

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
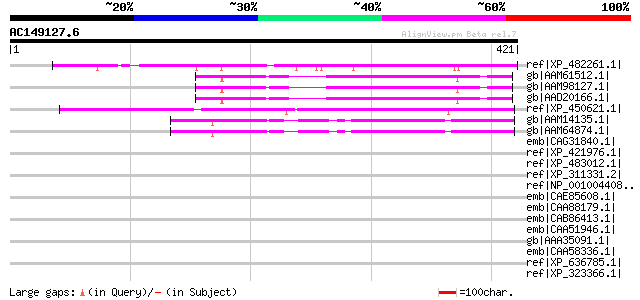
Query= AC149127.6 - phase: 0
(421 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
ref|XP_482261.1| unknown protein [Oryza sativa (japonica cultiva... 182 2e-44
gb|AAM61512.1| unknown [Arabidopsis thaliana] 168 3e-40
gb|AAM98127.1| expressed protein [Arabidopsis thaliana] gi|25083... 168 3e-40
gb|AAD20166.1| expressed protein [Arabidopsis thaliana] gi|18407... 168 3e-40
ref|XP_450621.1| unknown protein [Oryza sativa (japonica cultiva... 150 9e-35
gb|AAM14135.1| unknown protein [Arabidopsis thaliana] gi|1581026... 147 6e-34
gb|AAM64874.1| unknown [Arabidopsis thaliana] 144 4e-33
emb|CAG31840.1| hypothetical protein [Gallus gallus] 45 0.006
ref|XP_421976.1| PREDICTED: similar to KIAA1604 protein [Gallus ... 45 0.006
ref|XP_483012.1| unknown protein [Oryza sativa (japonica cultiva... 44 0.009
ref|XP_311331.2| ENSANGP00000001657 [Anopheles gambiae str. PEST... 44 0.012
ref|NP_001004408.1| hypothetical protein LOC424547 [Gallus gallu... 44 0.012
emb|CAE85608.1| putative protein [Neurospora crassa] 43 0.021
emb|CAA88179.1| gar2 [Schizosaccharomyces pombe] 42 0.036
emb|CAB86413.1| gar2 [Schizosaccharomyces pombe] gi|19114443|ref... 42 0.036
emb|CAA51946.1| unnamed protein product [Saccharomyces cerevisia... 42 0.047
gb|AAA35091.1| selected as a weak suppressor of a mutant of the ... 42 0.047
emb|CAA58336.1| U87 [Human herpesvirus 6] gi|9628389|ref|NP_0429... 41 0.061
ref|XP_636785.1| hypothetical protein DDB0219334 [Dictyostelium ... 41 0.061
ref|XP_323366.1| predicted protein [Neurospora crassa] gi|289187... 41 0.061
>ref|XP_482261.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|37805969|dbj|BAC99384.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
gi|37572976|dbj|BAC98668.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 481
Score = 182 bits (461), Expect = 2e-44
Identities = 136/429 (31%), Positives = 208/429 (47%), Gaps = 57/429 (13%)
Query: 36 SDSTDSDMKWWLHVKTNLAGDTDYTCQHLNTLESKL------DAFSTRLHGDNVSIGSDQ 89
+ S D++WWL ++ NL G + +HL L ++ D+ +H + +
Sbjct: 67 NSSLPPDIRWWLQLQPNLGGQKNLAGEHLYFLGREISDKEVEDSAQKNIHDEPLFCEMFD 126
Query: 90 TVKNFDAVSFVGNAATAAIEQQWNVYPKYMK-NSDTRTSKIEASLNSDLYLTPKKKNEGE 148
T N + + V E W V MK +S+T ++ + K+N +
Sbjct: 127 T--NPEKIEDV-------FEPSWMVSTASMKYSSETGLQDLKNIGGYSQVPSKCKENASD 177
Query: 149 FWFSD----DATSFLISEHCKSTSSDFEPH--WLGAEKSQPWWRTTGKDDLASLVAKKSF 202
F+D D +F DF+ + W G E+SQPWW+ T +++LA LVA+++
Sbjct: 178 CLFNDKEFLDFKNFNPPPSKNPQKDDFDMNAPWKGGERSQPWWQITDENELALLVAERAM 237
Query: 203 EYVENCDLPEPNIKPFRKIHTLQPRETDKEENQV---SSLNQKLEMCSSDSNGC------ 253
+++ENCDLP P +I +Q E+ EN S M D+ C
Sbjct: 238 QHIENCDLPRPT-----QIVRVQGTESRSHENMGRYRGSSGPAGTMSYPDTGQCEHIECS 292
Query: 254 --TSTT----LTSGCSFQDSDRTFSSSESKDSDSSCN---KNSKVNSESAAKAELLKALC 304
T++T LTS +Q +R + S+++D N + + A +A+LL+ALC
Sbjct: 293 YSTASTDEVDLTSDGVWQQQERNVARSDAQDFSRGINTEPRGKRTYQNPAEQAQLLEALC 352
Query: 305 RSQTRAREAEKAAQEACDEKEHILSLFFKQASQLFAYKQWLHVLQLENLCLQYKSKNQPL 364
SQTRAREAE A ++A EK+ ++ LFF+QAS LFA KQWL +LQLEN+CLQ K + +
Sbjct: 353 HSQTRAREAEMAGKKAQSEKDDVIKLFFRQASHLFACKQWLKMLQLENICLQLKHREHQI 412
Query: 365 LNNL--FPY---------EGKKHRKNRRKVKNSRRGIGKC-IFAFAVGLALAGAGLLLGW 412
+ P+ + +K R+K + + C FAVGL LAGAGLLLGW
Sbjct: 413 ATMIPDIPWITLKKRTTPDHEKEDWTRKKGRRHKNAGSFCDALLFAVGLGLAGAGLLLGW 472
Query: 413 TIGCMFPSF 421
T G + F
Sbjct: 473 TFGWLLAKF 481
>gb|AAM61512.1| unknown [Arabidopsis thaliana]
Length = 397
Score = 168 bits (425), Expect = 3e-40
Identities = 109/281 (38%), Positives = 149/281 (52%), Gaps = 55/281 (19%)
Query: 155 ATSFLISEHCKSTSSDFEPH--W--LGAEKSQPWWRTTGKDDLASLVAKKSFEYVENCDL 210
+T + S K + F+P W L +EK+ PWWRTT KD+LASLVA++S +YVENCDL
