

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149127.7 - phase: 0
(360 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
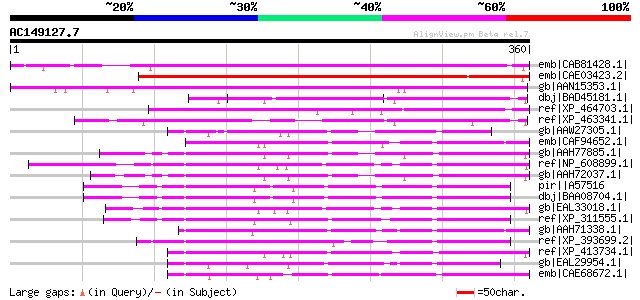
Score E
Sequences producing significant alignments: (bits) Value
emb|CAB81428.1| putative calcium binding protein [Arabidopsis th... 289 1e-76
emb|CAE03423.2| OSJNBa0032F06.6 [Oryza sativa (japonica cultivar... 221 3e-56
gb|AAN15353.1| Unknown protein [Arabidopsis thaliana] gi|9759352... 196 9e-49
dbj|BAD45181.1| calcium binding protein-like [Oryza sativa (japo... 123 7e-27
ref|XP_464703.1| calcium-binding EF hand family protein-like [Or... 118 2e-25
ref|XP_463341.1| B1129G05.13 [Oryza sativa (japonica cultivar-gr... 99 2e-19
gb|AAW27305.1| unknown [Schistosoma japonicum] 92 2e-17
emb|CAF94652.1| unnamed protein product [Tetraodon nigroviridis] 83 1e-14
gb|AAH77885.1| Rcn2-prov protein [Xenopus laevis] 82 3e-14
ref|NP_608899.1| CG31650-PC, isoform C [Drosophila melanogaster]... 79 2e-13
gb|AAH72037.1| MGC78878 protein [Xenopus laevis] 77 1e-12
pir||A57516 DNA supercoiling factor - silkworm 76 2e-12
dbj|BAA08704.1| DNA supercoiling factor [Bombyx mori] 76 2e-12
gb|EAL33018.1| GA16367-PA [Drosophila pseudoobscura] 75 2e-12
ref|XP_311555.1| ENSANGP00000010221 [Anopheles gambiae str. PEST... 75 3e-12
gb|AAH71338.1| Zgc:86646 [Danio rerio] gi|50344972|ref|NP_001002... 75 3e-12
ref|XP_393699.2| PREDICTED: similar to CG31650-PC, isoform C [Ap... 74 5e-12
ref|XP_413734.1| PREDICTED: similar to Reticulocalbin 2 precurso... 72 3e-11
gb|EAL29954.1| GA21575-PA [Drosophila pseudoobscura] 72 3e-11
emb|CAE68672.1| Hypothetical protein CBG14576 [Caenorhabditis br... 71 4e-11
>emb|CAB81428.1| putative calcium binding protein [Arabidopsis thaliana]
gi|4972110|emb|CAB43967.1| putative calcium binding
protein [Arabidopsis thaliana]
gi|15234272|ref|NP_194508.1| calcium-binding EF hand
family protein [Arabidopsis thaliana]
gi|7488067|pir||T09018 probable calcium-binding protein
T27E11.30 - Arabidopsis thaliana
Length = 345
Score = 289 bits (739), Expect = 1e-76
Identities = 155/370 (41%), Positives = 225/370 (59%), Gaps = 35/370 (9%)
Query: 1 MSKAVVYTVIATATLLIFIVLS--PLNLEESKGRLNNRRFGYKILERAPTFDPLVTKIER 58
M+K VVYT++ TAT++ I+L+ N +S L RR G ++ P FDPLVT+IER
Sbjct: 1 MAKVVVYTIL-TATIIFLILLAHKKQNQTQSIEGLITRRIGRRL--EMPVFDPLVTRIER 57
Query: 59 ESEQKNQQHKNDFDNNKNVAPRTGLGSTTTVSEIKETY-----EYLTSGGTLNTTLRLII 113
S +K T TV KE EY LNTT+R+
Sbjct: 58 LSHEKE-------------------AGTKTVEAAKEEKDDMFEEYFAQERRLNTTMRIKF 98
Query: 114 LFPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYLPDLS 173
LFPLLD P+DGFV EL++W+ Q+ + + Y T ELE +DK+ D ++F EYLP S
Sbjct: 99 LFPLLDASPRDGFVSLKELQTWMMQQTEDNMVYRTAKELELQDKDKDGVITFEEYLPQFS 158
Query: 174 EKDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVNDKF 233
++DIEK HGEAGW ME+F +D+DHNG L+ E +FLHPEDS+N + +W++ ++
Sbjct: 159 KQDIEKNEKGHGEAGWWMEQFKNSDFDHNGSLDIEEFNNFLHPEDSRNGDTQRWVLKERM 218
Query: 234 NMDDYEHDGKINFNQFEDNVYVTYESYVDFETNGEGDIPTAKDKFAELDVNKDQFLSPEE 293
D DGK+ + +F N Y Y+ + FE + ++PT + FAE+D +KD+FL +E
Sbjct: 219 TGMDTNGDGKLEYKEFVKNAYEMYKEFAKFEKEEDENVPTPQLLFAEMDRDKDRFLVADE 278
Query: 294 LFPIIPYVYPGELAYAKYYTSYLMNEADDNEDRKLTLDEMLDHEFAFFNTVHADGHVEID 353
L PI+ Y+ PGE++YAK+Y+++L +EAD+++D KL+L+EML HE F+ VH H ++D
Sbjct: 279 LRPILQYLQPGEMSYAKFYSTFLCHEADEDKDGKLSLEEMLHHEDVFYKAVH---HEDLD 335
Query: 354 DD---DHDEL 360
D+ DHDEL
Sbjct: 336 DEDYFDHDEL 345
>emb|CAE03423.2| OSJNBa0032F06.6 [Oryza sativa (japonica cultivar-group)]
Length = 358
Score = 221 bits (562), Expect = 3e-56
Identities = 114/275 (41%), Positives = 171/275 (61%), Gaps = 5/275 (1%)
