

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC149080.7 - phase: 0
(165 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
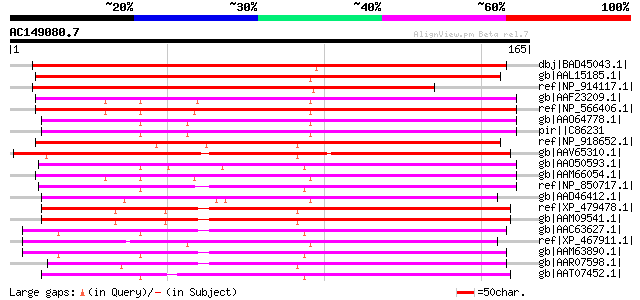
Score E
Sequences producing significant alignments: (bits) Value
dbj|BAD45043.1| putative ER6 protein [Oryza sativa (japonica cul... 193 2e-48
gb|AAL15185.1| unknown protein [Arabidopsis thaliana] gi|1433494... 158 6e-38
ref|NP_914117.1| B1146F03.27 [Oryza sativa (japonica cultivar-gr... 145 4e-34
gb|AAF23209.1| unknown protein [Arabidopsis thaliana] gi|1099813... 112 3e-24
ref|NP_566406.1| universal stress protein (USP) family protein [... 112 3e-24
gb|AAO64778.1| At1g09740 [Arabidopsis thaliana] gi|30681321|ref|... 111 8e-24
pir||C86231 hypothetical protein [imported] - Arabidopsis thalia... 110 2e-23
ref|NP_918652.1| P0520B06.18 [Oryza sativa (japonica cultivar-gr... 109 2e-23
gb|AAV65310.1| universal stress protein [Hordeum vulgare subsp. ... 109 2e-23
gb|AAO50593.1| unknown protein [Arabidopsis thaliana] gi|2839326... 108 4e-23
gb|AAM66054.1| ethylene-responsive protein, putative [Arabidopsi... 107 9e-23
ref|NP_850717.1| universal stress protein (USP) family protein [... 107 9e-23
gb|AAD46412.1| ER6 protein [Lycopersicon esculentum] 107 1e-22
ref|XP_479478.1| universal stress protein USP1-like protein [Ory... 106 3e-22
gb|AAM09541.1| putative universal stress protein USP1 [Oryza sat... 105 3e-22
gb|AAC63627.1| expressed protein [Arabidopsis thaliana] gi|20148... 104 8e-22
ref|XP_467911.1| putative ethylene-responsive protein [Oryza sat... 102 4e-21
gb|AAM63890.1| unknown [Arabidopsis thaliana] 102 4e-21
gb|AAR07598.1| fiber protein Fb19 [Gossypium barbadense] 100 1e-20
gb|AAT07452.1| putative universal stress protein [Mirabilis jalapa] 99 6e-20
>dbj|BAD45043.1| putative ER6 protein [Oryza sativa (japonica cultivar-group)]
gi|52075680|dbj|BAD44900.1| putative ER6 protein [Oryza
sativa (japonica cultivar-group)]
Length = 182
Score = 193 bits (490), Expect = 2e-48
Identities = 91/153 (59%), Positives = 119/153 (77%), Gaps = 2/153 (1%)
Query: 8 GRRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTGYLFSSDI 67
GRRI+VA+DE EES +ALTWCL N+V + D L+LL+ + PR VY+A D +GY+ +SD+
Sbjct: 30 GRRIVVAVDESEESTHALTWCLANVVSSSGGDTLVLLHARRPRPVYAAMDSSGYMMTSDV 89
Query: 68 TATMEKYSQQVADCVLEKAKIVCNDVQNV--ETRIENGDPRDVICQAVQKMGVDILVMGS 125
A+M+KY+ V+ + KAK +C +V ET +E+GDPRDVIC A +KM D+LVMG+
