

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148819.4 - phase: 0
(1009 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
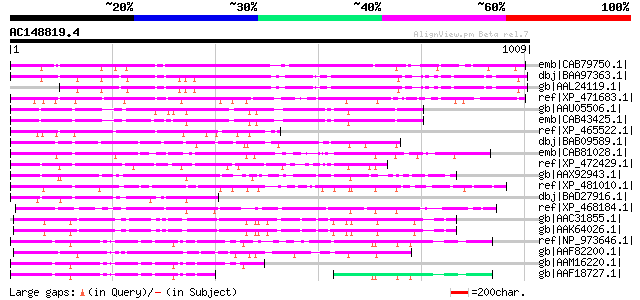
Score E
Sequences producing significant alignments: (bits) Value
emb|CAB79750.1| putative protein [Arabidopsis thaliana] gi|49144... 684 0.0
dbj|BAA97363.1| unnamed protein product [Arabidopsis thaliana] 650 0.0
gb|AAL24119.1| unknown protein [Arabidopsis thaliana] gi|2839384... 532 e-149
ref|XP_471683.1| OSJNBa0035B13.5 [Oryza sativa (japonica cultiva... 421 e-116
gb|AAU05506.1| At3g52490 [Arabidopsis thaliana] 362 5e-98
emb|CAB43425.1| putative protein [Arabidopsis thaliana] gi|15231... 361 8e-98
ref|XP_465522.1| heat shock protein-related-like [Oryza sativa (... 347 9e-94
dbj|BAB09589.1| 101 kDa heat shock protein; HSP101-like protein ... 334 1e-89
emb|CAB81028.1| putative protein [Arabidopsis thaliana] gi|15234... 328 6e-88
ref|XP_472429.1| OJ991214_12.16 [Oryza sativa (japonica cultivar... 290 1e-76
gb|AAX92943.1| transposon protein, putative, mutator sub-class [... 262 4e-68
ref|XP_481010.1| 101 kDa heat shock protein; HSP101-like protein... 256 2e-66
dbj|BAD27916.1| heat shock protein-related-like [Oryza sativa (j... 249 3e-64
ref|XP_468184.1| 101 kDa heat shock protein-like [Oryza sativa (... 223 3e-56
gb|AAC31855.1| expressed protein [Arabidopsis thaliana] gi|74860... 207 1e-51
gb|AAK64026.1| unknown protein [Arabidopsis thaliana] 207 2e-51
ref|NP_973646.1| heat shock protein-related [Arabidopsis thaliana] 189 3e-46
gb|AAF82200.1| Strong similarity to an unknown protein F23F1.11 ... 180 2e-43
gb|AAM16220.1| At2g40130/T7M7.2 [Arabidopsis thaliana] gi|154507... 164 1e-38
gb|AAF18727.1| hypothetical protein [Arabidopsis thaliana] gi|25... 161 9e-38
>emb|CAB79750.1| putative protein [Arabidopsis thaliana] gi|4914416|emb|CAB43667.1|
putative protein [Arabidopsis thaliana]
gi|15233760|ref|NP_194721.1| heat shock protein-related
[Arabidopsis thaliana] gi|7486224|pir||T08553
hypothetical protein F27B13.160 - Arabidopsis thaliana
Length = 1017
Score = 684 bits (1764), Expect = 0.0
Identities = 452/1078 (41%), Positives = 613/1078 (55%), Gaps = 140/1078 (12%)
Query: 1 MRSGAYTLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSL-RNACLKS 59
MR+GAYT+ +TLT EAAS+LK SL LARRRGH+Q+TPLHVA+TLL+ S+L R ACLKS
Sbjct: 1 MRTGAYTVHQTLTPEAASVLKQSLTLARRRGHSQVTPLHVASTLLTSSRSNLFRRACLKS 60
Query: 60 Q------QQNNHP-LQCRALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQ 112
+Q HP L CRALELCFNV+LNRLPT +PL Q Q PSLSNAL+AALKRAQ
Sbjct: 61 NPFTALGRQMAHPSLHCRALELCFNVSLNRLPTNP-NPLFQTQ--PSLSNALVAALKRAQ 117
Query: 113 AHQRRGCIEQNQQQQQQPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENS 172
AHQRRGC+EQ Q QQ QP L+VKVEL+QL++SILDDPSVSRVMREAG SS SVK+N+E+
Sbjct: 118 AHQRRGCVEQQQSQQNQPFLAVKVELEQLVVSILDDPSVSRVMREAGLSSVSVKSNIEDD 177
Query: 173 STLIN-----SSSVFHSSPSPLSHNHFLSSYGYGSV-----------LFSSQKKEQVVYH 216
S++++ SSS SP S + ++ G G++ L + EQ +
Sbjct: 178 SSVVSPVFYGSSSSVGVFSSPCSPSSSENNQGGGTLSPNPSKIWHAHLTNHHSFEQNPFF 237
Query: 217 PFLKSSESN-------KEDINLVFDVLLRK---KKKNTVIVGDTVSLTEGLVSEIMKRFE 266
F K +ED N V +VLL K KK+NTVIVGD+VSLTEG+V+++M R E
Sbjct: 238 HFPKGKTFTPDQAFPVREDANPVIEVLLGKKNNKKRNTVIVGDSVSLTEGVVAKLMGRIE 297
Query: 267 RGEVPDEMKTTHFVKFHGLSSVSLKYMKKEEVEMNVIRVLKRKVSDYVAL-GVGAIFYVG 325
RGEVPD++K THF+KF S V L +MKKE++E +R LKRK+ + + G G I +G
Sbjct: 298 RGEVPDDLKQTHFIKFQ-FSQVGLNFMKKEDIE-GQVRELKRKIDSFTSWGGKGVIVCLG 355
Query: 326 DLKWIV--DDNDGSLNEKEVVDYVVEEIGKLFGEEGNKNGKIWLVATASYQSYMRCQMRV 383
DL W V N S + D++VEEIG+L + N K+WL+ TASYQ+YMRCQM+
Sbjct: 356 DLDWAVWGGGNSASSSNYSAADHLVEEIGRLVYDYSNTGAKVWLLGTASYQTYMRCQMKQ 415
Query: 384 PTFENQWCLQAVPVPSGGLGLSLHSSSVHDSKMSISQNPSPMLESKFFSNKEEHE-KLNC 442
P + W LQAV +PSGGL L+LH+SS + + P + E + + +EE E KLN
Sbjct: 416 PPLDVHWALQAVSIPSGGLSLTLHASSSEMASQVMEMKPFRVKEEEEGAREEEEEDKLNF 475
Query: 443 CEECVSNYEKEAQLFKPDQKNLLPSWLQSH--STEARQKDELTQLNKKWNRLCQCLHQNK 500
C EC NYEKEA+ F Q +LP WLQ H + QKDEL+ L KKWNR CQ LH K
Sbjct: 476 CGECAFNYEKEAKAFISAQHKILPPWLQPHGDNNNINQKDELSGLRKKWNRFCQALHHKK 535
Query: 501 QPQNHWSNNHSSNAKIYPYNSSYPYWPNQGSSILPDTSSSISFADSATKP-AYSSNLIPR 559
W Q SS+LP S DS+ K + +S+ + +
Sbjct: 536 PSMTAWR-------------------AEQSSSVLPG-----SLMDSSLKQNSRASSSVAK 571
Query: 560 FRRGQQSCTIEFNFNDEKAQKNQVTATLELDSLKGM--EGTKEVKTTLALGNSTF---SV 614
FRR Q SCTIEF+F + + + T L LD K EG K K TLALG+S F S
Sbjct: 572 FRR-QNSCTIEFSFGSNRQEGLKKTDELSLDGFKSNNDEGVK-TKITLALGHSPFPSDSE 629
Query: 615 SDQKRMENLTLQRDHIYKVLQENIPWHCETVSSIAEALVDS-KSSKECATWLFLQGNDSV 673
+ ++ ++ + + L ENIPW + + SI EA+ +S K SK W+ + GND
Sbjct: 630 NSEEEEPEKAIKMSKLLEKLHENIPWQKDVLPSIVEAMEESVKRSKRKDAWMLVSGNDVT 689
Query: 674 GKKRLALAIAESVFGSVEMFSHVDMMKRENSETPFSEKVVGPLKNNEKFVVLVENADFGD 733
K+RLA+ + S+FGS E +++ + SE E++ LK E+ V+L+E D D
Sbjct: 690 AKRRLAITLTTSLFGSHENMLKINLRTSKASEA--CEELKNALKKKEEVVILIERVDLAD 747
Query: 734 TLIRKMLADEFEIAKF----GTLGQKIFILSNGGSMVSEDQKKDSVMKLVLKISETEKKP 789
+L D FE G Q IF+L+ E++ V+ +VL
Sbjct: 748 AQFMNILVDRFEAGDLDGFQGKKSQIIFLLTREDDECVENE--HFVIPMVL--------- 796
Query: 790 TFELSPSSSSSSKSPCLGNKRSAELD---LFSKIKIPRIEENE---------GNKKREFS 837
+ + S S + NKR E D K K PRIEE++ N K+E
Sbjct: 797 -------NCNKSGSGLVNNKRKPEYDAAPTMIKKKNPRIEEDDDESNVACDISNIKKE-- 847
Query: 838 FSRQSSF-NNTLDLNMKADEEDNEDYDEGENSPISSDLTRETLGEHLISNESLDSIENLF 896
FSRQ F +N LDLN++ D +++E+ + + ISS LDSI+N F
Sbjct: 848 FSRQLKFESNALDLNLRVDADEDEEEEAKPATEISSG----------FEERFLDSIQNRF 897
Query: 897 EFNQSPAKNKEMTQMFMSRVKESFEEVLGNVK----FSVQDKVIEEIGVGCGSFTNNMFE 952
+F + ++++T+ F++++K+S EE+LG + F+V ++IE+ GCG F N +FE
Sbjct: 898 DF--TVLSDEDITKFFVTKIKDSCEEILGQREERFGFTVDAELIEKFYKGCGFFANGLFE 955
Query: 953 KWLKGIFQTSLERVNGGDKNGI-VYTLCWGG------KEDRKWDSGFMGSCLPKNIQI 1003
+W+K +FQ L V G K GI V LC GG E + + GFMG+CLP I +
Sbjct: 956 EWVKEVFQRGLVTVKNGGKEGISVINLCLGGIDMIDQGEVYEEEEGFMGTCLPNRIHV 1013
>dbj|BAA97363.1| unnamed protein product [Arabidopsis thaliana]
Length = 1028
Score = 650 bits (1676), Expect = 0.0
Identities = 462/1105 (41%), Positives = 619/1105 (55%), Gaps = 177/1105 (16%)
Query: 1 MRSGAYTLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKS- 59
MR+G YT+Q+TLT EAAS+LKHSL LARRRGHAQ+TPLHVAATLLS +S LR AC+KS
Sbjct: 1 MRTGGYTIQQTLTTEAASVLKHSLTLARRRGHAQVTPLHVAATLLSSRTSLLRRACIKSH 60
Query: 60 ------------------QQQNNHPLQCRALELCFNVALNRLPTTTTSPLLQPQHVPSLS 101
NHPLQCRALELCFNVALNRLP T P+ Q PSL+
Sbjct: 61 PGFSTNYQFAPSRLQHHHHHNQNHPLQCRALELCFNVALNRLP-TVPGPMFHGQ--PSLA 117
Query: 102 NALIAALKRAQAHQRRGCIEQNQQQQQQP------LLSVKVELDQLILSILDDPSVSRVM 155
NAL+AALKRAQAHQRRGCIEQ QQ Q P LL+VKVEL+QL++SILDDPSVSRVM
Sbjct: 118 NALVAALKRAQAHQRRGCIEQQQQTQTHPQTQQTQLLAVKVELEQLVISILDDPSVSRVM 177
Query: 156 REAGFSSPSVKNNLENSSTLI------------------------NSSSVFHSSPSPLSH 191
REAGF+S +VK+ +E+ S NS + H +P
Sbjct: 178 REAGFNSTAVKSCVEDCSVSSVFYGGSAVGVFSSPNSPDQQQQHHNSINRLHHYQNPKDF 237
Query: 192 NHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKE---------DINLVFDVLLRK--K 240
N ++ F +Q +Q +P L SS ++ D+ LV DVL+RK K
Sbjct: 238 NFINPNFPLWQTHFLNQSPDQ---NPLLLSSSASHHHQQQRLREIDLKLVVDVLMRKKTK 294
Query: 241 KKNTVIVGDTVSLTEGLVSEIMKRFERGEVPD--EMKTTHFVKFHGLSSVSLKYMKKEEV 298
KKN VIVGD++S TEG VSE+M + ERGE+ E+K THFVKFH S ++ K+M++E+V
Sbjct: 295 KKNPVIVGDSISFTEGFVSELMAKLERGEIDQTGELKQTHFVKFH-FSPMASKFMRREDV 353
Query: 299 EMNVIRVLKRKVSDYVALGVGAIFYVGDLKW----IVDDNDGSLNE----KEVVDYVVEE 350
E+N I+ L++KV G AI + GDLKW I ++N G +NE +D++VEE
Sbjct: 354 ELN-IKELRKKVLSLTTSGKNAIIFTGDLKWTVKEITNNNSGGINEISSSYSPLDHLVEE 412
Query: 351 IGKLFGE-------EGNKNGKIWLVATASYQSYMRCQMRVPTFENQWCLQAVPVP-SGGL 402
IGKL E + K K+W++ TAS+Q+YMRCQMR P+ E W L V VP S L
Sbjct: 413 IGKLITECNDDGDDDDCKTRKVWVMGTASFQTYMRCQMRQPSLETLWALHPVSVPSSANL 472
Query: 403 GLSLHSSSVHDSKMSISQNPSPMLESKFFSNKEE--HEKLNCCEECVSNYEKEAQLFKPD 460
GLSLH++S H+++ + N + L + +EE L+CC ECV+++++EA+ K +
Sbjct: 473 GLSLHATSGHEARNMSTVNATKSLSGYDKAEEEETISHVLSCCPECVTSFDREAKSLKAN 532
Query: 461 QKNLLPSWLQSHSTE-ARQKDELTQLNKKWNRLCQCLHQNKQPQNHWSNNHSSNAKIYPY 519
Q LLPSWLQSH + + QKDEL L +KWNR C+ LH + N YPY
Sbjct: 533 QDKLLPSWLQSHDADSSSQKDELMGLKRKWNRFCETLHNQTGQLSMMGN--------YPY 584
Query: 520 NSSYPYWPNQGSSILPDTSSSISFADS-ATKP-AYSSNLIPRFRRGQQSCTIEFNFNDEK 577
Y GSS ++S S S DS KP ++N I +FRR Q SCTIEF+ +
Sbjct: 585 GLPY------GSS--HESSKSTSLIDSLGLKPNQRATNSIAKFRR-QNSCTIEFDLGGNE 635
Query: 578 AQKNQVTATLELDSLKGMEGTKEVKTTLALGNSTF---SVSDQKRMENLTLQRDHIYKVL 634
+K + E D KG E TL LG S F SV+D + L+ + K L
Sbjct: 636 HEKGESINEAEDD--KGNE-----TVTLDLGRSLFRSDSVTDTR------LKLSALVKAL 682
