

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148996.15 + phase: 0
(153 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
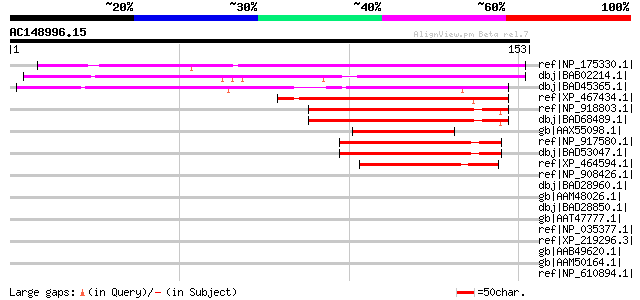
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_175330.1| expressed protein [Arabidopsis thaliana] gi|463... 112 1e-24
dbj|BAB02214.1| unnamed protein product [Arabidopsis thaliana] g... 97 1e-19
dbj|BAD45365.1| hypothetical protein [Oryza sativa (japonica cul... 71 6e-12
ref|XP_467434.1| unknown protein [Oryza sativa (japonica cultiva... 67 1e-10
ref|NP_918803.1| B1096D03.21 [Oryza sativa (japonica cultivar-gr... 50 8e-06
dbj|BAD68489.1| hypothetical protein [Oryza sativa (japonica cul... 50 8e-06
gb|AAX55098.1| hypothetical protein At1g71740 [Arabidopsis thali... 47 9e-05
ref|NP_917580.1| P0704D04.7 [Oryza sativa (japonica cultivar-gro... 47 9e-05
dbj|BAD53047.1| hypothetical protein [Oryza sativa (japonica cul... 47 9e-05
ref|XP_464594.1| hypothetical protein [Oryza sativa (japonica cu... 44 6e-04
ref|NP_908426.1| unknown protein [Oryza sativa (japonica cultiva... 42 0.003
dbj|BAD28960.1| hypothetical protein [Oryza sativa (japonica cul... 39 0.033
gb|AAM48026.1| unknown protein [Arabidopsis thaliana] gi|1825297... 37 0.096
dbj|BAD28850.1| unknown protein [Oryza sativa (japonica cultivar... 37 0.13
gb|AAT47777.1| AT12465p [Drosophila melanogaster] 35 0.36
ref|NP_035377.1| retinoblastoma binding protein 6 [Mus musculus]... 34 0.62
ref|XP_219296.3| PREDICTED: similar to PACT [Rattus norvegicus] 34 0.62
gb|AAB49620.1| PACT 34 0.62
gb|AAM50164.1| GH12664p [Drosophila melanogaster] gi|28381499|gb... 33 1.1
ref|NP_610894.1| CG6209-PA [Drosophila melanogaster] gi|7303284|... 33 1.1
>ref|NP_175330.1| expressed protein [Arabidopsis thaliana] gi|46359837|gb|AAS88782.1|
At1g49000 [Arabidopsis thaliana]
gi|45476547|gb|AAS65939.1| At1g49000 [Arabidopsis
thaliana] gi|7770334|gb|AAF69704.1| F27J15.21
[Arabidopsis thaliana]
Length = 156
Score = 112 bits (281), Expect = 1e-24
Identities = 64/149 (42%), Positives = 89/149 (58%), Gaps = 9/149 (6%)
Query: 9 SPNRHHSNGGFFVTISAILALLSRKANRLKEKAKSSSTTKPIRD-----EEWRFDLKTPP 63
SP+ + F V++ + L +R ANRL +K K S + E +R+++ +
Sbjct: 11 SPSHKRLSVSFLVSM---MVLCARHANRLSKKLKLKSKKRTCSGGEGGGERFRWNMISSS 67
Query: 64 KSPMAKPKKLLSNISNKALSQFGKKKQREEREKEGWGNGGVWQKEILMGGKCEPLDFSGV 123
S +PK+L + +SNKA++ +K EE+ G+WQ+EILMGGKCEPLDFSGV
Sbjct: 68 MSS-PRPKELFTTLSNKAMTMVRRKNPPEEKATAMEEEHGLWQREILMGGKCEPLDFSGV 126
Query: 124 IYYDINGKQTREVPIRSPRASPLPGYLTR 152
IYYD NG+ EVP RSPR +PLP Y TR
Sbjct: 127 IYYDSNGRLLNEVPPRSPRGTPLPSYPTR 155
>dbj|BAB02214.1| unnamed protein product [Arabidopsis thaliana]