Query: 90 SEIKETYEYLTSGGTLNTTLRLIILFPLLDRDPKDGFVGFNELESWVTQRALERLDYATQ 149
+E+ Y S G LN RL+ LFP+LDR PKDG V ELE+W+ ++A +RLD +
Sbjct: 84 AELGPLERYFGSDGELNVKERLLYLFPMLDRAPKDGGVSCGELEAWLRRQAADRLDAVAR 143
Query: 150 VELESKDKNGDLALSFREYLPDLSEKDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTE 209
EL+ DK+GD ++ REYL ++ I+ + HGE GW + KF AD DH+G ++F E
Sbjct: 144 RELKRHDKDGDGVVTLREYLAVDHDQHIDWTDTEHGEPGWWLHKFISADRDHSGAMDFIE 203
Query: 210 LRDFLHPEDSQNKEMLKWMVNDKFNMDDYEHDGKINFNQFEDNVY-VTYESYVDFETNGE 268
L DFLHPEDS +++ W++ DK + D++ DGK++ ++F + + + S V+ + +
Sbjct: 204 LNDFLHPEDSSQEKVKLWLLKDKLSGMDHDRDGKLSLDEFISQFHMIDHNSIVEHSADDD 263
Query: 269 GDIPTAKDKFAELDVNKDQFLSPEELFPIIPYVYPGELAYAKYYTSYLMNEADDNEDRKL 328
A+ KF ELD N D +L+ EE P+I + GE +YAK + LM +ADDN+D KL
Sbjct: 264 TSCAEAEKKFRELDSNNDGYLTVEEARPVIQSLISGEFSYAKSHAKLLM-KADDNKDNKL 322
Query: 329 TLDEMLDHEFAFFNTVHADGHVEIDD---DDHDEL 360
+L+EML+H +F+N V+ D H + DD + HDEL
Sbjct: 323 SLEEMLNHYLSFYNIVYMDDHYDYDDIGNNIHDEL 357
>gb|AAN15353.1| Unknown protein [Arabidopsis thaliana] gi|9759352|dbj|BAB10007.1|
unnamed protein product [Arabidopsis thaliana]
gi|18415883|ref|NP_568202.1| calcium-binding EF hand
family protein [Arabidopsis thaliana]
gi|16648869|gb|AAL24286.1| Unknown protein [Arabidopsis
thaliana]
Length = 391
Score = 196 bits (498), Expect = 9e-49
Identities = 127/389 (32%), Positives = 195/389 (49%), Gaps = 30/389 (7%)
Query: 1 MSKAVVYTVIATATLLIFIV-LSPLNLEESK----GRLNNRR------FGYKILERAPT- 48
MSKA V I L++F+V SP + G + R F +K P
Sbjct: 1 MSKASVILYITVGILVLFLVSYSPKKKGDHDHHHGGHNQHHRLKLRSSFNFKPTRHDPVP 60
Query: 49 FDPLVTKIERESEQKNQQ-----HKNDFDNNKNVAPRTGLG--------STTTVSEIKET 95
FDPLV +ER E K + H + + + TG G S E +
Sbjct: 61 FDPLVADMERRREDKEWERQYIEHSHPELVSHSQKETTGGGHEHAPGHESQPEWEEFMDA 120
Query: 96 YEYLTSGGTLNTTLRLIILFPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESK 155
+YL N T RLI+LFP +D P DGF+ +EL W Q + + + + TQ +L+
Sbjct: 121 EDYLNDEEKFNVTDRLILLFPKIDVSPADGFMTESELTEWTMQSSAKEVVHRTQRDLDVH 180
Query: 156 DKNGDLALSFREYLPDLSEKDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLH 215
D+N D +SF EY P + + + + W E F+ +D + +GLLN TE DFLH
Sbjct: 181 DRNKDGFISFSEYEPPSWVRKSDNNSFGYDMGWWKEEHFNASDANGDGLLNLTEFNDFLH 240
Query: 216 PEDSQNKEMLKWMVNDKFNMDDYEHDGKINFNQFEDNVYVTYESYVDFETNGE---GDIP 272
P D++N ++L W+ ++ D + DGKI+F +F ++ T +Y + N D+P
Sbjct: 241 PADTKNPKLLLWLCKEEVRERDSDKDGKISFEEFFHGLFDTVRNYEEDNHNSTHPYHDLP 300
Query: 273 --TAKDKFAELDVNKDQFLSPEELFPIIPYVYPGELAYAKYYTSYLMNEADDNEDRKLTL 330
AK F++LD N D +LS EL PII ++P E YAK Y++++AD ++DR+LTL
Sbjct: 301 EGPAKQLFSQLDKNDDGYLSDVELLPIISKIHPTEHYYAKQQADYIISQADSDKDRRLTL 360
Query: 331 DEMLDHEFAFFNTVHADGHVEIDDDDHDE 359
EM++H + F++ + + + D HDE
Sbjct: 361 AEMIEHPYVFYSAIFDEDDTDDDYGFHDE 389
>dbj|BAD45181.1| calcium binding protein-like [Oryza sativa (japonica
cultivar-group)] gi|52076317|dbj|BAD45102.1| calcium
binding protein-like [Oryza sativa (japonica
cultivar-group)]
Length = 207
Score = 123 bits (309), Expect = 7e-27
Identities = 70/213 (32%), Positives = 121/213 (55%), Gaps = 13/213 (6%)
Query: 152 LESKDKNGDLALSFREYLPDLSEKDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELR 211
+E DKNGD +S+ ++ +E E ++ G W E F+ +D D +G LN TE
Sbjct: 1 MELYDKNGDGVVSYGDFRAQHNESSGEVNSL--GFPWWKEEHFNASDADGHGFLNKTEFN 58
Query: 212 DFLHPEDSQNKEMLKWMVNDKFNMDDYEHDGKINFNQFEDNVYVTYESYVDFET---NGE 268
DFL+P DS+N +++ + + D + DGK+NF ++ ++ Y D+E +
Sbjct: 59 DFLNPSDSENPQIINLLCKQEIRQRDKDGDGKLNFEEYFHGLHDHIHGY-DYENADISHI 117
Query: 269 GDIPTAKDKFAELDVNKDQFLSPEELFPIIPYVYPGELAYAKYYTSYLMNEADDNEDRKL 328
G+ AK++F++LD + D F+S EL P++ ++ E YA+ ++ ++EAD + D +L
Sbjct: 118 GNNTVAKERFSKLDKDSDGFISEHELEPVLDKLHLSERYYARQQAAHAISEADKDHDGRL 177