Sbjct: 149 STGYDESSEKKLSELSFDPSSPWNLLSSEKAAPWWRTTDKDELASLVAQRSLDYVENCDL 208
Query: 211 PEPNIKPFRKIHTLQPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRT 270
P P + ++ + PR D + +D +
Sbjct: 209 PTP--QKMKRSYYGSPRGFDSD------------------------------GLRDYSVS 236
Query: 271 FSSSESKDSDSSCNKNSKVNSES-AAKAELLKALCRSQTRAREAEKAAQEACDEKEHILS 329
+ + SSC + +SES +K+ELL+AL RSQTRAREAE A+EA EKEH++
Sbjct: 237 GQTIKGTSKGSSCKNRPEASSESDLSKSELLEALRRSQTRAREAENMAKEAYAEKEHLVK 296
Query: 330 LFFKQASQLFAYKQWLHVLQLENLCLQYKSKNQPLLNNLFP-------------YEGKKH 376
+ KQA++LF YKQWL +LQLE L LQ K+K NN P EG+K
Sbjct: 297 ILLKQAAELFGYKQWLQLLQLEALYLQIKNKEIDNKNNDDPGVSIPCWSNGKARKEGRKR 356
Query: 377 RKNRRKVKNSRRGIGKCIFAFAVGLALAGAGLLLGWTIGCM 417
R R K ++ +G A+G++L GAGLLLGWT+G M
Sbjct: 357 RSKRGKPNGAKYAVG-----LALGMSLVGAGLLLGWTVGWM 392
>gb|AAM98127.1| expressed protein [Arabidopsis thaliana] gi|25083763|gb|AAN72116.1|
expressed protein [Arabidopsis thaliana]
Length = 397
Score = 168 bits (425), Expect = 3e-40
Identities = 109/281 (38%), Positives = 149/281 (52%), Gaps = 55/281 (19%)
Query: 155 ATSFLISEHCKSTSSDFEPH--W--LGAEKSQPWWRTTGKDDLASLVAKKSFEYVENCDL 210
+T + S K + F+P W L +EK+ PWWRTT KD+LASLVA++S +YVENCDL
Sbjct: 149 STGYDESSEKKLSELSFDPSSPWNLLSSEKAAPWWRTTDKDELASLVAQRSLDYVENCDL 208
Query: 211 PEPNIKPFRKIHTLQPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRT 270
P P + ++ + PR D + +D +
Sbjct: 209 PTP--QKMKRSYYGSPRGFDSD------------------------------GLRDYSVS 236
Query: 271 FSSSESKDSDSSCNKNSKVNSES-AAKAELLKALCRSQTRAREAEKAAQEACDEKEHILS 329
+ + SSC + +SES +K+ELL+AL RSQTRAREAE A+EA EKEH++
Sbjct: 237 GQTIKGTSKGSSCKNRPEASSESDLSKSELLEALRRSQTRAREAENMAKEAYAEKEHLVK 296
Query: 330 LFFKQASQLFAYKQWLHVLQLENLCLQYKSKNQPLLNNLFP-------------YEGKKH 376
+ KQA++LF YKQWL +LQLE L LQ K+K NN P EG+K
Sbjct: 297 ILLKQAAELFGYKQWLQLLQLEALYLQIKNKEIDNKNNDDPGVSIPCWSNGKARKEGRKR 356
Query: 377 RKNRRKVKNSRRGIGKCIFAFAVGLALAGAGLLLGWTIGCM 417
R R K ++ +G A+G++L GAGLLLGWT+G M
Sbjct: 357 RSKRGKPNGAKYAVG-----LALGMSLVGAGLLLGWTVGWM 392
>gb|AAD20166.1| expressed protein [Arabidopsis thaliana]
gi|18407105|ref|NP_566080.1| expressed protein
[Arabidopsis thaliana] gi|25370833|pir||C84904
hypothetical protein At2g46550 [imported] - Arabidopsis
thaliana
Length = 397
Score = 168 bits (425), Expect = 3e-40
Identities = 109/281 (38%), Positives = 149/281 (52%), Gaps = 55/281 (19%)
Query: 155 ATSFLISEHCKSTSSDFEPH--W--LGAEKSQPWWRTTGKDDLASLVAKKSFEYVENCDL 210
+T + S K + F+P W L +EK+ PWWRTT KD+LASLVA++S +YVENCDL
Sbjct: 149 STGYDESSEKKLSELSFDPSSPWNLLSSEKAAPWWRTTDKDELASLVAQRSLDYVENCDL 208
Query: 211 PEPNIKPFRKIHTLQPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRT 270
P P + ++ + PR D + +D +
Sbjct: 209 PTP--QKMKRSYYGSPRGFDSD------------------------------GLRDYSVS 236
Query: 271 FSSSESKDSDSSCNKNSKVNSES-AAKAELLKALCRSQTRAREAEKAAQEACDEKEHILS 329
+ + SSC + +SES +K+ELL+AL RSQTRAREAE A+EA EKEH++
Sbjct: 237 GQTIKGTSKGSSCKNRPEASSESDLSKSELLEALRRSQTRAREAENMAKEAYAEKEHLVK 296
Query: 330 LFFKQASQLFAYKQWLHVLQLENLCLQYKSKNQPLLNNLFP-------------YEGKKH 376
+ KQA++LF YKQWL +LQLE L LQ K+K NN P EG+K
Sbjct: 297 ILLKQAAELFGYKQWLQLLQLEALYLQIKNKEIDNKNNDDPGVSIPCWSNGKARKEGRKR 356
Query: 377 RKNRRKVKNSRRGIGKCIFAFAVGLALAGAGLLLGWTIGCM 417
R R K ++ +G A+G++L GAGLLLGWT+G M
Sbjct: 357 RSKRGKPNGAKYAVG-----LALGMSLVGAGLLLGWTVGWM 392
>ref|XP_450621.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|50726194|dbj|BAD33713.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
gi|48716731|dbj|BAD23412.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 463
Score = 150 bits (378), Expect = 9e-35
Identities = 112/396 (28%), Positives = 178/396 (44%), Gaps = 24/396 (6%)
Query: 42 DMKWWLHVKTNLAGDTDYTCQHLNTLESKLDAFSTRLHGDNVSIGSDQTVKNFDAVSFVG 101
D +WWL ++ N + LN + ++D + K D