Query: 635 QENIPWHCETVSSIAEALVDSKSSKECATWLFLQGNDSVGKKRLALAIAESVFGSVEMFS 694
+E+IP T+ IAE+L+D S K+ +W+ ++G D+ K+R+A ++ESVFGS E
Sbjct: 683 EESIPRQTVTMRLIAESLMDCVSKKK-DSWIIIEGRDTTAKRRVARTVSESVFGSFESLV 741
Query: 695 HVDMMKREN-SETPFSEKVVGPLKNNEKFVVLVENADFGDTLIRKMLADEFEIAKFGTLG 753
H+D+ K+ N S+ + + LKN EK V L+E+ D D+ K+LAD FE + G
Sbjct: 742 HIDLKKKGNESKASPATLLAYELKNPEKVVFLIEDIDLADSRFLKLLADRFEDKRRIKTG 801
Query: 754 ----QKIFILSNGGSMVSEDQKKDSVMKLVLKISETEKKPTFELSPSSSSSSKSPCLGNK 809
Q IFIL+ S + +DSV+++ L+I +++SP G K
Sbjct: 802 IDHRQAIFILTKEDS--RNVRNRDSVLQIGLEI-----------------TAQSP--GKK 840
Query: 810 RSAELDLFSKIKIPRIEENEGNKKREFSFSRQSSFNNT-LDLNMKADEEDNEDYDEGENS 868
R E DL IE KK SRQSSFN++ LDLN+KA++E+ EGE S
Sbjct: 841 RKPESDL-------SIENGFWMKKE--VCSRQSSFNSSYLDLNIKAEDEE----VEGEIS 887
Query: 869 PISSDLTRETLGEHLISNESLDSIENLFEFNQS--PAKNKEMTQMFMSRVKESFEEVLGN 926
PISSDLT E E S+ L+ I+N F N+S P K M + EE G
Sbjct: 888 PISSDLTGEEETEFSSSSNFLNRIQNRFVLNRSCEPGIEKGMITAAFREIFPEREEG-GG 946
Query: 927 VKFSVQDKVIEEIGVGCGSFTNNMFEKWLKGIFQTSLERV-NGGDKNGIVYTLCWGGKED 985
V+FSV+DK++EE+ N FE+WLK +FQT L V GG K+ V + +GG D
Sbjct: 947 VRFSVEDKLVEEL----YGIQNGAFERWLKEVFQTGLLTVKKGGKKDTGVIRMVFGGIVD 1002
Query: 986 RK----WDSGFMGSCLPKNIQIVNY 1006
K G+M + LP +Q+ +
Sbjct: 1003 NKGYGGGVGGYMATFLPNKVQVSKF 1027
>gb|AAL24119.1| unknown protein [Arabidopsis thaliana] gi|28393849|gb|AAO42332.1|
unknown protein [Arabidopsis thaliana]
gi|18423941|ref|NP_568849.1| expressed protein
[Arabidopsis thaliana]
Length = 920
Score = 532 bits (1370), Expect = e-149
Identities = 393/989 (39%), Positives = 542/989 (54%), Gaps = 155/989 (15%)
Query: 98 PSLSNALIAALKRAQAHQRRGCIEQNQQQQQQP------LLSVKVELDQLILSILDDPSV 151
PSL+NAL+AALKRAQAHQRRGCIEQ QQ Q P LL+VKVEL+QL++SILDDPSV
Sbjct: 6 PSLANALVAALKRAQAHQRRGCIEQQQQTQTHPQTQQTQLLAVKVELEQLVISILDDPSV 65
Query: 152 SRVMREAGFSSPSVKNNLENSSTLI------------------------NSSSVFHSSPS 187
SRVMREAGF+S +VK+ +E+ S NS + H +
Sbjct: 66 SRVMREAGFNSTAVKSCVEDCSVSSVFYGGSAVGVFSSPNSPDQQQQHHNSINRLHHYQN 125
Query: 188 PLSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKE---------DINLVFDVLLR 238
P N ++ F +Q +Q +P L SS ++ D+ LV DVL+R
Sbjct: 126 PKDFNFINPNFPLWQTHFLNQSPDQ---NPLLLSSSASHHHQQQRLREIDLKLVVDVLMR 182
Query: 239 K--KKKNTVIVGDTVSLTEGLVSEIMKRFERGEVPD--EMKTTHFVKFHGLSSVSLKYMK 294
K KKKN VIVGD++S TEG VSE+M + ERGE+ E+K THFVKFH S ++ K+M+
Sbjct: 183 KKTKKKNPVIVGDSISFTEGFVSELMAKLERGEIDQTGELKQTHFVKFH-FSPMASKFMR 241
Query: 295 KEEVEMNVIRVLKRKVSDYVALGVGAIFYVGDLKW----IVDDNDGSLNE----KEVVDY 346
+E+VE+N I+ L++KV G AI + GDLKW I ++N G +NE +D+
Sbjct: 242 REDVELN-IKELRKKVLSLTTSGKNAIIFTGDLKWTVKEITNNNSGGINEISSSYSPLDH 300
Query: 347 VVEEIGKLFGE-------EGNKNGKIWLVATASYQSYMRCQMRVPTFENQWCLQAVPVP- 398
+VEEIGKL E + K K+W++ TAS+Q+YMRCQMR P+ E W L V VP
Sbjct: 301 LVEEIGKLITECNDDGDDDDCKTRKVWVMGTASFQTYMRCQMRQPSLETLWALHPVSVPS 360
Query: 399 SGGLGLSLHSSSVHDSKMSISQNPSPMLESKFFSNKEE--HEKLNCCEECVSNYEKEAQL 456
S LGLSLH++S H+++ + N + L + +EE L+CC ECV+++++EA+
Sbjct: 361 SANLGLSLHATSGHEARNMSTVNATKSLSGYDKAEEEETISHVLSCCPECVTSFDREAKS 420
Query: 457 FKPDQKNLLPSWLQSHSTE-ARQKDELTQLNKKWNRLCQCLHQNKQPQNHWSNNHSSNAK 515
K +Q LLPSWLQSH + + QKDEL L +KWNR C+ LH + N
Sbjct: 421 LKANQDKLLPSWLQSHDADSSSQKDELMGLKRKWNRFCETLHNQTGQLSMMGN------- 473
Query: 516 IYPYNSSYPYWPNQGSSILPDTSSSISFADS-ATKP-AYSSNLIPRFRRGQQSCTIEFNF 573
YPY Y GSS ++S S S DS KP ++N I +FRR Q SCTIEF+
Sbjct: 474 -YPYGLPY------GSS--HESSKSTSLIDSLGLKPNQRATNSIAKFRR-QNSCTIEFDL 523
Query: 574 NDEKAQKNQVTATLELDSLKGMEGTKEVKTTLALGNSTF---SVSDQKRMENLTLQRDHI 630
+ +K + E D KG E TL LG S F SV+D + L+ +
Sbjct: 524 GGNEHEKGESINEAEDD--KGNE-----TVTLDLGRSLFRSDSVTDTR------LKLSAL 570
Query: 631 YKVLQENIPWHCETVSSIAEALVDSKSSKECATWLFLQGNDSVGKKRLALAIAESVFGSV 690
K L+E+IP T+ IAE+L+D S K+ +W+ ++G D+ K+R+A ++ESVFGS
Sbjct: 571 VKALEESIPRQTVTMRLIAESLMDCVSKKK-DSWIIIEGRDTTAKRRVARTVSESVFGSF 629
Query: 691 EMFSHVDMMKREN-SETPFSEKVVGPLKNNEKFVVLVENADFGDTLIRKMLADEFEIAKF 749
E H+D+ K+ N S+ + + LKN EK V L+E+ D D+ K+LAD FE +
Sbjct: 630 ESLVHIDLKKKGNESKASPATLLAYELKNPEKVVFLIEDIDLADSRFLKLLADRFEDKRR 689
Query: 750 GTLG----QKIFILSNGGSMVSEDQKKDSVMKLVLKISETEKKPTFELSPSSSSSSKSPC 805
G Q IFIL+ S + +DSV+++ L+I +++SP
Sbjct: 690 IKTGIDHRQAIFILTKEDS--RNVRNRDSVLQIGLEI-----------------TAQSP- 729
Query: 806 LGNKRSAELDLFSKIKIPRIEENEGNKKREFSFSRQSSFNNT-LDLNMKADEEDNEDYDE 864
G KR E DL IE KK SRQSSFN++ LDLN+KA++E+ E
Sbjct: 730 -GKKRKPESDL-------SIENGFWMKKE--VCSRQSSFNSSYLDLNIKAEDEE----VE 775
Query: 865 GENSPISSDLTRETLGEHLISNESLDSIENLFEFNQS--PAKNKEMTQMFMSRVKESFEE 922
GE SPISSDLT E E S+ L+ I+N F N+S P K M + EE
Sbjct: 776 GEISPISSDLTGEEETEFSSSSNFLNRIQNRFVLNRSCEPGIEKGMITAAFREIFPEREE 835
Query: 923 VLGNVKFSVQDKVIEEIGVGCGSFTNNMFEKWLKGIFQTSLERV-NGGDKNGIVYTLCWG 981
G V+FSV+DK++EE+ N FE+WLK +FQT L V GG K+ V + +G
Sbjct: 836 G-GGVRFSVEDKLVEEL----YGIQNGAFERWLKEVFQTGLLTVKKGGKKDTGVIRMVFG 890
Query: 982 GKEDRK----WDSGFMGSCLPKNIQIVNY 1006
G D K G+M + LP +Q+ +
Sbjct: 891 GIVDNKGYGGGVGGYMATFLPNKVQVSKF 919
>ref|XP_471683.1| OSJNBa0035B13.5 [Oryza sativa (japonica cultivar-group)]
gi|21741941|emb|CAD40432.1| OSJNBa0035B13.5 [Oryza sativa
(japonica cultivar-group)]
Length = 1064
Score = 421 bits (1082), Expect = e-116
Identities = 365/1135 (32%), Positives = 536/1135 (47%), Gaps = 204/1135 (17%)
Query: 1 MRSGAYTLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSL------------P 48
MR+GAYT+ ++LTAEAA++LK +LG+ARRRGHAQ+TPLHVA LLS P
Sbjct: 1 MRAGAYTIHQSLTAEAAAVLKLALGIARRRGHAQVTPLHVAFALLSPACSPPQQQPAPPP 60
Query: 49 SSSLRNACLKSQQQNN-----HPLQCRALELCFNVALNRLPTTT---------------- 87
L+ ACL+S HPLQCRALELCFNVALNRLPT+
Sbjct: 61 YGLLKRACLRSHPSAAAAVAAHPLQCRALELCFNVALNRLPTSAPHSPPPSSSAPSGAVA 120
Query: 88 ---TSPLLQPQHVPSLSNALIAALKRAQAHQRRGCIEQNQQ----------------QQQ 128
S L+QP P+LSNAL+AALKRAQA+QRRGC+E QQ QQQ
Sbjct: 121 PPFASSLIQPN--PTLSNALVAALKRAQANQRRGCVELQQQPPPPPPPPPPPVAATAQQQ 178
Query: 129 QPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSSTLINSSSVFHSSPSP 188
QPLL++KVELDQLI+SILDDPSVSRVMREAGFSS +VK+NLE S L+ S+S S P P
Sbjct: 179 QPLLAIKVELDQLIISILDDPSVSRVMREAGFSSSTVKSNLEGESALMMSTS--SSPPPP 236
Query: 189 LSHNHFL--SSYGYGSVLFSSQKKEQVVYHPFLKSS----ESNKEDINLVFDVLLRK--K 240
HF S G G + PFL S S KED+ V +V++RK +
Sbjct: 237 AIPPHFFLDPSIGVGGNGGGGGGGFMLWPAPFLSSPGMAVPSCKEDVRAVLEVMVRKQGR 296
Query: 241 KKNTVIVGDTVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFHGLSSVSLKYMKKEEVEM 300
+ N V+VGD+VS+ E + E+++R E G+VPDE+ H +K LS V ++ M + +V+
Sbjct: 297 RTNPVVVGDSVSMAEAVAGELLRRLEGGDVPDELAGAHLLKLQ-LSYVHVRLMSRADVDA 355
Query: 301 NVIRVLKRKVSDYVALGVGAIFYVGDLKWIVD-----------DNDGSLNEKEVVDYVVE 349
+ R+ D V G G + YVGDL+W +D D+ + + V+++V
Sbjct: 356 KAAEL--RRSVDAVKRG-GLVVYVGDLRWALDEDHHHHHHPGADHHNTASSYSPVEHMVA 412
Query: 350 EIGKLFGE---EGNKNGKIWLVATASYQSYMRCQMRVPTFENQWCLQAVPVPSG-GLGLS 405
E+G+L G+ G++WLVATASYQ+YMRC+ R P+ E+ W LQAV VP+G G GL+
Sbjct: 413 ELGRLLGDLRASAPPRGRVWLVATASYQTYMRCRRRRPSLESAWALQAVVVPTGAGTGLA 472
Query: 406 LHS-----------SSVHDSKMSIS-QNPSPMLESKFFS--------NKEEHEKLNCCEE 445
L++ V + ++ + Q L S F + N+ + + L C E
Sbjct: 473 LNNLHAVATTTSNGEPVQQAMVATNHQQQQQQLASPFVAMAAEPAARNELDDKLLVLCTE 532
Query: 446 CVSNYEKEAQLFK---------PDQKNLLPSWLQSHSTEARQKDELTQLNKKWNRLCQCL 496
C NYE+EA K P LP WL E +++ L +L +KW+RLC+ L
Sbjct: 533 CSHNYEREASAVKAEAAADEEGPRAAGNLPGWL---VPEPPKENYLIELKRKWSRLCRKL 589
Query: 497 HQNKQPQNHWSNNHSSNAKIYPYNSSYPYWPNQGSSILPDTSSSISFADSATKPAYSSNL 556
H + + A + N SS+LP S+S + KP+ + L
Sbjct: 590 HLCGGGDPCSGQSFGAGA-----------YGNGPSSLLPWWSASCLLPNGGGKPSIAGFL 638
Query: 557 IPRFRRGQQSCTIEFNFNDEKAQKNQVTATLELDSLKGMEGTKEVKTTLALGNSTF--SV 614
G ++ + A L SL+ E ++V T LALG+ S
Sbjct: 639 ------GMEAL---------RWSPPAAAALPSLSSLREPE-CQDVTTALALGSLPLSDSA 682
Query: 615 SDQKRMENLTLQRDHIYKVLQENIPWHCETVSSIAEALV----DSKSSKECATWLFLQGN 670
S + + L++N+PW V+ IA+A+ +K WL L+G+
Sbjct: 683 SSSGGGGGDGAAARELERRLRKNVPWQRAAVAEIADAVAAGARSGNGTKGAGVWLLLKGS 742
Query: 671 DSVGKKRLALAIAESVFGSVEMFSHVDM-MKRENSETPFSEKVV-----GPLKNNEKFVV 724
D +R+A IAE+ GS + V + F VV K V+
Sbjct: 743 DHAAVRRVAAVIAETHCGSADRVVVVSADPNKFGCADDFRSDVVARASMAAAAGGNKLVL 802
Query: 725 LVENADFGDTLIRKMLADEFEIAKFGTLGQKIFILSNGGSMVSEDQKKDSVMKLVLKISE 784
+V++ + + + L ++ G L K GG + D V+ K+++
Sbjct: 803 VVDDVERAPQHVVECLV---AASRSGALKDKF-----GGQEL--DLSGSVVVMTTSKLAD 852
Query: 785 TEKKPTFELSPSSSSSSKSPCLGNKRSAELDLFSKIKIPRIEENEGNKKREFSFSRQSSF 844
L +S +SP G+ K K P ++KR + +R+S+
Sbjct: 853 AAVSGVISL--RLYTSEQSPPSGD---------LKRKTPTSSPPTSDRKR--ARARRSAG 899
Query: 845 N-NTLDLNMKADEEDNEDYDEG----ENSPISSDLTRETLGE---------HLISNESLD 890
N ++LDLN+ D++D D G ++ + SD+T E G+ H + ES+