Sbjct: 90 MASMDKYAAAVSAAAVGKAKHICAAFPHVTVETMVESGDPRDVICDATEKMAADLLVMGT 149
Query: 126 HGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPK 158
HGYG+I+RAFLGSVSNHCAQN KCPVLIVK+PK
Sbjct: 150 HGYGLIQRAFLGSVSNHCAQNCKCPVLIVKRPK 182
>gb|AAL15185.1| unknown protein [Arabidopsis thaliana] gi|14334946|gb|AAK59650.1|
unknown protein [Arabidopsis thaliana]
gi|15228790|ref|NP_191814.1| universal stress protein
(USP) family protein [Arabidopsis thaliana]
Length = 162
Score = 158 bits (399), Expect = 6e-38
Identities = 77/151 (50%), Positives = 107/151 (69%), Gaps = 3/151 (1%)
Query: 9 RRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTGYLFSSDIT 68
R+I+VA+DE EES+ AL+W L NL S + LILLYVKPP VYS+ D G++ + D
Sbjct: 7 RKIVVAVDESEESMEALSWSLDNLFPYGSNNTLILLYVKPPLPVYSSLDAAGFIVTGDPV 66
Query: 69 ATMEKYSQQVADCVLEKAKIVCNDVQ---NVETRIENGDPRDVICQAVQKMGVDILVMGS 125
A ++KY ++ + V+ +++ V D + N+E R+ GD ++VIC AVQK+ VD+LVMG+
Sbjct: 67 AALKKYEYELVESVMARSRTVYQDYESDINIERRVGRGDAKEVICNAVQKLRVDMLVMGT 126
Query: 126 HGYGVIKRAFLGSVSNHCAQNVKCPVLIVKK 156
H YG KRA LGSVS +CA+ VKCPV+IVKK
Sbjct: 127 HDYGFFKRALLGSVSEYCAKRVKCPVVIVKK 157
>ref|NP_914117.1| B1146F03.27 [Oryza sativa (japonica cultivar-group)]
Length = 164
Score = 145 bits (366), Expect = 4e-34
Identities = 68/130 (52%), Positives = 96/130 (73%), Gaps = 2/130 (1%)
Query: 8 GRRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFDGTGYLFSSDI 67
GRRI+VA+DE EES +ALTWCL N+V + D L+LL+ + PR VY+A D +GY+ +SD+
Sbjct: 30 GRRIVVAVDESEESTHALTWCLANVVSSSGGDTLVLLHARRPRPVYAAMDSSGYMMTSDV 89
Query: 68 TATMEKYSQQVADCVLEKAKIVCNDVQN--VETRIENGDPRDVICQAVQKMGVDILVMGS 125
A+M+KY+ V+ + KAK +C + VET +E+GDPRDVIC A +KM D+LVMG+
Sbjct: 90 MASMDKYAAAVSAAAVGKAKHICAAFPHVTVETMVESGDPRDVICDATEKMAADLLVMGT 149
Query: 126 HGYGVIKRAF 135
HGYG+I+R +
Sbjct: 150 HGYGLIQRYY 159
>gb|AAF23209.1| unknown protein [Arabidopsis thaliana] gi|10998131|dbj|BAB03102.1|
unnamed protein product [Arabidopsis thaliana]
gi|16323234|gb|AAL15351.1| AT3g11930/MEC18.3
[Arabidopsis thaliana] gi|16226626|gb|AAL16217.1|
At3g11930/MEC18.3 [Arabidopsis thaliana]
gi|15215662|gb|AAK91376.1| MEC18.3/MEC18.3 [Arabidopsis
thaliana] gi|13926250|gb|AAK49598.1| MEC18.3/MEC18.3
[Arabidopsis thaliana] gi|30681955|ref|NP_850562.1|
universal stress protein (USP) family protein
[Arabidopsis thaliana]
Length = 200
Score = 112 bits (281), Expect = 3e-24
Identities = 65/167 (38%), Positives = 100/167 (58%), Gaps = 14/167 (8%)
