPPSDELARYKNDPNISKMLGTRIWPPE---GQ 242
Query: 426 LLEGEP--VGKGRQDSCRCQVQGSVECVRFHIAEKSAKLKLEIGVAFYQWNLDKAGEEVR 483
+L+ + G+GR +SC C GS C + H +L+ E+G AF +W D GEEV
Sbjct: 243 VLQTDKKIAGQGRMESCNCSYPGSFFCRQHHTDAARDQLRCELGRAFTEWRFDSMGEEVS 302
Query: 484 RCWTAEEEKKFKDVVKSNPASLDRCFWDDIFTTFPKKSRESLVSYYFNVFLLQRRGYQNR 543
+ WT EE+ KF + + P + FW K+R LV YY NVFL++R Q R
Sbjct: 303 KMWTREEQLKFNALERLVPVMDHKTFWAVASKHLASKTRIELVRYYLNVFLMRRVLSQCR 362
Query: 544 HTPDNIDSDDEESE 557
IDSD++E+E
Sbjct: 363 LNLLEIDSDEDETE 376
>ref|NP_173980.1| expressed protein [Arabidopsis thaliana] gi|45773908|gb|AAS76758.1|
At1g26580 [Arabidopsis thaliana] gi|25518777|pir||H86392
hypothetical protein T1K7.5 - Arabidopsis thaliana
gi|44681378|gb|AAS47629.1| At1g26580 [Arabidopsis
thaliana] gi|9797742|gb|AAF98560.1| Contains similarity
to a putative MYB family transcription factor gene
T4M8.10 gi|4335752 from Arabidopsis thaliana BAC T4M8
gb|AC006284 and contains a Myb-like DNA-binding PF|00249
domain. ESTs gb|T75914, gb|T45901 come from this gene
Length = 493
Score = 112 bits (281), Expect = 3e-23
Identities = 81/244 (33%), Positives = 124/244 (50%), Gaps = 27/244 (11%)
Query: 378 ILDAPPTVNIPLGPNHQAEVPEWTGT-------------THKSD----SKWLGTQIWPLQ 420
+LD +P+GP HQAE+PEW G+ H S K GT + P+
Sbjct: 126 LLDQRAKKQVPIGPGHQAEIPEWEGSQTGNIETSGMSVQNHISGCADGEKLFGTSVIPMP 185
Query: 421 IVKSKLLEGEPVGKGRQDSCRCQVQGSVECVRFHIAEKSAKLKLEIG-VAFYQWNLDKAG 479
+ + + VGKGR+ C C+ + SV CV HI E +L G F + L + G
Sbjct: 186 GLTTVAHIDDIVGKGRK-FCVCRDRDSVRCVCQHIKEAREELVKTFGNETFKELGLCEMG 244
Query: 480 EEVRRCWTAEEEKKFKDVVKSNPASLDRCFWDDIFTTFPKKSRESLVSYYFNVFLLQRRG 539
E+ W+ E+ + F +VV SNP +L + FW + F ++++ +VS+YFNVF+L+RR
Sbjct: 245 EKGALKWSDEDAQLFHEVVYSNPVTLGQNFWRHLEAAFCSRTQKEIVSFYFNVFVLRRRA 304
Query: 540 YQNRHTPDNIDSDDEESE--FTPLKGSFGHQTDKSNSFTLLSP-----KKPQPTPHTKGR 592
QNR +IDSDD+E + G+ + D+ +S + SP KK P H +G
Sbjct: 305 IQNRAFILDIDSDDDEWHGCYGGSSGTRYVEEDEEDS-AIESPLHQGTKKVYPLHHEEGE 363
Query: 593 QLVN 596
+ V+
Sbjct: 364 EDVS 367
>gb|AAP21247.1| At1g13880 [Arabidopsis thaliana] gi|8778387|gb|AAF79395.1| F16A14.9
[Arabidopsis thaliana] gi|15222974|ref|NP_172841.1| ELM2
domain-containing protein [Arabidopsis thaliana]
Length = 424
Score = 111 bits (278), Expect = 6e-23
Identities = 67/186 (36%), Positives = 103/186 (55%), Gaps = 20/186 (10%)
Query: 383 PTVNIPLGPNHQAEVPEWTGTTHKSDS---------KWLGTQIWPLQIVKSKLLEGEPVG 433
P +P+G ++QA++PE S + G + P+ ++++ + +G
Sbjct: 124 PRKTVPIGSDYQADIPECVKEEANDQSGQGVGYDEEQVTGKCVIPMPDCETEVCK---IG 180
Query: 434 KGRQDSCRCQVQGSVECVRFHIAEKSAKLKLEIGVAFYQWNLD----KAGEEVRRCWTAE 489