Sbjct: 900 NGHSLDLNLNLFAHDDDDNDAGDVDDDDDGVPSDITHEGGGDDSGEHGHSHHRLLLESIA 959
Query: 891 SIENLFEFNQSPAKNKEMTQMFMSRVKESFEEVL--GNVKFSVQDKVIEEIGVGCGSFTN 948
+ + + A + V+E L G + V + + G F +
Sbjct: 960 TRVVTLDGDHHGA---------AAAVRERLSGRLDGGGRELRVDGEAAAALAAASGHFVD 1010
Query: 949 NMFEKWLKGIFQTSLERVNGGDKNGIVYTLCWGGKEDRKWDSGFMGSCLPKNIQI 1003
+ E+W+ +F+ + V G K ++ GG GFMGS LP + +
Sbjct: 1011 EVMERWVAEVFEPAAATVKNGGKAVVLGVGPSGGGAHE--SVGFMGSVLPSRVHV 1063
>gb|AAU05506.1| At3g52490 [Arabidopsis thaliana]
Length = 837
Score = 362 bits (928), Expect = 5e-98
Identities = 268/849 (31%), Positives = 422/849 (49%), Gaps = 112/849 (13%)
Query: 1 MRSGAYTLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQ 60
MR+G T+++ LTA+AA+++K ++GLARRRGHAQ+TPLHVA+T+LS P+ LR ACL
Sbjct: 1 MRAGGCTVEQALTADAANVVKQAMGLARRRGHAQVTPLHVASTMLSAPTGLLRTACL--- 57
Query: 61 QQNNHPLQCRALELCFNVALNRLPTTTTSPLL--QPQHVPSLSNALIAALKRAQAHQRRG 118
Q + HPLQCRALELCFNVALNRLPT+T SP+L PS+SNAL AA KRAQAHQRRG
Sbjct: 58 QSHTHPLQCRALELCFNVALNRLPTSTGSPMLGVPTSPFPSISNALGAAFKRAQAHQRRG 117
Query: 119 CIEQNQQQQQQPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSSTLINS 178
IE QQQP+L+VK+E++QLI+SILDDPSVSRVMREAGFSSP VK +E + +L
Sbjct: 118 SIE----SQQQPILAVKIEVEQLIISILDDPSVSRVMREAGFSSPQVKTKVEQAVSLEIC 173
Query: 179 SSVFHSSPSPLSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKEDINLVFDVLLR 238
S SS+ KE + P ED+ V + L+
Sbjct: 174 S----------------------KTTSSSKPKEGKLLTPV------RNEDVMNVINNLVD 205
Query: 239 KKKKNTVIVGDTVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKF----HGLSSVSLKYMK 294
KK++N VIVG+ ++ +G+V +M++ ++ +VP+ +K F+ G S + K
Sbjct: 206 KKRRNFVIVGECLATIDGVVKTVMEKVDKKDVPEVLKDVKFITLSFSSFGQPSRADVERK 265
Query: 295 KEEVEMNVIRVL-------KRKVSDYVAL-----GVGAIFYVGDLKWIVDD--NDGSLNE 340
EE+E +V R L +RK+ + L G G I +GDL W V+ SL
Sbjct: 266 LEELEADVERKLEELEADVERKLEELETLVKSCVGKGVILNLGDLNWFVESRTRGSSLYN 325
Query: 341 KE----VVDYVVEEIGKL-FGEEGNKNGKIWLVATASYQSYMRCQMRVPTFENQWCLQAV 395
VV++++ EIGKL G +G+ WL+ A+ Q+Y+RC+ P+ E+ WCL +
Sbjct: 326 NNDSYCVVEHMIMEIGKLACGLVMGDHGRFWLMGLATSQTYVRCKSGQPSLESLWCLTTL 385
Query: 396 PVPSGGLGLSLHSSSVHDSKMSISQNPSPMLESKFFSNKEEHEKLNCCEECVSNYEKEAQ 455
+P+ L L S + ++ S+N S L+ + ++L+ CEEC +E EA+
Sbjct: 386 TIPATSNSLRLSLVSESELEVKKSENVSLQLQ-------QSSDQLSFCEECSVKFESEAR 438
Query: 456 LFKPDQKNL----LPSWLQSHSTEAR----QKDELTQLNKKWNRLCQCLHQNKQPQNHWS 507
K N+ LP+WLQ + E + D + +L KWN +C +H+ +
Sbjct: 439 FLKSSNSNVTTVALPAWLQQYKKENQNSHTDSDSIKELVVKWNSICDSIHKRPSLKTLTL 498
Query: 508 NNHSSNAKIYPYNSSYPYWPNQGSSILPDTSSSISFADSATKPAYSSNLIPRFRRGQQSC 567
++ +S+ S P S+ + P +N ++
Sbjct: 499 SSPTSSF---------------SGSTQPSISTLHHLQTNGDWPVIETNTHRHHSVVHETS 543
Query: 568 TIEFNFNDEKAQKNQVTATLELDSLKGMEGTKEVKTTLALGNSTFSVSDQKRMENLTLQR 627
+ + +++ +S E + L +S F + ENL
Sbjct: 544 HLRLFIPEHDSEQKTELVCSNPNSTMNSEASSSDAMELEHASSRFK---EMNAENLAT-- 598
Query: 628 DHIYKVLQENIPWHCETVSSIAEALVDSKS-----------SKECATWLFLQGNDSVGKK 676
+ L+ +PW + V +A+ ++ +S K+ TW+F QG D K+
Sbjct: 599 --LCAALESKVPWQKDLVPELAKTVLKCRSGSSTRKINGNEDKKEDTWMFFQGLDVDAKE 656
Query: 677 RLALAIAESVFGSVEMFSHVDMMKRENSETPFSEKVVGPLKNNEKFVVLVENADFGDTL- 735
++A +A+ VFGS + F + + ++ + +E + +E+ + +E +L
Sbjct: 657 KIARELAKLVFGSQDSFVSICLSSFSSTRSDSAEDLRNKRLRDEQSLSYIERFSEAVSLD 716
Query: 736 -IRKMLADEFEIAKFGTLGQKIFILSNGGSMVSEDQKKDSVMKLVLKISETEKKPTFELS 794
R +L ++ E A + L Q F + V +++ +K + I E+ + +
Sbjct: 717 PNRVILVEDIEQADY--LSQVGFKRAVERGRVCNSSGEEASLKDAIVILSCERFRSRSRA 774
Query: 795 PSSSSSSKS 803
S S+ KS
Sbjct: 775 CSPPSNQKS 783
>emb|CAB43425.1| putative protein [Arabidopsis thaliana]
gi|15231233|ref|NP_190817.1| heat shock protein-related
[Arabidopsis thaliana] gi|44917467|gb|AAS49058.1|
At3g52490 [Arabidopsis thaliana] gi|7486001|pir||T08450
hypothetical protein F22O6.130 - Arabidopsis thaliana
Length = 815
Score = 361 bits (926), Expect = 8e-98
Identities = 266/833 (31%), Positives = 417/833 (49%), Gaps = 102/833 (12%)
Query: 1 MRSGAYTLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQ 60
MR+G T+++ LTA+AA+++K ++GLARRRGHAQ+TPLHVA+T+LS P+ LR ACL
Sbjct: 1 MRAGGCTVEQALTADAANVVKQAMGLARRRGHAQVTPLHVASTMLSAPTGLLRTACL--- 57
Query: 61 QQNNHPLQCRALELCFNVALNRLPTTTTSPLL--QPQHVPSLSNALIAALKRAQAHQRRG 118
Q + HPLQCRALELCFNVALNRLPT+T SP+L PS+SNAL AA KRAQAHQRRG
Sbjct: 58 QSHTHPLQCRALELCFNVALNRLPTSTGSPMLGVPTSPFPSISNALGAAFKRAQAHQRRG 117
Query: 119 CIEQNQQQQQQPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSSTLINS 178
IE QQQP+L+VK+E++QLI+SILDDPSVSRVMREAGFSSP VK +E + +L
Sbjct: 118 SIE----SQQQPILAVKIEVEQLIISILDDPSVSRVMREAGFSSPQVKTKVEQAVSLEIC 173
Query: 179 SSVFHSSPSPLSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKEDINLVFDVLLR 238
S SS+ KE + P ED+ V + L+
Sbjct: 174 S----------------------KTTSSSKPKEGKLLTPV------RNEDVMNVINNLVD 205
Query: 239 KKKKNTVIVGDTVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFHGLSSVSLKYMKKEEV 298
KK++N VIVG+ ++ +G+V +M++ ++ +VP+ +K VKF LS S + +V
Sbjct: 206 KKRRNFVIVGECLATIDGVVKTVMEKVDKKDVPEVLKD---VKFITLSFSSFGQPSRADV 262
Query: 299 EMNVIRVLKRKVSDYVALGVGAIFYVGDLKWIVDD--NDGSLNEKE----VVDYVVEEIG 352
E + L+ V V G G I +GDL W V+ SL VV++++ EIG
Sbjct: 263 ERK-LEELETLVKSCV--GKGVILNLGDLNWFVESRTRGSSLYNNNDSYCVVEHMIMEIG 319
Query: 353 KL-FGEEGNKNGKIWLVATASYQSYMRCQMRVPTFENQWCLQAVPVPSGGLGLSLHSSSV 411
KL G +G+ WL+ A+ Q+Y+RC+ P+ E+ WCL + +P+ L L S
Sbjct: 320 KLACGLVMGDHGRFWLMGLATSQTYVRCKSGQPSLESLWCLTTLTIPATSNSLRLSLVSE 379
Query: 412 HDSKMSISQNPSPMLESKFFSNKEEHEKLNCCEECVSNYEKEAQLFKPDQKNL----LPS 467
+ ++ S+N S L+ + ++L+ CEEC +E EA+ K N+ LP+
Sbjct: 380 SELEVKKSENVSLQLQ-------QSSDQLSFCEECSVKFESEARFLKSSNSNVTTVALPA 432
Query: 468 WLQSHSTEAR----QKDELTQLNKKWNRLCQCLHQNKQPQNHWSNNHSSNAKIYPYNSSY 523
WLQ + E + D + +L KWN +C +H+ + ++ +S+
Sbjct: 433 WLQQYKKENQNSHTDSDSIKELVVKWNSICDSIHKRPSLKTLTLSSPTSSF--------- 483
Query: 524 PYWPNQGSSILPDTSSSISFADSATKPAYSSNLIPRFRRGQQSCTIEFNFNDEKAQKNQV 583
S P S+ + P +N ++ + + +++
Sbjct: 484 ------SGSTQPSISTLHHLQTNGDWPVIETNTHRHHSVVHETSHLRLFIPEHDSEQKTE 537
Query: 584 TATLELDSLKGMEGTKEVKTTLALGNSTFSVSDQKRMENLTLQRDHIYKVLQENIPWHCE 643
+S E + L +S F + ENL + L+ +PW +
Sbjct: 538 LVCSNPNSTMNSEASSSDAMELEHASSRFK---EMNAENLAT----LCAALESKVPWQKD 590
Query: 644 TVSSIAEALVDSKS-----------SKECATWLFLQGNDSVGKKRLALAIAESVFGSVEM 692
V +A+ ++ +S K+ TW+F QG D K+++A +A+ VFGS +
Sbjct: 591 LVPELAKTVLKCRSGSSTRKINGNEDKKEDTWMFFQGLDVDAKEKIARELAKLVFGSQDS 650
Query: 693 FSHVDMMKRENSETPFSEKVVGPLKNNEKFVVLVENADFGDTL--IRKMLADEFEIAKFG 750
F + + ++ + +E + +E+ + +E +L R +L ++ E A +
Sbjct: 651 FVSICLSSFSSTRSDSAEDLRNKRLRDEQSLSYIERFSEAVSLDPNRVILVEDIEQADY- 709
Query: 751 TLGQKIFILSNGGSMVSEDQKKDSVMKLVLKISETEKKPTFELSPSSSSSSKS 803
L Q F + V +++ +K + I E+ + + S S+ KS
Sbjct: 710 -LSQVGFKRAVERGRVCNSSGEEASLKDAIVILSCERFRSRSRACSPPSNQKS 761
>ref|XP_465522.1| heat shock protein-related-like [Oryza sativa (japonica
cultivar-group)] gi|47496876|dbj|BAD19840.1| heat shock
protein-related-like [Oryza sativa (japonica
cultivar-group)]
Length = 778
Score = 347 bits (891), Expect = 9e-94
Identities = 246/622 (39%), Positives = 330/622 (52%), Gaps = 122/622 (19%)
Query: 1 MRSGAYTLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSS--------- 51
MR+G YT+ + L+A+AA++LK +L LARRRGHAQ+TPLHVA TLL SSS
Sbjct: 1 MRAGGYTVHQALSADAAAVLKLALALARRRGHAQLTPLHVAFTLLRSSSSSSSSSSPSDP 60
Query: 52 -----------------LRNACLKSQQQ----------NNHPLQCRALELCFNVALNRLP 84
LR AC+++ +HPL+CRALELCFNVALNRLP
Sbjct: 61 PPFACSGGEPSCCAHGLLRRACVRAHPAVAACAPAAAAASHPLRCRALELCFNVALNRLP 120
Query: 85 TTTT-----------SPLLQPQHVPSLSNALIAALKRAQAHQRRGCIEQNQ--------Q 125
T S L+ P P+LSNAL+AALKRAQA+QRRGCIE Q
Sbjct: 121 ATNAMADCGRACSPASSLVPPD--PTLSNALVAALKRAQANQRRGCIELQSLQPPQHALQ 178
Query: 126 QQQQPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSSTLINS---SSVF 182
QQQPLL++KVELDQLI+SILDDPSVSRVMREAGFSS +VK+ LE ++ S V
Sbjct: 179 PQQQPLLAIKVELDQLIISILDDPSVSRVMREAGFSSAAVKSTLEEGGAMLPSLGGHHVC 238
Query: 183 HSSPSPLSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKEDINLVFDVLLRKK-- 240
+SS SP H + G + Q ++ P SS +ED+ + +V++RK+
Sbjct: 239 YSSSSPEPHIDLDAHAASGG---GAPWPAQFLHRPDTGSS-CKEEDVRAILEVMVRKQWA 294
Query: 241 KKNTVIVGDTVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFHGLSSVSLKYMKKEEVEM 300
+ N V+VGD+VS+ E V+E+M+R E G+VP E++ H ++ H LS V L+ M + +V+
Sbjct: 295 RPNPVVVGDSVSVAEASVAELMRRLETGDVPGELRGAHVLRLH-LSRVHLRLMTRADVDA 353
Query: 301 NVIRVLKRKVSDYV--ALGVGAIFYVGDLKWIVDDND----GSLNEKEV-VDYVVEEIGK 353
V L+R + V A G + YVGD++W VDD+D +L E D++V E+ +
Sbjct: 354 QVAE-LRRTANSIVVDAKAAGLVIYVGDVRWAVDDDDHHHHHALAEYSAPEDHMVAELAR 412
Query: 354 LFGE-EGNKNGKIWLVATASYQSYMRCQM-----RVPTFENQWCLQAVPVPSG-----GL 402