Query: 9 RRIMVAIDEGEESIYALTWCL---KNLVFQNSKDH-----LILLYVKPPRVVYSAFDG-- 58
+R++VAIDE + S YAL W + NL+ + L +++V+ P ++AF
Sbjct: 33 KRMVVAIDESDSSFYALQWVIDHFSNLLLTTAAAEAESGMLTVIHVQSPFNHFAAFPAGP 92
Query: 59 ---TGYLFSSDITATMEKYSQQVADCVLEKAKIVCNDVQ-NVETRIENGDPRDVICQAVQ 114
T SS + +++K Q+ + +L +A +C Q ET + G+ +++IC+AV+
Sbjct: 93 GGATAVYASSSMIESVKKAQQETSAALLSRALQMCRAKQIRTETLVLEGEAKEMICEAVE 152
Query: 115 KMGVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPKSTT 161
KM VD+LV+GS G G IKRAFLGSVS++CA + CP+LIVK PK T
Sbjct: 153 KMHVDLLVVGSRGLGKIKRAFLGSVSDYCAHHANCPILIVKPPKEMT 199
>ref|NP_566406.1| universal stress protein (USP) family protein [Arabidopsis
thaliana]
Length = 199
Score = 112 bits (281), Expect = 3e-24
Identities = 65/166 (39%), Positives = 100/166 (60%), Gaps = 13/166 (7%)
Query: 9 RRIMVAIDEGEESIYALTWCL---KNLVFQNSKDH-----LILLYVKPPRVVYSAFD--- 57
+R++VAIDE + S YAL W + NL+ + L +++V+ P ++AF
Sbjct: 33 KRMVVAIDESDSSFYALQWVIDHFSNLLLTTAAAEAESGMLTVIHVQSPFNHFAAFPAGP 92
Query: 58 -GTGYLFSSDITATMEKYSQQVADCVLEKAKIVCNDVQ-NVETRIENGDPRDVICQAVQK 115
G SS + +++K Q+ + +L +A +C Q ET + G+ +++IC+AV+K
Sbjct: 93 GGATVYASSSMIESVKKAQQETSAALLSRALQMCRAKQIRTETLVLEGEAKEMICEAVEK 152
Query: 116 MGVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPKSTT 161
M VD+LV+GS G G IKRAFLGSVS++CA + CP+LIVK PK T
Sbjct: 153 MHVDLLVVGSRGLGKIKRAFLGSVSDYCAHHANCPILIVKPPKEMT 198
>gb|AAO64778.1| At1g09740 [Arabidopsis thaliana] gi|30681321|ref|NP_172445.2|
ethylene-responsive protein, putative [Arabidopsis
thaliana]
Length = 171
Score = 111 bits (277), Expect = 8e-24
Identities = 61/161 (37%), Positives = 95/161 (58%), Gaps = 10/161 (6%)
Query: 11 IMVAIDEGEESIYALTWCLKNLVFQNSKDH--LILLYVKPPRVVYSA-------FDGTGY 61
++VA+D E S+ AL W L NL +S ++L+V+P V + F G
Sbjct: 10 VVVAVDGSEVSMEALRWALDNLKLSSSSSDSSFVVLHVQPSPSVAAGVSPGTIPFGGPSG 69
Query: 62 LFSSDITATMEKYSQQVADCVLEKAKIVCNDVQ-NVETRIENGDPRDVICQAVQKMGVDI 120
L TA +E++ +++ D +LE A +C + NV+T++ GDP+ IC+AV+ + D+
Sbjct: 70 LEVPAFTAAIEQHQKRITDTILEHASQICAEKSVNVKTQVVIGDPKYKICEAVENLHADL 129
Query: 121 LVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPKSTT 161
LVMGS YG IKR FLGSVSN+C + CPV+I+K + ++
Sbjct: 130 LVMGSRAYGRIKRMFLGSVSNYCTNHAHCPVVIIKPKEDSS 170
>pir||C86231 hypothetical protein [imported] - Arabidopsis thaliana
gi|2160182|gb|AAB60745.1| ESTs
gb|ATTS1236,gb|T43334,gb|N97019,gb|AA395203 come from
this gene. [Arabidopsis thaliana]
Length = 174
Score = 110 bits (274), Expect = 2e-23