gi|15809742|gb|AAL06799.1| AT3g18560/K24M9_5
[Arabidopsis thaliana] gi|14190489|gb|AAK55725.1|
AT3g18560/K24M9_5 [Arabidopsis thaliana]
gi|18401929|ref|NP_566614.1| expressed protein
[Arabidopsis thaliana]
Length = 195
Score = 96.7 bits (239), Expect = 1e-19
Identities = 67/183 (36%), Positives = 96/183 (51%), Gaps = 40/183 (21%)
Query: 5 TSPESPNRHHSNGGFFVTISAILALLSRKANRLKEKAKSSSTTKPIRDEEWRFDLKT--- 61
++PE + H +S ++ L +R A+R+ +K K T K E++ K+
Sbjct: 4 STPEHVSSAHKRISVSFLVS-LMVLCARHASRVSKKLKPKKTRKQTHLEDYLESPKSNGN 62
Query: 62 --------------PPK--SPM-AKPKKLLSNISNKALSQFGKKKQR------------- 91
P + SPM +PK+L + +SNKA++ G+K +
Sbjct: 63 GSEDGRGGGRFGWSPARTFSPMRVRPKELYTTLSNKAMTMVGRKNKAYDGGPTKKTAVEM 122
Query: 92 --EEREKEGWGNGGVWQKEILMGGKCEPLDFSGVIYYDINGKQTREVPIRSPRASPLPGY 149
EE E+E GVWQ+EILMGGKCEPLD+SGVIYYD +G Q ++VP RSPRAS +P
Sbjct: 123 VMEEDEEEY----GVWQREILMGGKCEPLDYSGVIYYDCSGHQLKQVPPRSPRASLVPER 178
Query: 150 LTR 152
TR
Sbjct: 179 PTR 181
>dbj|BAD45365.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
Length = 171
Score = 70.9 bits (172), Expect = 6e-12
Identities = 49/147 (33%), Positives = 76/147 (51%), Gaps = 13/147 (8%)
Query: 3 KITSPESPNRHHSNGGFFVTISAILALLSRKANRLKEKAKSSSTTKPIRDEEWRFDLKTP 62
K T+ + +R GG + + A+ AL+S RL+ A+ S+ ++
Sbjct: 23 KATTAAASDRKVMAGGASL-LHAVAALMSTCTRRLQRAARRVSSAAAGAGKQGASSRAVV 81
Query: 63 P-KSPMAKPKKLLSNISNKALSQFGKKKQREEREKEGWGNGGVWQKEILMGGKCEPLDFS 121
P + ++ P + + A + R +EG +GG+W+KEILMG +C+PLDFS
Sbjct: 82 PWRKALSLPAAATAKVKAAAAAA---------RREEG-DSGGLWRKEILMGERCQPLDFS 131
Query: 122 GVIYYDINGKQ-TREVPIRSPRASPLP 147
GVIYYD +G++ P RSP SPLP
Sbjct: 132 GVIYYDADGRRLAHPPPPRSPMRSPLP 158
>ref|XP_467434.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|41052652|dbj|BAD07500.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 138
Score = 66.6 bits (161), Expect = 1e-10
Identities = 34/69 (49%), Positives = 46/69 (66%), Gaps = 2/69 (2%)
Query: 80 KALSQFGKKKQREEREKEGWGNGGVWQKEILMGGKCEPLDFSGVIYYDINGKQTRE-VPI 138
KA+S +++++ E GVW+KEILMG +C+PLDFSGVIYYD G++ + P
Sbjct: 56 KAISS-SRRRRKAGAELSFRAEDGVWRKEILMGERCQPLDFSGVIYYDAEGRRLEQPPPP 114
Query: 139 RSPRASPLP 147
RSP SPLP
Sbjct: 115 RSPLRSPLP 123
>ref|NP_918803.1| B1096D03.21 [Oryza sativa (japonica cultivar-group)]
Length = 918
Score = 50.4 bits (119), Expect = 8e-06
Identities = 28/62 (45%), Positives = 39/62 (62%), Gaps = 5/62 (8%)
Query: 89 KQREEREKEGWGNGGVWQKEILMGGKCEPLDFSGVIYYDINGKQTREVPIRSPRA---SP 145
++R +E GV +KEILM +C+ LDFSG+IYYD+ G++ + P PRA SP
Sbjct: 846 RRRRRQELSFRVEDGVCRKEILMEERCQSLDFSGMIYYDVAGRRLEQPP--PPRALLHSP 903
Query: 146 LP 147
LP
Sbjct: 904 LP 905
>dbj|BAD68489.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