Query: 329 TLDEMLDHEFAFFNTVHADGHVEIDDDD--HDE 359
TLDEM+++ +AF+ +V DD+D HDE
Sbjct: 178 TLDEMIENPYAFYGSVFLS-----DDEDYFHDE 205
Score = 47.4 bits (111), Expect = 7e-04
Identities = 34/143 (23%), Positives = 61/143 (41%), Gaps = 8/143 (5%)
Query: 125 GFVGFNELESWVTQRALER---LDYATQVELESKDKNGDLALSFREYLPDLSEK----DI 177
GF+ E ++ E ++ + E+ +DK+GD L+F EY L + D
Sbjct: 50 GFLNKTEFNDFLNPSDSENPQIINLLCKQEIRQRDKDGDGKLNFEEYFHGLHDHIHGYDY 109
Query: 178 EKKNMAH-GEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVNDKFNMD 236
E +++H G E+F D D +G ++ EL L + + +
Sbjct: 110 ENADISHIGNNTVAKERFSKLDKDSDGFISEHELEPVLDKLHLSERYYARQQAAHAISEA 169
Query: 237 DYEHDGKINFNQFEDNVYVTYES 259
D +HDG++ ++ +N Y Y S
Sbjct: 170 DKDHDGRLTLDEMIENPYAFYGS 192
>ref|XP_464703.1| calcium-binding EF hand family protein-like [Oryza sativa (japonica
cultivar-group)] gi|46806523|dbj|BAD17636.1|
calcium-binding EF hand family protein-like [Oryza
sativa (japonica cultivar-group)]
gi|46806504|dbj|BAD17628.1| calcium-binding EF hand
family protein-like [Oryza sativa (japonica
cultivar-group)]
Length = 392
Score = 118 bits (296), Expect = 2e-25
Identities = 80/277 (28%), Positives = 136/277 (48%), Gaps = 18/277 (6%)
Query: 97 EYLTSGGTLNTTLRLIILFPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKD 156
+Y+ N T R+ LFP +D DP DG V EL +W A + + T EL+ D
Sbjct: 120 DYINDAARFNLTRRVEALFPKIDVDPADGAVTPAELTAWNLASARREVMHRTARELDLHD 179
Query: 157 KNGDLALSFREY-LPDLSEKDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRD--- 212
++ D ++F EY P + + + + G W E F+ +D D L+ F +L
Sbjct: 180 RDHDGRIAFSEYERPSWAWRFDDHNSSNDGVGWWKEEHFNASDMDVFSLVYFVQLLTSSR 239
Query: 213 FLHPEDSQNKEMLKWMVNDKFNMD------DYEHDGKINFNQFEDNVYVT---YESYVDF 263
+ P+ ++ + V D D ++DGK+NF +F + ++ + ++
Sbjct: 240 YYEPKANKLVVQRRSQVGDSTGYSLESRERDKDNDGKLNFQEFYNGLFYSIRHFDEEAST 299
Query: 264 ETNGEGDIPTAKDKFAELDVNKDQFLSPEELFPIIPYVYPGELAYAKYYTSYLMNEADDN 323
+ + D P A+ F LD++ D LS +EL PII ++P E YAK Y++ +AD N
Sbjct: 300 DDSNASDAP-ARKSFTHLDLDNDGLLSADELKPIIGNLHPPEHFYAKQQADYVITQADTN 358
Query: 324 EDRKLTLDEMLDHEFAFFNTVHADGHVEIDDDDHDEL 360
+D +L+L EM+++ + F++ + E D HDEL
Sbjct: 359 KDGQLSLQEMIENPYVFYSAL----FTEDDYGFHDEL 391
>ref|XP_463341.1| B1129G05.13 [Oryza sativa (japonica cultivar-group)]
Length = 381
Score = 99.0 bits (245), Expect = 2e-19
Identities = 82/338 (24%), Positives = 151/338 (44%), Gaps = 65/338 (19%)
Query: 46 APTFDPLVTKIERESEQKNQQHKNDFDNNKNVAPRTGLGSTTTVSE------IKETYEYL 99
A FDP + ++ER E K + ++ + G G ++E +++
Sbjct: 83 AAPFDPEIAELERRLEDKEWEREH-----YRILHGDGGGGEADEHMREWEEFLREDEDFI 137
Query: 100 TSGGTLNTTLRLIILFPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNG 159
N R+ LFP +D P+DGF +EL W +++ + + E+E DKNG
Sbjct: 138 NDDERFNLGDRIRALFPKIDLAPRDGFASLDELTRWNLEQSRADQLHRSAREMELYDKNG 197
Query: 160 DLALSFREYLPDLSEKDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDS 219
D +S+ ++ H E+ + +F+ A +S
Sbjct: 198 DGVVSYGDF------------RAQHNESSGVQNEFNKAG------------------SES 227
Query: 220 QNKEMLKWMVND-KFNMDDYEHDGKINFNQFEDNVYVTYESYVDFET---NGEGDIPTAK 275
+ + ++ND + D + DGK+NF ++ ++ Y D+E + G+ AK
Sbjct: 228 MRQFGVSVLINDLQERQRDKDGDGKLNFEEYFHGLHDHIHGY-DYENADISHIGNNTVAK 286
Query: 276 DKFAELDVNKDQFLSPEELFPIIPYVYPGELAYAKYYTSYLMNE------------ADDN 323
++F++LD + D F+S EL P++ ++ E YA+ ++ ++E AD +
Sbjct: 287 ERFSKLDKDSDGFISEHELEPVLDKLHLSERYYARQQAAHAISEDDMFNWLIQKLQADKD 346
Query: 324 EDRKLTLDEMLDHEFAFFNTVHADGHVEIDDDD--HDE 359
D +LTLDEM+++ +AF+ +V DD+D HDE
Sbjct: 347 HDGRLTLDEMIENPYAFYGSVFLS-----DDEDYFHDE 379
>gb|AAW27305.1| unknown [Schistosoma japonicum]
Length = 281
Score = 92.0 bits (227), Expect = 2e-17
Identities = 73/236 (30%), Positives = 120/236 (49%), Gaps = 30/236 (12%)