Sbjct: 72 DTQWWLQLQPNFGCQNVLASEDLNYMCGEVDVKKVESFAPVSKLEDINPKKTADPFEPPW 131
Query: 102 NAATAAIEQQWNVYPKYMKNSDTRTSKIEASLNSDLYLTPKKKNEGEFWFSDDATSFLIS 161
+TA ++Q + + +K+ + S YL K++ F L
Sbjct: 132 IVSTAFMKQTYETGFEELKSLPAYSEMTLKCRGSATYLHEDKEHMDFKTFDP-----LYP 186
Query: 162 EHCKSTSSDFEPHWLGAEKSQPWWRTTGKDDLASLVAKKSFEYVENCDLPEPNIKPF-RK 220
+ ++ + + W KS+PWW+ D LAS+VA+ V +LP P + K
Sbjct: 187 KKPQTACYEMDAPWQENRKSRPWWQVAEADGLASVVAESEMHNVGKNELPRPTQRAHGSK 246
Query: 221 IHTLQPRE-----TDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSE 275
++ + ++ T KE V + L S S T+++ G Q +D + +
Sbjct: 247 LNNHENKDDYGPYTGKESPPVQ-YDTMLCSYSISSTNETNSSDGGGWQHQRNDARGGTQD 305
Query: 276 SKDSDSSCNKNSKVNSESAAKAELLKALCRSQTRAREAEKAAQEACDEKEHILSLFFKQA 335
S SD + +A + +LL AL SQTRAREAE AA++A DEK+H++ L F+QA
Sbjct: 306 SCSSDDRTPGSKPTYRSAAERTQLLDALRHSQTRAREAEMAAKKAYDEKDHVIKLLFRQA 365
Query: 336 SQLFAYKQWLHVLQLENLCLQYKSKNQP------------LLNNLFPYEGKKHRKNRRKV 383
S LFA KQWL +LQLEN+CLQ + K L + P + +K ++
Sbjct: 366 SHLFACKQWLKMLQLENICLQLRLKEHQIAAMFPELPWIMLKEKVTPGQERKDGTRKKGR 425
Query: 384 KNSRRGIGKCIFAFAVGLALAGAGLLLGWTIGCMFP 419
K ++ + FAVG+ + GAGLLLGWT+G + P
Sbjct: 426 KQNKDSHLRKAVVFAVGVGIVGAGLLLGWTLGWLLP 461
>gb|AAM14135.1| unknown protein [Arabidopsis thaliana] gi|15810265|gb|AAL07020.1|
unknown protein [Arabidopsis thaliana]
gi|42571289|ref|NP_973735.1| expressed protein
[Arabidopsis thaliana] gi|18378816|ref|NP_563623.1|
expressed protein [Arabidopsis thaliana]
gi|42571291|ref|NP_973736.1| expressed protein
[Arabidopsis thaliana] gi|16226611|gb|AAL16213.1|
At1g01240/F6F3_11 [Arabidopsis thaliana]
gi|25511619|pir||F86142 F633.5 protein - Arabidopsis
thaliana gi|9665136|gb|AAF97320.1| Unknown protein
[Arabidopsis thaliana]
Length = 331
Score = 147 bits (371), Expect = 6e-34
Identities = 104/301 (34%), Positives = 157/301 (51%), Gaps = 41/301 (13%)
Query: 134 NSDLYLTPKKKNEGEFWFSDDAT-SFLISEHCKST-----------SSDFEPHWLGAEKS 181
+SD K +G F ++ T SFL CK D+ A+ +
Sbjct: 56 SSDFSCDTKWWLKGSTGFDEEVTNSFLEDTKCKKLHEFVDLIGIREEEDYSFISKKADAT 115
Query: 182 QPWWR-TTGKDDLASLVAKKSFEY-VENCDLPEPNIKPFRKIHTLQPRETDKEENQVSSL 239
PWWR TT KD+LA +VA KS ++ ++NCDLP P K + IH+ +
Sbjct: 116 TPWWRSTTDKDELALMVATKSVDHNIQNCDLPPPQ-KLHKSIHSSSGEK----------- 163
Query: 240 NQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAAKAEL 299
K + S G S+ S+ESK++ + S S+ +K +L
Sbjct: 164 GFKTAVKSPWKQGVWKDRFERSLSYN------GSTESKNT----SPMSSPRSDDLSKGQL 213
Query: 300 LKALCRSQTRAREAEKAAQEACDEKEHILSLFFKQASQLFAYKQWLHVLQLENLCLQYKS 359
L+AL SQTRAREAE+AA+EAC EK+ ++++ KQASQ+ AYKQWL +L++E L LQ K
Sbjct: 214 LEALRHSQTRAREAERAAREACAEKDRVITILLKQASQMLAYKQWLKLLEMEALYLQMKK 273
Query: 360 KNQPLLNNLFPYEGKKHRKNRRKVKNSRRG-IGKCIFAFAVGLALAGAGLLLGWTIGCMF 418
+ + +G +K +++ + ++G G+ + AFA+G +L GAGLLLGWT+G +
Sbjct: 274 EEE----QEEQVKGMNLKKRKQRGEKKKKGETGRYMMAFALGFSLIGAGLLLGWTVGWLL 329
Query: 419 P 419
P
Sbjct: 330 P 330
>gb|AAM64874.1| unknown [Arabidopsis thaliana]
Length = 331
Score = 144 bits (364), Expect = 4e-33
Identities = 103/301 (34%), Positives = 156/301 (51%), Gaps = 41/301 (13%)
Query: 134 NSDLYLTPKKKNEGEFWFSDDAT-SFLISEHCKST-----------SSDFEPHWLGAEKS 181
+SD K +G F ++ T SFL CK D+ A+ +
Sbjct: 56 SSDFSCDTKWWLKGSTGFDEEVTNSFLEDTKCKKLHEFVDLIGIREEEDYSFISKKADAT 115
Query: 182 QPWWR-TTGKDDLASLVAKKSFEY-VENCDLPEPNIKPFRKIHTLQPRETDKEENQVSSL 239
PWWR TT KD+LA +VA KS ++ ++NCDLP P K + IH+ +
Sbjct: 116 TPWWRSTTDKDELALMVATKSVDHNIQNCDLPPPQ-KLHKSIHSSSGEK----------- 163
Query: 240 NQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAAKAEL 299
K + S G S+ S+ESK++ + S S+ +K +L
Sbjct: 164 GFKTAVKSPWKQGVWKDRFERSLSYN------GSTESKNT----SPMSSPRSDDLSKGQL 213
Query: 300 LKALCRSQTRAREAEKAAQEACDEKEHILSLFFKQASQLFAYKQWLHVLQLENLCLQYKS 359
L+AL SQTRA EAE+AA+EAC EK+ ++++ KQASQ+ AYKQWL +L++E L LQ K
