

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
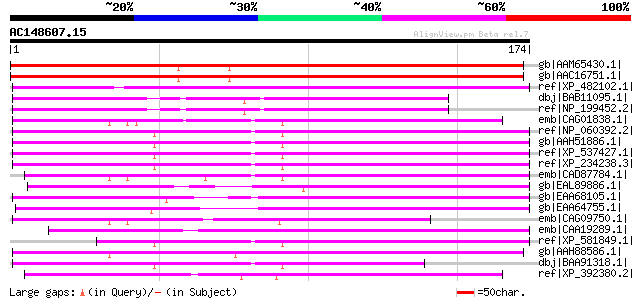
Query= AC148607.15 - phase: 0
(174 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
gb|AAM65430.1| unknown [Arabidopsis thaliana] gi|20259567|gb|AAM... 209 3e-53
gb|AAC16751.1| Contains similarity to pre-mRNA processing protei... 209 3e-53
ref|XP_482102.1| unknown protein [Oryza sativa (japonica cultiva... 95 7e-19
dbj|BAB11095.1| unnamed protein product [Arabidopsis thaliana] 90 3e-17
ref|NP_199452.2| expressed protein [Arabidopsis thaliana] 90 3e-17
emb|CAG01838.1| unnamed protein product [Tetraodon nigroviridis] 86 4e-16
ref|NP_060392.2| PRP39 pre-mRNA processing factor 39 homolog [Ho... 84 2e-15
gb|AAH51886.1| PRPF39 protein [Homo sapiens] 84 2e-15
ref|XP_537427.1| PREDICTED: similar to PRP39 pre-mRNA processing... 84 2e-15
ref|XP_234238.3| PREDICTED: similar to PRP39 pre-mRNA processing... 84 2e-15
emb|CAD87784.1| novel protein similar to pre-mRNA processing pro... 82 8e-15
gb|EAL89886.1| conserved hypothetical protein [Aspergillus fumig... 74 1e-12
gb|EAA68105.1| hypothetical protein FG01244.1 [Gibberella zeae P... 73 4e-12
gb|EAA64755.1| hypothetical protein AN1635.2 [Aspergillus nidula... 72 6e-12
emb|CAG09750.1| unnamed protein product [Tetraodon nigroviridis] 71 1e-11
emb|CAA19289.1| SPBC4B4.09 [Schizosaccharomyces pombe] gi|191132... 70 2e-11
ref|XP_581849.1| PREDICTED: similar to PRP39 pre-mRNA processing... 67 2e-10
gb|AAH88586.1| LOC496864 protein [Xenopus tropicalis] 66 4e-10
dbj|BAA91318.1| unnamed protein product [Homo sapiens] 64 2e-09
ref|XP_392380.2| PREDICTED: similar to novel protein similar to ... 63 3e-09
>gb|AAM65430.1| unknown [Arabidopsis thaliana] gi|20259567|gb|AAM14126.1| unknown
protein [Arabidopsis thaliana]
gi|15810565|gb|AAL07170.1| unknown protein [Arabidopsis
thaliana] gi|18379230|ref|NP_563700.1|
hydroxyproline-rich glycoprotein family protein
[Arabidopsis thaliana]
Length = 768
Score = 209 bits (531), Expect = 3e-53
Identities = 109/182 (59%), Positives = 131/182 (71%), Gaps = 10/182 (5%)
Query: 1 MQQDWARLAMIYTRILENPNQQLDRYFNSFKELASNRPLSELRTADEAAAVAGVV----- 55
MQQDW+R+A+IYTRILENP Q LDRYF+SFKELA RPLSELR+A+E+AA A V
Sbjct: 220 MQQDWSRVALIYTRILENPIQNLDRYFSSFKELAETRPLSELRSAEESAAAAVAVAGDAS 279
Query: 56 ----SEGIDQGVEGEVHPDGA-DNSPKPASAGLTEAEELEKYIAIREEMYKKAKEFDSKI 110
SE ++ EG DG+ + SPK SA TE EEL+KY+ IRE MY K+KEF+SKI
Sbjct: 280 ESAASESGEKADEGRSQVDGSTEQSPKLESASSTEPEELKKYVGIREAMYIKSKEFESKI 339
Query: 111 IGFETTIRRPYFHVRPLNIGELENWHNYLDFIEREGDLSKVTKLLSYKGTFCSIFSCYRL 170
IG+E IRRPYFHVRPLN+ ELENWHNYLDFIER+GD +KV KL C+ + Y +
Sbjct: 340 IGYEMAIRRPYFHVRPLNVAELENWHNYLDFIERDGDFNKVVKLYERCVVTCANYPEYWI 399
Query: 171 TY 172
Y
Sbjct: 400 RY 401
>gb|AAC16751.1| Contains similarity to pre-mRNA processing protein PRP39 gb|L29224
from S. cerevisiae. ESTs gb|R64908 and gb|T88158,
gb|N38703 and gb|AA651043 come from this gene.