Identities = 61/164 (37%), Positives = 95/164 (57%), Gaps = 13/164 (7%)
Query: 11 IMVAIDEGEESIYALTWCLKNLVFQNSKDH--LILLYVKPPRVVYSA-------FDGTGY 61
++VA+D E S+ AL W L NL +S ++L+V+P V + F G
Sbjct: 10 VVVAVDGSEVSMEALRWALDNLKLSSSSSDSSFVVLHVQPSPSVAAGVSPGTIPFGGPSG 69
Query: 62 LFSSDITATMEKYSQQVADCVLEKAKIVCNDVQ----NVETRIENGDPRDVICQAVQKMG 117
L TA +E++ +++ D +LE A +C + NV+T++ GDP+ IC+AV+ +
Sbjct: 70 LEVPAFTAAIEQHQKRITDTILEHASQICAEKSVSRVNVKTQVVIGDPKYKICEAVENLH 129
Query: 118 VDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPKSTT 161
D+LVMGS YG IKR FLGSVSN+C + CPV+I+K + ++
Sbjct: 130 ADLLVMGSRAYGRIKRMFLGSVSNYCTNHAHCPVVIIKPKEDSS 173
>ref|NP_918652.1| P0520B06.18 [Oryza sativa (japonica cultivar-group)]
gi|14495190|dbj|BAB60909.1| putative ER6 protein [Oryza
sativa (japonica cultivar-group)]
gi|20804499|dbj|BAB92194.1| putative ER6 protein [Oryza
sativa (japonica cultivar-group)]
Length = 167
Score = 109 bits (273), Expect = 2e-23
Identities = 62/154 (40%), Positives = 97/154 (62%), Gaps = 6/154 (3%)
Query: 9 RRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLY--VKPPRVVYSAFDGTGY---LF 63
++ MVA+DE E S +AL W L+NL + L+L + P V +A G+ +
Sbjct: 12 QKAMVAVDESEFSHHALEWALRNLAPTIAPPLLVLTVQPLLPLGYVSAASFGSPLGTPVV 71
Query: 64 SSDITATMEKYSQQVADCVLEKAKIVC-NDVQNVETRIENGDPRDVICQAVQKMGVDILV 122
+ ++ M++ QQ++ +L+KAK +C VET I+ GDP+++ICQA ++ VD+L+
Sbjct: 72 APELIKAMQEQQQQLSQALLDKAKQICAQHGVAVETMIKVGDPKEMICQAAEESKVDLLI 131
Query: 123 MGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKK 156
+GSH G ++R FLGSVSN+C + KCPVL+VKK
Sbjct: 132 VGSHSRGPVQRLFLGSVSNYCMHHSKCPVLVVKK 165
>gb|AAV65310.1| universal stress protein [Hordeum vulgare subsp. vulgare]
Length = 173
Score = 109 bits (273), Expect = 2e-23
Identities = 61/164 (37%), Positives = 99/164 (60%), Gaps = 9/164 (5%)
Query: 2 AGITENGRR----IMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSAFD 57
A E+G R ++VA+D+ + S AL W ++++ + L++++ KPP +F
Sbjct: 11 AAEAEDGSRKKTVVLVAVDDSDHSYRALEWAVRHVATTGAAAELVVVHAKPPASSVVSFG 70
Query: 58 GTGYLFSSDITATMEKYSQQVADCVLEKAKIVC--NDVQNVETRIENGDPRDVICQAVQK 115
+ D+ ++ ++ A+ V+++A+ +C N V + IE G+PR V+C AV K
Sbjct: 71 SPAA--AGDLVRVVDADLRKRAEDVVDRARRLCVANSVHALIEVIE-GEPRHVLCSAVDK 127
Query: 116 MGVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPKS 159
D+L +GSHGYG IKRAFLGSVS++CA + C V+IVK+PKS
Sbjct: 128 HHADLLAVGSHGYGAIKRAFLGSVSDYCAHHAHCSVMIVKQPKS 171
>gb|AAO50593.1| unknown protein [Arabidopsis thaliana] gi|28393267|gb|AAO42062.1|
