

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149204.4 - phase: 0 /pseudo
(640 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
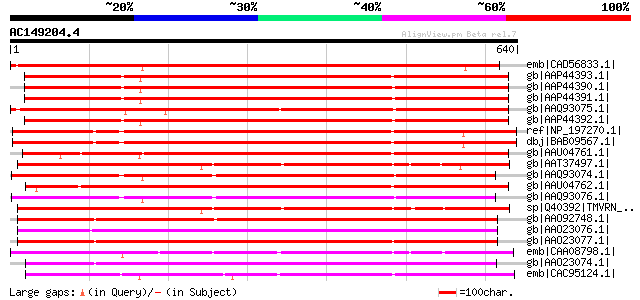
Score E
Sequences producing significant alignments: (bits) Value
emb|CAD56833.1| putative resistance gene analogue protein [Lens ... 805 0.0
gb|AAP44393.1| nematode resistance-like protein [Solanum tuberosum] 509 e-142
gb|AAP44390.1| nematode resistance protein [Solanum tuberosum] 506 e-142
gb|AAP44391.1| nematode resistance-like protein [Solanum tuberosum] 505 e-141
gb|AAQ93075.1| TIR-NBS-LRR type R protein 7 [Malus baccata] 504 e-141
gb|AAP44392.1| nematode resistance-like protein [Solanum tuberosum] 502 e-140
ref|NP_197270.1| disease resistance protein (TIR-NBS-LRR class),... 491 e-137
dbj|BAB09567.1| disease resistance protein-like [Arabidopsis tha... 488 e-136
gb|AAU04761.1| MRGH13 [Cucumis melo] 469 e-130
gb|AAT37497.1| N-like protein [Nicotiana tabacum] 468 e-130
gb|AAQ93074.1| putative TIR-NBS type R protein 4 [Malus baccata] 459 e-127
gb|AAU04762.1| MRGH21 [Cucumis melo] 456 e-126
gb|AAQ93076.1| putative TIR-NBS type R protein 4 [Malus baccata] 454 e-126
sp|Q40392|TMVRN_NICGU TMV resistance protein N gi|558887|gb|AAA5... 453 e-126
gb|AAO92748.1| candidate disease-resistance protein SR1 [Glycine... 453 e-126
gb|AAO23076.1| R 1 protein [Glycine max] 453 e-126
gb|AAO23077.1| R 8 protein [Glycine max] 452 e-125
emb|CAA08798.1| NL27 [Solanum tuberosum] 450 e-125
gb|AAO23074.1| R 10 protein [Glycine max] 450 e-125
emb|CAC95124.1| TIR/NBS/LRR protein [Populus deltoides] 449 e-124
>emb|CAD56833.1| putative resistance gene analogue protein [Lens culinaris]
Length = 810
Score = 805 bits (2079), Expect = 0.0
Identities = 410/685 (59%), Positives = 508/685 (73%), Gaps = 70/685 (10%)
Query: 2 EASSSSSCRSSSTTSLCTYHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQ 61
EASSSSS S TS T HVF+SFR EDTR+GFTDHL A+LER+GI TFKDD DL+RG+
Sbjct: 9 EASSSSS--PSILTSKWTNHVFVSFRSEDTRQGFTDHLFASLERRGIKTFKDDHDLKRGE 66
Query: 62 VISEKLINAIKDSMFAITILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDV 121
VIS +L AI++SMFAI ILSP+YASSTWCLDELQ I+ECS + P+F+GVDPSDV
Sbjct: 67 VISVELNKAIQESMFAIIILSPNYASSTWCLDELQKIVECSKSSGQTFFPIFHGVDPSDV 126
Query: 122 RHQRGCFEEAFRKHQEKFGQHSDRVDRWRDAFTQVASYSGWDSKG--------------- 166
RHQRG F +AFRKH+EK + ++++RWRDA +VASYSGWDSKG
Sbjct: 127 RHQRGSFAKAFRKHEEKLRKDRNKIERWRDALREVASYSGWDSKGWLVEMFMLISFYLEF 186
Query: 167 -------------------------------QHEASLVENIAQHIHRKLVPKLPSCTENL 195
+ EASLVE IA+HIH+KL+PKLP C +NL
Sbjct: 187 PKHETIITCFLYRLVALFTYRLMQVSFPSLCRKEASLVETIAEHIHKKLIPKLPVCKDNL 246
Query: 196 VGIVSKVEEVNKFLGMGLNDVRFIGIWGMGGIGKSTIARAVYETIRCEFELTCFLENVRE 255
VGI S++EE+ LGM L+DVRFIGIWGMGGIGK+TIAR+VY+ I+ EF+++CFL ++RE
Sbjct: 247 VGIDSRIEEIYSLLGMRLSDVRFIGIWGMGGIGKTTIARSVYDAIKDEFQVSCFLADIRE 306
Query: 256 -ISETNGLVHLQRQLLSHLSISRNDFHDLYDGKKTIQNSLCRKKVLLVLDDVNELNQLEN 314
IS TNGLV +Q +LLSHL+I NDF++++DGKK + NS KKVLLVLDDV+EL+QLE+
Sbjct: 307 TISRTNGLVRIQTELLSHLTIRSNDFYNIHDGKKILANSFRNKKVLLVLDDVSELSQLES 366
Query: 315 LVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGDKPQEG 374
L GKQ+WFG G RVIIT+RDKHLLMTHGV++TYK L K++AL LFCLKAFK ++P+E
Sbjct: 367 LAGKQEWFGSGIRVIITSRDKHLLMTHGVNETYKAKGLVKNEALKLFCLKAFKQNQPKEE 426
Query: 375 YLDLSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKLRSFPHPRVQDNLKISYDSL 434
YL L KEVV+Y GLPLALEVLGS+ +GR ++VWHSA++++R+ PH ++ D LKISYDSL
Sbjct: 427 YLSLCKEVVEYARGLPLALEVLGSHFHGRTVEVWHSALEQMRNVPHSKIHDTLKISYDSL 486
Query: 435 DTMEKDIFLDIACFFKGMKGDKVIDILESCGYFPQIGIQILIERSLITLDSVNNKLGMHD 494
ME+++FLDIACFFKGM D V++ILE CGY+P+IGI ILIERSL++ D + KL MHD
Sbjct: 487 QPMERNMFLDIACFFKGMDIDGVMEILEDCGYYPKIGIDILIERSLVSFDRGDRKLWMHD 546
Query: 495 LLQEMGRDIVFQESPNDPCRRSRLWSQEDIDRVLTKNKGTEAINSIDMKLLQPYEAHWNT 554
LL+EMGR+IV QESPNDP +RSRLWSQ+DID+VLTKNKGT+ I I + L+QPYEA WN
Sbjct: 547 LLEEMGRNIVCQESPNDPGKRSRLWSQKDIDQVLTKNKGTDKIQGIALNLVQPYEAGWNI 606
Query: 555 EAFSKTSQLKFLSLCEMQLP---------------------LGLSCLPSSLKVLHWRGCP 593
EAFS+ SQL+ L LCE++LP GL C PSSLKVL WRGCP
Sbjct: 607 EAFSRLSQLRLLKLCEIKLPRGSRHELSASPLGTQYVNKTSRGLGCFPSSLKVLDWRGCP 666
Query: 594 LKTLPITTQLDELVDITLSHSKIEQ 618
LKT P T DE+V++ L HSKIE+
Sbjct: 667 LKTPPQTNHFDEIVNLKLFHSKIEK 691
>gb|AAP44393.1| nematode resistance-like protein [Solanum tuberosum]
Length = 1136
Score = 509 bits (1311), Expect = e-142
Identities = 276/616 (44%), Positives = 394/616 (63%), Gaps = 9/616 (1%)
Query: 19 TYHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQVISEKLINAIKDSMFAI 78
+Y VFLSFRGED RK F DHL AL++K I TFKDD+ LE+G+ IS +L+++I++S A+
Sbjct: 17 SYDVFLSFRGEDVRKTFVDHLYLALQQKCINTFKDDEKLEKGKFISPELVSSIEESRIAL 76
Query: 79 TILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVRHQRGCFEEAFRKHQEK 138
I S +YA+STWCLDEL IMEC + V+PVFY VDPS VR Q+ F EAF KH+ +
Sbjct: 77 IIFSKNYANSTWCLDELTKIMECKNVKGQIVVPVFYDVDPSTVRKQKSIFGEAFSKHEAR 136
Query: 139 FGQHSDRVDRWRDAFTQVASYSGWD---SKGQHEASLVENIAQHIHRKL-VPKLPSCTEN 194
F + D+V +WR A + A+ SGWD + HEA ++E IA+ I +L + S N
Sbjct: 137 FQE--DKVQKWRAALEEAANISGWDLPNTSNGHEARVMEKIAEDIMARLGSQRHASNARN 194
Query: 195 LVGIVSKVEEVNKFLGMGLNDVRFIGIWGMGGIGKSTIARAVYETIRCEFELTCFLENVR 254
LVG+ S + +V K LG+G V F+GI GM G+GK+T+AR +Y+ IR +F+ CFL VR
Sbjct: 195 LVGMESHMHQVYKMLGIGSGGVHFLGILGMSGVGKTTLARVIYDNIRSQFQGACFLHEVR 254
Query: 255 EISETNGLVHLQRQLLSH-LSISRNDFHDLYDGKKTIQNSLCRKKVLLVLDDVNELNQLE 313
+ S GL LQ LLS L + + +D ++G + L KKVLLVLDDV+ ++QL
Sbjct: 255 DRSAKQGLERLQEILLSEILVVKKLRINDSFEGANMQKQRLQYKKVLLVLDDVDHIDQLN 314
Query: 314 NLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGDKPQE 373
L G+++WFG GSR+IITT+DKHLL+ + K Y+ L +++L LF AFK ++P +
Sbjct: 315 ALAGEREWFGDGSRIIITTKDKHLLVKYETEKIYRMKTLNNYESLQLFKQHAFKKNRPTK 374
Query: 374 GYLDLSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKLRSFPHPRVQDNLKISYDS 433
+ DLS +V+ + GLPLAL+VLGS+LYGR +D W S V++L+ P + L+ S+
Sbjct: 375 EFEDLSAQVIKHTDGLPLALKVLGSFLYGRGLDEWISEVERLKQIPENEILKKLEQSFTG 434
Query: 434 LDTMEKDIFLDIACFFKGMKGDKVIDILESCGYFPQIGIQILIERSLITLDSVNNKLGMH 493
L E+ IFLDIACFF G K D V ILES + P IGI++L+E+ LIT ++ ++ +H
Sbjct: 435 LHNTEQKIFLDIACFFSGKKKDSVTRILESFHFCPVIGIKVLMEKCLIT--TLQGRITIH 492
Query: 494 DLLQEMGRDIVFQESPNDPCRRSRLWSQEDIDRVLTKNKGTEAINSIDMKLLQPYEAHWN 553
L+Q+MG IV +E+ +DP SRLW +EDI VL +N GT+ I + + L E ++