L E G+ WLVA ASYQ+Y+RCQ R P+ E W LQAV VP+G G
Sbjct: 413 LMSELRAASRGRAWLVAAASYQTYVRCQQRRRRRRAPSLEATWSLQAVVVPAGAGADAGT 472
Query: 403 GLSLHSSSVHDSKMSISQNPSPMLESKFFSNKEEHEKLN------------CCEECVSNY 450
GLSL + + PS + E + E L+ C EC Y
Sbjct: 473 GLSL------GRRAPPAPPPSRVAEDDQIAKLGEIPTLDLALGGDDGGVPALCAECADGY 526
Query: 451 EKEAQLF--KPDQKNL----LPSWLQSHSTEARQKDELTQLNKKWNRLCQCLHQNKQPQN 504
EKEA K D L P W ++ + K EL +L +KW LCQ +H
Sbjct: 527 EKEASQVRAKADGTTLALTYFPGWPHANEPQTSHKAELMELRRKWGILCQRVH------- 579
Query: 505 HWSNNHSSNAKIYPYNSSYPYW 526
S +H+ A + S P+W
Sbjct: 580 --SRSHNDQASV---PSPMPWW 596
>dbj|BAB09589.1| 101 kDa heat shock protein; HSP101-like protein [Arabidopsis
thaliana] gi|15242850|ref|NP_200579.1| heat shock
protein-related [Arabidopsis thaliana]
Length = 990
Score = 334 bits (856), Expect = 1e-89
Identities = 262/810 (32%), Positives = 413/810 (50%), Gaps = 97/810 (11%)
Query: 1 MRSGAYTLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQ 60
MR+G T+Q+TLT EAA++L S+ A RR H Q TPLHVAATLL+ P+ LR AC++S
Sbjct: 1 MRAGLSTIQQTLTPEAATVLNQSIAEAARRNHGQTTPLHVAATLLASPAGFLRRACIRSH 60
Query: 61 QQNNHPLQCRALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQRRGCI 120
++HPLQCRALELCF+VAL RLPT TT+ P + P +SNAL+AALKRAQAHQRRGC
Sbjct: 61 PNSSHPLQCRALELCFSVALERLPTATTT----PGNDPPISNALMAALKRAQAHQRRGCP 116
Query: 121 EQNQQQQQQPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSSTLINSSS 180
E QQQQPLL+VKVEL+QLI+SILDDPSVSRVMREA FSSP+VK +E S +N+S
Sbjct: 117 E----QQQQPLLAVKVELEQLIISILDDPSVSRVMREASFSSPAVKATIEQS---LNNSV 169
Query: 181 VFHSSPSPLSHNHFLSSYGYGSVLFSSQKKEQVVYH-PFLKSSESNKEDINLVFDVLLRK 239
PS S G G + +S ++ + ++S S +D+ V D+L R
Sbjct: 170 TPTPIPSVSSVGLNFRPGGGGPMTRNSYLNPRLQQNASSVQSGVSKNDDVERVMDILGRA 229
Query: 240 KKKNTVIVGDTVSLTEGLVSEIMKRFERGEVPD-EMKTTHFVKFHGLSSVSLKYMKKEEV 298
KKKN V+VGD S ++ EI+K+ E GEV + +K + V +SS +K
Sbjct: 230 KKKNPVLVGD--SEPGRVIREILKKIEVGEVGNLAVKNSKVVSLEEISSDKALRIK---- 283
Query: 299 EMNVIRVLKRKVSDYVALGVGAIFYVGDLKWIVDDNDGSLNEKEVVDYVVEEIGKLFGEE 358
E++ + + K SD + G G I +GDLKW+V+ + + V EIG+ E
Sbjct: 284 ELDGLLQTRLKNSDPIG-GGGVILDLGDLKWLVEQPSST----QPPATVAVEIGRTAVVE 338
Query: 359 GNK-----NGKIWLVATASYQSYMRCQMRVPTFENQWCLQAVPVPSGGLGLSLHSSSVHD 413
+ G++W + TA+ ++Y+RCQ+ P+ E W LQAV V + +S V
Sbjct: 339 LRRLLEKFEGRLWFIGTATCETYLRCQVYHPSVETDWDLQAVSVAA-----KAPASGVFP 393
Query: 414 SKMSISQNPSPMLESKFFSNKEEHEKLNCCEECVSNYEKEA----QLFKPD------QKN 463
+ ++ +P L+S +N+ L CC +C+ +YE+E + P+ Q
Sbjct: 394 RLANNLESFTP-LKSFVPANR----TLKCCPQCLQSYERELAEIDSVSSPEVKSEVAQPK 448
Query: 464 LLPSW-LQSHSTEARQKDELTQLNKKWNRLCQCLHQNKQPQNHWSNNHSSNAKIYPYN-- 520
LP W L++ + + ++ ++ KKWN C LH + H+ N +I P
Sbjct: 449 QLPQWLLKAKPVDRLPQAKIEEVQKKWNDACVRLH---------PSFHNKNERIVPIPVP 499
Query: 521 ---SSYPYWPNQ--GSSILPDTSSSISFADSA-TKPAYSSNLIPRFRRGQQSCTIEFNFN 574
++ PY PN + P + + KP ++ ++ +
Sbjct: 500 ITLTTSPYSPNMLLRQPLQPKLQPNRELRERVHLKPMSPLVAEQAKKKSPPGSPVQTDLV 559
Query: 575 DEKAQKNQVTATLELDSLKGMEGTKEVKTTLALGNSTFSVSDQKRMENLTLQRDHIYKVL 634
+A+ ++ +++ G ++ V+ N+ SV ++ + N +L D K+L
Sbjct: 560 LGRAEDSEKAGDVQVRDFLGCISSESVQ-----NNNNISVLQKENLGN-SLDIDLFKKLL 613
Query: 635 Q---ENIPWHCETVSSIAEALVDSKSS--------KECATWLFLQGNDSVGKKRLALAIA 683
+ E + W + +++A + K + WL G D VGK+++ A++
Sbjct: 614 KGMTEKVWWQNDAAAAVAATVSQCKLGNGKRRGVLSKGDVWLLFSGPDRVGKRKMVSALS 673
Query: 684 ESVFGS----VEMFSHVDMMKRENS--ETPFSEKVVGPLKNNEKFVVLVENADFGDTLIR 737
V+G+ +++ S D +S +K+ +K + V+L+E+ D D L+R
Sbjct: 674 SLVYGTNPIMIQLGSRQDAGDGNSSFRGKTALDKIAETVKRSPFSVILLEDIDEADMLVR 733
Query: 738 KMLADEFEIAKFG-------TLGQKIFILS 760
+ + + +LG IF+++
Sbjct: 734 GSIKQAMDRGRIRDSHGREISLGNVIFVMT 763
>emb|CAB81028.1| putative protein [Arabidopsis thaliana]
gi|15234709|ref|NP_194764.1| heat shock protein-related
[Arabidopsis thaliana] gi|25407627|pir||H85354
hypothetical protein AT4g30350 [imported] - Arabidopsis
thaliana
Length = 924
Score = 328 bits (841), Expect = 6e-88
Identities = 308/1004 (30%), Positives = 470/1004 (46%), Gaps = 183/1004 (18%)
Query: 1 MRSGAYTLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQ 60
MR+ T+Q+TLT EAA++L S+ A RR H TPLHVAATLLS S LR AC+KS
Sbjct: 1 MRADLITIQQTLTPEAATVLNQSIAEATRRNHGHTTPLHVAATLLSSSSGYLRQACIKSH 60
Query: 61 QQNNHPLQCRALELCFNVALNRLPTTTT-------SPLLQPQHV--PSLSNALIAALKRA 111
++HPLQCRALELCF+VAL RLPTT+T S P P LSNAL AALKRA
Sbjct: 61 PNSSHPLQCRALELCFSVALERLPTTSTTTTTTSSSSSSSPSQTQEPLLSNALTAALKRA 120
Query: 112 QAHQRRGCIEQNQQQQQQPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLEN 171
QAHQRRGC E QQQQPLL+VKVEL+QLI+SILDDPSVSRVMREA FSSP+VK+ +E
Sbjct: 121 QAHQRRGCPE----QQQQPLLAVKVELEQLIISILDDPSVSRVMREASFSSPAVKSAIE- 175
Query: 172 SSTLINSSSVFHSSPSPLSHNHFLSSYGYGSV-------LFSSQKKEQVVYHPFLKSSES 224
S + NS S + SP N +GY SV L+ + + +Q
Sbjct: 176 QSLIGNSVSNSRQTGSPGIINPSAIGFGYRSVPAPVNRNLYLNPRLQQPGVGMQSGMMIQ 235
Query: 225 NKEDINLVFDVLLRKKKKNTVIVGDTVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFHG 284
++ V ++++R +K+N V+VGD S LV EI+++ E GE D
Sbjct: 236 RTDEAKRVIEIMIRTRKRNPVLVGD--SEPHILVKEILEKIENGEFSD----------GA 283
Query: 285 LSSVSLKYMKKEEVEMNVIRVLKRKVSDYVAL---GVGAIFYVGDLKWIVDD---NDGSL 338
L + + ++KE V R+ ++S V G G + +GDLKW+V+ N G+
Sbjct: 284 LRNFQVIRLEKELVSQLATRL--GEISGLVETRIGGGGVVLDLGDLKWLVEHPAANGGA- 340
Query: 339 NEKEVVDYVVEEIGKLFGEEGNKNGKIWLVATASYQSYMRCQMRVPTFENQWCLQAVPVP 398
V E+ KL G++ + TA+ ++Y+RCQ+ P+ EN W LQA+P+
Sbjct: 341 ---------VVEMRKLL---ERYKGRLCFIGTATCETYLRCQVYYPSMENDWDLQAIPIA 388
Query: 399 SGGLGLSLH---SSSVHDSKMSISQNPSPMLE-SKFFSNKEEHEKLNCCEECVSNYEKEA 454
+ ++ S+ +++ M +S N + S S + K++CC C+ +YE +
Sbjct: 389 AKSSLPAIFPRLGSNNNNNAMLLSNNIISIESISPTRSFQIPMSKMSCCSRCLQSYENDV 448
Query: 455 QLFKP----DQKNLLPSWLQSHST------EARQKDELTQLNKKWNRLCQCLHQNKQPQN 504
+ D +++LP WLQ+ + + ++ +L KKWN LC LH N+
Sbjct: 449 AKVEKDLTGDNRSVLPQWLQNAKANDDGDKKLTKDQQIVELQKKWNDLCLRLHPNQSVSE 508
Query: 505 HWSNNHSSNAKIYPYNSSYPYWPNQGSSILPDTSSSISFADSATKPAYSSNLIPRFRRGQ 564
+ + S KI N S I P S P + ++ R RG
Sbjct: 509 RIAPSTLSMMKI-----------NTRSDITPPGS-----------PVGTDLVLGRPNRGL 546
Query: 565 QSCTIEFNFNDEKAQKNQVTATLELDSLKGMEGTKEVKTTLALGNSTFSVSDQKRMENLT 624
S EK T+E + LG+S F + K+
Sbjct: 547 SS--------PEKK-------------------TREARFG-KLGDS-FDIDLFKK----- 572
Query: 625 LQRDHIYKVLQENIPWHCETVSSIAEALVDSK---SSKECATWLFLQGNDSVGKKRLALA 681
+ K L +++ W + SS+A A+ + K + WL G D GK ++A A
Sbjct: 573 -----LLKGLAKSVWWQHDAASSVAAAITECKHGNGKSKGDIWLMFTGPDRAGKSKMASA 627
Query: 682 IAESVFGSVEMFSHVDMMKRENSETPFS-----EKVVGPLKNNEKFVVLVENADFGDTLI 736
+++ V GS + + R + ++ ++ N V+++E+ D D L+
Sbjct: 628 LSDLVSGSQPITISLGSSSRMDDGLNIRGKTALDRFAEAVRRNPFAVIVLEDIDEADILL 687
Query: 737 RKMLADEFEIAK----FG---TLGQKIFILSNGGSMVSEDQKKDSVMKLVLKISETEKKP 789
R + E + +G +LG I IL+ S+ S K V I ET +
Sbjct: 688 RNNVKIAIERGRICDSYGREVSLGNVIIILTANSSLGS--------AKNVASIDETRLES 739
Query: 790 T----FELSPSSSSSSKSPCLGNKRSAELDLFSKIKIPRIEENEGNKKRE---FSFSRQS 842
+EL S +SSK+ KR L+S +N+ K+R+ F + +
Sbjct: 740 LVNKGWELRLSVCNSSKT----RKRKPNW-LYS--------DNDQTKQRKEICFDLNEAA 786
Query: 843 SFNNTLDLNMKADEEDNEDY---------DEGENSPISSDLTRETLGEHLISNESLDSIE 893
F+++ D+ ++ D+EDN + D P+ D + E L S +
Sbjct: 787 EFDSSSDVTVEHDQEDNGNLVHKLVGLVDDAILFRPVDFDSIKSKTAESLKKRFSNGLAD 846
Query: 894 NLFEFNQSPAKNKEMTQMFMSRV--KESFEEVLGNVKFSVQDKV 935
L + A + +++S++ +E EE +G+ SV+ +V
Sbjct: 847 GLTVEIEDDALERIAGAIWLSKISLEEWLEEAMGSSLNSVKSRV 890
>ref|XP_472429.1| OJ991214_12.16 [Oryza sativa (japonica cultivar-group)]
gi|39545713|emb|CAD40931.3| OSJNBa0033G16.3 [Oryza
sativa (japonica cultivar-group)]
gi|38344035|emb|CAE01527.2| OJ991214_12.16 [Oryza sativa
(japonica cultivar-group)]
Length = 877
Score = 290 bits (743), Expect = 1e-76
Identities = 259/831 (31%), Positives = 384/831 (46%), Gaps = 170/831 (20%)
Query: 1 MRSGAYTLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQ 60
MR+G T+Q+ LTAEAA+++K ++ LARRRG+AQ+TPLHVA+ +L+ P LR ACL+S
Sbjct: 1 MRAGGCTVQQALTAEAAAVVKQAVTLARRRGNAQVTPLHVASAMLAPPGGLLRAACLRS- 59
Query: 61 QQNNHPLQCRALELCFNVALNRLPTTT---TSPLLQPQ--------HVPSLSNALIAALK 109
++HPLQC+ALELCFNVALNRLP + +SPLL + PSLSNAL+AA K
Sbjct: 60 --HSHPLQCKALELCFNVALNRLPASAAVASSPLLGGHGHGHHGHYYPPSLSNALVAAFK 117
Query: 110 RAQAHQRRGCIEQNQQQQQQPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNL 169
RAQAHQRRG +E QQQP+L+VK+EL+QL++SILDDPSVSRVMREAGFSS VK N+
Sbjct: 118 RAQAHQRRGSVET----QQQPVLAVKIELEQLVVSILDDPSVSRVMREAGFSSTQVKANV 173
Query: 170 ENS-STLINSSSVFHSSPSPLSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKED 228
E + ST +S+ + +P+P ++ S + K ++ L + ED
Sbjct: 174 EQACSTTTATSAPPNQNPNPSCTGAAATATATTSPAHPPEIKAKLPLLDML----ARDED 229