[Arabidopsis thaliana] gi|7485869|pir||T00964
hypothetical protein F20D22.14 - Arabidopsis thaliana
Length = 1345
Score = 209 bits (531), Expect = 3e-53
Identities = 109/182 (59%), Positives = 131/182 (71%), Gaps = 10/182 (5%)
Query: 1 MQQDWARLAMIYTRILENPNQQLDRYFNSFKELASNRPLSELRTADEAAAVAGVV----- 55
MQQDW+R+A+IYTRILENP Q LDRYF+SFKELA RPLSELR+A+E+AA A V
Sbjct: 220 MQQDWSRVALIYTRILENPIQNLDRYFSSFKELAETRPLSELRSAEESAAAAVAVAGDAS 279
Query: 56 ----SEGIDQGVEGEVHPDGA-DNSPKPASAGLTEAEELEKYIAIREEMYKKAKEFDSKI 110
SE ++ EG DG+ + SPK SA TE EEL+KY+ IRE MY K+KEF+SKI
Sbjct: 280 ESAASESGEKADEGRSQVDGSTEQSPKLESASSTEPEELKKYVGIREAMYIKSKEFESKI 339
Query: 111 IGFETTIRRPYFHVRPLNIGELENWHNYLDFIEREGDLSKVTKLLSYKGTFCSIFSCYRL 170
IG+E IRRPYFHVRPLN+ ELENWHNYLDFIER+GD +KV KL C+ + Y +
Sbjct: 340 IGYEMAIRRPYFHVRPLNVAELENWHNYLDFIERDGDFNKVVKLYERCVVTCANYPEYWI 399
Query: 171 TY 172
Y
Sbjct: 400 RY 401
>ref|XP_482102.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|40253684|dbj|BAD05627.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
gi|40253455|dbj|BAD05406.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 1161
Score = 95.1 bits (235), Expect = 7e-19
Identities = 52/173 (30%), Positives = 96/173 (55%), Gaps = 3/173 (1%)
Query: 2 QQDWARLAMIYTRILENPNQQLDRYFNSFKELASNRPLSELRTADEAAAVAGVVSEGIDQ 61
Q+ +LA IY L+ P ++L RY+ SF++L + L E A + + + + +
Sbjct: 134 QKQLIQLATIYIDTLKFPTKKLRRYYESFRKLVT---LMEHEAAGAERSSENLRTLEVIK 190
Query: 62 GVEGEVHPDGADNSPKPASAGLTEAEELEKYIAIREEMYKKAKEFDSKIIGFETTIRRPY 121
+ EV ++ +G A+ +++Y+ E +Y+++ + D +I FE +I+RP+
Sbjct: 191 AEDSEVDASIKISALLDEHSGHLRADAVKQYLLSGESLYQRSSKIDKEISCFEASIKRPF 250
Query: 122 FHVRPLNIGELENWHNYLDFIEREGDLSKVTKLLSYKGTFCSIFSCYRLTYSK 174
FHV+PL+ +LENWH YLDF+E++GD KL C+ +S + + Y++
Sbjct: 251 FHVKPLDDDQLENWHRYLDFVEKKGDFDWAVKLYERCLIPCANYSEFWIRYAE 303
>dbj|BAB11095.1| unnamed protein product [Arabidopsis thaliana]
Length = 1022
Score = 89.7 bits (221), Expect = 3e-17
Identities = 55/154 (35%), Positives = 84/154 (53%), Gaps = 15/154 (9%)