Sbjct: 493 QLIQDMGWHIVRREATDDPRMCSRLWKREDICPVLERNLGTDKIEGMSLHLTNEEEVNFG 552
Query: 554 TEAFSKTSQLKFLSLCEMQLPLGLSCLPSSLKVLHWRGCPLKTLPITTQLDELVDITLSH 613
+AF + ++L+FL + G LP L+ L W G P K+LP + + D+LV + L
Sbjct: 553 GKAFMQMTRLRFLKFQNAYVCQGPEFLPDELRWLDWHGYPSKSLPNSFKGDQLVSLKLKK 612
Query: 614 SKIEQLWQGVKVIDSL 629
S+I QLW+ K + L
Sbjct: 613 SRIIQLWKTSKDLGKL 628
>gb|AAP44390.1| nematode resistance protein [Solanum tuberosum]
Length = 1136
Score = 506 bits (1303), Expect = e-142
Identities = 275/616 (44%), Positives = 393/616 (63%), Gaps = 9/616 (1%)
Query: 19 TYHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQVISEKLINAIKDSMFAI 78
+Y VFLSFRGED RK F DHL ALE+K I TFKDD+ LE+G+ IS +L+++I++S A+
Sbjct: 17 SYDVFLSFRGEDVRKTFVDHLYLALEQKCINTFKDDEKLEKGKFISPELVSSIEESRIAL 76
Query: 79 TILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVRHQRGCFEEAFRKHQEK 138
I S +YA+STWCLDEL IMEC + V+PVFY VDPS VR Q+ F EAF KH+ +
Sbjct: 77 IIFSKNYANSTWCLDELTKIMECKNVKGQIVVPVFYDVDPSTVRKQKSIFGEAFSKHEAR 136
Query: 139 FGQHSDRVDRWRDAFTQVASYSGWD---SKGQHEASLVENIAQHIHRKL-VPKLPSCTEN 194
F + D+V +WR A + A+ SGWD + HEA ++E IA+ I +L + S N
Sbjct: 137 FQE--DKVQKWRAALEEAANISGWDLPNTANGHEARVMEKIAEDIMARLGSQRHASNARN 194
Query: 195 LVGIVSKVEEVNKFLGMGLNDVRFIGIWGMGGIGKSTIARAVYETIRCEFELTCFLENVR 254
LVG+ S + +V K LG+G V F+GI GM G+GK+T+AR +Y+ IR +F+ CFL VR
Sbjct: 195 LVGMESHMHKVYKMLGIGSGGVHFLGILGMSGVGKTTLARVIYDNIRSQFQGACFLHEVR 254
Query: 255 EISETNGLVHLQRQLLSH-LSISRNDFHDLYDGKKTIQNSLCRKKVLLVLDDVNELNQLE 313
+ S GL LQ LLS L + + +D ++G + L KKVLLVLDDV+ ++QL
Sbjct: 255 DRSAKQGLERLQEILLSEILVVKKLRINDSFEGANMQKQRLQYKKVLLVLDDVDHIDQLN 314
Query: 314 NLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGDKPQE 373
L G+++WFG GSR+IITT+DKHLL+ + K Y+ L +++L LF AFK ++P +
Sbjct: 315 ALAGEREWFGDGSRIIITTKDKHLLVKYETEKIYRMKTLNNYESLQLFKQHAFKKNRPTK 374
Query: 374 GYLDLSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKLRSFPHPRVQDNLKISYDS 433
+ DLS +V+ + GLPLAL+VLGS+LYGR +D W S V++L+ P + L+ S+
Sbjct: 375 EFEDLSAQVIKHTDGLPLALKVLGSFLYGRGLDEWISEVERLKQIPENEILKKLEQSFTG 434
Query: 434 LDTMEKDIFLDIACFFKGMKGDKVIDILESCGYFPQIGIQILIERSLITLDSVNNKLGMH 493
L E+ IFLDIACFF G K D V ILES + P IGI++L+E+ LIT+ + ++ +H
Sbjct: 435 LHNTEQKIFLDIACFFSGKKKDSVTRILESFHFCPVIGIKVLMEKCLITI--LQGRITIH 492
Query: 494 DLLQEMGRDIVFQESPNDPCRRSRLWSQEDIDRVLTKNKGTEAINSIDMKLLQPYEAHWN 553
L+Q+MG IV +E+ +DP SR+W +EDI VL +N GT+ + + L E ++
Sbjct: 493 QLIQDMGWHIVRREATDDPRMCSRMWKREDICPVLERNLGTDKNEGMSLHLTNEEEVNFG 552
Query: 554 TEAFSKTSQLKFLSLCEMQLPLGLSCLPSSLKVLHWRGCPLKTLPITTQLDELVDITLSH 613
+AF + ++L+FL + G LP L+ L W G P K+LP + + D+LV + L
Sbjct: 553 GKAFMQMTRLRFLKFRNAYVCQGPEFLPDELRWLDWHGYPSKSLPNSFKGDQLVGLKLKK 612
Query: 614 SKIEQLWQGVKVIDSL 629
S+I QLW+ K + L
Sbjct: 613 SRIIQLWKTSKDLGKL 628
>gb|AAP44391.1| nematode resistance-like protein [Solanum tuberosum]
Length = 1121
Score = 505 bits (1300), Expect = e-141
Identities = 275/616 (44%), Positives = 394/616 (63%), Gaps = 9/616 (1%)
Query: 19 TYHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQVISEKLINAIKDSMFAI 78
+Y VFLSFRGE+ RK F DHL ALE+K I TFKDD+ LE+G+ IS +L+++I++S A+
Sbjct: 17 SYDVFLSFRGENVRKTFVDHLYLALEQKCINTFKDDEKLEKGKFISPELMSSIEESRIAL 76
Query: 79 TILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVRHQRGCFEEAFRKHQEK 138
I S +YA+STWCLDEL I+EC + V+PVFY VDPS VR Q+ F EAF KH+ +
Sbjct: 77 IIFSKNYANSTWCLDELTKIIECKNVKGQIVVPVFYDVDPSTVRRQKNIFGEAFSKHEAR 136
Query: 139 FGQHSDRVDRWRDAFTQVASYSGWD---SKGQHEASLVENIAQHIHRKL-VPKLPSCTEN 194
F + D+V +WR A + A+ SGWD + HEA ++E I + I +L + S N
Sbjct: 137 FEE--DKVKKWRAALEEAANISGWDLPNTSNGHEARVIEKITEDIMVRLGSQRHASNARN 194
Query: 195 LVGIVSKVEEVNKFLGMGLNDVRFIGIWGMGGIGKSTIARAVYETIRCEFELTCFLENVR 254
+VG+ S + +V K LG+G VRF+GI GM G+GK+T+AR +Y+ I+ +FE CFL VR
Sbjct: 195 VVGMESHMHQVYKMLGIGSGGVRFLGILGMSGVGKTTLARVIYDNIQSQFEGACFLHEVR 254
Query: 255 EISETNGLVHLQRQLLSH-LSISRNDFHDLYDGKKTIQNSLCRKKVLLVLDDVNELNQLE 313
+ S GL HLQ LLS L + + +D ++G + L KKVLLVLDDV+ ++QL
Sbjct: 255 DRSAKQGLEHLQEILLSEILVVKKLRINDSFEGANMQKQRLQYKKVLLVLDDVDHIDQLN 314
Query: 314 NLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGDKPQE 373
L G+++WFG GSR+IITT+DKHLL+ + K Y+ G L K+++L LF AFK + +
Sbjct: 315 ALAGEREWFGDGSRIIITTKDKHLLVKYETEKIYRMGTLDKYESLQLFKQHAFKKNHSTK 374
Query: 374 GYLDLSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKLRSFPHPRVQDNLKISYDS 433
+ DLS +V+++ GGLPLAL+VLGS+LYGR +D W S V++L+ P + L+ S+
Sbjct: 375 EFEDLSAQVIEHTGGLPLALKVLGSFLYGRGLDEWISEVERLKQIPQNEILKKLEPSFTG 434
Query: 434 LDTMEKDIFLDIACFFKGMKGDKVIDILESCGYFPQIGIQILIERSLITLDSVNNKLGMH 493
L+ +E+ IFLDIACFF G K D V ILES + P IGI++L+E+ LIT+ + ++ +H
Sbjct: 435 LNNIEQKIFLDIACFFSGKKKDSVTRILESFHFSPVIGIKVLMEKCLITI--LKGRITIH 492
Query: 494 DLLQEMGRDIVFQESPNDPCRRSRLWSQEDIDRVLTKNKGTEAINSIDMKLLQPYEAHWN 553
L+QEMG IV +E+ +P SRLW +EDI VL +N T+ I + + L E ++
Sbjct: 493 QLIQEMGWHIVRREASYNPRICSRLWKREDICPVLEQNLCTDKIEGMSLHLTNEEEVNFG 552
Query: 554 TEAFSKTSQLKFLSLCEMQLPLGLSCLPSSLKVLHWRGCPLKTLPITTQLDELVDITLSH 613
+A + + L+FL + G LP L+ L W G P K LP + + D+LV + L
Sbjct: 553 GKALMQMTSLRFLKFRNAYVYQGPEFLPDELRWLDWHGYPSKNLPNSFKGDQLVSLKLKK 612
Query: 614 SKIEQLWQGVKVIDSL 629
S+I QLW+ K + L
Sbjct: 613 SRIIQLWKTSKDLGKL 628
>gb|AAQ93075.1| TIR-NBS-LRR type R protein 7 [Malus baccata]
Length = 1095
Score = 504 bits (1297), Expect = e-141
Identities = 275/644 (42%), Positives = 398/644 (61%), Gaps = 24/644 (3%)
Query: 2 EASSSSSCRSSSTTSLCTYHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQ 61
EASSSSS S + L Y +FLSFRGEDTR GFT HL AAL+ +G + D DL RG+
Sbjct: 9 EASSSSS----SMSKLWNYDLFLSFRGEDTRNGFTGHLHAALKDRGYQAYMDQDDLNRGE 64
Query: 62 VISEKLINAIKDSMFAITILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDV 121
I E+L AI+ S +I + S YA S+WCLDEL IMEC SK HVLP+FY VDPS V
Sbjct: 65 EIKEELFRAIEGSRISIIVFSKRYADSSWCLDELVKIMECRSKLGRHVLPIFYHVDPSHV 124
Query: 122 RHQRGCFEEAFRKHQEKFGQHSD---------RVDRWRDAFTQVASYSGWDSKGQHEASL 172
R Q G EAF KH+E G+ +D RV +W+ A T+ A+ SG D +
Sbjct: 125 RKQDGDLAEAFLKHEEGIGEGTDGKKREAKQERVKQWKKALTEAANLSGHDLRITDNGRE 184
Query: 173 VENIAQHIHRKLVPKLPSCTENL------VGIVSKVEEVNKFLGMGLNDVRFIGIWGMGG 226
+ I ++ K T L VGI S+++++ L G ++V +GIWGMGG
Sbjct: 185 ANLCPREIVDNIITKWLMSTNKLRVAKHQVGINSRIQDIISRLSSGGSNVIMVGIWGMGG 244
Query: 227 IGKSTIARAVYETIRCEFELTCFLENVREISETNGLVHLQRQLLSHLSISRNDFHDLYDG 286
+GK+T A+A+Y I EF+ FL +V + +GLV+LQ++L+ + +++ + +G