Query: 229 INLVFDVLLRK---------KKKNTVIVGDTVSLTEGLVSEIMKRFERGEVP--DEMKTT 277
I V D L ++ V+V ++ + E + R RGE D ++
Sbjct: 230 IAAVLDCLAPAAAGGRGGGGSRRRVVVVAESTAAAEATARAAVDRVRRGEAKQHDALRGA 289
Query: 278 HFVKFHGLSSVSLKYMKKEEVEMNV--IRVLKRKVSDYVALGVGAIFYVGDLKWIVD--- 332
V L S + M +EE E + +R L + + G + V DLKW D
Sbjct: 290 QVV---SLRVSSFRDMPREEAERRLAELRCLVK------SRGARVLLVVEDLKWAADFWA 340
Query: 333 ---------DNDGSLNEKEVVDYVVEEIGKLFGEEGNKNGKIWLVATASYQSYMRCQMRV 383
+ G V++VV E+ L + +G IWLV +YQ+YM+C+
Sbjct: 341 AAHAGARRVGSGGGGGYYCSVEHVVTEVRAL----ASCDGGIWLVGFGTYQTYMKCRAGH 396
Query: 384 PTFENQWCLQAVPVPSGGLGLSLHSSSVHDSKMSISQN------------PSPMLESKFF 431
P+ E+ W LQ + VP+G L LSL + + +++Q+ P+P
Sbjct: 397 PSLESMWGLQTLAVPAGSLALSLTCAFDDSALGAVNQSMKASPHTTDGNRPAPSCGPLLG 456
Query: 432 SNKEEHEKLNC----CEECVSNYEKEAQLFKPD---QKNLLPSWLQ-SHSTEARQKDELT 483
+ H C C + +E + + P + LPSWLQ + ++
Sbjct: 457 GS---HLLSRCCGGDCSAATTTHEHDTKASLPRSFVSSSSLPSWLQHCRDQQLQESTHFA 513
Query: 484 QLNKKWNRLCQCLHQNKQPQNHWSNNHSSNAKIYPYNSSYPYWPNQ----GSSILPDTSS 539
L K W +C +++ H+S S + I Y + + Q S +L D
Sbjct: 514 DLGKTWGSICG--KPSQRMTLHFSAPVSPASSISSYEHGHGHQQQQHQPHHSWLLADLD- 570
Query: 540 SISFADSATKPAYSSNLIPRFRRGQQSCTIEFNFNDEKAQKNQVTATLELDSLKGMEGTK 599
A KP + +DEKA+ + D G+
Sbjct: 571 ----AKHPWKPKREDD------------------DDEKAKSHD-------DCSGASNGSV 601
Query: 600 EVKTTLALGNSTFSVSDQKRMENLTLQRDHIYKVLQENIPWHCETVSSIAEALVDSKS-- 657
EV+ S F + ENL L + L++ +PW E V +A A++ +S
Sbjct: 602 EVEC-----RSRFK---ELNAENLKL----LCAALEKEVPWQKEIVPEVASAVLQCRSGI 649
Query: 658 -----------SKECATWLFLQGNDSVGKKRLALAIAESVFGSVEMFSHVDM-------- 698
+KE TWLF G D GK+R+A +A VFGS + F V +
Sbjct: 650 AKRRDRSRSTEAKE-ETWLFFLGGDGHGKERVARELAGLVFGSRKSFLSVKLGASSSSPS 708
Query: 699 ----------MKRENSETPFS------EKVVGPLKNNEKFVVLVENADFGD 733
KR + T S E++ + N V+L+E+ + GD
Sbjct: 709 ASGSTEDHHRSKRPRTTTTSSASEAYLERLYDAVSENPHRVILIEDVEQGD 759
>gb|AAX92943.1| transposon protein, putative, mutator sub-class [Oryza sativa
(japonica cultivar-group)]
Length = 845
Score = 262 bits (670), Expect = 4e-68
Identities = 266/922 (28%), Positives = 405/922 (43%), Gaps = 175/922 (18%)
Query: 1 MRSGAYTLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQ 60
MR+G +Q+ L AEAA +++ ++ LARRRGHAQ+TPLHVA+ +LS + LR ACL+S
Sbjct: 1 MRAGGCAVQQALAAEAAGVVRQAVTLARRRGHAQVTPLHVASAMLSA-AGLLRAACLQS- 58
Query: 61 QQNNHPLQCRALELCFNVALNRLPTTTTSPLL-----QPQHV-------PSLSNALIAAL 108
++HPLQC+ALELCFNVALNRLPT + P H P+LSNAL+AA
Sbjct: 59 --HSHPLQCKALELCFNVALNRLPTAGPAAAAAIFHHHPHHPGGGGGHHPALSNALVAAF 116
Query: 109 KRAQAHQRRGCIEQNQQQQQ--QPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVK 166
KRAQAHQRRG +E Q P+++ KVEL+QLI+SILDDPSVSRVMREAGFSS VK
Sbjct: 117 KRAQAHQRRGSVEGQPPPQPPPSPVVASKVELEQLIISILDDPSVSRVMREAGFSSSQVK 176
Query: 167 NNLENSSTLINSSSVFHSSPSPLSHNHFLS-SYGYGSVLFSSQKKEQVVYHPFLKSSESN 225
N+E + +S+ H++ + H S + G+G ++KE SS +
Sbjct: 177 ANVEKAVV----ASLDHANAAGGGGGHAGSPNSGHG-----GRRKES-------SSSRAR 220
Query: 226 KEDINL-VFDVLLRKKKKNTVIV-GDTVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFH 283
+D + V D + K+ V+V G+ + E +V +M R + E+ + ++F
Sbjct: 221 VDDDAMRVLDCMASGTKRCVVVVGGEGAAAAEAVVKAVMDRVSKAELHHRHERLKNLQFV 280
Query: 284 GLSSVSLKYMKKEEVEMNVIRVLKRKVSDYVALGVGAIFYVGDLKWIVDDNDGSLNEKE- 342
LS S +EEVE L+ V A G G + + DL + D + N +
Sbjct: 281 PLSIASFHGAPREEVEAKA-SDLRALVRSGCAAGKGVVLVLEDLAYAADAWAAASNTRRR 339
Query: 343 -----------VVDYVVEEIGKLFGEEGNKNGKIWLVATASYQSYMRCQMR-VPTFENQW 390
+++ V E+ L G + W++ SYQ YM+C+ P E W
Sbjct: 340 AAAAAGGQSYCPMEHAVMEVSSLVSGGGGGGERFWVLGFGSYQVYMKCRAAGQPPLEAVW 399
Query: 391 CLQAVPVPSGGLGLSLHSSSVHDSKMSISQNPSPMLESKFFSNKEEHEKLNCCEECVSNY 450
L V VP GGL LSL S ++ + Q +P F +
Sbjct: 400 ELHPVVVPDGGLALSLTCS---EASQATHQAAAPTAGWPFVNGAG--------------- 441
Query: 451 EKEAQLFKPDQKNLLPSWLQ-----SHSTEARQKDELTQLNKKWNRLCQCLHQNKQPQNH 505
E A P +P WL+ H+T A L Q+ WN P +
Sbjct: 442 EAAATTASP----TIPPWLRRYQDPDHATPASCGTGL-QIQDLWN-----------PMRN 485
Query: 506 WSNNHSSNAKIYPYNSSYPYWPNQGSSILPDTSSSISFADSATKPAYSSNLIPRFRRGQQ 565
S H ++ ++S P SSI TS Y++N + + Q
Sbjct: 486 GSAPHHTSELTLSFSSPSP------SSISGYTS------------CYNNNNMMSSKPWQL 527
Query: 566 SCTIEFNFNDEKAQKNQVTATLE---LDSLKGMEGTKEVKTTLALGNSTFSVSDQKRMEN 622
+ + + Q+ + + + LD+ E + V +L G + + EN
Sbjct: 528 EARQPWPIHGHEGQRMAMASYHDHHPLDTNPSPE-SNSVSNSLDGGETRRPKFIELNAEN 586
Query: 623 LTLQRDHIYKVLQENIPWHCETVSSIAEALVDSKSSKE-------------CATWLFLQG 669
L + + L+ +P H V IA ++ +S + TWL QG
Sbjct: 587 LKI----LCNALESRVPQHSNIVPDIASTVLQCRSGMKKMKLRHKEIIKASSTTWLLFQG 642
Query: 670 NDSVGKKRLALAIAESVFGSVEMFSHVDMMKRENSETPFSEKVVGPLKNNEKFVVLVENA 729
D GKK +A +A+ VFGS FS + + +P+S+ G L
Sbjct: 643 RDVDGKKAMAQELAKLVFGSSTEFSSISF---DELTSPYSDSSSGEL------------- 686
Query: 730 DFGDTLIRKMLADEFEIAKFGTLGQKIFILSNGGSMVSEDQKKDSVMKLVLKISETEKKP 789
TL R+ AD E + Q++ +VS++ + V+ I + ++
Sbjct: 687 ----TLKRQRSADSNE----HSFAQRLC------EIVSKNPHQVIVIN---DIEQLDQDS 729
Query: 790 TFELSPSSSSSSKSPCLGNKRSAELDLFSKIKIPRIEENEGNKKREFSFSR--QSSFNNT 847
+ + ++ C G E+D I + EE ++ R S R Q NN
Sbjct: 730 EISIKKAIANGRMRGCTGE----EVDFEDAIIVLSYEEEFDSRSRASSSPRVKQRLMNNN 785
Query: 848 LDLNMKADEEDNEDYDEGENSP 869
D+E++ ++G+NSP
Sbjct: 786 -------DDEESSSTEKGDNSP 800
>ref|XP_481010.1| 101 kDa heat shock protein; HSP101-like protein [Oryza sativa
(japonica cultivar-group)] gi|40253563|dbj|BAD05509.1|
101 kDa heat shock protein; HSP101-like protein [Oryza
sativa (japonica cultivar-group)]
Length = 1041
Score = 256 bits (655), Expect = 2e-66
Identities = 290/1086 (26%), Positives = 451/1086 (40%), Gaps = 209/1086 (19%)
Query: 1 MRSGAYTLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQ 60
MR+ T+Q+TLT EAA+ L ++ A RR H Q TPLHVAA LL+ P+ LR AC ++
Sbjct: 1 MRADLSTIQQTLTPEAAAALARAMDEAGRRRHGQTTPLHVAAALLAAPAGLLRQACARAA 60
Query: 61 QQNN---------------HPLQCRALELCFNVALNRLPTTTTSPLLQ--PQHVPSLSNA 103
HPL CRALELCF+VAL+RLP + P +SNA
Sbjct: 61 SAAGVGGGGGAAAGAGAGAHPLHCRALELCFSVALDRLPAAAAAAAAAHGAGASPPVSNA 120
Query: 104 LIAALKRAQAHQRRGCIEQNQQQQQQPLLSVKVELDQLILSILDDPSVSRVMREAGFSSP 163
L+AALKRAQA QRRGC E QQPLL+VKVEL+QL+LSILDDPSVSRVMREA FSS
Sbjct: 121 LVAALKRAQAQQRRGCPEA----AQQPLLAVKVELEQLVLSILDDPSVSRVMREASFSSA 176
Query: 164 SVKNNLENS------------STLINSSSVFHSSPSPLSHNHFLSSYGYGSVLFSSQKKE 211
+VK+ +E S ST SPSPL ++Y + ++
Sbjct: 177 AVKSIIEQSLSAPSPCPSAAASTTTAGPGPLSPSPSPLPRAGAANAYLNPRLAAAAAVAS 236
Query: 212 QVVYHPFLKSSESNKEDINLVFDVLLRKKKKNTVIVGDTVSLTEGLVSEIMKRFERGEVP 271
+D V DV+L+ ++N V+VGD + ++ E ++R P
Sbjct: 237 G--------GGGGGGDDARKVIDVMLKPTRRNPVLVGDAGP--DAVLKEAIRRIPTAGFP 286
Query: 272 DEMKTTHFVKFHGLSSVSLKYMKKEEVEMNVIRVLKRKVSDYVALGVGAIFYVGDLKWIV 331
K L + K + I L V + G + +GDLKW+V
Sbjct: 287 ----ALAGAKVLPLEAELAKLAGDKAAMAARIGDLGAVVERLLGEHGGVVLDLGDLKWLV 342
Query: 332 DDNDGSLNEKEVVDYVVEEIGKLFGEEGNKNGKIWLVATASYQSYMRCQMRVPTFENQWC 391
D + +E V E+G+L G +W V TA+ +Y+RC++ P E +W
Sbjct: 343 DGPAAAASEGGKA--AVAEMGRLLRRFGRAG--VWAVCTAACTTYLRCKVYHPGMEAEWD 398
Query: 392 LQAVPVPSGGLGLSLHS---------SSVHDSKMSI---SQNPSPMLESKFF-----SNK 434
L AVP+ GG ++ + S + +S M + + P P+ + S++
Sbjct: 399 LHAVPIARGGAPIAAAAAGSALRPGGSGILNSSMGMLSPALRPMPVTPTALRWPPPGSDQ 458
Query: 435 EEHEKLNCCEECVSNYEKEAQLFKPDQ-----------KNLLPSWLQSHSTEARQKDE-- 481
K C C +YE+E + +Q K LP WLQ + + + K++
Sbjct: 459 SPAAKPAMCLLCKGSYERELAKLEAEQTDKPASRPEAAKPGLPHWLQLSNDQNKAKEQEL 518
Query: 482 -----LTQLNKKWNRLCQCLHQNKQPQNHWSNNHSSNAKIYPYNSSYPYWPNQGSSILPD 536
+L +KW C +H S+ P P
Sbjct: 519 KLKRSKDELERKWRETCARIH-----------------------SACPMAP--------- 546
Query: 537 TSSSISFADSATKPAYSSNLIPRFRRGQQSCTIEFNFNDEKAQKNQVTATLELDSLKGME 596
+ S+ A +P L RG T++ N + EK V TLEL +
Sbjct: 547 -ALSVPLATFTPRPPVEPKL--GVARGAAVPTLKMNPSWEKPS---VAPTLEL---RKSP 597
Query: 597 GTKEVKTTLAL-----GNSTFSVSDQKRM-ENLT-LQRDHI------------YKVLQEN 637
VKT L L G + ++QK E LT LQ+ I K L E
Sbjct: 598 PASPVKTDLVLCRLDPGTNPAVENEQKESCEGLTALQKAKIAGISDIESFKRLLKGLTEK 657
Query: 638 IPWHCETVSSIAEALVDSKSS----KECAT----WLFLQGNDSVGKKRLALAIAESVFGS 689
+ W + S+IA ++ +S + T WL G D GK+++ A++E + +
Sbjct: 658 VSWQSDAASAIAAVVIQCRSGSGKRRNVGTRGDMWLLFVGPDQAGKRKMVNALSELMANT 717
Query: 690 ---VEMFSHVDMMKRENSETPFS--------EKVVGPLKNNEKFVVLVENADFGDTLIRK 738
V F + R ++ P ++V ++ N V+++E D D ++
Sbjct: 718 RPVVVNFGGDSRLGRVGNDGPNMGFWGKTALDRVTEAVRQNPFSVIVLEGIDQVDVVVHG 777
Query: 739 MLADEFEIAKFG-------TLGQKIFILSNGGSMVSEDQKKDSVMKLVLKISETEKKPTF 791
+ E + +LG IF+L+ + V E+ K +V L+ +
Sbjct: 778 KIKRAMETGRLPDSRGREVSLGNVIFVLTT--NWVPEELKGSNVETLL-------RGEER 828
Query: 792 ELSPSSSSSSKSPCLGNKRSAELDLFSKIKIPRIEENEGNKKREFSFSRQSSFNNTLDLN 851
L +SSS +G+K+ +K + + + + SS +LDLN
Sbjct: 829 MLESTSSSWQLELSIGDKQ---------VKHRADWLCDDVRPAKLAKELSSSHGLSLDLN 879