Query: 110 RLIILFPLLDRDPKDGFVGFNELESWV--TQRALERLDYATQVELESKDKNGDLALSFRE 167
RL I F +D + +GF+ ++EL SW+ T +L+R ++A + +L D N D +S E
Sbjct: 43 RLHIYFKKIDTN-NNGFIEYDELTSWIFKTYESLDR-EHAEK-QLVKYDTNKDAKVSLDE 99
Query: 168 YLPDLSEKDIEKKNMAHGE--AGWLME-------KFDVADYDHNGLLNFTELRDFLHPED 218
Y+ E E+ N + + + +++E +F+ AD D +GLL+ E FL PE+
Sbjct: 100 YISQTYETSEEELNHSKDDQSSNFILESLKNERSRFNFADKDCDGLLSLEEFTLFLRPEN 159
Query: 219 SQNKEMLKWMVNDKFNMDDYEHDGKINFNQFEDNVYVTYESYVDFETNGEGDIPTAKDKF 278
+ +M + + F+ D DG VTY+ + +F G ++F
Sbjct: 160 YE--DMANYELQKSFSSFDQNGDG-----------VVTYDEFTNFSYRGVSQQNYLHEQF 206
Query: 279 AELDVNKDQFLSPEELFPIIPYVYPGELAYAKYYTSYLMNEADDNEDRKLTLDEML 334
LDV+K+ L+ +EL P ++ P A AK ++LMN D N D +LTLDE+L
Sbjct: 207 KSLDVDKNNLLTLDELRP---WLLPSLKAAAKSEATWLMNLTDSNHDGQLTLDEIL 259
>emb|CAF94652.1| unnamed protein product [Tetraodon nigroviridis]
Length = 306
Score = 82.8 bits (203), Expect = 1e-14
Identities = 70/248 (28%), Positives = 112/248 (44%), Gaps = 23/248 (9%)
Query: 123 KDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYLPD-----LSEK-- 175
KDG+V EL W+ R ++ + D+N D + + EY L E+
Sbjct: 72 KDGYVSHTELHHWIKHRQRRYIEENVNKNWKDYDQNQDGKIGWEEYKNTTYGYYLGEEFS 131
Query: 176 DIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVNDKFNM 235
D+E K +F AD D +G+ E FLHPE+ M +V +
Sbjct: 132 DVEDKATYQAMLARDNRRFKYADQDRDGIATREEFTAFLHPEEFDY--MKDVVVQETMED 189
Query: 236 DDYEHDGKINFNQFEDNVYVTY--ESYVDFETNGEGDIPTAKDKFAEL-DVNKDQFLSPE 292
D + DGKIN +++ ++Y ES D+ + T K +F+E D NKD +L
Sbjct: 190 IDKDGDGKINLDEYIGDMYTPENDESEPDW-------VQTEKKQFSEFRDTNKDGYLDAG 242
Query: 293 ELFPIIPYVYPGELAYAKYYTSYLMNEADDNEDRKLTLDEMLDHEFAFFNTVHADGHVEI 352
E + ++ PGE+ +A +L++E D ++D K+T E+L + F + A + E
Sbjct: 243 E---VAHWILPGEVDHADNEAKHLIHETDTDKDEKITKKEILANWNMFVGS-QATNYGED 298
Query: 353 DDDDHDEL 360
HDEL
Sbjct: 299 LTKRHDEL 306
Score = 44.7 bits (104), Expect = 0.004
Identities = 47/189 (24%), Positives = 72/189 (37%), Gaps = 47/189 (24%)
Query: 183 AHGEAGWLMEKFDVADYDHNGLLNFTELRDF--LHPEDSQNK------------------ 222
AH +AG YDH L E + F L PE+S+ K
Sbjct: 26 AHNDAGGFQ-------YDHEAFLGKEEAKTFDQLTPEESKEKLAKIVNGIDTNKDGYVSH 78
Query: 223 -EMLKWM-----------VNDKFNMDDYEHDGKINFNQFEDNVYVTY--ESYVDFETNGE 268
E+ W+ VN + D DGKI + ++++ Y Y E + D E
Sbjct: 79 TELHHWIKHRQRRYIEENVNKNWKDYDQNQDGKIGWEEYKNTTYGYYLGEEFSDVEDKAT 138
Query: 269 GDIPTAKD--KFAELDVNKDQFLSPEELFPIIPYVYPGELAYAK-YYTSYLMNEADDNED 325
A+D +F D ++D + EE +++P E Y K M + D + D
Sbjct: 139 YQAMLARDNRRFKYADQDRDGIATREE---FTAFLHPEEFDYMKDVVVQETMEDIDKDGD 195
Query: 326 RKLTLDEML 334
K+ LDE +
Sbjct: 196 GKINLDEYI 204
>gb|AAH77885.1| Rcn2-prov protein [Xenopus laevis]
Length = 313
Score = 82.0 bits (201), Expect = 3e-14
Identities = 80/317 (25%), Positives = 139/317 (43%), Gaps = 43/317 (13%)
Query: 63 KNQQHKNDFDNNKNVAPRTGLGSTTTVSEIKETYEYLTSGGTLNTTLRLIILFPLLDRDP 122
++ +H + ++N + LG E E L+S L RL ++ +D D
Sbjct: 21 QHDEHYENGEHNAEYDKKLFLGGEEEAEEFTE----LSSEDQLK---RLKLIIRRIDTD- 72
Query: 123 KDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYLPDLSEK--DIEKK 180
DG++ EL SW+ + + T+ DK+GD +++ EY L ++ D ++
Sbjct: 73 SDGYLTEEELSSWIQKSFRHYILEDTKEHFADIDKDGDGIVTWDEYNMHLYDRIIDYDEN 132
Query: 181 NMAHGEAGWLME--------KFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVNDK 232
+ E +FD AD D LN TE DF HPE++ + M ++++
Sbjct: 133 TVLEDEEEESFRLIHMKDKRRFDHADTDKIPGLNLTEFTDFEHPEETDH--MSEFVIEGA 190
Query: 233 FNMDDYEHDGKINFNQFEDNVYVTYESYVDFETNGEGDIP------TAKDKFA-ELDVNK 285
D + DG +V+ E Y+ T G + KD+F + D +
Sbjct: 191 LEEHDEDGDG-----------FVSLEEYLGDYTQDSGAVEDPHWLIVEKDRFVNDYDKDG 239
Query: 286 DQFLSPEELFPIIPYVYPGELAYAKYYTSYLMNEADDNEDRKLTLDEMLDHEFAFFNTVH 345