KGR++ C C +GS+ CV+ HI E L IG Y LD + GEEV T +
Sbjct: 181 KGRKE-CICLDKGSIRCVQQHIMENREDLFATIG---YDRCLDIGLCEMGEEVAARLTED 236
Query: 490 EEKKFKDVVKSNPASLDRCFWDDIFTTFPKKSRESLVSYYFNVFLLQRRGYQNRHTPDNI 549
EE F ++V SNP S+DR FW + + FP ++ + +VSYYFNVF+L+RR QNR ++
Sbjct: 237 EEDLFHEIVYSNPVSMDRDFWKHLKSAFPSRTMKEIVSYYFNVFILRRRAIQNRSKSLDV 296
Query: 550 DSDDEE 555
DSDD+E
Sbjct: 297 DSDDDE 302
>ref|XP_468766.1| expressed protein [Oryza sativa (japonica cultivar-group)]
gi|41469370|gb|AAS07212.1| expressed protein [Oryza
sativa (japonica cultivar-group)]
Length = 520
Score = 109 bits (272), Expect = 3e-22
Identities = 71/212 (33%), Positives = 101/212 (47%), Gaps = 35/212 (16%)
Query: 374 VPESILDAPPTVNIPLGPNHQAEVPEW-------------------------TGTTHKSD 408
+ I D P + +GP+HQA++PEW +G+ + +
Sbjct: 201 IDSPIHDHLPRKAVAIGPDHQADIPEWRPRISMTVPYGSGSCADLSYSSVSTSGSAPRDE 260
Query: 409 S----KWLGTQIWPLQIVKSKLLEGEPVGKGRQDSCRCQVQGSVECVRFHIAEKSAKLKL 464
KW+ + + S G+ GR C C +GS+ CVR H+ E LK
Sbjct: 261 DSESDKWIKHCVIEMPSSCSVAWVGD---HGRD--CGCSDEGSIRCVRRHVLESRENLKR 315
Query: 465 EIGV-AFYQWNLDKAGEEVRRCWTAEEEKKFKDVVKSNPASLDRCFWDDIFTTFPKKSRE 523
G F + L GE++ + WT EEE F VV SNP SL + FW + P K+
Sbjct: 316 IFGEDKFRELGLCDMGEDIAQRWTDEEESLFYRVVYSNPPSLGKNFWHFLPRALPGKTSM 375
Query: 524 SLVSYYFNVFLLQRRGYQNRHTPDNIDSDDEE 555
LVSYYFNVF+L++R QNR P ++DSDD+E
Sbjct: 376 ELVSYYFNVFMLRKRAQQNRSEPLHVDSDDDE 407
>gb|EAA52186.1| hypothetical protein MG04878.4 [Magnaporthe grisea 70-15]
gi|39940724|ref|XP_359899.1| hypothetical protein
MG04878.4 [Magnaporthe grisea 70-15]
Length = 1755
Score = 47.8 bits (112), Expect = 0.001
Identities = 26/77 (33%), Positives = 43/77 (55%), Gaps = 4/77 (5%)
Query: 5 EQTLDLYKLFMVVKDKGGYDVVCKNELWDLVGEEYGLGVKV----GSSVKLVYSKYLSGL 60
++ LDLYKL V+++GG+D VCK++ W +G + G K+ +S+K Y K+L
Sbjct: 188 KKPLDLYKLKKAVENRGGFDKVCKSKKWAEIGRDLGYSGKIMSSLSTSLKNSYQKFLCPY 247
Query: 61 ETPLKNVVDGEFPKRDL 77
E L+ G + +L
Sbjct: 248 EEYLRAARPGVHQQLEL 264
>gb|EAL85962.1| PHD transcription factor (Rum1), putative [Aspergillus fumigatus
Af293]
Length = 1748
Score = 45.1 bits (105), Expect = 0.007
Identities = 25/70 (35%), Positives = 39/70 (55%), Gaps = 4/70 (5%)
Query: 5 EQTLDLYKLFMVVKDKGGYDVVCKNELWDLVGEEYGLGVKV----GSSVKLVYSKYLSGL 60
++ LDLYKL V+ +GG+D VCK + W +G + G K+ +S+K Y ++L
Sbjct: 192 KRPLDLYKLKKAVEIRGGFDQVCKMKKWAEIGRDLGYSGKIMSSLSTSLKNSYQRWLQPY 251
Query: 61 ETPLKNVVDG 70
E L+ V G
Sbjct: 252 EEYLRVVKPG 261
>emb|CAA89909.1| unknown [Saccharomyces cerevisiae] gi|6323830|ref|NP_013901.1|
Non-essential protein of unknown function, contains
ATP/GTP-binding site motif A; null mutant exhibits