Sbjct: 245 LGKTTAAKAIYNQIHHEFQFKSFLPDVGNAASKHGLVYLQKELIYDILKTKSKISSVDEG 304
Query: 287 KKTIQNSLCRKKVLLVLDDVNELNQLENLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKT 346
I++ ++VL+++D+++E+ QL+ +VG DWFGPGSR+IITTRD+HLL V KT
Sbjct: 305 IGLIEDQFRHRRVLVIMDNIDEVGQLDAIVGNPDWFGPGSRIIITTRDEHLLKQ--VDKT 362
Query: 347 YKTGMLCKHDALVLFCLKAFKGDKPQEGYLDLSKEVVDYCGGLPLALEVLGSYLYGRNID 406
Y L + +AL LF AF + P E YL+LS++VV YCGGLPLALEVLGS+L+ R I
Sbjct: 363 YVAQKLDEREALELFSWHAFGNNWPNEEYLELSEKVVSYCGGLPLALEVLGSFLFKRPIA 422
Query: 407 VWHSAVKKLRSFPHPRVQDNLKISYDSLDTMEKDIFLDIACFFKGMKGDKVIDILESCGY 466
W S ++KL+ P ++ +L+IS++ LD +K IFLDI+CFF G D V +L+ CG+
Sbjct: 423 EWKSQLEKLKRTPEGKIIKSLRISFEGLDDAQKAIFLDISCFFIGEDKDYVAKVLDGCGF 482
Query: 467 FPQIGIQILIERSLITLDSVNNKLGMHDLLQEMGRDIVFQESPNDPCRRSRLWSQEDIDR 526
+ IGI +L ER L+T++ +NKL MHDLL+EM + I+ ++SP DP + SRLW + ++
Sbjct: 483 YATIGISVLRERCLVTVE--HNKLNMHDLLREMAKVIISEKSPGDPGKWSRLWDKREVIN 540
Query: 527 VLTKNKGTEAINSIDMKLLQPYEAHWNTEAFSKTSQLKFLSLCEMQLPLGLSCLPSSLKV 586
VLT GTE + + + ++ ++TEAF+ +L+ L LC ++L LP L
Sbjct: 541 VLTNKSGTEEVEGLALPWGYRHDTAFSTEAFANLKKLRLLQLCRVELNGEYKHLPKELIW 600
Query: 587 LHWRGCPLKTLPIT-TQLDELVDITLSHSKIEQLWQGVKVIDSL 629
LHW CPLK++P D+LV + + SK+ Q+W+G K + +L
Sbjct: 601 LHWFECPLKSIPDDFFNQDKLVVLEMQWSKLVQVWEGSKSLHNL 644
>gb|AAP44392.1| nematode resistance-like protein [Solanum tuberosum]
Length = 1136
Score = 502 bits (1293), Expect = e-140
Identities = 274/616 (44%), Positives = 393/616 (63%), Gaps = 9/616 (1%)
Query: 19 TYHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQVISEKLINAIKDSMFAI 78
+Y VFLSFRGED RK F DHL AL++K I TFKDD+ LE+G+ IS +L+++I++S A+
Sbjct: 17 SYDVFLSFRGEDVRKTFVDHLYLALQQKCINTFKDDEKLEKGKFISPELMSSIEESRIAL 76
Query: 79 TILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVRHQRGCFEEAFRKHQEK 138
I S +YA+STWCLDEL IMEC + V+PVFY VDPS VR Q+ F EAF KH+ +
Sbjct: 77 IIFSKNYANSTWCLDELTKIMECKNVKGQIVVPVFYDVDPSTVRKQKSIFGEAFSKHEAR 136
Query: 139 FGQHSDRVDRWRDAFTQVASYSGWD---SKGQHEASLVENIAQHIHRKL-VPKLPSCTEN 194
F + D+V +WR A + A+ SGWD + HEA ++E IA+ I +L + S N
Sbjct: 137 FQE--DKVQKWRAALEEAANISGWDLPNTSNGHEARVMEKIAEDIMARLGSQRHASNARN 194
Query: 195 LVGIVSKVEEVNKFLGMGLNDVRFIGIWGMGGIGKSTIARAVYETIRCEFELTCFLENVR 254
LVG+ S + +V K LG+G V F+GI GM G+GK+T+AR +Y+ IR +F+ CFL VR
Sbjct: 195 LVGMESHMLKVYKMLGIGSGGVHFLGILGMSGVGKTTLARVIYDNIRSQFQGACFLHEVR 254
Query: 255 EISETNGLVHLQRQLLSH-LSISRNDFHDLYDGKKTIQNSLCRKKVLLVLDDVNELNQLE 313
+ S GL LQ LLS L + + ++ ++G + L KKVLLVLDDV+ ++QL
Sbjct: 255 DRSAKQGLERLQEILLSEILVVKKLRINNSFEGANMQKQRLQYKKVLLVLDDVDHIDQLN 314
Query: 314 NLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGDKPQE 373
L G+++WFG GSR+IITT+DKHLL+ + K Y+ L +++L LF AFK ++P +
Sbjct: 315 ALAGEREWFGDGSRIIITTKDKHLLVKYETEKIYRMKTLNNYESLQLFKQHAFKKNRPTK 374
Query: 374 GYLDLSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKLRSFPHPRVQDNLKISYDS 433
+ DLS +V+ + GLPLAL+VLGS+LYGR +D W S V++L+ P + L+ S+
Sbjct: 375 EFEDLSAQVIKHTDGLPLALKVLGSFLYGRGLDEWISEVERLKQIPENEILKKLEQSFTG 434
Query: 434 LDTMEKDIFLDIACFFKGMKGDKVIDILESCGYFPQIGIQILIERSLITLDSVNNKLGMH 493
L E+ IFLDIACFF G K D V ILES + P IGI++L+E+ LIT+ + ++ +H
Sbjct: 435 LHNTEQKIFLDIACFFSGKKKDSVTRILESFHFCPVIGIKVLMEKCLITI--LQGRITIH 492
Query: 494 DLLQEMGRDIVFQESPNDPCRRSRLWSQEDIDRVLTKNKGTEAINSIDMKLLQPYEAHWN 553
L+Q+MG IV +E+ +DP SRLW +EDI VL +N GT+ + + L E ++
Sbjct: 493 QLIQDMGWHIVRREATDDPRMCSRLWKREDICPVLERNLGTDKNEGMSLHLTNEEEVNFG 552
Query: 554 TEAFSKTSQLKFLSLCEMQLPLGLSCLPSSLKVLHWRGCPLKTLPITTQLDELVDITLSH 613
+AF + ++L+FL + G LP L+ L W G P K+LP + + D+LV + L
Sbjct: 553 GKAFMQMTRLRFLKFRNAYVCQGPEFLPDELRWLDWHGYPSKSLPNSFKGDQLVGLKLKK 612
Query: 614 SKIEQLWQGVKVIDSL 629
S+I QLW+ K + L
Sbjct: 613 SRIIQLWKTSKDLGKL 628
>ref|NP_197270.1| disease resistance protein (TIR-NBS-LRR class), putative
[Arabidopsis thaliana]
Length = 1294
Score = 491 bits (1264), Expect = e-137
Identities = 265/646 (41%), Positives = 398/646 (61%), Gaps = 20/646 (3%)
Query: 4 SSSSSCRSSSTTSLCTYHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQVI 63
+S S SSS++++ VF+SFRGED RK F HL +R GI F+DD DL+RG+ I
Sbjct: 2 ASLPSSSSSSSSTVWKTDVFVSFRGEDVRKTFVSHLFCEFDRMGIKAFRDDLDLQRGKSI 61
Query: 64 SEKLINAIKDSMFAITILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVRH 123
S +LI+AIK S FAI ++S +YA+S+WCLDEL IMEC+ ++P+FY VDPSDVR
Sbjct: 62 SPELIDAIKGSRFAIVVVSRNYAASSWCLDELLKIMECNKDT---IVPIFYEVDPSDVRR 118
Query: 124 QRGCFEEAFRKHQEKFGQHSDRVDRWRDAFTQVASYSGWDSKGQHEASLVENIAQHIHRK 183
QRG F E H +K ++V +W++A ++A+ SG DS+ ++ L++ I + I K
Sbjct: 119 QRGSFGEDVESHSDK-----EKVGKWKEALKKLAAISGEDSRNWDDSKLIKKIVKDISDK 173
Query: 184 LVPKLPSCTENLVGIVSKVEEVNKFLGMGLNDVRFIGIWGMGGIGKSTIARAVYETIRCE 243
LV ++ L+G+ S ++ + + + DVR +GIWGMGG+GK+TIA+ +Y + +
Sbjct: 174 LVSTSWDDSKGLIGMSSHMDFLQSMISIVDKDVRMLGIWGMGGVGKTTIAKYLYNQLSGQ 233
Query: 244 FELTCFLENVREISETNGLVHLQRQLLSHLSISRN-DFHDLYDGKKTIQNSLCRKKVLLV 302
F++ CF+ENV+E+ G+ LQ + L + R+ + I+ K V +V
Sbjct: 234 FQVHCFMENVKEVCNRYGVRRLQVEFLCRMFQERDKEAWSSVSCCNIIKERFRHKMVFIV 293
Query: 303 LDDVNELNQLENLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFC 362
LDDV+ QL LV + WFGPGSR+I+TTRD+HLL++HG++ YK L K +AL LFC
Sbjct: 294 LDDVDRSEQLNELVKETGWFGPGSRIIVTTRDRHLLLSHGINLVYKVKCLPKKEALQLFC 353
Query: 363 LKAFKGDKP-QEGYLDLSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKLRSFPHP 421
AF+ + G+ +LS + V+Y GLPLAL VLGS+LY R+ W S + +L+++PH
Sbjct: 354 NYAFREEIILPHGFEELSVQAVNYASGLPLALRVLGSFLYRRSQIEWESTLARLKTYPHS 413
Query: 422 RVQDNLKISYDSLDTMEKDIFLDIACFFKGMKGDKVIDILESCGYFPQIGIQILIERSLI 481
+ + L++SYD LD EK IFL I+CF+ + D V +L+ CGY +IGI IL E+SLI
Sbjct: 414 DIMEVLRVSYDGLDEQEKAIFLYISCFYNMKQVDYVRKLLDLCGYAAEIGITILTEKSLI 473
Query: 482 TLDSVNNKLGMHDLLQEMGRDIVFQESPNDPCRRSRLWSQEDIDRVLTKNKGTEAINSID 541
N + +HDLL++MGR++V Q++ N+P +R LW EDI +L++N GT+ + I
Sbjct: 474 V--ESNGCVKIHDLLEQMGRELVRQQAVNNPAQRLLLWDPEDICHLLSENSGTQLVEGIS 531
Query: 542 MKLLQPYEAHWNTEAFSKTSQLKFLSLCEM--------QLPLGLSCLPSSLKVLHWRGCP 593
+ L + E + AF S LK L+ ++ LP GLS LP L+ L W G P
Sbjct: 532 LNLSEISEVFASDRAFEGLSNLKLLNFYDLSFDGETRVHLPNGLSYLPRKLRYLRWDGYP 591
Query: 594 LKTLPITTQLDELVDITLSHSKIEQLWQGVKVIDSLTKNSIKAYHY 639
LKT+P + LV++ +S+S +E+LW G++ + +L K + Y
Sbjct: 592 LKTMPSRFFPEFLVELCMSNSNLEKLWDGIQPLRNLKKMDLSRCKY 637
>dbj|BAB09567.1| disease resistance protein-like [Arabidopsis thaliana]
Length = 1295
Score = 488 bits (1255), Expect = e-136