Sbjct: 214 LEALRHSQTRAGEAERAAREACAEKDRVITILLKQASQMLAYKQWLKLLEMEALYLQMKK 273
Query: 360 KNQPLLNNLFPYEGKKHRKNRRKVKNSRRG-IGKCIFAFAVGLALAGAGLLLGWTIGCMF 418
+ + +G +K +++ + ++G G+ + AFA+G +L GAGLLLGWT+G +
Sbjct: 274 EEE----QEEQVKGMNLKKRKQRGEKKKKGETGRYMMAFALGFSLIGAGLLLGWTVGWLL 329
Query: 419 P 419
P
Sbjct: 330 P 330
>emb|CAG31840.1| hypothetical protein [Gallus gallus]
Length = 854
Score = 44.7 bits (104), Expect = 0.006
Identities = 31/90 (34%), Positives = 52/90 (57%), Gaps = 7/90 (7%)
Query: 239 LNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAAKAE 298
+ QK ++ SSDS+ S++ T S DSD + SSSES S S + +S + A+KA+
Sbjct: 586 MTQKQDVESSDSS---SSSETDSSSDSDSDSSSSSSESSSSSDSSSSSSDSSDSDASKAK 642
Query: 299 LLKALCRSQTRAREAEKAAQEACDEKEHIL 328
R+Q + RE++KA+++ ++K L
Sbjct: 643 KR----RTQKKNRESDKASRKKQEKKRKSL 668
Score = 37.7 bits (86), Expect = 0.67
Identities = 31/111 (27%), Positives = 55/111 (48%), Gaps = 12/111 (10%)
Query: 224 LQPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSS- 282
+ ++ D E + SS ++ SDS+ +S++ +S SD + SSS+S DSD+S
Sbjct: 585 IMTQKQDVESSDSSSSSETDSSSDSDSDSSSSSSESSS----SSDSSSSSSDSSDSDASK 640
Query: 283 ------CNKNSKVNSESAAKAE-LLKALCRSQTRAREAEKAAQEACDEKEH 326
KN + + S K E K+L + R R+ +++ E+ E+ H
Sbjct: 641 AKKRRTQKKNRESDKASRKKQEKKRKSLEKKTGRRRQEDRSDSESKSERNH 691
>ref|XP_421976.1| PREDICTED: similar to KIAA1604 protein [Gallus gallus]
Length = 859
Score = 44.7 bits (104), Expect = 0.006
Identities = 31/90 (34%), Positives = 52/90 (57%), Gaps = 7/90 (7%)
Query: 239 LNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAAKAE 298
+ QK ++ SSDS+ S++ T S DSD + SSSES S S + +S + A+KA+
Sbjct: 591 MTQKQDVESSDSS---SSSETDSSSDSDSDSSSSSSESSSSSDSSSSSSDSSDSDASKAK 647
Query: 299 LLKALCRSQTRAREAEKAAQEACDEKEHIL 328
R+Q + RE++KA+++ ++K L
Sbjct: 648 KR----RTQKKNRESDKASRKKQEKKRKSL 673
Score = 37.7 bits (86), Expect = 0.67
Identities = 31/111 (27%), Positives = 55/111 (48%), Gaps = 12/111 (10%)
Query: 224 LQPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSS- 282
+ ++ D E + SS ++ SDS+ +S++ +S SD + SSS+S DSD+S
Sbjct: 590 IMTQKQDVESSDSSSSSETDSSSDSDSDSSSSSSESSS----SSDSSSSSSDSSDSDASK 645
Query: 283 ------CNKNSKVNSESAAKAE-LLKALCRSQTRAREAEKAAQEACDEKEH 326
KN + + S K E K+L + R R+ +++ E+ E+ H
Sbjct: 646 AKKRRTQKKNRESDKASRKKQEKKRKSLEKKTGRRRQEDRSDSESKSERNH 696
>ref|XP_483012.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|42408159|dbj|BAD09297.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 215
Score = 43.9 bits (102), Expect = 0.009
Identities = 33/113 (29%), Positives = 55/113 (48%), Gaps = 5/113 (4%)
Query: 299 LLKALCRSQTRAREAEKAAQEACDEKEHILSLFFKQASQLFAYKQWLHVLQLENLCLQYK 358
L++AL SQ+RAREAE+ A + +L + + L A+++W+ +L+ EN L+
Sbjct: 105 LMRALRLSQSRAREAEEKLAAAGASNGELSALLVRDSVVLSAHRRWVMMLEAENSGLRGA 164
Query: 359 SKNQPLLNNLFPYEGKKHRKNRRKVKNSRRGIGKCIFAFAVGLALAGAGLLLG 411
+ EG ++ +RRG A AV + +AG GL +G
Sbjct: 165 AGAAGSAK-----EGVGEDEDEDDDGGARRGAAAWWLALAVCVGIAGIGLAMG 212
>ref|XP_311331.2| ENSANGP00000001657 [Anopheles gambiae str. PEST]
gi|55243135|gb|EAA06779.3| ENSANGP00000001657 [Anopheles
gambiae str. PEST]
Length = 269
Score = 43.5 bits (101), Expect = 0.012
Identities = 26/90 (28%), Positives = 42/90 (45%), Gaps = 5/90 (5%)
Query: 237 SSLNQKLEMCSSDSNGCTSTT-----LTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNS 291
SS + CSS + CTS+T TS CS S R+ S++ S SSC+ + +S
Sbjct: 53 SSCSSATSSCSSGTTSCTSSTSSSNSCTSSCSSGTSSRSSSTTSSSSGTSSCSSGTTSSS 112
Query: 292 ESAAKAELLKALCRSQTRAREAEKAAQEAC 321
A+ + C S T + + ++ +C
Sbjct: 113 SCASSCSSSTSSCSSGTSSCRSGTSSCSSC 142
Score = 36.6 bits (83), Expect = 1.5
Identities = 23/80 (28%), Positives = 35/80 (43%), Gaps = 2/80 (2%)
Query: 229 TDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSK 288
T + +S + CSS ++ C+S T S CS + T S+S S SSC+ +