D L+P EL + ++ P L ++ +LM E D NED++L+ +E+L ++ F +
Sbjct: 240 DGRLNPTEL---LSWIVPNNLGISQEEAIHLMTEMDKNEDQRLSEEEILQNKDIFLTSEA 296
Query: 346 ADGHVEIDDDD--HDEL 360
D ++ D HDEL
Sbjct: 297 TDYGRQLQDKQFYHDEL 313
>ref|NP_608899.1| CG31650-PC, isoform C [Drosophila melanogaster]
gi|24581831|ref|NP_723049.1| CG31650-PB, isoform B
[Drosophila melanogaster] gi|24581829|ref|NP_723048.1|
CG31650-PA, isoform A [Drosophila melanogaster]
gi|22945629|gb|AAN10522.1| CG31650-PC, isoform C
[Drosophila melanogaster] gi|7296934|gb|AAF52207.1|
CG31650-PB, isoform B [Drosophila melanogaster]
gi|22945628|gb|AAN10521.1| CG31650-PA, isoform A
[Drosophila melanogaster] gi|16769482|gb|AAL28960.1|
LD34388p [Drosophila melanogaster]
Length = 342
Score = 79.3 bits (194), Expect = 2e-13
Identities = 83/360 (23%), Positives = 161/360 (44%), Gaps = 41/360 (11%)
Query: 14 TLLIFIVLSPLNLEESKGRLNNRRFGYKILERAPTFDPLVTKIERESEQKNQQHKNDFDN 73
TL +L+ + + G + N K L + D + + ++ +H +FD+
Sbjct: 11 TLCAVALLAAVGPMPAHGAVANSHKHEKHLSKERVKDGIYAPRDAHHHGEDGEHNVEFDH 70
Query: 74 NKNVAPRTGLGSTTTVSEIKETYEYLTSGGTLNTTLRLIILFPLLDRDPKDGFVGFNELE 133
+ KE E+ S + RL+IL ++D + KD F+ +EL+
Sbjct: 71 E------------AIIGNTKEAQEF-DSLSPDESKRRLLILIKMMDLN-KDEFIDRHELK 116
Query: 134 SWVTQRALERLDYATQVELESKDKNGDLALSFREYLPD---LSEKDIEKKNMAH----GE 186
+W+ + + + E D++ D ++++EYL D + ++D +K+ + + E
Sbjct: 117 AWILRSFKKLSEEEAADRFEEIDQDADERITWKEYLQDTYAMEDEDFKKETIDYDSYEDE 176
Query: 187 AGWL---MEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVNDKFNMDDYEHDGK 243
+ E F+ AD + +G+L E F +PE ++ +ML ++ D +HDGK
Sbjct: 177 QKMIKQDKEMFNAADTNKDGVLTLEEFVLFQNPE--EHPQMLPILLEHTMQDKDADHDGK 234
Query: 244 INFNQFEDNVYVTYESYVDFETNGEGDIPTAKDKF-AELDVNKDQFLSPEELFPIIPYVY 302
INF +F + S+ D E + T K++F + D N D L+ +E ++ ++
Sbjct: 235 INFQEFVGDA----ASHHDKEW-----LITEKERFDKDHDSNGDGVLTGDE---VLSWIV 282
Query: 303 PGELAYAKYYTSYLMNEADDNEDRKLTLDEMLDHEFAFFNTVHADGHVEIDDDDH--DEL 360
P A A +L D++ D +L+ E+L++ F + D + + +H DEL
Sbjct: 283 PSNTAIANDEVDHLFVSTDEDHDDRLSYLEILNNYDTFVGSEATDYGDHLQNINHLSDEL 342
>gb|AAH72037.1| MGC78878 protein [Xenopus laevis]
Length = 313
Score = 76.6 bits (187), Expect = 1e-12
Identities = 79/323 (24%), Positives = 139/323 (42%), Gaps = 49/323 (15%)
Query: 57 ERESEQKNQQHKNDFDNNKNVAPRTGLGSTTTVSEIKETYEYLTSGGTLNTTLRLIILFP 116
+ + +N +H +FD + LG E E L+S L RL +
Sbjct: 21 QHDEHYENGEHNAEFDK------KLFLGGEEEAEEFTE----LSSEDQLK---RLKSIIR 67
Query: 117 LLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYLPDLSEK- 175
+D D DG++ EL SW+ + + T+ DK+ + +++ EY + ++
Sbjct: 68 KIDTD-SDGYLTEEELSSWIQKSFKHYILDDTKEHFAEIDKDANDIVTWDEYNMHMYDRI 126
Query: 176 -DIEKKNMAHGEAGWLME--------KFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLK 226
D ++ + E +FD AD D LN +E DF HPE++ + M +
Sbjct: 127 IDYDENTVLEDEEEESFRQIHLKDKRRFDHADRDEISGLNLSEFTDFEHPEETDH--MSE 184
Query: 227 WMVNDKFNMDDYEHDGKINFNQFEDNVYVTYESYVDFETNGEGDIP------TAKDKFA- 279
+++ D + DG +V+ E Y+ T G + KD+F
Sbjct: 185 FVIEGALEEHDKDGDG-----------FVSLEEYLGDYTQDPGTVEDPHWLIVEKDRFVN 233
Query: 280 ELDVNKDQFLSPEELFPIIPYVYPGELAYAKYYTSYLMNEADDNEDRKLTLDEMLDHEFA 339
+ D + D L+P EL + ++ P L ++ S+LM E D NED++L+ +E+L +
Sbjct: 234 DYDKDGDGRLNPTEL---LSWIVPNNLGISQEEASHLMEEMDKNEDQRLSEEEILQSKDI 290
Query: 340 FFNTVHADGHVEIDDDD--HDEL 360
F ++ D ++ D HDEL
Sbjct: 291 FLSSEATDYGRQLQDKHFYHDEL 313
>pir||A57516 DNA supercoiling factor - silkworm
Length = 322
Score = 75.9 bits (185), Expect = 2e-12
Identities = 74/310 (23%), Positives = 137/310 (43%), Gaps = 39/310 (12%)
Query: 52 LVTKIERESEQKNQQHKNDFDNNKNVAPRTGLGSTTTVSEIKETYEYLTSGGTLNTTLRL 111
L+ + +N+ H FD++ + + +T++ L+ + RL
Sbjct: 27 LMDHLSDAEHYRNEHHNKQFDHDAFLG-----------EDQAKTFDQLSPE---ESKRRL 72