Identities = 265/647 (40%), Positives = 399/647 (60%), Gaps = 21/647 (3%)
Query: 4 SSSSSCRSSSTTSLCTYHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQVI 63
+S S SSS++++ VF+SFRGED RK F HL +R GI F+DD DL+RG+ I
Sbjct: 2 ASLPSSSSSSSSTVWKTDVFVSFRGEDVRKTFVSHLFCEFDRMGIKAFRDDLDLQRGKSI 61
Query: 64 SEKLINAIKDSMFAITILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVRH 123
S +LI+AIK S FAI ++S +YA+S+WCLDEL IMEC+ ++P+FY VDPSDVR
Sbjct: 62 SPELIDAIKGSRFAIVVVSRNYAASSWCLDELLKIMECNKDT---IVPIFYEVDPSDVRR 118
Query: 124 QRGCFEEAFRKHQEKFGQHSDRVDRWRDAFTQVASYSGWDSKG-QHEASLVENIAQHIHR 182
QRG F E H +K ++V +W++A ++A+ SG DS+ + ++ L++ I + I
Sbjct: 119 QRGSFGEDVESHSDK-----EKVGKWKEALKKLAAISGEDSRNWRDDSKLIKKIVKDISD 173
Query: 183 KLVPKLPSCTENLVGIVSKVEEVNKFLGMGLNDVRFIGIWGMGGIGKSTIARAVYETIRC 242
KLV ++ L+G+ S ++ + + + DVR +GIWGMGG+GK+TIA+ +Y +
Sbjct: 174 KLVSTSWDDSKGLIGMSSHMDFLQSMISIVDKDVRMLGIWGMGGVGKTTIAKYLYNQLSG 233
Query: 243 EFELTCFLENVREISETNGLVHLQRQLLSHLSISRN-DFHDLYDGKKTIQNSLCRKKVLL 301
+F++ CF+ENV+E+ G+ LQ + L + R+ + I+ K V +
Sbjct: 234 QFQVHCFMENVKEVCNRYGVRRLQVEFLCRMFQERDKEAWSSVSCCNIIKERFRHKMVFI 293
Query: 302 VLDDVNELNQLENLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLF 361
VLDDV+ QL LV + WFGPGSR+I+TTRD+HLL++HG++ YK L K +AL LF
Sbjct: 294 VLDDVDRSEQLNELVKETGWFGPGSRIIVTTRDRHLLLSHGINLVYKVKCLPKKEALQLF 353
Query: 362 CLKAFKGDKP-QEGYLDLSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKLRSFPH 420
C AF+ + G+ +LS + V+Y GLPLAL VLGS+LY R+ W S + +L+++PH
Sbjct: 354 CNYAFREEIILPHGFEELSVQAVNYASGLPLALRVLGSFLYRRSQIEWESTLARLKTYPH 413
Query: 421 PRVQDNLKISYDSLDTMEKDIFLDIACFFKGMKGDKVIDILESCGYFPQIGIQILIERSL 480
+ + L++SYD LD EK IFL I+CF+ + D V +L+ CGY +IGI IL E+SL
Sbjct: 414 SDIMEVLRVSYDGLDEQEKAIFLYISCFYNMKQVDYVRKLLDLCGYAAEIGITILTEKSL 473
Query: 481 ITLDSVNNKLGMHDLLQEMGRDIVFQESPNDPCRRSRLWSQEDIDRVLTKNKGTEAINSI 540
I N + +HDLL++MGR++V Q++ N+P +R LW EDI +L++N GT+ + I
Sbjct: 474 IV--ESNGCVKIHDLLEQMGRELVRQQAVNNPAQRLLLWDPEDICHLLSENSGTQLVEGI 531
Query: 541 DMKLLQPYEAHWNTEAFSKTSQLKFLSLCEM--------QLPLGLSCLPSSLKVLHWRGC 592
+ L + E + AF S LK L+ ++ LP GLS LP L+ L W G
Sbjct: 532 SLNLSEISEVFASDRAFEGLSNLKLLNFYDLSFDGETRVHLPNGLSYLPRKLRYLRWDGY 591
Query: 593 PLKTLPITTQLDELVDITLSHSKIEQLWQGVKVIDSLTKNSIKAYHY 639
PLKT+P + LV++ +S+S +E+LW G++ + +L K + Y
Sbjct: 592 PLKTMPSRFFPEFLVELCMSNSNLEKLWDGIQPLRNLKKMDLSRCKY 638
>gb|AAU04761.1| MRGH13 [Cucumis melo]
Length = 1024
Score = 469 bits (1207), Expect = e-130
Identities = 258/619 (41%), Positives = 384/619 (61%), Gaps = 11/619 (1%)
Query: 17 LCTYHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQV---ISEKLINAIKD 73
L Y VFLS R +DT + F L AL +GI F+DD D E G+ + EK+ A+++
Sbjct: 35 LRNYDVFLSHRAKDTGQSFAADLHEALTSQGIVVFRDDVDEEDGEKPYGVEEKM-KAVEE 93
Query: 74 SMFAITILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVRHQRGCFEEAFR 133
S +I + S +Y S C+ E+ I C + VLP+FY +DP +VR Q G FE+ F
Sbjct: 94 SRSSIVVFSENYGSFV-CMKEVGKIAMCKELMDQLVLPIFYKIDPGNVRKQEGNFEKYFN 152
Query: 134 KHQEKFGQHSDRVDRWRDAFTQVASYSGW---DSKGQHEASLVENIAQHIHRKLVPKLPS 190
+H+ + V+ WR + QV SGW DS+ + E S+++ + +HI KL P L
Sbjct: 153 EHEANPKIDIEEVENWRYSMNQVGHLSGWHVQDSQSE-EGSIIDEVVKHIFNKLRPDLFR 211
Query: 191 CTENLVGIVSKVEEVNKFLGMGLNDVRFIGIWGMGGIGKSTIARAVYETIRCEFELTCFL 250
+ LVGI ++ ++N LG+GL+DVRF+GIWGMGGIGK+T+AR +Y+++ F+ FL
Sbjct: 212 YDDKLVGITPRLHQINMLLGIGLDDVRFVGIWGMGGIGKTTLARIIYKSVSHLFDGCYFL 271
Query: 251 ENVREISETNGLVHLQRQLLSHLSISRNDFHDLYDGKKTIQNSLCRKKVLLVLDDVNELN 310
+NV+E + + LQ++L++ + RN DG I+ + + K L++LDDVN L+
Sbjct: 272 DNVKEALKKEDIASLQQKLITGTLMKRNIDIPNADGATLIKRRISKIKALIILDDVNHLS 331
Query: 311 QLENLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGDK 370
QL+ L G DWFG GSRVI+TTRD+HLL++HG+ + Y +L + L LF KAF +
Sbjct: 332 QLQKLAGGLDWFGSGSRVIVTTRDEHLLISHGIERRYNVEVLKIEEGLQLFSQKAFGEEH 391
Query: 371 PQEGYLDLSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKLRSFPHPRVQDNLKIS 430
P+E Y DL +VV+Y GGLPLA+EVLGS L+ + ++ W +AV+KL + + LKIS
Sbjct: 392 PKEEYFDLCSQVVNYAGGLPLAIEVLGSSLHNKPMEDWINAVEKLWEVRDKEIIEKLKIS 451
Query: 431 YDSLDTMEKDIFLDIACFFKGMKGDKVIDILESCGYFPQIGIQILIERSLITLDSVNNKL 490
Y L+ E+ IFLDIACFFK ++ I+ILES G+ +G++IL E+ LIT + ++KL
Sbjct: 452 YYMLEESEQKIFLDIACFFKRKSKNQAIEILESFGFPAVLGLEILEEKCLIT--APHDKL 509
Query: 491 GMHDLLQEMGRDIVFQESPNDPCRRSRLWSQEDIDRVLTKNKGTEAINSIDMKLLQPYEA 550
+HDL+QEMG++IV PN+P +R+RLW +EDI+ L++++GTEAI I M + E+
Sbjct: 510 QIHDLIQEMGQEIVRHTFPNEPEKRTRLWLREDINLALSRDQGTEAIEGIMMDFDEEGES 569
Query: 551 HWNTEAFSKTSQLKFLSLCEMQLPLGLSCLPSSLKVLHWRGCPLKTLPITTQLDELVDIT 610
H N +AFS + L+ L L + L + L L+ L+W G PLKTLP L+++
Sbjct: 570 HLNAKAFSSMTNLRVLKLNNVHLCEEIEYLSDQLRFLNWHGYPLKTLPSNFNPTNLLELE 629
Query: 611 LSHSKIEQLWQGVKVIDSL 629
L +S I LW K +++L
Sbjct: 630 LPNSSIHLLWTTSKSMETL 648
>gb|AAT37497.1| N-like protein [Nicotiana tabacum]
Length = 941
Score = 468 bits (1204), Expect = e-130
Identities = 259/633 (40%), Positives = 395/633 (61%), Gaps = 21/633 (3%)
Query: 11 SSSTTSLCTYHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQVISEKLINA 70
SSS++S +Y VFLSFRGEDTRK FT HL L+ +GI TF+D+K LE G I E+L A
Sbjct: 3 SSSSSSRWSYDVFLSFRGEDTRKTFTSHLYEVLKDRGIKTFQDEKRLEYGATIPEELCKA 62
Query: 71 IKDSMFAITILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVRHQRGCFEE 130
I++S FAI + S +YA+S WCL+EL IMEC ++ ++P+FY VDPS VR+Q+ F +
Sbjct: 63 IEESQFAIVVFSENYATSRWCLNELVKIMECKTQFRQTIIPIFYDVDPSHVRNQKESFAK 122
Query: 131 AFRKHQEKFGQHSDRVDRWRDAFTQVASYSG-WDSKGQHEASLVENIAQHIHRKLVPKLP 189
AF +H+ K+ + + RWR A A+ G D++ + +A + I I KL
Sbjct: 123 AFEEHETKYKDDVEGIQRWRTALNAAANLKGSCDNRDKTDADCIRQIVDQISSKLSKISL 182
Query: 190 SCTENLVGIVSKVEEVNKFLGMGLNDVRFIGIWGMGGIGKSTIARAVYETI------RCE 243
S +N+VGI + +EE+ LG+G+NDVR +GIWGMGG+GK+TIARA+++T+ +
Sbjct: 183 SYLQNIVGIDTHLEEIESLLGIGINDVRIVGIWGMGGVGKTTIARAMFDTLLGRRDSSYQ 242
Query: 244 FELTCFLENVREISETNGLVHLQRQLLSHLSISRNDFHDLYDGKKTIQNSLCRKKVLLVL 303
F+ CFL++++E G+ LQ LL L ++++ DGK + + L KKVL+VL
Sbjct: 243 FDGACFLKDIKE--NKRGMHSLQNTLLFELLRENANYNNEDDGKHQMASRLRSKKVLIVL 300
Query: 304 DDVNELNQ-LENLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFC 362
DD+++ + LE L G DWFG GSR+I+TTRDKHL+ + + Y+ L H+A+ LF
Sbjct: 301 DDIDDKDHYLEYLAGDLDWFGNGSRIIVTTRDKHLIGKNDI--IYEVTALPDHEAIQLFY 358
Query: 363 LKAFKGDKPQEGYLDLSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKLRSFPHPR 422
AFK + P E + +LS EVV++ GLPLAL+V GS L+ R+I VW SA+++++ P+ +