Sbjct: 31 TSSGSSGTTSCSSCTSSCSSGTSSCSSAT--SSCSSGTTSCTSSTSSSNSCTSSCSSGTS 88
Query: 289 VNSESAAKAELLKALCRSQT 308
S S + + C S T
Sbjct: 89 SRSSSTTSSSSGTSSCSSGT 108
>ref|NP_001004408.1| hypothetical protein LOC424547 [Gallus gallus]
gi|3123014|sp|P87498|VIT1_CHICK Vitellogenin I precursor
(Minor vitellogenin) [Contains: Lipovitellin I (LVI);
Phosvitin (PV); Lipovitellin II (LVII); YGP42]
gi|7442003|pir||T29088 vitellogenin I precursor
[validated] - chicken gi|1871444|dbj|BAA13973.1|
vitellogenin I [Gallus gallus]
Length = 1912
Score = 43.5 bits (101), Expect = 0.012
Identities = 33/106 (31%), Positives = 52/106 (48%), Gaps = 9/106 (8%)
Query: 229 TDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNK-NS 287
+D + SS + SS S+ +S+ S S+R+ SSS SKDS SS +K NS
Sbjct: 1167 SDSSSSSRSSSSSDSSSSSSSSSSSSSSKSKSSSRSSKSNRSSSSSNSKDSSSSSSKSNS 1226
Query: 288 KVNSESAAKAELLKALCRSQTR--------AREAEKAAQEACDEKE 325
K +S S++KA + + Q++ A E E++ E E +
Sbjct: 1227 KGSSSSSSKASGTRQKAKKQSKTTSFPHASAAEGERSVHEQKQETQ 1272
>emb|CAE85608.1| putative protein [Neurospora crassa]
Length = 412
Score = 42.7 bits (99), Expect = 0.021
Identities = 55/220 (25%), Positives = 96/220 (43%), Gaps = 16/220 (7%)
Query: 102 NAATAAIEQQWNVYPKYMKNSDTRTSKIEAS-LNSDLYLTPKKKNEGEFWFSDDATSFLI 160
+A+ +I Q W PK + + EA + T K + E S +S
Sbjct: 60 DASLVSIFQTWEAKPKQAALATPAPGEDEAKKTKAAAPATNPLKRKAE---SSSESSSSS 116
Query: 161 SEHCKSTSSDFEPHWLGAEKSQPWWRTTGKDDLASLVAKKSFEYVENCDLPEPNIKPFRK 220
S+ S SS E A+K + ++ + +S + S E+ D + KP K
Sbjct: 117 SDSSSSDSSSSEDEAPKAKKQKVAESSSSSSESSSDSSSDSSSSSESEDDKK---KPAAK 173
Query: 221 IHTLQPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSD 280
P++ + + SS + + SSDS+ +S+ +S S DSD + S S+S SD
Sbjct: 174 --AASPKKAAAAKKEESSDSSSSDSSSSDSSSESSSDSSSSDSSSDSDSSSSDSDS-SSD 230
Query: 281 SSCNKNSKVNSESAAKAELLKALCRSQTRAREAEKAAQEA 320
SS + +S +S+S + + S+ +E +K A++A
Sbjct: 231 SSSDSDSSSDSDSDSDSS------ESEAEKKEVKKEAKKA 264
>emb|CAA88179.1| gar2 [Schizosaccharomyces pombe]
Length = 500
Score = 42.0 bits (97), Expect = 0.036
Identities = 40/145 (27%), Positives = 65/145 (44%), Gaps = 25/145 (17%)
Query: 199 KKSFEYVENCDLPEPNIKPFRKIHTLQPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTL 258
KKS + + PEP+ K +K Q + KEE SS + E SS+S +S +
Sbjct: 52 KKSKKEAKRASSPEPSKKSVKK----QKKSKKKEE---SSSESESESSSSESESSSSESE 104
Query: 259 TSGCSFQDSDRTFSSSESKD-------------SDSSCNKNSKVNSESAAKAELLK---- 301
+S + S SSSES++ S+SS + S+ E+ K E K
Sbjct: 105 SSSSESESSSSESSSSESEEEVIVKTEEKKESSSESSSSSESEEEEEAVVKIEEKKESSS 164
Query: 302 -ALCRSQTRAREAEKAAQEACDEKE 325
+ S + E+E ++ E+ +E+E
Sbjct: 165 DSSSESSSSESESESSSSESEEEEE 189
Score = 38.1 bits (87), Expect = 0.51
Identities = 56/233 (24%), Positives = 92/233 (39%), Gaps = 46/233 (19%)
Query: 113 NVYPKYMKNSDTRTSKIEASLNSDLYLTPKKKNEGEFWFSDDATSFLISEHCKSTSSDFE 172
+V PK K R S E S S + +KK++ + S ++ S S +S+SS+ E
Sbjct: 48 DVSPKKSKKEAKRASSPEPSKKS---VKKQKKSKKKEESSSESESESSSSESESSSSESE 104
Query: 173 PHWLGAEKSQPWWRTTGKDDLASLVAKKSFEYVENCDLPEPNIKPFRKIHTLQPRETDKE 232
+E S + + V K+ E E+ + E+++E
Sbjct: 105 SSSSESESSSS---ESSSSESEEEVIVKTEEKKESSS------------ESSSSSESEEE 149
Query: 233 ENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDR----------------------- 269
E V + +K E S S+ +S+ S S +S+
Sbjct: 150 EEAVVKIEEKKESSSDSSSESSSSESESESSSSESEEEEEVVEKTEEKKEGSSESSSDSE 209
Query: 270 --TFSSSESKDSDSSCNKNSKVNSESAAKAELLKALCRSQTRAREAEKAAQEA 320
+ SSSES DSDSS + S+ +SE K KA S+ R + K +Q++
Sbjct: 210 SSSDSSSESGDSDSSSDSESESSSEDEKKR---KAEPASEERPAKITKPSQDS 259
>emb|CAB86413.1| gar2 [Schizosaccharomyces pombe] gi|19114443|ref|NP_593531.1|
hypothetical protein SPAC140.02 [Schizosaccharomyces
pombe 972h-] gi|6226864|sp|P41891|GAR2_SCHPO Protein
gar2
Length = 500
Score = 42.0 bits (97), Expect = 0.036
Identities = 40/145 (27%), Positives = 65/145 (44%), Gaps = 25/145 (17%)