unknown protein [Arabidopsis thaliana]
gi|30694811|ref|NP_191404.2| universal stress protein
(USP) family protein [Arabidopsis thaliana]
Length = 204
Score = 108 bits (271), Expect = 4e-23
Identities = 63/164 (38%), Positives = 99/164 (59%), Gaps = 12/164 (7%)
Query: 10 RIMVAIDEGEESIYALTWCLKNLVFQNSKDH--------LILLYVKPP--RVVYSAFDGT 59
++MVAIDE + S AL W + +L S + L LL+V P + +Y +
Sbjct: 31 KVMVAIDESKNSFDALEWAVDHLRVVISAEPETGQEGGLLTLLHVHPTYLQYIYPSGGTA 90
Query: 60 GYLFSSD-ITATMEKYSQQVADCVLEKAKIVCND-VQNVETRIENGDPRDVICQAVQKMG 117
++++D + M K ++ + +A +C + ET I GDP+++ICQAV++
Sbjct: 91 SAVYATDSVPEPMRKAREESTTNLFTRALEICRGKMVKTETMILEGDPKEMICQAVEQTH 150
Query: 118 VDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPKSTT 161
VD+LV+GS G G+IKRAFLGSVS++CAQ+ KCP+LIV+ P+ T+
Sbjct: 151 VDLLVVGSRGLGMIKRAFLGSVSDYCAQHAKCPILIVRPPRETS 194
>gb|AAM66054.1| ethylene-responsive protein, putative [Arabidopsis thaliana]
Length = 199
Score = 107 bits (268), Expect = 9e-23
Identities = 64/166 (38%), Positives = 99/166 (59%), Gaps = 13/166 (7%)
Query: 9 RRIMVAIDEGEESIYALTWCL---KNLVFQNSKDH-----LILLYVKPPRVVYSAFD--- 57
+R++VAIDE + S YAL + NL+ + L +++V+ P ++AF
Sbjct: 33 KRMVVAIDESDSSFYALQLVIDHFSNLLLTTAAAEAESGMLTVIHVQSPFNHFAAFPAGP 92
Query: 58 -GTGYLFSSDITATMEKYSQQVADCVLEKAKIVCNDVQ-NVETRIENGDPRDVICQAVQK 115
G SS + +++K Q+ + +L +A +C Q ET + G+ +++IC+AV+K
Sbjct: 93 GGATVYASSSMIESVKKAQQETSAALLSRALQMCRAKQIRTETLVLEGEAKEMICEAVEK 152
Query: 116 MGVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPKSTT 161
M VD+LV+GS G G IKRAFLGSVS++CA + CP+LIVK PK T
Sbjct: 153 MHVDLLVVGSRGLGKIKRAFLGSVSDYCAHHANCPILIVKPPKEMT 198
>ref|NP_850717.1| universal stress protein (USP) family protein [Arabidopsis
thaliana]
Length = 197
Score = 107 bits (268), Expect = 9e-23
Identities = 63/161 (39%), Positives = 94/161 (58%), Gaps = 13/161 (8%)
Query: 10 RIMVAIDEGEESIYALTWCLKNLVFQNSKDH--------LILLYVKPPRVVYSAFDGTGY 61
++MVAIDE + S AL W + +L S + L LL+V P + Y G
Sbjct: 31 KVMVAIDESKNSFDALEWAVDHLRVVISAEPETGQEGGLLTLLHVHPTYLQYIYPSGG-- 88
Query: 62 LFSSDITATMEKYSQQVADCVLEKAKIVCND-VQNVETRIENGDPRDVICQAVQKMGVDI 120
+ + M K ++ + +A +C + ET I GDP+++ICQAV++ VD+
Sbjct: 89 --TDSVPEPMRKAREESTTNLFTRALEICRGKMVKTETMILEGDPKEMICQAVEQTHVDL 146
Query: 121 LVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPKSTT 161
LV+GS G G+IKRAFLGSVS++CAQ+ KCP+LIV+ P+ T+
Sbjct: 147 LVVGSRGLGMIKRAFLGSVSDYCAQHAKCPILIVRPPRETS 187
>gb|AAD46412.1| ER6 protein [Lycopersicon esculentum]