Length = 138
Score = 50.4 bits (119), Expect = 8e-06
Identities = 28/62 (45%), Positives = 39/62 (62%), Gaps = 5/62 (8%)
Query: 89 KQREEREKEGWGNGGVWQKEILMGGKCEPLDFSGVIYYDINGKQTREVPIRSPRA---SP 145
++R +E GV +KEILM +C+ LDFSG+IYYD+ G++ + P PRA SP
Sbjct: 66 RRRRRQELSFRVEDGVCRKEILMEERCQSLDFSGMIYYDVAGRRLEQPP--PPRALLHSP 123
Query: 146 LP 147
LP
Sbjct: 124 LP 125
>gb|AAX55098.1| hypothetical protein At1g71740 [Arabidopsis thaliana]
gi|15217551|ref|NP_177319.1| hypothetical protein
[Arabidopsis thaliana] gi|7239501|gb|AAF43227.1|
F14O23.12 [Arabidopsis thaliana] gi|25406118|pir||G96739
F14O23.12 [imported] - Arabidopsis thaliana
Length = 130
Score = 47.0 bits (110), Expect = 9e-05
Identities = 20/30 (66%), Positives = 24/30 (79%)
Query: 102 GGVWQKEILMGGKCEPLDFSGVIYYDINGK 131
G +WQK ILMGGKC+ DFSGVI YD +G+
Sbjct: 86 GPLWQKNILMGGKCQLPDFSGVILYDADGQ 115
>ref|NP_917580.1| P0704D04.7 [Oryza sativa (japonica cultivar-group)]
Length = 135
Score = 47.0 bits (110), Expect = 9e-05
Identities = 21/48 (43%), Positives = 30/48 (61%), Gaps = 2/48 (4%)
Query: 98 GWGNGGVWQKEILMGGKCEPLDFSGVIYYDINGKQTREVPIRSPRASP 145
G G G+W++ ILMG +CEPL F G I+YD G++ + R +A P
Sbjct: 52 GGGGEGLWRRAILMGERCEPLSFPGAIHYDSRGRRLSQP--RRAKAKP 97
>dbj|BAD53047.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
Length = 505
Score = 47.0 bits (110), Expect = 9e-05
Identities = 21/48 (43%), Positives = 30/48 (61%), Gaps = 2/48 (4%)
Query: 98 GWGNGGVWQKEILMGGKCEPLDFSGVIYYDINGKQTREVPIRSPRASP 145
G G G+W++ ILMG +CEPL F G I+YD G++ + R +A P
Sbjct: 422 GGGGEGLWRRAILMGERCEPLSFPGAIHYDSRGRRLSQP--RRAKAKP 467
>ref|XP_464594.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|49387925|dbj|BAD25025.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 299
Score = 44.3 bits (103), Expect = 6e-04
Identities = 23/41 (56%), Positives = 26/41 (63%), Gaps = 2/41 (4%)
Query: 104 VWQKEILMGGKCEPLDFSGVIYYDINGKQTREVPIRSPRAS 144
VWQK ILMGGKC+ +FSGVI YD G P PRA+
Sbjct: 98 VWQKNILMGGKCQLPEFSGVINYDAAGNIV--APSGRPRAA 136
>ref|NP_908426.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|14587253|dbj|BAB61171.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
gi|13486649|dbj|BAB39887.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 175
Score = 42.0 bits (97), Expect = 0.003
Identities = 19/41 (46%), Positives = 24/41 (58%)
Query: 104 VWQKEILMGGKCEPLDFSGVIYYDINGKQTREVPIRSPRAS 144
+W+K I+MG KC PL FSG I YD +G Q I A+
Sbjct: 126 LWRKTIMMGDKCRPLQFSGHIAYDSDGNQLPATTISKEAAN 166
>dbj|BAD28960.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
Length = 202
Score = 38.5 bits (88), Expect = 0.033
Identities = 17/31 (54%), Positives = 22/31 (70%)