Query: 199 KKSFEYVENCDLPEPNIKPFRKIHTLQPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTL 258
KKS + + PEP+ K +K Q + KEE SS + E SS+S +S +
Sbjct: 52 KKSKKEAKRASSPEPSKKSVKK----QKKSKKKEE---SSSESESESSSSESESSSSESE 104
Query: 259 TSGCSFQDSDRTFSSSESKD-------------SDSSCNKNSKVNSESAAKAELLK---- 301
+S + S SSSES++ S+SS + S+ E+ K E K
Sbjct: 105 SSSSESESSSSESSSSESEEEVIVKTEEKKESSSESSSSSESEEEEEAVVKIEEKKESSS 164
Query: 302 -ALCRSQTRAREAEKAAQEACDEKE 325
+ S + E+E ++ E+ +E+E
Sbjct: 165 DSSSESSSSESESESSSSESEEEEE 189
Score = 38.1 bits (87), Expect = 0.51
Identities = 56/233 (24%), Positives = 92/233 (39%), Gaps = 46/233 (19%)
Query: 113 NVYPKYMKNSDTRTSKIEASLNSDLYLTPKKKNEGEFWFSDDATSFLISEHCKSTSSDFE 172
+V PK K R S E S S + +KK++ + S ++ S S +S+SS+ E
Sbjct: 48 DVSPKKSKKEAKRASSPEPSKKS---VKKQKKSKKKEESSSESESESSSSESESSSSESE 104
Query: 173 PHWLGAEKSQPWWRTTGKDDLASLVAKKSFEYVENCDLPEPNIKPFRKIHTLQPRETDKE 232
+E S + + V K+ E E+ + E+++E
Sbjct: 105 SSSSESESSSS---ESSSSESEEEVIVKTEEKKESSS------------ESSSSSESEEE 149
Query: 233 ENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDR----------------------- 269
E V + +K E S S+ +S+ S S +S+
Sbjct: 150 EEAVVKIEEKKESSSDSSSESSSSESESESSSSESEEEEEVVEKTEEKKEGSSESSSDSE 209
Query: 270 --TFSSSESKDSDSSCNKNSKVNSESAAKAELLKALCRSQTRAREAEKAAQEA 320
+ SSSES DSDSS + S+ +SE K KA S+ R + K +Q++
Sbjct: 210 SSSDSSSESGDSDSSSDSESESSSEDEKKR---KAEPASEERPAKITKPSQDS 259
>emb|CAA51946.1| unnamed protein product [Saccharomyces cerevisiae]
gi|6322945|ref|NP_013018.1| Nucleolar, serine-rich
protein with a role in preribosome assembly or
transport; may function as a chaperone of small
nucleolar ribonucleoprotein particles (snoRNPs);
immunologically and structurally to rat Nopp140; Srp40p
[Saccharomyces cerevisiae] gi|486581|emb|CAA82171.1|
SRP40 [Saccharomyces cerevisiae]
gi|548976|sp|P32583|SRP40_YEAST Suppressor protein SRP40
Length = 406
Score = 41.6 bits (96), Expect = 0.047
Identities = 43/175 (24%), Positives = 80/175 (45%), Gaps = 13/175 (7%)
Query: 121 NSDTRTSKIEASLNSDLYLTPKKKNEGEFWFSDDATSFLISEHCKSTSSDFEPHWLGAEK 180
+S + +S E+S +S + + + S+ ++S S S+SSD E
Sbjct: 42 SSSSSSSSGESSSSSSSSSSSSSSDSSDSSDSESSSSSSSSSSSSSSSSDSE------SS 95
Query: 181 SQPWWRTTGKDDLASLVAKKSFEYVENCDLPEPNIKPFRKIHTLQPRETDKEENQVSSLN 240
S+ ++G +S + +S E+ D + K R+ +ET K + + S +
Sbjct: 96 SESDSSSSGSSSSSSSSSDESSSESESEDETK---KRARESDNEDAKETKKAKTEPESSS 152
Query: 241 QKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAA 295
S S+G +S++ + S DSD + SSS S DS+S +S+ +S S++
Sbjct: 153 SS----ESSSSGSSSSSESESGSESDSDSSSSSSSSSDSESDSESDSQSSSSSSS 203
Score = 37.0 bits (84), Expect = 1.1
Identities = 23/71 (32%), Positives = 40/71 (55%), Gaps = 1/71 (1%)
Query: 228 ETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNS 287
E+D E + SS + SSDS+ +S + + S S + S S+S DSDSS + +S
Sbjct: 188 ESDSESDSQSSSSSSSSDSSSDSDSSSSDSSSDSDSSSSSSSSSSDSDS-DSDSSSDSDS 246
Query: 288 KVNSESAAKAE 298
+S+S++ ++
Sbjct: 247 SGSSDSSSSSD 257
Score = 37.0 bits (84), Expect = 1.1
Identities = 25/78 (32%), Positives = 43/78 (55%), Gaps = 1/78 (1%)
Query: 220 KIHTLQPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDS 279
++ L +E + EE SS + SS S+ +S++ +SG S S + SSS S S
Sbjct: 10 EVPKLSVKEKEIEEKSSSSSSSSSSSSSSSSSSSSSSS-SSGESSSSSSSSSSSSSSDSS 68
Query: 280 DSSCNKNSKVNSESAAKA 297
DSS +++S +S S++ +
Sbjct: 69 DSSDSESSSSSSSSSSSS 86
Score = 35.4 bits (80), Expect = 3.3
Identities = 23/72 (31%), Positives = 40/72 (54%), Gaps = 1/72 (1%)
Query: 228 ETDKEENQVSSLNQKLEMCS-SDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKN 286
E+D + + SS + E S SDS +S++ + S DS + SSS+S S SS + +
Sbjct: 172 ESDSDSSSSSSSSSDSESDSESDSQSSSSSSSSDSSSDSDSSSSDSSSDSDSSSSSSSSS 231
Query: 287 SKVNSESAAKAE 298
S +S+S + ++
Sbjct: 232 SDSDSDSDSSSD 243
Score = 34.3 bits (77), Expect = 7.4