Length = 168
Score = 107 bits (267), Expect = 1e-22
Identities = 58/160 (36%), Positives = 96/160 (59%), Gaps = 15/160 (9%)
Query: 11 IMVAIDEGEESIYALTWCLKNLVFQ-------NSKDHLILLYVKPPRVVYSAFDGTGYLF 63
++V++D EES+ AL W L N+ + S+ +++L+V+ P + + + F
Sbjct: 6 VIVSVDGSEESMNALNWTLDNIKLKPHDPDSPESQGFIVILHVQSPPSIAAGLNPGAIPF 65
Query: 64 S--SDI-----TATMEKYSQQVADCVLEKAKIVCNDVQ-NVETRIENGDPRDVICQAVQK 115
SD+ TA +E + +++ +L+ A +C NV+T++ GDP++ IC AV++
Sbjct: 66 GGPSDVEVPAFTAAIEAHQKRITQAILDHALGICAKKNANVKTQVVIGDPKEKICDAVEE 125
Query: 116 MGVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVK 155
M D+LVMGS +G IKR FLGSVSN+C + +CPV+IVK
Sbjct: 126 MNADLLVMGSRAFGPIKRMFLGSVSNYCTNHAQCPVIIVK 165
>ref|XP_479478.1| universal stress protein USP1-like protein [Oryza sativa (japonica
cultivar-group)] gi|22831145|dbj|BAC16006.1| universal
stress protein USP1-like protein [Oryza sativa (japonica
cultivar-group)]
Length = 171
Score = 106 bits (264), Expect = 3e-22
Identities = 59/154 (38%), Positives = 95/154 (61%), Gaps = 8/154 (5%)
Query: 11 IMVAIDEGEESIYALTWCLKNL---VFQNSKDHLILLYVKP-PRVVYSAFDGTGYLFSSD 66
++V +D+ E S YAL W +++L + + L++++ KP P V G G S +
Sbjct: 13 VVVGVDDSEHSNYALEWTMQHLASGMAGSGGAELVIVHAKPSPSSVVGFGAGPG---SGE 69
Query: 67 ITATMEKYSQQVADCVLEKAKIVC-NDVQNVETRIENGDPRDVICQAVQKMGVDILVMGS 125
+ +E ++ A+ V+EKA+ +C + + + G+PR V+C AV+K +LV+GS
Sbjct: 70 VVRYVEADLRKTAEDVVEKARRLCIANAMHALIEVIEGEPRYVLCNAVEKHSAGLLVVGS 129
Query: 126 HGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPKS 159
HGYG IKRAFLGSVS++CA + C V+IVK+PK+
Sbjct: 130 HGYGAIKRAFLGSVSDYCAHHAHCSVMIVKQPKA 163
>gb|AAM09541.1| putative universal stress protein USP1 [Oryza sativa (indica
cultivar-group)]
Length = 171
Score = 105 bits (263), Expect = 3e-22
Identities = 59/154 (38%), Positives = 94/154 (60%), Gaps = 8/154 (5%)
Query: 11 IMVAIDEGEESIYALTWCLKNL---VFQNSKDHLILLYVKP-PRVVYSAFDGTGYLFSSD 66
++V +D+ E S YAL W +++L + L++++ KP P V G G S +
Sbjct: 13 VVVGVDDSEHSNYALEWTMQHLASGMAGGGGAELVIVHAKPSPSSVVGFGAGPG---SGE 69
Query: 67 ITATMEKYSQQVADCVLEKAKIVC-NDVQNVETRIENGDPRDVICQAVQKMGVDILVMGS 125
+ +E ++ A+ V+EKA+ +C + + + G+PR V+C AV+K +LV+GS
Sbjct: 70 VVRYVEADLRKTAEDVVEKARRLCIANAMHALIEVIEGEPRYVLCNAVEKHSAGLLVVGS 129
Query: 126 HGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPKS 159
HGYG IKRAFLGSVS++CA + C V+IVK+PK+
Sbjct: 130 HGYGAIKRAFLGSVSDYCAHHAHCSVMIVKQPKA 163
>gb|AAC63627.1| expressed protein [Arabidopsis thaliana] gi|20148413|gb|AAM10097.1|