Query: 112 IILFPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREY--- 168
+ +D D +DGF+ EL+ W+ +D + ++ N + +++ Y
Sbjct: 73 GEIADKIDSD-QDGFITLVELKDWIRYTQKRYIDEDVERHWRQQNPNNEEFVTWEAYRKN 131
Query: 169 ----LPDLSEKDIEKKNMAHGEAGWLMEKFD-----VADYDHNGLLNFTELRDFLHPEDS 219
+ D+ EK+++ N + G ++K D AD D N LN TE FLHPED
Sbjct: 132 VYGFMDDMDEKELKAPN-SEGFTYSNLQKRDRRRWTYADADQNDALNRTEFAAFLHPED- 189
Query: 220 QNKEMLKWMVNDKFNMDDYEHDGKINFNQFEDNVYVTYESYVDFETNGEGD-IPTAKDKF 278
+ M +V + D + DGK++ +++ ++Y + D E E D + +++F
Sbjct: 190 -HSSMRDVVVLETLEDIDKDQDGKVSLDEYIGDMY----NAGDGEDEEEPDWVKQEREQF 244
Query: 279 AEL-DVNKDQFLSPEELFPIIPYVYPGELAYAKYYTSYLMNEADDNEDRKLTLDEMLDHE 337
D NKD F+ E + ++ P E +A+ +L+ EAD + D KLT E++D
Sbjct: 245 TGYRDTNKDGFMDEHE---VKDWIAPPEFDHAEAEARHLVFEADADADEKLTKAEIIDKY 301
Query: 338 FAFFNTVHAD 347
F + D
Sbjct: 302 DLFVGSQATD 311
>dbj|BAA08704.1| DNA supercoiling factor [Bombyx mori]
Length = 322
Score = 75.9 bits (185), Expect = 2e-12
Identities = 74/310 (23%), Positives = 137/310 (43%), Gaps = 39/310 (12%)
Query: 52 LVTKIERESEQKNQQHKNDFDNNKNVAPRTGLGSTTTVSEIKETYEYLTSGGTLNTTLRL 111
L+ + +N+ H FD++ + + +T++ L+ + RL
Sbjct: 27 LMDHLSDAEHYRNEHHNKQFDHDAFLG-----------EDQAKTFDQLSPE---ESKRRL 72
Query: 112 IILFPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREY--- 168
+ +D D +DGF+ EL+ W+ +D + ++ N + +++ Y
Sbjct: 73 GEIADKIDSD-QDGFITLVELKDWIRYTQKRYIDEDVERHWRQQNPNNEEFVTWEAYRKN 131
Query: 169 ----LPDLSEKDIEKKNMAHGEAGWLMEKFD-----VADYDHNGLLNFTELRDFLHPEDS 219
+ D+ EK+++ N + G ++K D AD D N LN TE FLHPED
Sbjct: 132 VYGFMDDMDEKELKAPN-SEGFTYSNLQKRDRRRWTYADADQNDALNRTEFAAFLHPED- 189
Query: 220 QNKEMLKWMVNDKFNMDDYEHDGKINFNQFEDNVYVTYESYVDFETNGEGD-IPTAKDKF 278
+ M +V + D + DGK++ +++ ++Y + D E E D + +++F
Sbjct: 190 -HSSMRDVVVLETLEDIDKDQDGKVSLDEYIGDMY----NAGDGEDEEEPDWVKQEREQF 244
Query: 279 AEL-DVNKDQFLSPEELFPIIPYVYPGELAYAKYYTSYLMNEADDNEDRKLTLDEMLDHE 337
D NKD F+ E + ++ P E +A+ +L+ EAD + D KLT E++D
Sbjct: 245 TGYRDTNKDGFMDEHE---VKDWIAPPEFDHAEAEARHLVFEADADADEKLTKAEIIDKY 301
Query: 338 FAFFNTVHAD 347
F + D
Sbjct: 302 DLFVGSQATD 311
>gb|EAL33018.1| GA16367-PA [Drosophila pseudoobscura]
Length = 345
Score = 75.5 bits (184), Expect = 2e-12
Identities = 76/307 (24%), Positives = 144/307 (46%), Gaps = 35/307 (11%)
Query: 67 HKNDFDNNKNVAPRTGLGSTTTVSEIKETYEYLTSGGTLNTTLRLIILFPLLDRDPKDGF 126
H D ++N +G+T KE E+ T + RL++L L+D + KD F
Sbjct: 61 HGEDGEHNVEFDHEAIIGNT------KEAQEFDTLSPE-ESKRRLLVLVKLMDLN-KDEF 112
Query: 127 VGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYLPD---LSEKDIEKKNM- 182
V +EL++W+ + + + + D+ D ++++EYL D + +++ +K+ +
Sbjct: 113 VDRHELKAWILRSFKKLSEEEAADRFDEIDQETDERITWKEYLQDTYSMEDENFKKETID 172
Query: 183 --AHGEAGWLM----EKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVNDKFNMD 236
+ E ++ E F+ AD + +G+L+ E F +PE ++ +ML ++
Sbjct: 173 FDNYEEEQKMIKQDKEMFNAADINKDGVLSLEEFVYFHNPE--EHPQMLPILLEHTMQDK 230
Query: 237 DYEHDGKINFNQFEDNVYVTYESYVDFETNGEGDIPTAKDKF-AELDVNKDQFLSPEELF 295
D HDGKINF +F S+ D E + T K++F + D+N D L+ E
Sbjct: 231 DLNHDGKINFQEFVGEA----ASHHDKEW-----LLTEKERFDKDHDINGDGVLTGNE-- 279
Query: 296 PIIPYVYPGELAYAKYYTSYLMNEADDNEDRKLTLDEMLDHEFAFFNTVHADGHVEIDDD 355
++ ++ P A A +L D++ D +L+ E+L++ F + D + +
Sbjct: 280 -VLSWIVPSNTAIASDEVDHLFVSTDEDHDDRLSYLEILNNYETFVGSEATDYGDHLQNI 338
Query: 356 DH--DEL 360
+H DEL
Sbjct: 339 NHLADEL 345
>ref|XP_311555.1| ENSANGP00000010221 [Anopheles gambiae str. PEST]
gi|21295095|gb|EAA07240.1| ENSANGP00000010221 [Anopheles
gambiae str. PEST]
Length = 328
Score = 75.1 bits (183), Expect = 3e-12
Identities = 70/295 (23%), Positives = 135/295 (45%), Gaps = 33/295 (11%)