Sbjct: 359 QHAFKKEVPDECFKELSLEVVNHAKGLPLALKVWGSSLHKRDITVWKSAIEQMKINPNSK 418
Query: 423 VQDNLKISYDSLDTMEKDIFLDIACFFKGMKGDKVIDILESCGYFPQIGIQILIERSLIT 482
+ + LKISYD L++M++++FLDIACFF+G + D ++ +L+SC + + G+ +LIE+SL+
Sbjct: 419 IVEKLKISYDGLESMQQEMFLDIACFFRGRQKDYIMQVLKSCHFGAEYGLDVLIEKSLVF 478
Query: 483 LDSVNNKLGMHDLLQEMGRDIVFQESPNDPCRRSRLWSQEDIDRVLTKNKGTEAINSIDM 542
+ S N++ MHDL+Q+MG+ IV DP RSRLW ED++ V+ N GT ++ I +
Sbjct: 479 I-SEYNQVEMHDLIQDMGKYIV--NFKKDPGERSRLWLAEDVEEVMNNNAGTMSVEVIWV 535
Query: 543 KLLQPYEAHWNTEAFSKTSQLKFLS----LCEMQLPLGLSCLPSSLKVLHWRGCPLKTLP 598
+ +++ +A +L+ L L + LPS+L+ P ++LP
Sbjct: 536 H--YDFGLYFSNDAMKNMKRLRILHIKGYLSSTSHDGSIEYLPSNLRWFVLDDYPWESLP 593
Query: 599 ITTQLDELVDITLSHSKIEQLWQGVKVIDSLTK 631
T L LV + LS S + LW K + SL +
Sbjct: 594 STFDLKMLVHLELSRSSLHYLWTETKHLPSLRR 626
>gb|AAQ93074.1| putative TIR-NBS type R protein 4 [Malus baccata]
Length = 726
Score = 459 bits (1180), Expect = e-127
Identities = 260/616 (42%), Positives = 372/616 (60%), Gaps = 15/616 (2%)
Query: 3 ASSSSSCRSSSTTSLCTYHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQV 62
+SSSS+ S+S++ Y VF+SFRGEDTRK FT HL AL + GI F DD++L RG+
Sbjct: 108 SSSSSASPSASSSKGSLYEVFISFRGEDTRKNFTGHLHEALTKAGINAFIDDEELRRGED 167
Query: 63 ISEKLINAIKDSMFAITILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVR 122
I+ +L+ AI+ S +I + S YA S+WCL+EL IMEC VLP+FY VDPS+VR
Sbjct: 168 ITTELVQAIQGSRISIIVFSRRYADSSWCLEELVKIMECRRTLGQLVLPIFYDVDPSNVR 227
Query: 123 HQRGCFEEAFRKHQEKFGQHSDRVDRWRDAFTQVASYSGWDSKG---QHEASLVENIAQH 179
G F ++F KH ++ +V+RWR A T+ ++ SGWD K +HEA + I
Sbjct: 228 KLTGSFAQSFLKHTDE-----KKVERWRAALTEASNLSGWDLKNTLDRHEAKFIRMITNQ 282
Query: 180 IHRKLVPKLPSCTENLVGIVSKVEEVNKFLGMG-LNDVRFIGIWGMGGIGKSTIARAVYE 238
+ KL + + VGI ++V ++ +LG+G +DVR IGI GMGGIGK+TI +A+Y
Sbjct: 283 VTVKLNNRYFNVAPYQVGIDTRVLNISNYLGIGDSDDVRVIGISGMGGIGKTTIVKAIYN 342
Query: 239 TIRCEFELTCFLENVREISETNGLVHLQRQLLSHLSISRNDFHDLYDGKKTIQNSLCRKK 298
FE FLE VRE LV LQ+QLL + ++ + G + R +
Sbjct: 343 EFYERFEGKSFLEKVRE----KKLVKLQKQLLFDILQTKTKVSSVAVGTALVGERFRRLR 398
Query: 299 VLLVLDDVNELNQLENLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDAL 358
VL+++DDV+++ QL LVG FGPGSR+IITTR++ +L V + Y+ + + +AL
Sbjct: 399 VLVIVDDVDDVKQLRELVGNCHSFGPGSRIIITTRNERVLKEFAVDEIYRENGMDQEEAL 458
Query: 359 VLFCLKAFKGDKPQEGYLDLSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKLRSF 418
L AFK YL L++EVV+YCGGLPLALEVLGS ++ R+++ W S + +L+
Sbjct: 459 ELLSWHAFKSSWCPSQYLVLTREVVNYCGGLPLALEVLGSTIFKRSVNEWRSILDELKMI 518
Query: 419 PHPRVQDNLKISYDSL-DTMEKDIFLDIACFFKGMKGDKVIDILESCGYFPQIGIQILIE 477
P +Q LKISYD L D ++ IFLDIA FF GM + V+ IL+ CG++ GI++L++
Sbjct: 519 PRGEIQAQLKISYDGLNDHYKRQIFLDIAFFFIGMDKNDVMQILDGCGFYATTGIEVLLD 578
Query: 478 RSLITLDSVNNKLGMHDLLQEMGRDIVFQESPNDPCRRSRLWSQEDIDRVLTKNKGTEAI 537
R L+T+ NK+ MHDLL++MGRDIV E+P P RSRLW +D+ VL GTE I
Sbjct: 579 RCLVTIGR-KNKIMMHDLLRDMGRDIVHAENPGFPRERSRLWHPKDVHDVLIDKSGTEKI 637
Query: 538 NSIDMKLLQPYEAHWNTEAFSKTSQLKFLSLCEMQLPLGLSCLPSSLKVLHWRGCPLKTL 597
+ + L E ++T+AF +L+ L L ++L G CL L+ L W G PL+ +
Sbjct: 638 EGLALNLPSLEETSFSTDAFRNMKRLRLLQLNYVRLTGGYRCLSKKLRWLCWHGFPLEFI 697
Query: 598 PITTQLDELVDITLSH 613
PI +V I + +
Sbjct: 698 PIELCQPNIVAIDMQY 713
>gb|AAU04762.1| MRGH21 [Cucumis melo]
Length = 1020
Score = 456 bits (1172), Expect = e-126
Identities = 254/619 (41%), Positives = 372/619 (60%), Gaps = 13/619 (2%)
Query: 20 YHVFLSFRGEDTR------KGFTDHLCAALERKGITTFKDDKDLERGQVISEKLINAIKD 73
Y VFLS R +D R + F L AL +GI F D +D E G + + A+ +
Sbjct: 34 YDVFLSHRAKDHRANNDTGRSFISDLHEALTSQGIVVFIDKEDEEDGGKPLTEKMKAVDE 93
Query: 74 SMFAITILSPDYASSTW-CLDELQMIMECSSKNNLHVLPVFYGVDPSDVRHQRG-CFEEA 131
S +I + S +Y S W C+ E++ I C + VLP+FY VDP DVR Q G +
Sbjct: 94 SRSSIVVFSENYGS--WVCMKEIRKIRMCQKSRDQLVLPIFYKVDPGDVRKQEGESLVKF 151
Query: 132 FRKHQEKFGQHSDRVDRWRDAFTQVASYSGWDSK-GQHEASLVENIAQHIHRKLVPKLPS 190
F +H+ + V +WR + +V + SGW + Q E +++ + HI KL P L
Sbjct: 152 FNEHEANPNISIEEVKKWRKSMNKVGNLSGWHLQDSQFEEGIIKEVVDHIFNKLRPDLFR 211
Query: 191 CTENLVGIVSKVEEVNKFLGMGLNDVRFIGIWGMGGIGKSTIARAVYETIRCEFELTCFL 250
+ LVGI ++ E+NK +G+GL+DVRFIGIWGM GIGK+TIAR +Y+++ F+ FL
Sbjct: 212 YDDKLVGISRRLHEINKLMGIGLDDVRFIGIWGMSGIGKTTIARIIYKSVSHLFDGCYFL 271
Query: 251 ENVREISETNGLVHLQRQLLSHLSISRNDFHDLYDGKKTIQNSLCRKKVLLVLDDVNELN 310
+NV+E + G+ LQ++LL+ + RN DG I+ + K L++LDDV+ ++
Sbjct: 272 DNVKEALKKEGIASLQQKLLTGALMKRNIDIPNADGATLIKRRISNIKALIILDDVDNVS 331
Query: 311 QLENLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGDK 370
QL L G DWFG GSRVI+TT+ + +L++HG+ + Y +L + + LF KAF D
Sbjct: 332 QLRQLAGSLDWFGSGSRVIVTTKHEDILVSHGIERRYNVEVLKIDEGIQLFSQKAFGEDY 391
Query: 371 PQEGYLDLSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKLRSFPHPRVQDNLKIS 430
P+EGY DL +VVDY GGLPLA+EVLGS L + ++ W AVKKL + + LKIS
Sbjct: 392 PKEGYFDLCSQVVDYAGGLPLAIEVLGSSLRNKPMEDWIDAVKKLWEVRDKEINEKLKIS 451
Query: 431 YDSLDTMEKDIFLDIACFFKGMKGDKVIDILESCGYFPQIGIQILIERSLITLDSVNNKL 490
Y L+ +++IFLDIACFFK + I+ILES G+ +G+ IL E+SLIT + + K+
Sbjct: 452 YYMLENDDREIFLDIACFFKRKSKRRAIEILESFGFPAVLGLDILKEKSLIT--TPHEKI 509
Query: 491 GMHDLLQEMGRDIVFQESPNDPCRRSRLWSQEDIDRVLTKNKGTEAINSIDMKLLQPYEA 550
MHDL+QEMG+ IV +E P++P +RSRLW +EDI+R L++++GTE I I M L + E+
Sbjct: 510 QMHDLIQEMGQKIVNEEFPDEPEKRSRLWLREDINRALSRDQGTEEIEGIMMDLDEEGES 569
Query: 551 HWNTEAFSKTSQLKFLSLCEMQLPLGLSCLPSSLKVLHWRGCPLKTLPITTQLDELVDIT 610
H N ++FS + L+ L L + L + L L+ L+W G PLKTLP L+++
Sbjct: 570 HLNAKSFSSMTNLRVLKLNNVHLCEEIEYLSDQLRFLNWHGYPLKTLPSNFNPTNLLELE 629
Query: 611 LSHSKIEQLWQGVKVIDSL 629
L +S I LW K +++L
Sbjct: 630 LPNSSIHLLWTTSKSMETL 648
>gb|AAQ93076.1| putative TIR-NBS type R protein 4 [Malus baccata]
Length = 726
Score = 454 bits (1169), Expect = e-126
Identities = 258/616 (41%), Positives = 370/616 (59%), Gaps = 15/616 (2%)
Query: 3 ASSSSSCRSSSTTSLCTYHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQV 62
+SSSS+ +S++ Y VF+SFRGEDTRK FT HL AL + GI F DD++L RG+
Sbjct: 108 SSSSSASPPASSSKGSLYEVFISFRGEDTRKNFTGHLHEALTKAGINAFIDDEELRRGED 167
Query: 63 ISEKLINAIKDSMFAITILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVR 122
I+ +L+ AI+ S +I + S YA S+WCL+EL IMEC VLP+FY VDPS+VR
Sbjct: 168 ITTELVQAIQGSRISIIVFSRRYADSSWCLEELVKIMECRRTLGQLVLPIFYDVDPSNVR 227
Query: 123 HQRGCFEEAFRKHQEKFGQHSDRVDRWRDAFTQVASYSGWDSKG---QHEASLVENIAQH 179
G F ++F KH ++ +V+RWR A T+ ++ SGWD K +HEA + I
Sbjct: 228 KLTGSFAQSFLKHTDE-----KKVERWRAALTEASNLSGWDLKNTLDRHEAKFIRMITNQ 282