Query: 2 QQDWARLAMIYTRILENPNQQLDRYFNSFKELASNRPLSELRTADEAAAVAGVVSEGIDQ 61
QQ W+ LA +Y R L+ P+++LD Y+ +F+++A++ D V G +S D
Sbjct: 167 QQQWSSLANVYLRTLKYPSKKLDLYYKNFRKIAASLKEKIKCRID----VNGDLSS--DP 220
Query: 62 GVEGEVHPDGADNSPK--------PASAGLTEAEELEKYIAIREEMYKKAKEFDSKIIGF 113
E VH D P+S+ ++ L Y++I E+ Y+ +++ KI F
Sbjct: 221 MEEDLVHTRHTDEEISIVVRELMGPSSSSAV-SKALHTYLSIGEQFYQDSRQLMEKISCF 279
Query: 114 ETTIRRPYFHVRPLNIGELENWHNYLDFIEREGD 147
ET IRRPYFHV+PL+ +L+NWH YL F E GD
Sbjct: 280 ETQIRRPYFHVKPLDTNQLDNWHAYLSFGETYGD 313
>ref|NP_199452.2| expressed protein [Arabidopsis thaliana]
Length = 1036
Score = 89.7 bits (221), Expect = 3e-17
Identities = 55/154 (35%), Positives = 84/154 (53%), Gaps = 15/154 (9%)
Query: 2 QQDWARLAMIYTRILENPNQQLDRYFNSFKELASNRPLSELRTADEAAAVAGVVSEGIDQ 61
QQ W+ LA +Y R L+ P+++LD Y+ +F+++A++ D V G +S D
Sbjct: 167 QQQWSSLANVYLRTLKYPSKKLDLYYKNFRKIAASLKEKIKCRID----VNGDLSS--DP 220
Query: 62 GVEGEVHPDGADNSPK--------PASAGLTEAEELEKYIAIREEMYKKAKEFDSKIIGF 113
E VH D P+S+ ++ L Y++I E+ Y+ +++ KI F
Sbjct: 221 MEEDLVHTRHTDEEISIVVRELMGPSSSSAV-SKALHTYLSIGEQFYQDSRQLMEKISCF 279
Query: 114 ETTIRRPYFHVRPLNIGELENWHNYLDFIEREGD 147
ET IRRPYFHV+PL+ +L+NWH YL F E GD
Sbjct: 280 ETQIRRPYFHVKPLDTNQLDNWHAYLSFGETYGD 313
>emb|CAG01838.1| unnamed protein product [Tetraodon nigroviridis]
Length = 338
Score = 85.9 bits (211), Expect = 4e-16
Identities = 63/174 (36%), Positives = 88/174 (50%), Gaps = 12/174 (6%)
Query: 2 QQDWARLAMIYTRILENPNQQLDRYFNSFKE-LASNRP---LSE-----LRTADEAAAVA 52
QQ A + IY RIL P Q ++F FKE + SN P LSE LR +++A
Sbjct: 159 QQKLANVTAIYDRILGIPTQLYSQHFQKFKEHVQSNHPKHFLSEEEFVKLRVELSKSSLA 218
Query: 53 GVVSEGIDQGVEGEVHPDGADNSPKPASAGLTEAEELE-KYIAIREEMYKKAKEFDSKII 111
G++SE D V E P G ++ PA +TE E + K I IR+E++ + SK
Sbjct: 219 GMISED-DAAVAQEELPPGTEDLADPAKR-VTEIENMRHKVIEIRQELFNHNEHEVSKRW 276
Query: 112 GFETTIRRPYFHVRPLNIGELENWHNYLDFIEREGDLSKVTKLLSYKGTFCSIF 165
FE I+RPYFHV+ L +L NW YL+F G +V L C+++