unknown protein [Arabidopsis thaliana]
gi|15451080|gb|AAK96811.1| Unknown protein [Arabidopsis
thaliana] gi|25409055|pir||F84918 hypothetical protein
At2g47710 [imported] - Arabidopsis thaliana
gi|18407428|ref|NP_566108.1| universal stress protein
(USP) family protein [Arabidopsis thaliana]
Length = 162
Score = 104 bits (260), Expect = 8e-22
Identities = 59/160 (36%), Positives = 90/160 (55%), Gaps = 9/160 (5%)
Query: 5 TENGRRIMVA-IDEGEESIYALTWCLKNLVFQNSKDH---LILLYVKPPRVVYSAFDGTG 60
T +G+ +MV +D+ E+S YAL W L + ++ L +++ KP V G G
Sbjct: 3 TGDGKSVMVVGVDDSEQSTYALEWTLDRFFAPYAPNYPFKLFIVHAKPNAVSAVGLAGPG 62
Query: 61 YLFSSDITATMEKYSQQVADCVLEKAKIVCND--VQNVETRIENGDPRDVICQAVQKMGV 118
++++ ++ + A V+EKAK +C V + GD R+++C+ V K
Sbjct: 63 ---TAEVVPYVDADLKHTAAKVVEKAKAICQSRSVHGAVIEVFEGDARNILCEVVDKHHA 119
Query: 119 DILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPK 158
ILV+GSHGYG IKRA LGS S++CA + C V+IVKKPK
Sbjct: 120 SILVVGSHGYGAIKRAVLGSTSDYCAHHAHCSVMIVKKPK 159
>ref|XP_467911.1| putative ethylene-responsive protein [Oryza sativa (japonica
cultivar-group)] gi|47497367|dbj|BAD19406.1| putative
ethylene-responsive protein [Oryza sativa (japonica
cultivar-group)]
Length = 162
Score = 102 bits (254), Expect = 4e-21
Identities = 55/159 (34%), Positives = 90/159 (56%), Gaps = 9/159 (5%)
Query: 5 TENGRRIMVAIDEGEESIYALTWCLKNLVFQNSKDHLILLYVKPPRVVYSA-------FD 57
T N ++VA+D EES+ AL W L NL + L++L+V+PP + + F
Sbjct: 3 TGNLASVVVAVDGSEESMNALRWALDNLRLRPD-GALVVLHVQPPPSIAAGLNPGPIPFG 61
Query: 58 GTGYLFSSDITATMEKYSQQVADCVLEKAKIVCNDVQ-NVETRIENGDPRDVICQAVQKM 116
G + T +E + +++ +L+ A +C++ V+T + GDP++ IC+ +
Sbjct: 62 GPSEVEVPAFTQAIEAHQRRITQAILDHALKICSEKNVEVKTDVVVGDPKEKICEVTANL 121
Query: 117 GVDILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVK 155
D+LVMG +G +KR FLGSVSN+C NV CPV+++K
Sbjct: 122 KADLLVMGCRAFGPLKRMFLGSVSNYCINNVVCPVVVIK 160
>gb|AAM63890.1| unknown [Arabidopsis thaliana]
Length = 162
Score = 102 bits (254), Expect = 4e-21
Identities = 58/160 (36%), Positives = 89/160 (55%), Gaps = 9/160 (5%)
Query: 5 TENGRRIMVA-IDEGEESIYALTWCLKNLVFQNSKDH---LILLYVKPPRVVYSAFDGTG 60
T +G+ +MV +D+ E+S YAL W L + ++ L +++ KP V G G
Sbjct: 3 TGDGKSVMVVGVDDSEQSTYALEWTLDRFFAPYAPNYPFKLFIVHAKPNAVSAVGLAGPG 62
Query: 61 YLFSSDITATMEKYSQQVADCVLEKAKIVCND--VQNVETRIENGDPRDVICQAVQKMGV 118
++++ ++ + A V+EKAK +C V + GD R+++C+ V K
Sbjct: 63 ---TAEVVPYVDADLKHTAAKVVEKAKAICQSRSVHRAVIEVFEGDARNILCEVVDKHHA 119