Query: 180 IHRKLVPKLPSCTENLVGIVSKVEEVNKFLGMG-LNDVRFIGIWGMGGIGKSTIARAVYE 238
+ KL + + VGI ++V ++ +LG+G +DVR IGI G GGIGK+TI +A+Y
Sbjct: 283 VTVKLNNRYFNVAPYQVGIDTRVLNISNYLGIGDSDDVRVIGISGSGGIGKTTIVKAIYN 342
Query: 239 TIRCEFELTCFLENVREISETNGLVHLQRQLLSHLSISRNDFHDLYDGKKTIQNSLCRKK 298
FE FLE VRE LV LQ+QLL + ++ + G + R +
Sbjct: 343 EFYERFEGKSFLEKVRE----KKLVKLQKQLLFDILQTKTKVSSVAVGTALVGERFRRLR 398
Query: 299 VLLVLDDVNELNQLENLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDAL 358
VL+++DDV+++ QL LVG FGPGSR+IITTR++ +L V + Y+ + + +AL
Sbjct: 399 VLVIVDDVDDVKQLRELVGNCHSFGPGSRIIITTRNERVLKEFAVDEIYRENGMDQEEAL 458
Query: 359 VLFCLKAFKGDKPQEGYLDLSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKLRSF 418
L AFK YL L++EVV+YCGGLPLALEVLGS ++ R+++ W S + +L+
Sbjct: 459 ELLSWHAFKSSWCPSQYLVLTREVVNYCGGLPLALEVLGSTIFKRSVNEWRSILDELKMI 518
Query: 419 PHPRVQDNLKISYDSL-DTMEKDIFLDIACFFKGMKGDKVIDILESCGYFPQIGIQILIE 477
P +Q LKISYD L D ++ IFLDIA FF GM + V+ IL+ CG++ GI++L++
Sbjct: 519 PRGEIQAQLKISYDGLNDHYKRQIFLDIAFFFIGMDKNDVMQILDGCGFYATTGIEVLLD 578
Query: 478 RSLITLDSVNNKLGMHDLLQEMGRDIVFQESPNDPCRRSRLWSQEDIDRVLTKNKGTEAI 537
R L+T+ NK+ MHDLL++MGRDIV E+P P RSRLW +D+ VL GTE I
Sbjct: 579 RCLVTIGR-KNKIMMHDLLRDMGRDIVHAENPGFPRERSRLWHPKDVHDVLIDKSGTEKI 637
Query: 538 NSIDMKLLQPYEAHWNTEAFSKTSQLKFLSLCEMQLPLGLSCLPSSLKVLHWRGCPLKTL 597
+ + L E ++T+AF +L+ L L ++L G CL L+ L W G PL+ +
Sbjct: 638 EGLALNLPSLEETSFSTDAFRNMKRLRLLQLNYVRLTGGYRCLSKKLRWLCWHGFPLEFI 697
Query: 598 PITTQLDELVDITLSH 613
PI +V I + +
Sbjct: 698 PIELCQPNIVAIDMQY 713
>sp|Q40392|TMVRN_NICGU TMV resistance protein N gi|558887|gb|AAA50763.1| N
Length = 1144
Score = 453 bits (1165), Expect = e-126
Identities = 253/630 (40%), Positives = 387/630 (61%), Gaps = 18/630 (2%)
Query: 11 SSSTTSLCTYHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQVISEKLINA 70
SSS++S +Y VFLSFRGEDTRK FT HL L KGI TF+DDK LE G I +L A
Sbjct: 3 SSSSSSRWSYDVFLSFRGEDTRKTFTSHLYEVLNDKGIKTFQDDKRLEYGATIPGELCKA 62
Query: 71 IKDSMFAITILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVRHQRGCFEE 130
I++S FAI + S +YA+S WCL+EL IMEC ++ V+P+FY VDPS VR+Q+ F +
Sbjct: 63 IEESQFAIVVFSENYATSRWCLNELVKIMECKTRFKQTVIPIFYDVDPSHVRNQKESFAK 122
Query: 131 AFRKHQEKFGQHSDRVDRWRDAFTQVASYSG-WDSKGQHEASLVENIAQHIHRKLVPKLP 189
AF +H+ K+ + + RWR A + A+ G D++ + +A + I I KL
Sbjct: 123 AFEEHETKYKDDVEGIQRWRIALNEAANLKGSCDNRDKTDADCIRQIVDQISSKLCKISL 182
Query: 190 SCTENLVGIVSKVEEVNKFLGMGLNDVRFIGIWGMGGIGKSTIARAVYETI------RCE 243
S +N+VGI + +E++ L +G+N VR +GIWGMGG+GK+TIARA+++T+ +
Sbjct: 183 SYLQNIVGIDTHLEKIESLLEIGINGVRIMGIWGMGGVGKTTIARAIFDTLLGRMDSSYQ 242
Query: 244 FELTCFLENVREISETNGLVHLQRQLLSHLSISRNDFHDLYDGKKTIQNSLCRKKVLLVL 303
F+ CFL++++E G+ LQ LLS L + ++++ DGK + + L KKVL+VL
Sbjct: 243 FDGACFLKDIKE--NKRGMHSLQNALLSELLREKANYNNEEDGKHQMASRLRSKKVLIVL 300
Query: 304 DDV-NELNQLENLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFC 362
DD+ N+ + LE L G DWFG GSR+IITTRDKHL+ + + Y+ L H+++ LF
Sbjct: 301 DDIDNKDHYLEYLAGDLDWFGNGSRIIITTRDKHLIEKNDI--IYEVTALPDHESIQLFK 358
Query: 363 LKAFKGDKPQEGYLDLSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKLRSFPHPR 422
AF + P E + LS EVV+Y GLPLAL+V GS L+ + W SA++ +++ +
Sbjct: 359 QHAFGKEVPNENFEKLSLEVVNYAKGLPLALKVWGSLLHNLRLTEWKSAIEHMKNNSYSG 418
Query: 423 VQDNLKISYDSLDTMEKDIFLDIACFFKGMKGDKVIDILESCGYFPQIGIQILIERSLIT 482
+ D LKISYD L+ ++++FLDIACF +G + D ++ ILESC + G++ILI++SL+
Sbjct: 419 IIDKLKISYDGLEPKQQEMFLDIACFLRGEEKDYILQILESCHIGAEYGLRILIDKSLVF 478
Query: 483 LDSVNNKLGMHDLLQEMGRDIV-FQESPNDPCRRSRLWSQEDIDRVLTKNKGTEAINSID 541
+ S N++ MHDL+Q+MG+ IV FQ+ DP RSRLW ++++ V++ N GT A+ +I
Sbjct: 479 I-SEYNQVQMHDLIQDMGKYIVNFQK---DPGERSRLWLAKEVEEVMSNNTGTMAMEAIW 534
Query: 542 MKLLQPYEAHWNTEAFSKTSQLKFLSLCEMQLPLGLSCLPSSLKVLHWRGCPLKTLPITT 601
+ ++ +A +L+ ++ + LP++L+ P ++ P T
Sbjct: 535 VSSYSS-TLRFSNQAVKNMKRLRVFNMGRSSTHYAIDYLPNNLRCFVCTNYPWESFPSTF 593
Query: 602 QLDELVDITLSHSKIEQLWQGVKVIDSLTK 631
+L LV + L H+ + LW K + SL +
Sbjct: 594 ELKMLVHLQLRHNSLRHLWTETKHLPSLRR 623
>gb|AAO92748.1| candidate disease-resistance protein SR1 [Glycine max]
Length = 1137
Score = 453 bits (1165), Expect = e-126
Identities = 272/616 (44%), Positives = 374/616 (60%), Gaps = 14/616 (2%)
Query: 11 SSSTTSLCT-YHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQVISEKLIN 69
+++T SL + Y VFLSF G+DTR GFT +L AL+ +GI TF DD++L RG I L +
Sbjct: 2 AATTRSLASIYDVFLSFTGQDTRHGFTGYLYKALDDRGIYTFIDDQELPRGDEIKPALSD 61
Query: 70 AIKDSMFAITILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVRHQRGCFE 129
AI+ S AIT+LS +YA ST+CLDEL I+ C S+ L V+PVFY VDPS VRHQ+G +
Sbjct: 62 AIQGSRIAITVLSQNYAFSTFCLDELVTILHCKSEGLL-VIPVFYKVDPSHVRHQKGSYG 120
Query: 130 EAFRKHQEKFGQHSDRVDRWRDAFTQVASYSGWDSKG--QHEASLVENIAQHIHRKLVPK 187
EA KHQ++F + +++ +WR A QVA SG+ K +E +++I + + R++
Sbjct: 121 EAMAKHQKRFKANKEKLQKWRMALQQVADLSGYHFKDGDAYEYKFIQSIVEQVSREINRA 180
Query: 188 LPSCTENLVGIVSKVEEVNKFLGMGLNDV-RFIGIWGMGGIGKSTIARAVYETIRCEFEL 246
+ VG+ S+V EV K L +G +DV IGI GMGG+GK+T+A AVY I F+
Sbjct: 181 PLHVADYPVGLGSQVIEVRKLLDVGSDDVVHIIGIHGMGGLGKTTLAVAVYNLIAPHFDE 240
Query: 247 TCFLENVREISETNGLVHLQRQLLSHLSISRN-DFHDLYDGKKTIQNSLCRKKVLLVLDD 305
+CFL+NVRE S L HLQ LLS L ++ +G IQ+ L RKKVLL+LDD
Sbjct: 241 SCFLQNVREESN---LKHLQSSLLSKLLGEKDITLTSWQEGASMIQHRLRRKKVLLILDD 297
Query: 306 VNELNQLENLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKA 365
V++ QL+ +VGK DWFGPGSRVIITTRDKHLL H V +TY+ +L + AL L A
Sbjct: 298 VDKREQLKAIVGKPDWFGPGSRVIITTRDKHLLKYHEVERTYEVKVLNHNAALHLLTWNA 357
Query: 366 FKGDKPQEGYLDLSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKLRSFPHPRVQD 425
FK +K Y D+ VV Y GLPLALEV+GS LYG+ + W SA++ + P +
Sbjct: 358 FKREKIDPIYDDVLNRVVTYASGLPLALEVIGSNLYGKTVAEWESALETYKRIPSNEILK 417
Query: 426 NLKISYDSLDTMEKDIFLDIACFFKGMKGDKVIDILESC-GYFPQIGIQILIERSLITLD 484
L++S+D+L+ ++++FLDIAC FKG + +V DI + G + I +L+E+SLI +
Sbjct: 418 ILQVSFDALEEEQQNVFLDIACCFKGHEWTEVDDIFRALYGNGKKYHIGVLVEKSLIKYN 477
Query: 485 SVN-NKLGMHDLLQEMGRDIVFQESPNDPCRRSRLWSQEDIDRVLTKNKGTEAINSI--D 541
N + MH+L+Q+MGR+I Q SP +P +R RLWS +DI +VL N GT I I D
Sbjct: 478 RNNRGTVQMHNLIQDMGREIERQRSPEEPGKRKRLWSPKDIIQVLKHNTGTSKIEIICLD 537
Query: 542 MKLLQPYE-AHWNTEAFSKTSQLKFLSLCEMQLPLGLSCLPSSLKVLHWRGCPLKTLPIT 600
+ E WN AF K LK L + + +G + +P L+VL W P LP
Sbjct: 538 SSISDKEETVEWNENAFMKMENLKILIIRNGKFSIGPNYIPEGLRVLEWHRYPSNCLPSN 597
Query: 601 TQLDELVDITLSHSKI 616
LV L S I