Sbjct: 277 AFEEAIKRPYFHVKALEKTQLSNWREYLEFEIENGTPERVVVLYERCLIACALY 330
>ref|NP_060392.2| PRP39 pre-mRNA processing factor 39 homolog [Homo sapiens]
Length = 548
Score = 84.0 bits (206), Expect = 2e-15
Identities = 56/178 (31%), Positives = 84/178 (46%), Gaps = 6/178 (3%)
Query: 2 QQDWARLAMIYTRILENPNQQLDRYFNSFKELASNRPLSELRTADE----AAAVAGVVSE 57
Q + + IY RIL P Q +F FKE N +L T ++ +A V
Sbjct: 98 QGNLREVTAIYDRILGIPTQLYSHHFQRFKEHVQNNLPRDLLTGEQFIQLRRELASVNGH 157
Query: 58 GIDQGVEGEVHPDGADNSPKPASAGLTEAEELE-KYIAIREEMYKKAKEFDSKIIGFETT 116
D G G+ P G ++ PA +TE E + + I I +EM+ + SK FE
Sbjct: 158 SGDDGPPGDDLPSGIEDITDPAKL-ITEIENMRHRIIEIHQEMFNYNEHEVSKRWTFEEG 216
Query: 117 IRRPYFHVRPLNIGELENWHNYLDFIEREGDLSKVTKLLSYKGTFCSIFSCYRLTYSK 174
I+RPYFHV+PL +L+NW YL+F G +V L C+++ + + Y+K
Sbjct: 217 IKRPYFHVKPLEKAQLKNWKEYLEFEIENGTHERVVVLFERCVISCALYEEFWIKYAK 274
>gb|AAH51886.1| PRPF39 protein [Homo sapiens]
Length = 479
Score = 84.0 bits (206), Expect = 2e-15
Identities = 56/178 (31%), Positives = 84/178 (46%), Gaps = 6/178 (3%)
Query: 2 QQDWARLAMIYTRILENPNQQLDRYFNSFKELASNRPLSELRTADE----AAAVAGVVSE 57
Q + + IY RIL P Q +F FKE N +L T ++ +A V
Sbjct: 29 QGNLREVTAIYDRILGIPTQLYSHHFQRFKEHVQNNLPRDLLTGEQFIQLRRELASVNGH 88
Query: 58 GIDQGVEGEVHPDGADNSPKPASAGLTEAEELE-KYIAIREEMYKKAKEFDSKIIGFETT 116
D G G+ P G ++ PA +TE E + + I I +EM+ + SK FE
Sbjct: 89 SGDDGPPGDDLPSGIEDITDPAKL-ITEIENMRHRIIEIHQEMFNYNEHEVSKRWTFEEG 147
Query: 117 IRRPYFHVRPLNIGELENWHNYLDFIEREGDLSKVTKLLSYKGTFCSIFSCYRLTYSK 174
I+RPYFHV+PL +L+NW YL+F G +V L C+++ + + Y+K
Sbjct: 148 IKRPYFHVKPLEKAQLKNWKEYLEFEIENGTHERVVVLFERCVISCALYEEFWIKYAK 205
>ref|XP_537427.1| PREDICTED: similar to PRP39 pre-mRNA processing factor 39 homolog
[Canis familiaris]
Length = 712
Score = 84.0 bits (206), Expect = 2e-15
Identities = 56/178 (31%), Positives = 84/178 (46%), Gaps = 6/178 (3%)
Query: 2 QQDWARLAMIYTRILENPNQQLDRYFNSFKELASNRPLSELRTADE----AAAVAGVVSE 57
Q + + IY RIL P Q +F FKE N +L T ++ +A V
Sbjct: 262 QGNLREVTAIYDRILGIPTQLYSHHFQRFKEHVQNNLPRDLLTGEQFIQLRRELASVNGH 321