Identities = 43/202 (21%), Positives = 79/202 (38%), Gaps = 18/202 (8%)
Query: 120 KNSDTRTSKIEASLNSDLYLTPKKKNEGEFWFSDDATSFLISEHCKSTS-SDFEPHWLGA 178
++ D + S N D T K K E E S +++S S +S S S+ + +
Sbjct: 121 ESEDETKKRARESDNEDAKETKKAKTEPESSSSSESSSSGSSSSSESESGSESDSDSSSS 180
Query: 179 EKSQPWWRTTGKDDLASLVAKKSFEYVENCDLPEPNIKPFRKIHTLQPRETDKEENQVSS 238
S + + D S + S + + D +D + SS
Sbjct: 181 SSSSSDSESDSESDSQSSSSSSSSDSSSDSD----------------SSSSDSSSDSDSS 224
Query: 239 LNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAAKAE 298
+ SDS+ +S+ S S S + SSS+ S S + +S +S S+++ E
Sbjct: 225 SSSSSSSSDSDSDSDSSSDSDSSGSSDSSSSSDSSSDESTSSDSSDSDSDSDSGSSSELE 284
Query: 299 LLKALCRSQTRAREAEKAAQEA 320
+A +++A E ++ E+
Sbjct: 285 TKEATA-DESKAEETPASSNES 305
>gb|AAA35091.1| selected as a weak suppressor of a mutant of the subunit AC40 of
DNA dependant RNA polymerase I and III
Length = 406
Score = 41.6 bits (96), Expect = 0.047
Identities = 43/175 (24%), Positives = 80/175 (45%), Gaps = 13/175 (7%)
Query: 121 NSDTRTSKIEASLNSDLYLTPKKKNEGEFWFSDDATSFLISEHCKSTSSDFEPHWLGAEK 180
+S + +S E+S +S + + + S+ ++S S S+SSD E
Sbjct: 42 SSSSSSSSGESSSSSSSSSSSSSSDSSDSSDSESSSSSSSSSSSSSSSSDSE------SS 95
Query: 181 SQPWWRTTGKDDLASLVAKKSFEYVENCDLPEPNIKPFRKIHTLQPRETDKEENQVSSLN 240
S+ ++G +S + +S E+ D + K R+ +ET K + + S +
Sbjct: 96 SESDSSSSGSSSSSSSSSDESSSESESEDETK---KRARESDNEDAKETKKAKTEPESSS 152
Query: 241 QKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAA 295
S S+G +S++ + S DSD + SSS S DS+S +S+ +S S++
Sbjct: 153 SS----ESSSSGSSSSSESESGSESDSDSSSSSSSSSDSESDSESDSQSSSSSSS 203
Score = 37.0 bits (84), Expect = 1.1
Identities = 23/71 (32%), Positives = 40/71 (55%), Gaps = 1/71 (1%)
Query: 228 ETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNS 287
E+D E + SS + SSDS+ +S + + S S + S S+S DSDSS + +S
Sbjct: 188 ESDSESDSQSSSSSSSSDSSSDSDSSSSDSSSDSDSSSSSSSSSSDSDS-DSDSSSDSDS 246
Query: 288 KVNSESAAKAE 298
+S+S++ ++
Sbjct: 247 SGSSDSSSSSD 257
Score = 37.0 bits (84), Expect = 1.1
Identities = 25/78 (32%), Positives = 43/78 (55%), Gaps = 1/78 (1%)
Query: 220 KIHTLQPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDS 279
++ L +E + EE SS + SS S+ +S++ +SG S S + SSS S S
Sbjct: 10 EVPKLSVKEKEIEEKSSSSSSSSSSSSSSSSSSSSSSS-SSGESSSSSSSSSSSSSSDSS 68
Query: 280 DSSCNKNSKVNSESAAKA 297
DSS +++S +S S++ +
Sbjct: 69 DSSDSESSSSSSSSSSSS 86
Score = 35.4 bits (80), Expect = 3.3
Identities = 23/72 (31%), Positives = 40/72 (54%), Gaps = 1/72 (1%)
Query: 228 ETDKEENQVSSLNQKLEMCS-SDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKN 286
E+D + + SS + E S SDS +S++ + S DS + SSS+S S SS + +
Sbjct: 172 ESDSDSSSSSSSSSDSESDSESDSQSSSSSSSSDSSSDSDSSSSDSSSDSDSSSSSSSSS 231
Query: 287 SKVNSESAAKAE 298
S +S+S + ++
Sbjct: 232 SDSDSDSDSSSD 243
Score = 34.3 bits (77), Expect = 7.4
Identities = 43/202 (21%), Positives = 79/202 (38%), Gaps = 18/202 (8%)
Query: 120 KNSDTRTSKIEASLNSDLYLTPKKKNEGEFWFSDDATSFLISEHCKSTS-SDFEPHWLGA 178
++ D + S N D T K K E E S +++S S +S S S+ + +
Sbjct: 121 ESEDETKKRARESDNEDAKETKKAKTEPESSSSSESSSSGSSSSSESESGSESDSDSSSS 180
Query: 179 EKSQPWWRTTGKDDLASLVAKKSFEYVENCDLPEPNIKPFRKIHTLQPRETDKEENQVSS 238
S + + D S + S + + D +D + SS
Sbjct: 181 SSSSSDSESDSESDSQSSSSSSSSDSSSDSD----------------SSSSDSSSDSDSS 224
Query: 239 LNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAAKAE 298
+ SDS+ +S+ S S S + SSS+ S S + +S +S S+++ E
Sbjct: 225 SSSSSSSSDSDSDSDSSSDSDSSGSSDSSSSSDSSSDESTSSDSSDSDSDSDSGSSSELE 284
Query: 299 LLKALCRSQTRAREAEKAAQEA 320
+A +++A E ++ E+
Sbjct: 285 TKEATA-DESKAEETPASSNES 305
>emb|CAA58336.1| U87 [Human herpesvirus 6] gi|9628389|ref|NP_042980.1| possible
glycoprotein [Human herpesvirus 6]
Length = 830
Score = 41.2 bits (95), Expect = 0.061
Identities = 50/210 (23%), Positives = 89/210 (41%), Gaps = 23/210 (10%)