Sbjct: 598 FDPINLVICKLPDSSI 613
>gb|AAO23076.1| R 1 protein [Glycine max]
Length = 902
Score = 453 bits (1165), Expect = e-126
Identities = 274/616 (44%), Positives = 369/616 (59%), Gaps = 11/616 (1%)
Query: 11 SSSTTSLCT-YHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQVISEKLIN 69
+++T SL + Y VFL+FRGEDTR GFT +L AL KGI TF D+ L G I+ L
Sbjct: 2 AATTRSLASIYDVFLNFRGEDTRYGFTGNLYKALCDKGIHTFFDEDKLHSGDDITPALSK 61
Query: 70 AIKDSMFAITILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVRHQRGCFE 129
AI++S AIT+LS +YASS++CLDEL I+ C + L V+PVF+ VDPS VRH +G +
Sbjct: 62 AIQESRIAITVLSQNYASSSFCLDELVTILHCK-REGLLVIPVFHNVDPSAVRHLKGSYG 120
Query: 130 EAFRKHQEKFGQHSDRVDRWRDAFTQVASYSGWDSKG--QHEASLVENIAQHIHRKLVPK 187
EA KHQ++F +++ +WR A QVA SG+ K +E + NI + + RK+
Sbjct: 121 EAMAKHQKRFKAKKEKLQKWRMALHQVADLSGYHFKDGDAYEYKFIGNIVEEVSRKINCA 180
Query: 188 LPSCTENLVGIVSKVEEVNKFLGMGLND-VRFIGIWGMGGIGKSTIARAVYETIRCEFEL 246
+ VG+ S+V EV K L +G +D V IGI GMGG+GK+T+A AVY I F+
Sbjct: 181 PLHVADYPVGLGSQVIEVMKLLDVGSDDLVHIIGIHGMGGLGKTTLALAVYNFIALHFDE 240
Query: 247 TCFLENVREISETNGLVHLQRQLLSHLSISRN-DFHDLYDGKKTIQNSLCRKKVLLVLDD 305
+CFL+NVRE S +GL H Q LLS L ++ +G IQ+ L RKKVLL+LDD
Sbjct: 241 SCFLQNVREESNKHGLKHFQSILLSKLLGEKDITLTSWQEGASMIQHRLRRKKVLLILDD 300
Query: 306 VNELNQLENLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKA 365
V++ QLE +VG+ DWFGPGSRVIITTRDKHLL H V +TY+ +L + AL L A
Sbjct: 301 VDKREQLEAIVGRSDWFGPGSRVIITTRDKHLLKYHEVERTYEVKVLNHNAALQLLTWNA 360
Query: 366 FKGDKPQEGYLDLSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKLRSFPHPRVQD 425
FK +K Y D+ VV Y GLPLALEV+GS L+G+ + W SAV+ + P +
Sbjct: 361 FKREKIDPIYDDVLNRVVTYASGLPLALEVIGSDLFGKTVAEWESAVEHYKRIPSDEILK 420
Query: 426 NLKISYDSLDTMEKDIFLDIACFFKGMKGDKVIDILES-CGYFPQIGIQILIERSLITLD 484
LK+S+D+L +K++FLDIAC FKG K +V DIL + G + I +L+E+SLI L+
Sbjct: 421 ILKVSFDALGEEQKNVFLDIACCFKGYKWTEVDDILRAFYGNCKKHHIGVLVEKSLIKLN 480
Query: 485 SVNN-KLGMHDLLQEMGRDIVFQESPNDPCRRSRLWSQEDIDRVLTKNKGTEAINSI--D 541
++ + MHDL+Q+MGR+I Q SP +P + RLWS +DI +VL N GT I I D
Sbjct: 481 CYDSGTVEMHDLIQDMGREIERQRSPEEPWKCKRLWSPKDIFQVLKHNTGTSKIEIICLD 540
Query: 542 MKLLQPYE-AHWNTEAFSKTSQLKFLSLCEMQLPLGLSCLPSSLKVLHWRGCPLKTLPIT 600
+ E WN AF K LK L + + G + P L VL W P LP
Sbjct: 541 FSISDKEETVEWNENAFMKMENLKILIIRNGKFSKGPNYFPEGLTVLEWHRYPSNCLPYN 600
Query: 601 TQLDELVDITLSHSKI 616
+ L+ L S I
Sbjct: 601 FHPNNLLICKLPDSSI 616
>gb|AAO23077.1| R 8 protein [Glycine max]
Length = 892
Score = 452 bits (1164), Expect = e-125
Identities = 271/615 (44%), Positives = 371/615 (60%), Gaps = 11/615 (1%)
Query: 11 SSSTTSLC-TYHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQVISEKLIN 69
+++T SL Y VFLSF G+DTR+GFT +L AL +GI TF DD++L RG I L N
Sbjct: 2 AATTRSLAYNYDVFLSFTGQDTRQGFTGYLYKALCDRGIYTFIDDQELRRGDEIKPALSN 61
Query: 70 AIKDSMFAITILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVRHQRGCFE 129
AI++S AIT+LS +YASS++CLDEL I+ C S+ L V+PVFY VDPS VRHQ+G +
Sbjct: 62 AIQESRIAITVLSQNYASSSFCLDELVTILHCKSQGLL-VIPVFYKVDPSHVRHQKGSYG 120
Query: 130 EAFRKHQEKFGQHSDRVDRWRDAFTQVASYSGWDSKG--QHEASLVENIAQHIHRKLVPK 187
EA KHQ++F + +++ +WR A QVA SG+ K +E + +I + I RK
Sbjct: 121 EAMAKHQKRFKANKEKLQKWRMALHQVADLSGYHFKDGDSYEYEFIGSIVEEISRKFSRA 180
Query: 188 LPSCTENLVGIVSKVEEVNKFLGMGLNDV-RFIGIWGMGGIGKSTIARAVYETIRCEFEL 246
+ VG+ S+V EV K L +G +DV IGI GMGG+GK+T+A AV+ I F+
Sbjct: 181 SLHVADYPVGLESEVTEVMKLLDVGSHDVVHIIGIHGMGGLGKTTLALAVHNFIALHFDE 240
Query: 247 TCFLENVREISETNGLVHLQRQLLSHLSISRN-DFHDLYDGKKTIQNSLCRKKVLLVLDD 305
+CFL+NVRE S +GL HLQ LLS L ++ +G IQ+ L RKKVLL+LDD
Sbjct: 241 SCFLQNVREESNKHGLKHLQSILLSKLLGEKDITLTSWQEGASMIQHRLQRKKVLLILDD 300
Query: 306 VNELNQLENLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKA 365
V++ QL+ +VG+ DWFGPGSRVIITTRDKHLL H V +TY+ +L + AL L A
Sbjct: 301 VDKRQQLKAIVGRPDWFGPGSRVIITTRDKHLLKYHEVERTYEVKVLNQSAALQLLTWNA 360
Query: 366 FKGDKPQEGYLDLSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKLRSFPHPRVQD 425
FK +K Y D+ VV Y GLPLALEV+GS L+ + + W SA++ + P +Q+
Sbjct: 361 FKREKIDPSYEDVLNRVVTYASGLPLALEVIGSNLFEKTVAEWESAMEHYKRIPSDEIQE 420
Query: 426 NLKISYDSLDTMEKDIFLDIACFFKGMKGDKVIDILESC-GYFPQIGIQILIERSLITLD 484
LK+S+D+L +K++FLDIAC FKG + +V +IL G + I +L+E+SL+ +
Sbjct: 421 ILKVSFDALGEEQKNVFLDIACCFKGYEWTEVDNILRDLYGNCTKHHIGVLVEKSLVKV- 479
Query: 485 SVNNKLGMHDLLQEMGRDIVFQESPNDPCRRSRLWSQEDIDRVLTKNKGTEAINSI--DM 542
S + + MHD++Q+MGR+I Q SP +P + RL +DI +VL N GT I I D
Sbjct: 480 SCCDTVEMHDMIQDMGREIERQRSPEEPGKCKRLLLPKDIIQVLKDNTGTSKIEIICLDF 539
Query: 543 KLLQPYE-AHWNTEAFSKTSQLKFLSLCEMQLPLGLSCLPSSLKVLHWRGCPLKTLPITT 601
+ E WN AF K LK L + + G + P L+VL W P LP
Sbjct: 540 SISDKEETVEWNENAFMKMKNLKILIIRNCKFSKGPNYFPEGLRVLEWHRYPSNCLPSNF 599
Query: 602 QLDELVDITLSHSKI 616
LV L S I
Sbjct: 600 DPINLVICKLPDSSI 614
>emb|CAA08798.1| NL27 [Solanum tuberosum]
Length = 821
Score = 450 bits (1158), Expect = e-125
Identities = 256/641 (39%), Positives = 385/641 (59%), Gaps = 17/641 (2%)
Query: 1 MEASSSSSCRSSSTTSLCTYHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERG 60
M +SSSS S Y VFLSFRG DTR+ FT HL L+ +GI TF+DDK LE G
Sbjct: 1 MASSSSSFAIDSQYRLRWKYDVFLSFRGVDTRRTFTSHLYEGLKNRGIFTFQDDKRLENG 60
Query: 61 QVISEKLINAIKDSMFAITILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSD 120
I E+L+ AI++S A+ I S +YA+S WCL+EL IMEC + V+P+FY VDPS+
Sbjct: 61 DSIPEELLKAIEESQVALIIFSKNYATSRWCLNELVKIMECKEEKGQIVIPIFYDVDPSE 120
Query: 121 VRHQRGCFEEAFRKHQEKFG---QHSDRVDRWRDAFTQVASYSGWDSKGQHEASLVENIA 177
VR Q F EAF +H+ K+ + +V WR A + A G+D + E+ +++I
Sbjct: 121 VRKQTKSFAEAFTEHESKYANDIEGMQKVKGWRTALSDAADLKGYDISNRIESDYIQHIV 180
Query: 178 QHIHRKLVPKLPSCTENLVGIVSKVEEVNKFLG-MGLNDVRFIGIWGMGGIGKSTIARAV 236
HI L S +NLVGI + + + L + ++ V +GIWGM G+GK+TIARA+
Sbjct: 181 DHISVLCKGSL-SYIKNLVGIDTHFKNIRSLLAELQMSGVLIVGIWGMPGVGKTTIARAI 239
Query: 237 YETIRCEFELTCFLENVREISETNGLVHLQRQLLSHLSISR-NDFHDLYDGKKTIQNSLC 295
++ + +FE CFL +++E G+ LQ LLS L + N ++ DG+ + + L
Sbjct: 240 FDRLSYQFEAVCFLADIKE--NKCGMHSLQNILLSELLKEKDNCVNNKEDGRSLLAHRLR 297
Query: 296 RKKVLLVLDDVNELNQLENLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKH 355
KKVL+VLDD++ ++QL+ L G DWFG GSR+I TTRDKHL+ G + Y+ L H
Sbjct: 298 FKKVLVVLDDIDHIDQLDYLAGNLDWFGNGSRIIATTRDKHLI---GKNVVYELPTLHDH 354
Query: 356 DALVLFCLKAFKGDKPQEGYLDLSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKL 415
DA+ LF AFK + + +L+ EVV + GLPLAL+V G + + R+I W SA+K++