Query: 104 VWQKEILMGGKCEPLDFSGVIYYDINGKQTR 134
VWQ+ ILMG +CE FSG+I YD +G+ R
Sbjct: 138 VWQRRILMGMRCELPRFSGLILYDEHGRPIR 168
>gb|AAM48026.1| unknown protein [Arabidopsis thaliana] gi|18252977|gb|AAL62415.1|
unknown protein [Arabidopsis thaliana]
gi|22331082|ref|NP_188094.2| expressed protein
[Arabidopsis thaliana]
Length = 168
Score = 37.0 bits (84), Expect = 0.096
Identities = 31/98 (31%), Positives = 41/98 (41%), Gaps = 14/98 (14%)
Query: 44 SSTTKPIRDEEWRFDLKTPPKSPMAKPKKLLSNISNKALSQF-----GKKKQREERE--- 95
+S K + D EW + ++ K K S + + F K + E R
Sbjct: 42 NSRGKQVADAEWASETAFGSRTMEEKRKCKWSKVKRTLMGSFCWSSAAKWMEMETRRQPP 101
Query: 96 ----KEGWGNG--GVWQKEILMGGKCEPLDFSGVIYYD 127
KE N VWQ+ ILMG KCE FSG+I YD
Sbjct: 102 LLAVKERSLNAVDPVWQRPILMGEKCELPRFSGLILYD 139
>dbj|BAD28850.1| unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 143
Score = 36.6 bits (83), Expect = 0.13
Identities = 14/28 (50%), Positives = 21/28 (75%)
Query: 104 VWQKEILMGGKCEPLDFSGVIYYDINGK 131
+WQ+++LMG KC+ FSG+I YD G+
Sbjct: 89 IWQRKVLMGVKCQLPRFSGMILYDERGR 116
>gb|AAT47777.1| AT12465p [Drosophila melanogaster]
Length = 742
Score = 35.0 bits (79), Expect = 0.36
Identities = 29/138 (21%), Positives = 57/138 (41%), Gaps = 4/138 (2%)
Query: 20 FVTISAILALLSRKANRLK-EKAKSSSTTKPIRDEEWRFDLKTPPKSPMAKPKKLLSNIS 78
FV +L + R+ RLK + + D E FD P + + +P++L S+
Sbjct: 500 FVDKKRLLYHVRRECERLKIDSCEDELVLTDTSDSEDEFDFGVTPPAGLMRPERLKSSHV 559
Query: 79 NKALSQFGKKKQREEREKEGWGNGGVWQKEILMGGKCEPLDF---SGVIYYDINGKQTRE 135
+Q+ + + V + +MG + +P ++ GVI + + R
Sbjct: 560 KHTETQYNESDWVNPSLYPHMPDATVQYESCVMGEREKPFNWIYGKGVIQEEPKLPKMRN 619
Query: 136 VPIRSPRASPLPGYLTRG 153
+P + + P+PG + G
Sbjct: 620 IPKKPKKKKPVPGRVAGG 637
>ref|NP_035377.1| retinoblastoma binding protein 6 [Mus musculus]
gi|3858885|gb|AAC72432.1| proliferation
potential-related protein [Mus musculus]
gi|11360378|pir||T42727 proliferation potential-related
protein - mouse
Length = 1560
Score = 34.3 bits (77), Expect = 0.62
Identities = 23/67 (34%), Positives = 34/67 (50%), Gaps = 4/67 (5%)
Query: 32 RKANRLKEKAKSSSTTKPIRDEEWRFDLKTPPKS--PMAKPKKLLSNISNKALSQFGKKK 89
RKA+ + AK KP +DE+ + D KS P +K +K NK L G+K+
Sbjct: 860 RKAH--SKSAKEHQEAKPAKDEKVKKDCSKDIKSEKPASKDEKAKKPEKNKLLDSKGEKR 917
Query: 90 QREEREK 96
+R+ EK
Sbjct: 918 KRKTEEK 924
>ref|XP_219296.3| PREDICTED: similar to PACT [Rattus norvegicus]
Length = 1789
Score = 34.3 bits (77), Expect = 0.62
Identities = 22/67 (32%), Positives = 35/67 (51%), Gaps = 4/67 (5%)
Query: 32 RKANRLKEKAKSSSTTKPIRDEEWRFDLKTPPKS--PMAKPKKLLSNISNKALSQFGKKK 89
RKA+ + AK KP+++E+ + D KS P +K +K NK L G+K+
Sbjct: 1089 RKAH--SKSAKEHQEAKPVKEEKAKKDCSKDIKSEKPASKDEKAKKPEKNKLLESKGEKR 1146