Query: 115 YPKYMKNSDTRTSKIEASLNSDLYLTPKKKNEGEFWFSDDATSFLISEHCKSTSSDFEPH 174
YPK MK+S RT+ + + NS+ + K+ + +TS H S+SSD E
Sbjct: 450 YPK-MKHSPGRTTSKKNTTNSNGHQNFKEVSVKNVSGKATSTSPKSKTHHYSSSSDEEGQ 508
Query: 175 WLGAEKSQPWWRTTGKDDLASLVAKKSFEYVENCDLPEPNI--KPFRKIHTLQPRETDKE 232
+ K + +++ S C L P+I K K HT + +
Sbjct: 509 Y--------------KSPVKTIIQSPS----PYCKLKNPSIMDKNSAKNHTASADKNLTD 550
Query: 233 ENQV-SSLNQKLEMCSSDSNGCTSTTLTSG-CSFQDSDRTFSSSESKDSDSSCNKNSKVN 290
+ + S+LN S+ + T +T S C+ + + + +ESKD + +C KNS +
Sbjct: 551 NSPIRSNLNPTAFNKSNSNKSITDSTSNSDECTDRKPNCNSTKNESKDPNRTCGKNSDKH 610
Query: 291 SESAAKAELLKALCRSQTRAREAEKAAQEA 320
+ +A R+ +RA + + +A
Sbjct: 611 LSKSCTMASKRAPSRASSRASSRDSSRDQA 640
>ref|XP_636785.1| hypothetical protein DDB0219334 [Dictyostelium discoideum]
gi|60465182|gb|EAL63280.1| hypothetical protein
DDB0219334 [Dictyostelium discoideum]
Length = 1105
Score = 41.2 bits (95), Expect = 0.061
Identities = 54/249 (21%), Positives = 86/249 (33%), Gaps = 36/249 (14%)
Query: 81 DNVSIGSDQTVKNFDAVSFVGNAATAAIEQQWNVYPKYMKNSDTRTSKIEASLNSDLYLT 140
+N D N ++ + N E Q ++ Y + + S+ + Y
Sbjct: 181 NNDDNNDDNNNNNNNSKTTTKNRIIKKQENQEVLFSSYSPSEALKKSQHLFMVTPTFYAA 240
Query: 141 PKKKNEGEFWFSDDATSFLISEH--CKSTSSDFEPHW-----------LGAEKSQPWWRT 187
K + G F + + + + +EH K P+W L E+ P T
Sbjct: 241 KKNQKIGVFLYKNSNLNRICTEHPQLKLVLKPTSPYWIPSSFTRKVNRLNIEEYAPNQFT 300
Query: 188 TGKDDLASLVA-------KKSFEYVENCDL---PEPNIKP-------------FRKIHTL 224
T + DL + A KK+ Y NC + P PN + F T
Sbjct: 301 TLECDLTNGEAIFGPVRIKKTGWYYLNCSIELPPLPNAEQLLFKDFSFTFKLYFYGCPTK 360
Query: 225 QPRETDKEENQVSSLNQKLEMCSSDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCN 284
+E + SS + SS S+ S+T S +F D + SSS S +DS
Sbjct: 361 SKKEPSPSPSSSSSSSSSSSSSSSSSSSSPSSTSNSSTTFSDKSSSSSSSSSSSNDSITQ 420
Query: 285 KNSKVNSES 293
+ N S
Sbjct: 421 NDKSFNPSS 429
>ref|XP_323366.1| predicted protein [Neurospora crassa] gi|28918756|gb|EAA28426.1|
predicted protein [Neurospora crassa]
Length = 471
Score = 41.2 bits (95), Expect = 0.061
Identities = 57/253 (22%), Positives = 108/253 (42%), Gaps = 19/253 (7%)
Query: 68 ESKLDAFSTRLHGDNVSIGSDQTVKNFDAVSFVGNAATAAIEQQWNVYPKYMKNSDTRTS 127
ES + S D+ S D+ K A ++++++ + +S+
Sbjct: 90 ESSSSSESESSDSDSDSDSEDEKKKEAPAKKAESSSSSSSSSDSSSDSSSDSSSSEDEAK 149
Query: 128 KIEASLNSDLYLTPKKKNEGEFWFSDDATSFLISEHCKSTSSDFEPHWLGAEKSQPWWRT 187
K +A+ + L K ++ E +S S+ S SS E A+K + +
Sbjct: 150 KTKAAAPATNPLKRKAESSSE-------SSSSSSDSSSSDSSSSEDEAPKAKKQKVAESS 202
Query: 188 TGKDDLASLVAKKSFEYVENCDLPEPNIKPFRKIHTLQPRETDKEENQVSSLNQKLEMCS 247
+ + +S + S E+ D + KP K P++ + + SS + + S
Sbjct: 203 SSSSESSSDSSSDSSSSSESEDDKK---KPAAK--AASPKKAAAAKKEESSDSSSSDSSS 257
Query: 248 SDSNGCTSTTLTSGCSFQDSDRTFSSSESKDSDSSCNKNSKVNSESAAKAELLKALCRSQ 307
SDS+ +S+ +S S DSD + S S+S SDSS + +S +S+S + + S+
Sbjct: 258 SDSSSESSSDSSSSDSSSDSDSSSSDSDS-SSDSSSDSDSSSDSDSDSDSS------ESE 310
Query: 308 TRAREAEKAAQEA 320
+E +K A++A
Sbjct: 311 AEKKEVKKEAKKA 323
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.314 0.127 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 687,654,363
Number of Sequences: 2540612
Number of extensions: 28033980
Number of successful extensions: 106688
Number of sequences better than 10.0: 343
Number of HSP's better than 10.0 without gapping: 98
Number of HSP's successfully gapped in prelim test: 251
Number of HSP's that attempted gapping in prelim test: 104193
Number of HSP's gapped (non-prelim): 1655
length of query: 421
length of database: 863,360,394
effective HSP length: 131
effective length of query: 290
effective length of database: 530,540,222
effective search space: 153856664380
effective search space used: 153856664380
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 76 (33.9 bits)
Medicago: description of AC149127.6