Query: 66 QHKNDFDNNKNVAPRTGLGSTTTVSEIKETYEYLTSGGTLNTTLRLIILFPLLDRDPKDG 125
QH + ++NK LG E +T++ L + + RL ++ +DRD DG
Sbjct: 43 QHYQNDEHNKQYDHEAFLG------EDAKTFDQLEAD---ESRRRLGLIVDKIDRD-NDG 92
Query: 126 FVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREY-------LPDLSEKDIE 178
FV +EL++W+ +D + ++ + N + + Y L +L+ ++ +
Sbjct: 93 FVNMSELKAWIQYTQRRYIDDDVNRQWKTHNPNNTEKVHWDTYRKNVYGFLDELAAQEPD 152
Query: 179 KKNMAHGEAGWLMEK----FDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVNDKFN 234
+ H +M++ + +AD D + L E DFLHPE+S + M +V +
Sbjct: 153 HPSDEHFSYRTMMKRDRRRWSIADRDGDDELTREEFTDFLHPEESSH--MRDVVVTETIE 210
Query: 235 MDDYEHDGKINFNQFEDNVYVTYESYVDFETNGEGD-IPTAKDKFAEL-DVNKDQFLSPE 292
D + DGK++ ++ ++Y E + E D + ++ F D NKD F+ +
Sbjct: 211 DIDKDSDGKVSVEEYIGDMYRQGE-----QNEEEPDWVKHERETFTNFRDKNKDGFMDNQ 265
Query: 293 ELFPIIPYVYPGELAYAKYYTSYLMNEADDNEDRKLTLDEMLDHEFAFFNTVHAD 347
E + ++ P + +A+ +L+ EAD + D KLT +E+++ F + D
Sbjct: 266 E---VKDWITPADFDHAEAEARHLIYEADSDADEKLTKEEIIEKYDLFVGSQATD 317
>gb|AAH71338.1| Zgc:86646 [Danio rerio] gi|50344972|ref|NP_001002158.1| zgc:86646
[Danio rerio]
Length = 316
Score = 75.1 bits (183), Expect = 3e-12
Identities = 69/251 (27%), Positives = 111/251 (43%), Gaps = 20/251 (7%)
Query: 118 LDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFRE-------YLP 170
+D D KDGFV EL W+ R ++ D+N D + + E Y
Sbjct: 78 IDTD-KDGFVSHAELHHWIKHRQRRYIEENVDKHWNEYDQNKDGKIGWIEYKNTTYGYYI 136
Query: 171 DLSEKDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPEDSQNKEMLKWMVN 230
D D++ K +F AD D +G+ E FLHPE+ M ++
Sbjct: 137 DTEFDDVDDKATYKSMLNRDERRFKSADRDGDGVATREEFTAFLHPEEFD--FMRDIVIQ 194
Query: 231 DKFNMDDYEHDGKINFNQFEDNVYVTYESYVDFETNGEGDIPTAKDKFAEL-DVNKDQFL 289
+ D DGKI+ ++ ++Y + D ET + + T K +F+E D+NKD FL
Sbjct: 195 ETIEDIDKNGDGKIDLQEYIGDMY----NPEDGETEPDW-VTTEKKQFSEFRDMNKDGFL 249
Query: 290 SPEELFPIIPYVYPGELAYAKYYTSYLMNEADDNEDRKLTLDEMLDHEFAFFNTVHADGH 349
E + ++ P E+ +A +L++E D + D K+T E+L++ F + A +
Sbjct: 250 DATE---VSHWILPTEVDHADNEARHLIHETDKDNDDKITKKEILENWNMFVGS-QATNY 305
Query: 350 VEIDDDDHDEL 360
E HDEL
Sbjct: 306 GEDLTKRHDEL 316
>ref|XP_393699.2| PREDICTED: similar to CG31650-PC, isoform C [Apis mellifera]
Length = 331
Score = 74.3 bits (181), Expect = 5e-12
Identities = 72/262 (27%), Positives = 117/262 (44%), Gaps = 21/262 (8%)
Query: 89 VSEIKETYEYLTSGGTLNTTLRLIILFPLLDRDPKDGFVGFNELESWVTQRALERLDYAT 148
+ +KE E+ T + RL IL +D + D F+ NEL++W+ + +
Sbjct: 73 LGSVKEAEEF-DKLPTQESKRRLGILLTKMDLN-NDKFIERNELKAWILRSFSMLSAEES 130
Query: 149 QVELESKDKNGDLALSFREYLPDLSEKDIEKKNMAHGEAGWLMEKFDVADYDHNGLLNFT 208
Q LE D + D +S+ E L D D E + + F+ AD + +G L+
Sbjct: 131 QDRLEDTDTDEDGKVSWNEILQDTYGTDPEDLAVDDKLISDDKQTFEAADINKDGHLDKE 190
Query: 209 ELRDFLHPEDSQNKE--MLKWMVNDKFNMDDYEHDGKINFNQFEDNVYVTYESYVDFETN 266
E + + HPE++ +LK ++DK D + DG I+F +F N +
Sbjct: 191 EFKAYTHPEETPRMFPLLLKQALDDK----DTDKDGFISFQEFIGN---------RAKAE 237
Query: 267 GEGDIPTAKDKFA-ELDVNKDQFLSPEELFPIIPYVYPGELAYAKYYTSYLMNEADDNED 325
+ + KDKF E D N D L +E I+ ++ P A +L +DD+ D
Sbjct: 238 DKEWLLIEKDKFDYEHDKNGDGRLDSDE---ILSWLVPSNEEIASDEVDHLFAASDDDHD 294
Query: 326 RKLTLDEMLDHEFAFFNTVHAD 347
+L+ DE+LDH AF + D
Sbjct: 295 NRLSFDEILDHHDAFVGSEATD 316
>ref|XP_413734.1| PREDICTED: similar to Reticulocalbin 2 precursor (Calcium-binding
protein ERC-55) (Taipoxin-associated calcium-binding
protein-49) (TCBP-49) [Gallus gallus]
Length = 307
Score = 72.0 bits (175), Expect = 3e-11
Identities = 68/264 (25%), Positives = 122/264 (45%), Gaps = 24/264 (9%)
Query: 110 RLIILFPLLDRDPKDGFVGFNELESWVTQRALERLDYATQVELESKDKNGDLALSFREYL 169
RL + +D D DG + +EL SW+ Q + + DKNGD +S++EY
Sbjct: 55 RLKAIVRRIDAD-NDGLLSKDELSSWIQQSFKHYVTQEAKQHFHDYDKNGDGLVSWKEYN 113