Query: 852 MKADEEDNEDYDEGENSPISSDLTRETLGEH----------LISNESLDSIENLFEFNQS 901
+ D+ E S SSD++ E E ++ L+ +++ F
Sbjct: 880 LAVGA-----LDDTEGSHNSSDVSVEQEQEKGQLAVKRSTPAPGSDILELVDDAIVFR-- 932
Query: 902 PAKNKEMTQMFMSRVKESFEEVLG-NVKFSVQDKVIEEIGVGCGSFTNNMFEKWLKGIFQ 960
P + + FE V+G + F + + ++ + VG T+ E W + + +
Sbjct: 933 PVDFTPFRKTVTDCISAKFESVMGSSSSFRIDEDAVDWM-VGSVWLTDEKIEDWAEKVLK 991
Query: 961 TSLERV 966
S+ER+
Sbjct: 992 PSIERL 997
>dbj|BAD27916.1| heat shock protein-related-like [Oryza sativa (japonica
cultivar-group)] gi|50252658|dbj|BAD28827.1| heat shock
protein-related-like [Oryza sativa (japonica
cultivar-group)]
Length = 917
Score = 249 bits (636), Expect = 3e-64
Identities = 168/435 (38%), Positives = 240/435 (54%), Gaps = 59/435 (13%)
Query: 1 MRSGAYTLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSS-----LRNA 55
MR+G T+Q+ LTAEAA+++K ++ LARRRG+AQ+TPLHVA+ +L ++ LR A
Sbjct: 1 MRAGGCTVQQALTAEAAAVVKQAVSLARRRGNAQVTPLHVASAMLQAAGAAPAPGLLRAA 60
Query: 56 CLKSQQQNNHPLQCRALELCFNVALNRLPTTT-TSPLLQPQH------VPSLSNALIAAL 108
CL+S ++HPLQC+ALELCFNVALNRLP + SPLL H PSLSNAL+AA
Sbjct: 61 CLRS---HSHPLQCKALELCFNVALNRLPASAGASPLLGHGHGVGVYYPPSLSNALVAAF 117
Query: 109 KRAQAHQRRGCIEQNQQQQQQPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNN 168
KRAQAHQRRG +E QQQP+L+VK+EL+QL++SILDDPSVSRVMREAGFSS VK N
Sbjct: 118 KRAQAHQRRGSVET----QQQPVLAVKIELEQLVISILDDPSVSRVMREAGFSSTQVKAN 173
Query: 169 LENS-STLINSSSVFHSSPSPLSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKE 227
+E + S+ + +++ +P+P S + +++ S+ +QVV +E
Sbjct: 174 VEQAVSSSMEAATTKPQNPNPSSSSPPPAAHQEAK---PSRCIDQVVV---------REE 221
Query: 228 DINLVFDVLLRKKKKNTVIVGDTVSLTEGLVSEIMKRFERGEVPD----EMKTTHFVKFH 283
D+ + D L + KK ++V + + E + R RGE ++ T +F
Sbjct: 222 DVAAILDCLATRSKKRVMVVAECAAAAEAAARAAVDRIRRGEARQHAQAQVVTLAVSRFR 281
Query: 284 GLSSVSLKYMKKEEVEMNVIRVLKRKVSDYVALGVGAIFYVGDLKWIVDDNDGSLNEKE- 342
G +EE E R+ + + + G + V DL W + G
Sbjct: 282 G--------APREEAER---RLAELRCAVRGGGGRAVVLVVEDLAWAAEFWAGRRPPPSS 330
Query: 343 -----------VVDYVVEEIGKLFGEEGNKNGKIWLVATASYQSYMRCQMRVPTFENQWC 391
V++ V E+ L G G +WLV +YQ+ +RC+ P+ E W
Sbjct: 331 CGAGAGGYYYCAVEHAVAEVRALACGGGGGGGGVWLVGHGTYQTNIRCRTGHPSLETLWG 390
Query: 392 LQAVPVPSGGLGLSL 406
L + VP+G L LSL
Sbjct: 391 LHTLAVPAGSLALSL 405
Score = 38.9 bits (89), Expect = 0.88
Identities = 28/101 (27%), Positives = 45/101 (43%), Gaps = 17/101 (16%)
Query: 634 LQENIPWHCETVSSIAEALVDSKS----------SKECA-----TWLFLQGNDSVGKKRL 678
L++ +PW V IA ++ +S S C+ TW+ G D+ GK R+
Sbjct: 637 LEKEVPWQKVIVPEIASTVLRCRSGMAAPAMARRSSSCSSSKEHTWMLFLGGDADGKLRV 696
Query: 679 ALAIAESVFGSVEMFSHVDMMKRENSETPFSEKVVGPLKNN 719
A +A VFGS + F V + N+ P S P +++
Sbjct: 697 ARELASLVFGSSKSF--VSIGGAANASPPPSSSSSSPARSS 735
>ref|XP_468184.1| 101 kDa heat shock protein-like [Oryza sativa (japonica
cultivar-group)] gi|47497764|dbj|BAD19864.1| 101 kDa
heat shock protein-like [Oryza sativa (japonica
cultivar-group)]
Length = 965
Score = 223 bits (568), Expect = 3e-56
Identities = 247/985 (25%), Positives = 417/985 (42%), Gaps = 183/985 (18%)
Query: 12 LTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQQQNNHPLQCRA 71
LT +AA++L + A RR HA TPLH AA LLS P+ LR+AC+ + HPL+CRA
Sbjct: 23 LTPDAAAVLSRAAADASRRRHAHTTPLHAAAALLSGPAPLLRDACVAGLA-SPHPLRCRA 81
Query: 72 LELCFNVALNRLPTTTTSPLLQPQHV-PSLSNALIAALKRAQAHQRR---GCIEQNQQQQ 127
L+LCF VAL+RLPT+T Q H P LSNAL AALKRA AH RR G +E +
Sbjct: 82 LDLCFAVALDRLPTSTEH---QHHHAAPPLSNALAAALKRAYAHHRRIGSGVVEADDH-- 136
Query: 128 QQPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSSTLINSSSVFHSSPS 187
+V + L+L+ILDDPSV+RVMREA FSS +VK +++ S
Sbjct: 137 -------RVGVPHLVLAILDDPSVARVMREASFSSTAVK------------AAMLRSLSD 177
Query: 188 PLSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKEDINLVFDVLLRKKKKNTVIV 247
P + + V +++ + V H +E++N V +VL R KK+N V+V
Sbjct: 178 PAAPD--------SGVYVNARVLHRQVSH--------REEEVNKVVEVLKRGKKRNPVLV 221
Query: 248 GDTVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFHGLSSVSLKYMKKEEVEMNVIRVLK 307
GDTV + + +V E++ +R + + + +F L + + + E+ ++
Sbjct: 222 GDTVDV-DAVVQEVVTMIQRQRLGNARVISFQREFGDLVDLDRAELAAKIKELG--EAIR 278
Query: 308 RKVSDYVALGVGAIFYVGDLKWIVDDNDGSLNEKE-----VVDYV---VEEIGKLFGEEG 359
++ + G + +G+L+W+V++ + E+E V+D V E+ ++ + G
Sbjct: 279 SELLSPASRSAGVVVNLGNLQWLVEERCVAPGEQEKRRDVVLDTARAAVAEMARILRQSG 338
Query: 360 NKNGKIWLVATASYQSYMRCQMRVPTFENQWCLQAVPV-------PSGGLGLSLHSSSVH 412
+ ++W++ TA+ +Y++CQ+ P+ E++W LQAVP+ P LGLS SV+
Sbjct: 339 EREHRVWVIGTATCATYLKCQVYHPSLESEWDLQAVPITPRPPPPPPSSLGLS---PSVN 395
Query: 413 DSKMSISQNPSPMLESKFFSNKEEHEKLNCCEECVSNYEKEAQLFKPDQKNLLPSWLQSH 472
I + +L S ++ + + C C+ YE
Sbjct: 396 GVNRGILSSSVEVLSSAMTTSAMQSRSPSLCSACLDGYE--------------------- 434
Query: 473 STEARQKDELTQLNKKWNRLCQCLHQNKQPQNHWSNNHSSNAKIYPYNSSYPYWPNQGSS 532
R++ ++ + C LH +QP + W + ++ P++ + Q +
Sbjct: 435 ----RERADMAS-----SPGCGALHATEQPMSQWLQIGTPSSARPPFDRA------QDKA 479
Query: 533 ILPDTSSSISFADSATKPAYSSNLIPRFRRGQQSCTIEFNFNDEKAQKNQVTATLELDSL 592
D A ++ R E+N A + +
Sbjct: 480 READELRRRWLDRCAQLHSHGGGGCGGGRPSSMVTCSEWNGASVLANMQAIPVRPPPPAA 539
Query: 593 KGMEGTKEVKTTLALGNSTFSVSDQKRMENLTLQRDHIYKVLQENIPWHCETVSSIAEAL 652
V T LALG + + S + + K L E + W E +++A A+
Sbjct: 540 AAAPAAA-VDTDLALGPAASTAS--RPPAYCDTDEKLLVKRLTEAVRWQPEAAAAVAAAI 596
Query: 653 VDSKSSKE---------CATWLFLQGNDSVGKKRLALAIAESVFGSVEMFSHVDMMKREN 703
++S + TW+ G+D GK ++A A++ SVFG+ + V + N
Sbjct: 597 TKARSGERKRRGMGPTRADTWVLFSGHDVAGKTKMAEALSMSVFGT----NAVALRLAGN 652
Query: 704 SETPFS--------EKVVGPLKNNEKFVVLVENADF--GDTLIRKMLADEFEIAKF---- 749
P + + V ++ N V++++ D D +++ + E +
Sbjct: 653 GGEPIASCRGRTALDCVADAIRANPLRVIVLDGFDHHDDDRVVQASILRAVESGRLVDSR 712
Query: 750 ---GTLGQKIFI---LSNGGSMVSEDQKKDSVMKLVLKISETEKKPTFELSPSSSSSSKS 803
LG+ IF+ L + + Q DS L L++ +K E P +
Sbjct: 713 GRDVALGEAIFVVMSLDDTRRCQEDHQFTDSPWNLELRVRNNARKRRPEPQPLDGA---- 768
Query: 804 PCLGNKRSAELDLFSKIKIPRIEENEGNKKREFSFSRQSSFNNTLDLNMKADEEDNEDYD 863
G++R R+ S LDLN+ E+ +D D
Sbjct: 769 ---GDRRLK--------------------------PRKDSPPLHLDLNLSMCEDHTDDDD 799
Query: 864 EG--ENSPISSDLTRETLGEHLISNESLDSIENLFEFNQSPAKNKEMTQMFMSRVKESFE 921
G E+ SSDLT E E+ + +P+ E+T+ + V F+
Sbjct: 800 SGGEESRNSSSDLTVEHEQEYGQPAAAAAKF-------SAPSSFSELTKAVDATV--VFK 850
Query: 922 EV-LGNVKFSVQDKVIEEIGVGCGS 945
V G +K SV D V ++G G+
Sbjct: 851 PVDFGPLKRSVSDVVSAKLGDAAGA 875
>gb|AAC31855.1| expressed protein [Arabidopsis thaliana] gi|7486038|pir||T02488
hypothetical protein At2g29970 [imported] - Arabidopsis
thaliana gi|18402202|ref|NP_565689.1| heat shock
protein-related [Arabidopsis thaliana]
Length = 1002
Score = 207 bits (527), Expect = 1e-51
Identities = 259/937 (27%), Positives = 405/937 (42%), Gaps = 141/937 (15%)
Query: 7 TLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQQQNNHP 66
T ++ LT E A L ++ +ARRR HAQ T LH + LL++PSS LR C+ S+ +N P
Sbjct: 7 TARQCLTEETARALDDAVSVARRRSHAQTTSLHAVSGLLTMPSSILREVCI-SRAAHNTP 65
Query: 67 ----LQCRALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQRR----- 117
LQ RALELC V+L+RLP++ ++P + P +SN+L+AA+KR+QA QRR
Sbjct: 66 YSSRLQFRALELCVGVSLDRLPSSKSTPTTTVEEDPPVSNSLMAAIKRSQATQRRHPETY 125
Query: 118 --GCIEQNQQQQQQPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSSTL 175
I N + +L KVEL ILSILDDP VSRV EAGF S +K ++ +
Sbjct: 126 HLHQIHGNNNTETTSVL--KVELKYFILSILDDPIVSRVFGEAGFRSTDIKLDVLHPPVT 183
Query: 176 INSSSVFHS-SPSPLSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKEDINLVFD 234
SS F S S P L G V F PF E+ + + +
Sbjct: 184 SQFSSRFTSRSRIPPLFLCNLPESDSGRVRFG---------FPFGDLDENCRR----IGE 230
Query: 235 VLLRKKKKNTVIVGDTVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFHGLSSVSLKYMK 294
VL RK KKN ++VG ++ + R + G +P E+ GLS VS+K
Sbjct: 231 VLARKDKKNPLLVGVCGVEALKTFTDSINRGKFGFLPLEIS--------GLSVVSIKI-- 280
Query: 295 KEEVEMNVIRVLKRKVSDYVALGVGAIFYVGDLKWIVDDNDGSLNEKEVVDYVVEEIGKL 354
EV ++ R+ K D L G + +G+LK + D VD + + + KL
Sbjct: 281 -SEVLVDGSRI-DIKFDDLGRLKSGMVLNLGELKVLASDVFS-------VDVIEKFVLKL 331
Query: 355 FGEEGNKNGKIWLV-ATASYQSYMRCQMRVPTFENQWCLQAVPVPSGGLGLSLHSSSVHD 413
K+W + + +S ++Y++ R PT + W L +P+ S GL SS+
Sbjct: 332 ADLLKLHREKLWFIGSVSSNETYLKLIERFPTIDKDWNLHLLPITSSSQGL-YPKSSLMG 390
Query: 414 SKMSISQNPSPMLESKFFSNKEEHEKLNCCEECVSNYEKEAQLFK------PDQ-KNLLP 466
S + S + + S+ ++ L C C YE+E F DQ LP
Sbjct: 391 SFVPFGGFFSSTSDFRIPSSSSMNQTLPRCHLCNEKYEQEVTAFAKSGSMIDDQCSEKLP 450
Query: 467 SWLQSHSTE------ARQKDE-------LTQLNKKWNRLCQCLHQNK----------QPQ 503
SWL++ E + KD+ + L KKW+ +CQ +HQ +PQ
Sbjct: 451 SWLRNVEHEHEKGNLGKVKDDPNVLASRIPALQKKWDDICQRIHQTPAFPKLSFQPVRPQ 510