cellular volume up to four times greater than wild-type,
also large drooping buds with elongated necks; Ecm5p
[Saccharomyces cerevisiae]
gi|2497173|sp|Q03214|ECM5_YEAST Extracellular matrix
protein 5
Length = 1411
Score = 45.1 bits (105), Expect = 0.007
Identities = 21/57 (36%), Positives = 36/57 (62%), Gaps = 4/57 (7%)
Query: 5 EQTLDLYKLFMVVKDKGGYDVVCKNELWDLVGEEYGLGVKV----GSSVKLVYSKYL 57
++TLDLY+L VK +GG++ VC+ +LW +G E G ++ +S++ Y+K L
Sbjct: 215 KRTLDLYRLRSCVKLRGGFNAVCEKKLWAQIGRELGYSGRIMSSLSTSLRSAYAKIL 271
>gb|AAS53247.1| AFL127Cp [Ashbya gossypii ATCC 10895] gi|45198394|ref|NP_985423.1|
AFL127Cp [Eremothecium gossypii]
Length = 1521
Score = 44.7 bits (104), Expect = 0.009
Identities = 21/61 (34%), Positives = 37/61 (60%), Gaps = 4/61 (6%)
Query: 5 EQTLDLYKLFMVVKDKGGYDVVCKNELWDLVGEEYGLGVKV----GSSVKLVYSKYLSGL 60
++TLDLY+L+ V+ +GG+ VC+ +LW +G E G ++ +S++ YSK S
Sbjct: 263 KRTLDLYRLWNCVQLRGGFKTVCQKKLWAQLGRELGYSGRIMSSLSTSLRSAYSKVFSDF 322
Query: 61 E 61
+
Sbjct: 323 D 323
>emb|CAG03900.1| unnamed protein product [Tetraodon nigroviridis]
Length = 1638
Score = 44.7 bits (104), Expect = 0.009
Identities = 22/52 (42%), Positives = 32/52 (61%), Gaps = 2/52 (3%)
Query: 8 LDLYKLFMVVKDKGGYDVVCKNELWDLVGEEYGL--GVKVGSSVKLVYSKYL 57
LDLYKL +V D+GG+D+VC++ W + + G G VGS ++ Y K L
Sbjct: 111 LDLYKLNKLVADEGGFDIVCQDRRWTKIAVQMGFAPGKAVGSHLRGHYEKIL 162
>emb|CAH01775.1| unnamed protein product [Kluyveromyces lactis NRRL Y-1140]
gi|50305927|ref|XP_452924.1| unnamed protein product
[Kluyveromyces lactis]
Length = 1454
Score = 43.1 bits (100), Expect = 0.025
Identities = 18/45 (40%), Positives = 31/45 (68%)
Query: 5 EQTLDLYKLFMVVKDKGGYDVVCKNELWDLVGEEYGLGVKVGSSV 49
++TLDL++L V+ +GG+D VC+ +LW +G E G ++ SS+
Sbjct: 255 KRTLDLHRLRSCVQLRGGFDTVCQKKLWAQIGRELGYSGRIMSSL 299
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.314 0.133 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,076,712,790
Number of Sequences: 2540612
Number of extensions: 47492207
Number of successful extensions: 87226
Number of sequences better than 10.0: 286
Number of HSP's better than 10.0 without gapping: 85
Number of HSP's successfully gapped in prelim test: 205
Number of HSP's that attempted gapping in prelim test: 86978
Number of HSP's gapped (non-prelim): 383
length of query: 600
length of database: 863,360,394
effective HSP length: 134
effective length of query: 466
effective length of database: 522,918,386
effective search space: 243679967876
effective search space used: 243679967876
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 78 (34.7 bits)
Medicago: description of AC149204.7