Query: 170 PDLSEK--DIEKKNMAHGEAG------WLMEK--FDVADYDHNGLLNFTELRDFLHPEDS 219
+ ++ D ++ + + L EK F+ A+ D + LN E F HPE+
Sbjct: 114 LQMYDRVIDFDENTVLEDQEEESFRQLHLKEKKRFEKANRDDDPDLNVDEFIAFEHPEEV 173
Query: 220 QNKEMLKWMVNDKFNMDDYEHDGKINFNQFEDNVYVTYESYVDFETNGEGDIPTAKDKFA 279
+ M +++ + D + DG ++ +F + + D E I KD+F
Sbjct: 174 E--YMTDFVIEEALEEHDKDGDGFVSLEEFLGDYRRDPTAKEDPEW-----ILVEKDRFV 226
Query: 280 -ELDVNKDQFLSPEELFPIIPYVYPGELAYAKYYTSYLMNEADDNEDRKLTLDEMLDHEF 338
+ D + D L+P+EL + ++ P A+ +L+ E D N+D+KL+ E+L ++
Sbjct: 227 NDYDKDNDGKLNPQEL---LSWIVPNNQGIAQEEALHLIEEMDLNDDKKLSEAEVLKNQD 283
Query: 339 AFFNTVHADGHVEIDDDD--HDEL 360
F N+ D ++ D+ H+EL
Sbjct: 284 LFLNSEATDYGRQLHDERFYHEEL 307
>gb|EAL29954.1| GA21575-PA [Drosophila pseudoobscura]
Length = 328
Score = 72.0 bits (175), Expect = 3e-11
Identities = 68/242 (28%), Positives = 117/242 (48%), Gaps = 21/242 (8%)
Query: 110 RLIILFPLLDRDPKDGFVGFNELESWV--TQRALERLDYATQVELESKDKNGDLAL---- 163
RL ++F +D D KDG V +EL++W+ TQR D + D N ++
Sbjct: 79 RLGVIFDRIDED-KDGLVTLSELKNWISFTQRRYIEEDTGRLWRQHNPDNNATISWEAYR 137
Query: 164 -SFREYLPDLSEKDIEKKNMAHGEAGWLME---KFDVADYDHNGLLNFTELRDFLHPEDS 219
S +L DLS +++ ++ G L ++ VAD D + L E FLHPED
Sbjct: 138 DSVYSFLNDLSAEELAQEENGISYKGLLKRDRRRWAVADQDLDDSLTREEFTAFLHPED- 196
Query: 220 QNKEMLKWMVNDKFNMDDYEHDGKINFNQFEDNVYVTYESYVDFETNGEGDIPTAKDKFA 279
+ M ++ + D + DGKIN +++ ++Y E+ E E + + ++ FA
Sbjct: 197 -HPTMRDVVLQETVEDLDKDKDGKINEDEYIGDMYRPSEAN---EEEPEW-VASEREAFA 251
Query: 280 EL-DVNKDQFLSPEELFPIIPYVYPGELAYAKYYTSYLMNEADDNEDRKLTLDEMLDHEF 338
+ D + D +L+ E + ++ P + +++ +L+ EAD + D KLT E+LD
Sbjct: 252 KYRDTDGDGYLTETE---VRQWITPQDFDHSESEAKHLIFEADVDHDEKLTKAEVLDKYD 308
Query: 339 AF 340
AF
Sbjct: 309 AF 310
>emb|CAE68672.1| Hypothetical protein CBG14576 [Caenorhabditis briggsae]
Length = 314
Score = 71.2 bits (173), Expect = 4e-11
Identities = 71/263 (26%), Positives = 119/263 (44%), Gaps = 24/263 (9%)
Query: 110 RLIILFPLLDRDPKDGFVGFNELESWVT--QRALERLDYATQVELESKDKNGDLALSFRE 167
+L L P +D D DGF+ NEL+ + Q+ D + +K D + + +
Sbjct: 64 KLAKLVPKMDAD-SDGFIEENELKDHINFMQKRYVNNDVDRTWKNYKAEKIVDGKIKWED 122
Query: 168 YLP------DLSEKDIEK---KNMAHGEAGWLMEKFDVADYDHNGLLNFTELRDFLHPED 218
Y D + +++ K +A E W VADYD NG L+ TE F+HPED
Sbjct: 123 YREMVYGSADGAGQELSPEYAKMIARDEKRWA-----VADYDSNGALDRTEYGCFMHPED 177
Query: 219 SQNKEMLKWMVNDKFNMDDYEHDGKINFNQFEDNVYVTYESYVDFETNGEGDIPTAKDKF 278
+ M +V + + D DG ++ ++ ++Y E Y + + + ++ F
Sbjct: 178 CDH--MRDIVVAETVDDIDKNKDGSVDLEEYIGDMYRP-EDYPELAGKEPDWVQSEREMF 234
Query: 279 AE-LDVNKDQFLSPEELFPIIPYVYPGELAYAKYYTSYLMNEADDNEDRKLTLDEMLDHE 337
E D + D L+ EE+ ++ P +A+ +L+ ADDN+D KLTLDE++ H
Sbjct: 235 KEHRDKDGDGKLNQEEMRD---WIMPVGFDHAEAEARHLVGIADDNKDGKLTLDEIVAHY 291
Query: 338 FAFFNTVHADGHVEIDDDDHDEL 360
F + D ++ D EL
Sbjct: 292 DTFVGSQATDYGEQLQKHDPAEL 314
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.315 0.136 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 672,956,704
Number of Sequences: 2540612
Number of extensions: 31530394
Number of successful extensions: 79773
Number of sequences better than 10.0: 758
Number of HSP's better than 10.0 without gapping: 37
Number of HSP's successfully gapped in prelim test: 730
Number of HSP's that attempted gapping in prelim test: 78704
Number of HSP's gapped (non-prelim): 1208
length of query: 360
length of database: 863,360,394
effective HSP length: 129
effective length of query: 231
effective length of database: 535,621,446
effective search space: 123728554026
effective search space used: 123728554026
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 76 (33.9 bits)
Medicago: description of AC149127.7