Query: 504 NHWSNNHSSNAKIYPYNSSYPYWPNQGSSILPDTSSSISFADSATKPAYSSNLIPRFRRG 563
SS K+ + P + + S P + L + +
Sbjct: 511 FPLQLGSSSQTKMSLGS------PTEKIVCTRTSESFQGMVALPQNPPHQPGLSVKISKP 564
Query: 564 QQSCTIEFNFNDEKAQKNQVTATLELDSLKGMEGTKEVKTTLALGNSTFSVSDQKRMENL 623
+ T + + + + + VT L L ++ + +E T +++ F V +K++ L
Sbjct: 565 KH--TEDLSSSTTNSPLSFVTTDLGLGTIYASK-NQEPSTPVSVERRDFEVIKEKQL--L 619
Query: 624 TLQR-----DHIYKVLQENIPWHCETVSSIAEALV-----DSKSSKECAT----WLFLQG 669
+ R + ++L + + E V++I+E + + + AT WL L G
Sbjct: 620 SASRYCKDFKSLRELLSRKVGFQNEAVNAISEIVCGYRDESRRRNNHVATTSNVWLALLG 679
Query: 670 NDSVGKKRLALAIAESVFGSVEMFSHVDMMKRENSETPFSEKVV-----GPLKNNEKFVV 724
D GKK++ALA+AE G + F VD +++ + F K V G + VV
Sbjct: 680 PDKAGKKKVALALAEVFCGGQDNFICVDFKSQDSLDDRFRGKTVVDYIAGEVARRADSVV 739
Query: 725 LVENADFGDTLIRKMLADEFEIAKFGTLGQKIFILSN---GGSMVSEDQKKD-SVMKLVL 780
+EN + + + L++ K + + N ++ D+ D V++ +
Sbjct: 740 FIENVEKAEFPDQIRLSEAMRTGKLRDSHGREISMKNVIVVATISGSDKASDCHVLEEPV 799
Query: 781 KISE----TEKKPTFE--LSPSSSSSSKSPCLGNKRSAELDLFSKIKIPRIEENEGNKKR 834
K SE K T + L+ +S+ + P NKR R EE E
Sbjct: 800 KYSEERVLNAKNWTLQIKLADTSNVNKNGP---NKR-------------RQEEAETEVTE 843
Query: 835 EFSFSRQSSFNNTLDLNMKADE-EDNED--YDEGENS 868
+ Q SF LDLN+ DE E NED Y EN+
Sbjct: 844 LRALKSQRSF---LDLNLPVDEIEANEDEAYTMSENT 877
>gb|AAK64026.1| unknown protein [Arabidopsis thaliana]
Length = 1002
Score = 207 bits (526), Expect = 2e-51
Identities = 259/937 (27%), Positives = 405/937 (42%), Gaps = 141/937 (15%)
Query: 7 TLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQQQNNHP 66
T ++ LT E A L ++ +ARRR HAQ T LH + LL++PSS LR C+ S+ +N P
Sbjct: 7 TARQCLTEETARALDDAVSVARRRSHAQTTSLHAVSGLLTMPSSILREVCI-SRAAHNTP 65
Query: 67 ----LQCRALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQRR----- 117
LQ RALELC V+L+RLP++ ++P + P +SN+L+AA+KR+QA QRR
Sbjct: 66 YSSRLQFRALELCVGVSLDRLPSSKSTPTTTVEEDPPVSNSLMAAIKRSQATQRRHPETY 125
Query: 118 --GCIEQNQQQQQQPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSSTL 175
I N + +L KVEL ILSILDDP VSRV EAGF S +K ++ +
Sbjct: 126 HLHQIHGNNNTETTSVL--KVELKYFILSILDDPIVSRVFGEAGFRSTDIKLDVLHPPVT 183
Query: 176 INSSSVFHS-SPSPLSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKEDINLVFD 234
SS F S S P L G V F PF E+ + + +
Sbjct: 184 SQFSSRFTSRSRIPPLFLCNLPESDSGRVRFG---------FPFGDLDENCRR----IGE 230
Query: 235 VLLRKKKKNTVIVGDTVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFHGLSSVSLKYMK 294
VL RK KKN ++VG ++ + R + G +P E+ GLS VS+K
Sbjct: 231 VLARKDKKNPLLVGVCGVEALKTFTDSINRGKFGFLPLEIS--------GLSVVSIKI-- 280
Query: 295 KEEVEMNVIRVLKRKVSDYVALGVGAIFYVGDLKWIVDDNDGSLNEKEVVDYVVEEIGKL 354
EV ++ R+ K D L G + +G+LK + D VD + + + KL
Sbjct: 281 -SEVLVDGSRI-DIKFDDLGRLKSGMVLNLGELKVLASDVFS-------VDVIEKFVLKL 331
Query: 355 FGEEGNKNGKIWLV-ATASYQSYMRCQMRVPTFENQWCLQAVPVPSGGLGLSLHSSSVHD 413
K+W + + +S ++Y++ R PT + W L +P+ S GL SS+
Sbjct: 332 ADLLKLHREKLWFIGSVSSNETYLKLIERFPTIDKDWNLHLLPITSSSQGL-YPKSSLMG 390
Query: 414 SKMSISQNPSPMLESKFFSNKEEHEKLNCCEECVSNYEKEAQLFK------PDQ-KNLLP 466
S + S + + S+ ++ L C C YE+E F DQ LP
Sbjct: 391 SFVPFGGFFSSTSDFRIPSSSSMNQTLPRCHLCNEKYEQEVTAFAKSGSMIDDQCSEKLP 450
Query: 467 SWLQSHSTE------ARQKDE-------LTQLNKKWNRLCQCLHQNK----------QPQ 503
SWL++ E + KD+ + L KKW+ +CQ +HQ +PQ
Sbjct: 451 SWLRNVEHEHEKGNLGKVKDDPNVLASRIPALQKKWDDICQRIHQTPAFPKLSFQPVRPQ 510
Query: 504 NHWSNNHSSNAKIYPYNSSYPYWPNQGSSILPDTSSSISFADSATKPAYSSNLIPRFRRG 563
SS K+ + P + + S P + L + +
Sbjct: 511 FPLQLGSSSQTKMSLGS------PTEKIVCTRTSESFQGMVALPQNPPHQPGLSVKISKP 564
Query: 564 QQSCTIEFNFNDEKAQKNQVTATLELDSLKGMEGTKEVKTTLALGNSTFSVSDQKRMENL 623
+ T + + + + + VT L L ++ + +E T +++ F V +K++ L
Sbjct: 565 KH--TEDLSSSTTNSPLSFVTTDLGLGTIYASK-NQEPSTPVSVERRDFEVIKEKQL--L 619
Query: 624 TLQR-----DHIYKVLQENIPWHCETVSSIAEALV-----DSKSSKECAT----WLFLQG 669
+ R + ++L + + E V++I+E + + + AT WL L G
Sbjct: 620 SASRYCKDFKSLRELLSRKVGFQNEAVNAISEIVCGYRDESRRRNNHVATTSNVWLALLG 679
Query: 670 NDSVGKKRLALAIAESVFGSVEMFSHVDMMKRENSETPFSEKVV-----GPLKNNEKFVV 724
D GKK++ALA+AE G + F VD +++ + F K V G + VV
Sbjct: 680 PDKAGKKKVALALAEVFCGGQDNFICVDFKSQDSLDDRFRGKTVVDYIAGEVARRAGSVV 739
Query: 725 LVENADFGDTLIRKMLADEFEIAKFGTLGQKIFILSN---GGSMVSEDQKKD-SVMKLVL 780
+EN + + + L++ K + + N ++ D+ D V++ +
Sbjct: 740 FIENVEKAEFPDQIRLSEAMRTGKLRDSHGREISMKNVIVVATISGSDKASDCHVLEEPV 799
Query: 781 KISE----TEKKPTFE--LSPSSSSSSKSPCLGNKRSAELDLFSKIKIPRIEENEGNKKR 834
K SE K T + L+ +S+ + P NKR R EE E
Sbjct: 800 KYSEERVLNAKNWTLQIKLADTSNVNKNGP---NKR-------------RQEEAETEVTE 843
Query: 835 EFSFSRQSSFNNTLDLNMKADE-EDNED--YDEGENS 868
+ Q SF LDLN+ DE E NED Y EN+
Sbjct: 844 LRALKSQRSF---LDLNLPVDEIEANEDEAYTMSENT 877
>ref|NP_973646.1| heat shock protein-related [Arabidopsis thaliana]
Length = 910
Score = 189 bits (481), Expect = 3e-46
Identities = 248/988 (25%), Positives = 411/988 (41%), Gaps = 190/988 (19%)
Query: 1 MRSGAYTLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQ 60
M + ++ LTAEA+ L+ ++ +ARRRGH+Q T LH + LLSLP+S LR+AC + +
Sbjct: 1 MPTAVNVAKQCLTAEASYALEEAVNVARRRGHSQTTSLHAISALLSLPTSVLRDACARVR 60
Query: 61 QQNNHP-LQCRALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQR--- 116
P LQ +AL+LC +V+L+R+ + L P +SN+L+AA+KR+QAHQR
Sbjct: 61 NSAYSPRLQFKALDLCLSVSLDRI---QSGHQLGSDDSPPVSNSLMAAIKRSQAHQRRLP 117
Query: 117 ---RGCIEQNQQQQQQPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSS 173
R E +Q Q Q L VKVEL QLILSILDDP VSRV EAGF S +K ++
Sbjct: 118 ENFRIYQEMSQSQNQNSLSCVKVELRQLILSILDDPVVSRVFGEAGFRSSELKLSIIRPV 177
Query: 174 TLINSSSVFHSSPSPLSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKE-DINLV 232
+ + +SS PL FL + L + + V + + S N + D +
Sbjct: 178 PHL----LRYSSQQPL----FLCN------LTGNPEPNPVRWGFTVPSLNFNGDLDYRRI 223
Query: 233 FDVLLRKKKKNTVIVGDTVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFHGLSSVSLKY 292
V + K +N ++VG + G+++ + E+ + + T K HGL++V++
Sbjct: 224 SAVFTKDKGRNPLLVGVS---AYGVLTSYLNSLEKNQTDGMILPT---KLHGLTAVNIGS 277
Query: 293 MKKEEVEMNVIRV-LKRKVSDYVAL-----GVGAIFYVGDLKWIVDDNDGSLNEKEVVDY 346
+++ + + + D L G G + + GDL+ + + +G++ +Y
Sbjct: 278 EISDQISVKFDKTYTDTRFHDLGKLAEQGSGPGLLLHYGDLR-VFTNGEGNV---PAANY 333
Query: 347 VVEEIGKLFGEEGNKNGKIWLV-ATASYQSYMRCQMRVPTFENQWCLQAVPVPSGGLGLS 405
+V I +L G ++WL+ AT S + Y + R P E W LQ + + S L
Sbjct: 334 IVNRISELLRRHGR---RVWLIGATTSNEVYEKMMRRFPNVEKDWDLQLLTITSLKPCLP 390
Query: 406 LHSSSVHDSKMSISQNPSPMLESKFFSNKEEHEKLNCCEECVSNYEKE----AQLFKPDQ 461
+ SS+ S + FFS KL S ++ E
Sbjct: 391 HNKSSLIGSFVPF---------GGFFSTTPSELKLP-----FSGFKTEITGPVSSISDQT 436
Query: 462 KNLLPSWLQSHSTEARQKDELTQLNKKWNRLCQCLHQNKQPQNHWSNNHSSNAKIYPYNS 521
++ LP WLQ + T LN+K S+AK+
Sbjct: 437 QSTLPPWLQMTTR--------TDLNQK-----------------------SSAKVVQTKE 465
Query: 522 SYPYWPNQGSSILPDTSSSISFADSATKPAYSSNLIPRFRRGQQSCTIEFNFNDEKAQKN 581
++ F SA+ S+ +S T + N
Sbjct: 466 GL------------ESVCGNKFTSSASASTCSA----------KSVTTDLNL-------- 495
Query: 582 QVTATLELDSLKGMEGTKEVKTTLALGNSTFSVSDQKRMENLTLQRDHIYKVLQENIPWH 641
+V++ LK +K+ ++ + +F E+ + IY+ L + +
Sbjct: 496 RVSSVTTGSGLKKHLDSKDFSQPQSVSSYSFDNPRDLNAESFKI----IYRRLTDMVSGQ 551
Query: 642 CETVSSIAEALVD-SKSSKECATWLFLQGNDSVGKKRLALAIAESVFGSVEMFSHVDMMK 700
E I+ AL KS WL L G D+VGK+R++L +AE V+ S F VD+
Sbjct: 552 DEAARVISCALSQPPKSVTRRDVWLNLVGPDTVGKRRMSLVLAEIVYQSEHRFMAVDLGA 611
Query: 701 RENS----ETPFS-------EKVVGPLKNNEKFVVLVENADFGDTLIRKMLADEFEIAKF 749
E + P + + + N VV +EN + D ++ L+ E KF
Sbjct: 612 AEQGMGGCDDPMRLRGKTMVDHIFEVMCRNPFCVVFLENIEKADEKLQMSLSKAIETGKF 671
Query: 750 GT-------LGQKIFIL--SNGGSMVSEDQKKDSVMK-----LVLKISETEKKPTFE--L 793
+G IF++ S+ GS + ++ +++ + ++I P
Sbjct: 672 MDSHGREVGIGNTIFVMTSSSQGSATTTSYSEEKLLRVKGRQVEIRIETVSSLPMVRSVY 731
Query: 794 SPSSSSSSKSPCLGNKRSAELDLFSKIKIPRIEENEGNKKREFSFSRQSSFNNTLDLNMK 853
P+S + K LGN + + + S ++ R + N LDLN+
Sbjct: 732 GPTSVNKRKLMGLGNLQETKDTVESVKRLNR------------------TTNGVLDLNLP 773
Query: 854 ADE-EDNEDYDEGENSPISSDLTRETLGEHLISNESLDSIENLFEFNQSPAKNKEMTQMF 912
A E E E Y ENS + +L + + L E P + + +
Sbjct: 774 AQETEIEEKYHCEENSNVWL--------------MNLKNHKRLIEVPFKPFDFEGLAEKI 819
Query: 913 MSRVKESFEE-VLGNVKFSVQDKVIEEI 939
VKE+F++ V + V K+IE +
Sbjct: 820 KKSVKENFDKCVRSDCLLEVDPKIIERL 847
>gb|AAF82200.1| Strong similarity to an unknown protein F23F1.11 gi|7486038 from
Arabidopsis thaliana BAC F23F1 gb|AC004680. ESTs
gb|T75672, gb|N65732 and gb|AA404793 come from this gene
gi|25406961|pir||B86207 hypothetical protein [imported]
- Arabidopsis thaliana
Length = 979
Score = 180 bits (456), Expect = 2e-43
Identities = 217/840 (25%), Positives = 354/840 (41%), Gaps = 128/840 (15%)
Query: 7 TLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQQQN--- 63
T +E LT EAA L ++ +ARRR HAQ T LH + LL++PSS LR C+ ++