Sbjct: 355 DAIKLFERYAFKEQVSDKCFKELTLEVVSHAKGLPLALKVFGCFFHERDITEWRSAIKQI 414
Query: 416 RSFPHPRVQDNLKISYDSLDTMEKDIFLDIACFFKGMKGDKVIDILESCGYFPQIGIQIL 475
++ P+ + + LKISYD L+T+++ IFLDIACF +G + D V+ ILESC + IG+ +L
Sbjct: 415 KNNPNSEIVEKLKISYDGLETIQQSIFLDIACFLRGRRKDYVMQILESCDFGADIGLSVL 474
Query: 476 IERSLITLDSVNNKLGMHDLLQEMGRDIVFQESPNDPCRRSRLWSQEDIDRVLTKNKGTE 535
I++SL+++ S NN + MHDL+Q+MG+ +V + DP RSRLW +D + V+ N GT+
Sbjct: 475 IDKSLVSI-SGNNTIEMHDLIQDMGKYVV--KKQKDPGERSRLWLTKDFEEVMINNTGTK 531
Query: 536 AINSIDMKLLQPYEAHWNTEAFSKTSQLKFLSLCEMQ-LPLGLSCLPSSLKVLHWRGCPL 594
A+ +I + ++ EA + +L+ L + + L + LP+SL+ W P
Sbjct: 532 AVEAIWVPNFN--RPRFSKEAMTIMQRLRILCIHDSNCLDGSIEYLPNSLRWFVWNNYPC 589
Query: 595 KTLPITTQLDELVDITLSHSKIEQLWQGVKVIDSLTKNSIK 635
++LP + +LV + LS S + LW G K + L K ++
Sbjct: 590 ESLPENFEPQKLVHLDLSLSSLHHLWTGKKHLPFLQKLDLR 630
>gb|AAO23074.1| R 10 protein [Glycine max]
Length = 901
Score = 450 bits (1158), Expect = e-125
Identities = 265/604 (43%), Positives = 364/604 (59%), Gaps = 10/604 (1%)
Query: 20 YHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQVISEKLINAIKDSMFAIT 79
Y VFL+FRG DTR GFT +L AL KGI TF D+K L RG+ I+ L+ AI++S AIT
Sbjct: 12 YDVFLNFRGGDTRYGFTGNLYRALCDKGIHTFFDEKKLHRGEEITPALLKAIQESRIAIT 71
Query: 80 ILSPDYASSTWCLDELQMIMECSSKNNLHVLPVFYGVDPSDVRHQRGCFEEAFRKHQEKF 139
+LS +YASS++CLDEL I+ C S+ L V+PVFY VDPSDVRHQ+G + KHQ++F
Sbjct: 72 VLSKNYASSSFCLDELVTILHCKSEGLL-VIPVFYNVDPSDVRHQKGSYGVEMAKHQKRF 130
Query: 140 GQHSDRVDRWRDAFTQVASYSGWDSKG--QHEASLVENIAQHIHRKLVPKLPSCTENLVG 197
+++ +WR A QVA G+ K +E +++I + + R++ + VG
Sbjct: 131 KAKKEKLQKWRIALKQVADLCGYHFKDGDAYEYKFIQSIVEQVSREINRAPLHVADYPVG 190
Query: 198 IVSKVEEVNKFLGMGLNDV-RFIGIWGMGGIGKSTIARAVYETIRCEFELTCFLENVREI 256
+ S+V EV K L +G +DV IGI GMGG+GK+T+A AVY I F+ +CFL+NVRE
Sbjct: 191 LGSQVIEVRKLLDVGSHDVVHIIGIHGMGGLGKTTLALAVYNLIALHFDESCFLQNVREE 250
Query: 257 SETNGLVHLQRQLLSHLSISRN-DFHDLYDGKKTIQNSLCRKKVLLVLDDVNELNQLENL 315
S +GL HLQ LLS L ++ +G IQ+ L RKKVLL+LDDV++ QL+ +
Sbjct: 251 SNKHGLKHLQSILLSKLLGEKDITLTSWQEGASMIQHRLQRKKVLLILDDVDKREQLKAI 310
Query: 316 VGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGDKPQEGY 375
VG+ DWFGPGSRVIITTRDKHLL H V +TY+ +L + AL L AFK +K Y
Sbjct: 311 VGRPDWFGPGSRVIITTRDKHLLKYHEVERTYEVKVLNQSAALQLLKWNAFKREKIDPSY 370
Query: 376 LDLSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKLRSFPHPRVQDNLKISYDSLD 435
D+ VV Y GLPLALEV+GS L+G+ + W SA++ + P + + LK+S+D+L
Sbjct: 371 EDVLNRVVTYASGLPLALEVIGSNLFGKTVAEWESAMEHYKRIPSDEILEILKVSFDALG 430
Query: 436 TMEKDIFLDIACFFKGMKGDKVIDILESC-GYFPQIGIQILIERSLITLDSV-NNKLGMH 493
+K++FLDIAC F+G K +V DIL + G + I +L+E+SLI L+ + + MH
Sbjct: 431 EEQKNVFLDIACCFRGYKWTEVDDILRALYGNCKKHHIGVLVEKSLIKLNCYGTDTVEMH 490
Query: 494 DLLQEMGRDIVFQESPNDPCRRSRLWSQEDIDRVLTKNKGTEAINSI--DMKLLQPYE-A 550
DL+Q+M R+I + SP +P + RLW +DI +V N GT I I D + E
Sbjct: 491 DLIQDMAREIERKRSPQEPGKCKRLWLPKDIIQVFKDNTGTSKIEIICLDSSISDKEETV 550
Query: 551 HWNTEAFSKTSQLKFLSLCEMQLPLGLSCLPSSLKVLHWRGCPLKTLPITTQLDELVDIT 610
WN AF K LK L + + G + P L+VL W P LP + LV
Sbjct: 551 EWNENAFMKMENLKILIIRNDKFSKGPNYFPEGLRVLEWHRYPSNCLPSNFHPNNLVICK 610
Query: 611 LSHS 614
L S
Sbjct: 611 LPDS 614
>emb|CAC95124.1| TIR/NBS/LRR protein [Populus deltoides]
Length = 1147
Score = 449 bits (1156), Expect = e-124
Identities = 265/629 (42%), Positives = 366/629 (58%), Gaps = 19/629 (3%)
Query: 20 YHVFLSFRGEDTRKGFTDHLCAALERKGITTFKDDKDLERGQVISEKLINAIKDSMFAIT 79
Y VFLSFRGEDTRK FTDHL AL + GI TF+DD +L RG+ IS+ + AI++S +I
Sbjct: 39 YDVFLSFRGEDTRKTFTDHLYTALVQAGIHTFRDDDELPRGEEISDHFLRAIQESKISIA 98
Query: 80 ILSPDYASSTWCLDELQMIMECSS-KNNLHVLPVFYGVDPSDVRHQRGCFEEAFRKHQEK 138
+ S YASS WCL+EL I++C K VLP+FY +DPSDVR Q G F EAF KH+E+
Sbjct: 99 VFSKGYASSRWCLNELVEILKCKKRKTGQIVLPIFYDIDPSDVRKQNGSFAEAFVKHEER 158
Query: 139 FGQHSDRVDRWRDAFTQVASYSGW---DSKGQHEASLVENIAQHIHRKLVPKLPSCTENL 195
F V WR A + + SGW D HEA ++ I + + KL PK E+L
Sbjct: 159 F--EEKLVKEWRKALEEAGNLSGWNLNDMANGHEAKFIKEIIKVVLNKLEPKYLYVPEHL 216
Query: 196 VGIVSKVEEVNKFLGMGLNDVRFIGIWGMGGIGKSTIARAVYETIRCEFELTCFLENVRE 255
VG+ + FL +DVR +GI GM GIGK+TIA+AV+ + FE +CFL ++ E
Sbjct: 217 VGMDQLARNIFDFLSAATDDVRIVGIHGMPGIGKTTIAQAVFNQLCYGFEGSCFLSSINE 276
Query: 256 IS-ETNGLVHLQRQLLSHLSISRND---FHDLYDGKKTIQNSLCRKKVLLVLDDVNELNQ 311
S + NGLV LQ+QL H I + D F GK I+ L RK+VL+V DDV L Q
Sbjct: 277 RSKQVNGLVPLQKQL--HHDILKQDVANFDCADRGKVLIKERLRRKRVLVVADDVAHLEQ 334
Query: 312 LENLVGKQDWFGPGSRVIITTRDKHLLMTHGVHKTYKTGMLCKHDALVLFCLKAFKGDKP 371
L L+G + WFGPGSRVIITTRD +LL + Y+ L ++L LF AFK KP
Sbjct: 335 LNALMGDRSWFGPGSRVIITTRDSNLL--READQIYQIEELKPDESLQLFSRHAFKDSKP 392
Query: 372 QEGYLDLSKEVVDYCGGLPLALEVLGSYLYGRNIDVWHSAVKKLRSFPHPRVQDNLKISY 431
+ Y++LSK+ V YCGGLPLALEV+G+ LY +N S + L P+ +Q L ISY
Sbjct: 393 AQDYIELSKKAVGYCGGLPLALEVIGALLYRKNRGRCVSEIDNLSRIPNQDIQGKLLISY 452
Query: 432 DSLDTMEKDIFLDIACFFKGMKGDKVIDILES-CGYFPQIGIQILIERSLITLDSVNNKL 490
+LD + FLDIACFF G++ + V +L + C P++ ++ L ERSLI + +
Sbjct: 453 HALDGELQRAFLDIACFFIGIEREYVTKVLGARCRPNPEVVLETLSERSLIQV--FGETV 510
Query: 491 GMHDLLQEMGRDIVFQESPNDPCRRSRLWSQEDIDRVLTKNK--GTEAINSIDMKLLQPY 548
MHDLL++MGR++V + SP P +R+R+W+QED VL + K GT+ + + + +
Sbjct: 511 SMHDLLRDMGREVVCKASPKQPGKRTRIWNQEDAWNVLEQQKVRGTDVVKGLALDVRASE 570
Query: 549 EAHWNTEAFSKTSQLKFLSLCEMQLPLGLSCLPSSLKVLHWRGCPLKTLPITTQLDELVD 608
+ +F++ L L + + L L L + W CPLK LP LD L
Sbjct: 571 AKSLSAGSFAEMKCLNLLQINGVHLTGSLKLFSKELMWICWHECPLKYLPFDFTLDNLAV 630
Query: 609 ITLSHSKIEQLWQGVKVIDSLTKNSIKAY 637
+ + +S +++LW+G KV + L Y
Sbjct: 631 LDMQYSNLKELWKGKKVRNMLQSPKFLQY 659
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.321 0.137 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,096,246,560
Number of Sequences: 2540612
Number of extensions: 47049591
Number of successful extensions: 140115
Number of sequences better than 10.0: 2493
Number of HSP's better than 10.0 without gapping: 1251
Number of HSP's successfully gapped in prelim test: 1242
Number of HSP's that attempted gapping in prelim test: 134195
Number of HSP's gapped (non-prelim): 2995
length of query: 640
length of database: 863,360,394
effective HSP length: 134
effective length of query: 506
effective length of database: 522,918,386
effective search space: 264596703316
effective search space used: 264596703316
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 78 (34.7 bits)
Medicago: description of AC149204.4