Query: 119 DILVMGSHGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPK 158
ILV+GSHGYG I RA LGS S++CA + C V+IVKKPK
Sbjct: 120 SILVVGSHGYGAIXRAVLGSTSDYCAHHAHCSVMIVKKPK 159
>gb|AAR07598.1| fiber protein Fb19 [Gossypium barbadense]
Length = 151
Score = 100 bits (250), Expect = 1e-20
Identities = 57/151 (37%), Positives = 85/151 (55%), Gaps = 8/151 (5%)
Query: 13 VAIDEGEESIYALTWCLKNLVF---QNSKDHLILLYVKPPRVVYSAFDGTGYLFSSDITA 69
V ID+ E S YAL W L + N L++++ KP G G ++D+
Sbjct: 1 VGIDDSEHSTYALQWILDHFFAPFASNPPFKLVIVHAKPSASSAVGLAGPG---AADVLP 57
Query: 70 TMEKYSQQVADCVLEKAKIVC--NDVQNVETRIENGDPRDVICQAVQKMGVDILVMGSHG 127
++ +++A V+EKAK +C V + + GD +V+C AV+K IL +GSHG
Sbjct: 58 YVDADLRKIAARVVEKAKELCLSKSVHDAVVEVGEGDASNVLCDAVEKHHASILAVGSHG 117
Query: 128 YGVIKRAFLGSVSNHCAQNVKCPVLIVKKPK 158
YG IKRA LGSVS++C+ + C V+IVK+PK
Sbjct: 118 YGAIKRAVLGSVSDYCSHHAHCSVMIVKRPK 148
>gb|AAT07452.1| putative universal stress protein [Mirabilis jalapa]
Length = 170
Score = 98.6 bits (244), Expect = 6e-20
Identities = 58/154 (37%), Positives = 88/154 (56%), Gaps = 8/154 (5%)
Query: 11 IMVAIDEGEESIYALTWCLKNLVFQNSKDH---LILLYVKPPRVVYSAFDGTGYLFSSDI 67
++V IDE EE +YAL W L +L +H +L++ P S G +++I
Sbjct: 18 MIVGIDESEECMYALEWALNHLFLPYVPNHPFDFVLVHALPTA---SHAIGLAGPVAAEI 74
Query: 68 TATMEKYSQQVADCVLEKAKIVCND--VQNVETRIENGDPRDVICQAVQKMGVDILVMGS 125
+ ++ + +A V EKA +C + +V +GD R V+C AV+K +LV+GS
Sbjct: 75 SPYVDSDLKNIATRVKEKALELCRSKSLNDVTVETVDGDARKVLCDAVEKYNASMLVVGS 134
Query: 126 HGYGVIKRAFLGSVSNHCAQNVKCPVLIVKKPKS 159
G+G IKRA LGSVS++CA + C V+IVKKPK+
Sbjct: 135 RGHGAIKRAVLGSVSDYCAHHAHCSVIIVKKPKT 168
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.320 0.137 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 280,576,660
Number of Sequences: 2540612
Number of extensions: 10717759
Number of successful extensions: 23451
Number of sequences better than 10.0: 767
Number of HSP's better than 10.0 without gapping: 602
Number of HSP's successfully gapped in prelim test: 165
Number of HSP's that attempted gapping in prelim test: 22553
Number of HSP's gapped (non-prelim): 911
length of query: 165
length of database: 863,360,394
effective HSP length: 118
effective length of query: 47
effective length of database: 563,568,178
effective search space: 26487704366
effective search space used: 26487704366
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 70 (31.6 bits)
Medicago: description of AC149080.7