Sbjct: 7 TARECLTEEAARALDDAVVVARRRSHAQTTSLHAVSALLAMPSSILREVCVSRAARSVPY 66
Query: 64 NHPLQCRALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQRRGCIEQN 123
+ LQ RALELC V+L+RLP ++ SP + P +SN+L+AA+KR+QA+QRR +
Sbjct: 67 SSRLQFRALELCVGVSLDRLP-SSKSPATEED--PPVSNSLMAAIKRSQANQRRHPESYH 123
Query: 124 QQQQQQ--------PLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSSTL 175
QQ +KVEL ILSILDDP V+RV EAGF S +K ++ +
Sbjct: 124 LQQIHASNNGGGGCQTTVLKVELKYFILSILDDPIVNRVFGEAGFRSSEIKLDVLHPPVT 183
Query: 176 INSSSVFHSSPSPLSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKEDINLVFDV 235
SS PL FL + L +S + PF SS + E+ + +V
Sbjct: 184 QLSSRFSRGRCPPL----FLCN------LPNSDPNRE---FPFSGSSGFD-ENSRRIGEV 229
Query: 236 LLRKKKKNTVIVGDTVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFHGLSSVSLKYMKK 295
L RK KKN +++G+ + ++ + + G + ++ + S L K
Sbjct: 230 LGRKDKKNPLLIGNCANEALKTFTDSINSGKLGFLQMDISGLSLISIEKEISEILADGSK 289
Query: 296 EEVEMNV-IRVLKRKVSDYVALGVGAIFYVGDLKWIVDDNDGSLNEKEVVDYVVEEIGKL 354
E E+ + + L R V + G + +G+LK + + + +L + +V ++ L
Sbjct: 290 NEEEIRMKVDDLGRTVEQSGSKS-GIVLNLGELKVLTSEANAAL------EILVSKLSDL 342
Query: 355 FGEEGNKNGKIWLVATASYQSYMRCQMRVPTFENQWCLQAVPVPSGGLGLSLHSSSVHDS 414
E + I V +S ++Y + R PT E W L +P+ + + S+
Sbjct: 343 LKHESKQLSFIGCV--SSNETYTKLIDRFPTIEKDWDLHVLPITAS----TKPSTQGVYP 396
Query: 415 KMSISQNPSPMLESKFFSNKEE---------HEKLNCCEECVSNYEKEAQLFKPDQKNL- 464
K S+ + P FFS+ ++ L+ C C Y +E +L
Sbjct: 397 KSSLMGSFVPF--GGFFSSTSNFRVPLSSTVNQTLSRCHLCNEKYLQEVAAVLKAGSSLS 454
Query: 465 --------LPSWLQSHSTEA---------------RQKDELTQLNKKWNRLCQCLHQNKQ 501
L WL++ T+ + L KKW+ +CQ +H
Sbjct: 455 LADKCSEKLAPWLRAIETKEDKGITGSSKALDDANTSASQTAALQKKWDNICQSIH---- 510
Query: 502 PQNHWSNNHSSNAKIYPYNSSYPYWPNQGSSILPDTSSSISFADSATKPA--------YS 553
H+ + S P +P Q + +S + P +
Sbjct: 511 --------HTPAFPKLGFQSVSPQFPVQTEKSVRTPTSYLETPKLLNPPISKPKPMEDLT 562
Query: 554 SNLIPRFRRGQQSC-TIEFNFNDEKAQKNQVTATLELDSLKGMEGTKEVKTTLALGNSTF 612
+++ R SC T +F A KNQ + T T+E K L NS+
Sbjct: 563 ASVTNRTVSLPLSCVTTDFGLGVIYASKNQESKT-----------TRE-KPMLVTLNSSL 610
Query: 613 SVSDQKRMENLTLQRDHIYKVLQENIPWHCETVSSIAEALVDSKS-----SKECATWLFL 667
+ QK ++L ++L + W E V++I++ + K+ ++ WL L
Sbjct: 611 EHTYQKDFKSLR-------EILSRKVAWQTEAVNAISQIICGCKTDSTRRNQASGIWLAL 663
Query: 668 QGNDSVGKKRLALAIAESVFGSVEMFSHVDMMKRENS-ETPFSEK-----VVGPLKNNEK 721
G D VGKK++A+ ++E FG + VD S + F K V G L
Sbjct: 664 LGPDKVGKKKVAMTLSEVFFGGKVNYICVDFGAEHCSLDDKFRGKTVVDYVTGELSRKPH 723
Query: 722 FVVLVENADFGDTLIRKMLADEFEIAKFGTLGQKIFILSNGGSMVSEDQKKDSVMKLVLK 781
VVL+EN + + + L++ K L ++ + N +V+ KD+ V+K
Sbjct: 724 SVVLLENVEKAEFPDQMRLSEAVSTGKIRDLHGRVISMKNVIVVVTSGIAKDNATDHVIK 783
>gb|AAM16220.1| At2g40130/T7M7.2 [Arabidopsis thaliana] gi|15450731|gb|AAK96637.1|
At2g40130/T7M7.2 [Arabidopsis thaliana]
gi|18405278|ref|NP_030558.1| heat shock protein-related
[Arabidopsis thaliana]
Length = 491
Score = 164 bits (416), Expect = 1e-38
Identities = 155/515 (30%), Positives = 246/515 (47%), Gaps = 65/515 (12%)
Query: 1 MRSGAYTLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQ 60
M + ++ LTAEA+ L+ ++ +ARRRGH+Q T LH + LLSLP+S LR+AC + +
Sbjct: 1 MPTAVNVAKQCLTAEASYALEEAVNVARRRGHSQTTSLHAISALLSLPTSVLRDACARVR 60
Query: 61 QQNNHP-LQCRALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQR--- 116
P LQ +AL+LC +V+L+R+ + L P +SN+L+AA+KR+QAHQR
Sbjct: 61 NSAYSPRLQFKALDLCLSVSLDRI---QSGHQLGSDDSPPVSNSLMAAIKRSQAHQRRLP 117
Query: 117 ---RGCIEQNQQQQQQPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSS 173
R E +Q Q Q L VKVEL QLILSILDDP VSRV EAGF S +K ++
Sbjct: 118 ENFRIYQEMSQSQNQNSLSCVKVELRQLILSILDDPVVSRVFGEAGFRSSELKLSIIRPV 177
Query: 174 TLINSSSVFHSSPSPLSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKE-DINLV 232
+ + +SS PL FL + L + + V + + S N + D +
Sbjct: 178 PHL----LRYSSQQPL----FLCN------LTGNPEPNPVRWGFTVPSLNFNGDLDYRRI 223
Query: 233 FDVLLRKKKKNTVIVGDTVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFHGLSSVSLKY 292
V + K +N ++VG + G+++ + E+ + + T K HGL++V++
Sbjct: 224 SAVFTKDKGRNPLLVGVS---AYGVLTSYLNSLEKNQTDGMILPT---KLHGLTAVNIGS 277
Query: 293 MKKEEVEMNVIRV-LKRKVSDYVAL-----GVGAIFYVGDLKWIVDDNDGSLNEKEVVDY 346
+++ + + + D L G G + + GDL+ + + +G++ +Y
Sbjct: 278 EISDQISVKFDKTYTDTRFHDLGKLAEQGSGPGLLLHYGDLR-VFTNGEGNV---PAANY 333
Query: 347 VVEEIGKLFGEEGNKNGKIWLV-ATASYQSYMRCQMRVPTFENQWCLQAVPVPSGGLGLS 405
+V I +L G ++WL+ AT S + Y + R P E W LQ + + S L
Sbjct: 334 IVNRISELLRRHGR---RVWLIGATTSNEVYEKMMRRFPNVEKDWDLQLLTITSLKPCLP 390
Query: 406 LHSSSVHDSKMSISQNPSPMLESKFFSNKEEHEKLNCCEECVSNYEKE----AQLFKPDQ 461
+ SS+ S + FFS KL S ++ E
Sbjct: 391 HNKSSLIGSFVPF---------GGFFSTTPSELKLP-----FSGFKTEITGPVSSISDQT 436
Query: 462 KNLLPSWLQ-SHSTEARQKDELTQLNKK-WNRLCQ 494
++ LP WLQ + T+ QK KK WN+ +
Sbjct: 437 QSTLPPWLQMTTRTDLNQKSSAKCRPKKGWNQFVE 471
>gb|AAF18727.1| hypothetical protein [Arabidopsis thaliana] gi|25408669|pir||F84825
hypothetical protein At2g40130 [imported] - Arabidopsis
thaliana
Length = 907
Score = 161 bits (408), Expect = 9e-38
Identities = 132/414 (31%), Positives = 212/414 (50%), Gaps = 45/414 (10%)
Query: 1 MRSGAYTLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQ 60
M + ++ LTAEA+ L+ ++ +ARRRGH+Q T LH + LLSLP+S LR+AC + +
Sbjct: 1 MPTAVNVAKQCLTAEASYALEEAVNVARRRGHSQTTSLHAISALLSLPTSVLRDACARVR 60
Query: 61 QQNNHP-LQCRALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQR--- 116
P LQ +AL+LC +V+L+R+ + L P +SN+L+AA+KR+QAHQR
Sbjct: 61 NSAYSPRLQFKALDLCLSVSLDRI---QSGHQLGSDDSPPVSNSLMAAIKRSQAHQRRLP 117
Query: 117 ---RGCIEQNQQQQQQPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSS 173
R E +Q Q Q L VKVEL QLILSILDDP VSRV EAGF S +K ++
Sbjct: 118 ENFRIYQEMSQSQNQNSLSCVKVELRQLILSILDDPVVSRVFGEAGFRSSELKLSIIRPV 177
Query: 174 TLINSSSVFHSSPSPLSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKE-DINLV 232
+ + +SS PL FL + L + + V + + S N + D +
Sbjct: 178 PHL----LRYSSQQPL----FLCN------LTGNPEPNPVRWGFTVPSLNFNGDLDYRRI 223
Query: 233 FDVLLRKKKKNTVIVGDTVSLTEGLVSEIMKRFERGEVPDEMKTTHFVKFHGLSSVSLKY 292
V + K +N ++VG + G+++ + E+ + + T K HGL++V++
Sbjct: 224 SAVFTKDKGRNPLLVGVS---AYGVLTSYLNSLEKNQTDGMILPT---KLHGLTAVNIGS 277
Query: 293 MKKEEVEMNVIRV-LKRKVSDYVAL-----GVGAIFYVGDLKWIVDDNDGSLNEKEVVDY 346
+++ + + + D L G G + + GDL+ + + +G++ +Y
Sbjct: 278 EISDQISVKFDKTYTDTRFHDLGKLAEQGSGPGLLLHYGDLR-VFTNGEGNV---PAANY 333
Query: 347 VVEEIGKLFGEEGNKNGKIWLV-ATASYQSYMRCQMRVPTFENQWCLQAVPVPS 399
+V I +L G ++WL+ AT S + Y + R P E W LQ + + S
Sbjct: 334 IVNRISELLRRHGR---RVWLIGATTSNEVYEKMMRRFPNVEKDWDLQLLTITS 384
Score = 57.0 bits (136), Expect = 3e-06
Identities = 80/340 (23%), Positives = 135/340 (39%), Gaps = 62/340 (18%)
Query: 630 IYKVLQENIPWHCETVSSIAEALVDS-KSSKECATWLFLQGNDSVGKKRLALAIAESVFG 688
IY+ L + + E I+ AL KS WL L G D+VGK+R++L +AE V+
Sbjct: 537 IYRRLTDMVSGQDEAARVISCALSQPPKSVTRRDVWLNLVGPDTVGKRRMSLVLAEIVYQ 596
Query: 689 SVEMFSHVDMMKRENS----ETPFS-------EKVVGPLKNNEKFVVLVENADFGDTLIR 737
S F VD+ E + P + + + N VV +EN + D ++
Sbjct: 597 SEHRFMAVDLGAAEQGMGGCDDPMRLRGKTMVDHIFEVMCRNPFCVVFLENIEKADEKLQ 656
Query: 738 KMLADEFEIAKFGT-------LGQKIFIL--SNGGSMVSEDQKKDSVMK-----LVLKIS 783
L+ E KF +G IF++ S+ GS + ++ +++ + ++I
Sbjct: 657 MSLSKAIETGKFMDSHGREVGIGNTIFVMTSSSQGSATTTSYSEEKLLRVKGRQVEIRIE 716
Query: 784 ETEKKPTFE--LSPSSSSSSKSPCLGNKRSAELDLFSKIKIPRIEENEGNKKREFSFSRQ 841
P P+S + K LGN + + + S ++ R
Sbjct: 717 TVSSLPMVRSVYGPTSVNKRKLMGLGNLQETKDTVESVKRLNR----------------- 759
Query: 842 SSFNNTLDLNMKADE-EDNEDYDEGENSPISSDLTRETLGEHLISNESLDSIENLFEFNQ 900
+ N LDLN+ A E E E Y ENS + +L + + L E
Sbjct: 760 -TTNGVLDLNLPAQETEIEEKYHCEENSNVWL--------------MNLKNHKRLIEVPF 804
Query: 901 SPAKNKEMTQMFMSRVKESFEE-VLGNVKFSVQDKVIEEI 939
P + + + VKE+F++ V + V K+IE +
Sbjct: 805 KPFDFEGLAEKIKKSVKENFDKCVRSDCLLEVDPKIIERL 844
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.313 0.129 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,667,440,333
Number of Sequences: 2540612
Number of extensions: 72218234
Number of successful extensions: 239452
Number of sequences better than 10.0: 467
Number of HSP's better than 10.0 without gapping: 88
Number of HSP's successfully gapped in prelim test: 393
Number of HSP's that attempted gapping in prelim test: 238326
Number of HSP's gapped (non-prelim): 1178
length of query: 1009
length of database: 863,360,394
effective HSP length: 138
effective length of query: 871
effective length of database: 512,755,938
effective search space: 446610421998
effective search space used: 446610421998
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 80 (35.4 bits)
Medicago: description of AC148819.4