Query: 90 QREEREK 96
+R+ EK
Sbjct: 1147 KRKTEEK 1153
>gb|AAB49620.1| PACT
Length = 1587
Score = 34.3 bits (77), Expect = 0.62
Identities = 23/67 (34%), Positives = 34/67 (50%), Gaps = 4/67 (5%)
Query: 32 RKANRLKEKAKSSSTTKPIRDEEWRFDLKTPPKS--PMAKPKKLLSNISNKALSQFGKKK 89
RKA+ + AK KP +DE+ + D KS P +K +K NK L G+K+
Sbjct: 887 RKAH--SKSAKEHQEAKPAKDEKVKKDCSKDIKSEKPASKDEKAKKPEKNKLLDSKGEKR 944
Query: 90 QREEREK 96
+R+ EK
Sbjct: 945 KRKTEEK 951
>gb|AAM50164.1| GH12664p [Drosophila melanogaster] gi|28381499|gb|AAF57010.4|
CG31025-PA, isoform A [Drosophila melanogaster]
gi|28572009|ref|NP_733345.2| CG31025-PA, isoform A
[Drosophila melanogaster]
Length = 998
Score = 33.5 bits (75), Expect = 1.1
Identities = 28/138 (20%), Positives = 56/138 (40%), Gaps = 4/138 (2%)
Query: 20 FVTISAILALLSRKANRLK-EKAKSSSTTKPIRDEEWRFDLKTPPKSPMAKPKKLLSNIS 78
FV +L + R+ R K + + D E FD P + + +P++L S+
Sbjct: 756 FVDKKRLLYHVRRECERRKIDSCEDELVLTDTSDSEDEFDFGVTPPAGLMRPERLKSSHV 815
Query: 79 NKALSQFGKKKQREEREKEGWGNGGVWQKEILMGGKCEPLDF---SGVIYYDINGKQTRE 135
+Q+ + + V + +MG + +P ++ GVI + + R
Sbjct: 816 KHTETQYNESDWVNPSLYPHMPDATVQYESCVMGEREKPFNWIYGKGVIQEEPKPPKMRN 875
Query: 136 VPIRSPRASPLPGYLTRG 153
+P + + P+PG + G
Sbjct: 876 IPKKPKKKKPVPGRVAGG 893
>ref|NP_610894.1| CG6209-PA [Drosophila melanogaster] gi|7303284|gb|AAF58345.1|
CG6209-PA [Drosophila melanogaster]
Length = 607
Score = 33.5 bits (75), Expect = 1.1
Identities = 29/93 (31%), Positives = 41/93 (43%), Gaps = 10/93 (10%)
Query: 31 SRKANRLKEKAKSSSTTKPIRDEEWRFDLKTPPKSPMAKP--KKLLSNISNKALSQFGKK 88
+++ R K++ K TKP D+E K K K K+LL I K K
Sbjct: 257 AKRKEREKKEGKPLDCTKPSEDDETGAKNKAEKKKAKEKAEIKRLLKEIKGKC------K 310
Query: 89 KQREEREKEGWGNGGVWQ--KEILMGGKCEPLD 119
KQREE E++ + + KE KCE L+
Sbjct: 311 KQREEAERKKKEEAELNKKCKEAAAKKKCEELE 343
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.313 0.133 0.391
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 284,243,917
Number of Sequences: 2540612
Number of extensions: 11504722
Number of successful extensions: 28515
Number of sequences better than 10.0: 128
Number of HSP's better than 10.0 without gapping: 36
Number of HSP's successfully gapped in prelim test: 95
Number of HSP's that attempted gapping in prelim test: 28438
Number of HSP's gapped (non-prelim): 153
length of query: 153
length of database: 863,360,394
effective HSP length: 129
effective length of query: 24
effective length of database: 535,621,446
effective search space: 12854914704
effective search space used: 12854914704
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 67 (30.4 bits)
Medicago: description of AC148996.15