Query: 58 GIDQGVEGEVHPDGADNSPKPASAGLTEAEELE-KYIAIREEMYKKAKEFDSKIIGFETT 116
D G G+ P G ++ PA +TE E + + I I +EM+ + SK FE
Sbjct: 322 SGDDGPPGDDLPSGIEDITDPAKL-ITEIENMRHRIIEIHQEMFNYNEHEVSKRWTFEEG 380
Query: 117 IRRPYFHVRPLNIGELENWHNYLDFIEREGDLSKVTKLLSYKGTFCSIFSCYRLTYSK 174
I+RPYFHV+PL +L+NW YL+F G +V L C+++ + + Y+K
Sbjct: 381 IKRPYFHVKPLEKAQLKNWKEYLEFEIENGTHERVVVLFERCVISCALYEEFWIKYAK 438
>ref|XP_234238.3| PREDICTED: similar to PRP39 pre-mRNA processing factor 39 homolog
[Rattus norvegicus]
Length = 709
Score = 83.6 bits (205), Expect = 2e-15
Identities = 55/178 (30%), Positives = 84/178 (46%), Gaps = 6/178 (3%)
Query: 2 QQDWARLAMIYTRILENPNQQLDRYFNSFKELASNRPLSELRTADE----AAAVAGVVSE 57
Q + + +Y RIL P Q +F FKE N +L T ++ +A V
Sbjct: 261 QGNLREVTAVYDRILGIPTQLYSHHFQRFKEHVQNNLPRDLLTGEQFIQLRRELASVNGH 320
Query: 58 GIDQGVEGEVHPDGADNSPKPASAGLTEAEELE-KYIAIREEMYKKAKEFDSKIIGFETT 116
D G G+ P G ++ PA +TE E + + I I +EM+ + SK FE
Sbjct: 321 NGDDGPPGDDLPSGIEDITDPAKL-ITEIENMRHRIIEIHQEMFNYNEHEVSKRWTFEEG 379
Query: 117 IRRPYFHVRPLNIGELENWHNYLDFIEREGDLSKVTKLLSYKGTFCSIFSCYRLTYSK 174
I+RPYFHV+PL +L+NW YL+F G +V L C+++ + + Y+K
Sbjct: 380 IKRPYFHVKPLEKAQLKNWKEYLEFEIENGTHERVVVLFERCVISCALYEEFWIKYAK 437
>emb|CAD87784.1| novel protein similar to pre-mRNA processing proteins [Danio rerio]
gi|52218898|ref|NP_001004520.1| PRP39 pre-mRNA
processing factor 39 homolog [Danio rerio]
Length = 752
Score = 81.6 bits (200), Expect = 8e-15
Identities = 59/178 (33%), Positives = 84/178 (47%), Gaps = 10/178 (5%)
Query: 6 ARLAMIYTRILENPNQQLDRYFNSFKE-LASNRP---LSELRTADEAAAVAGVVSEGIDQ 61
A + IY R+L P Q ++F FK+ + SN P LSE +A D+
Sbjct: 294 ANVTAIYDRLLCIPTQLYSQHFQKFKDHVQSNNPKHFLSEEEFVSLRVELANANKPSGDE 353
Query: 62 GVE----GEVHPDGADNSPKPASAGLTEAEELE-KYIAIREEMYKKAKEFDSKIIGFETT 116
E GE P G ++ P PA +TE E + K I R+EM+ + SK FE
Sbjct: 354 DAETEAPGEELPPGTEDLPDPAKR-VTEIENMRHKVIETRQEMFNHNEHEVSKRWAFEEG 412
Query: 117 IRRPYFHVRPLNIGELENWHNYLDFIEREGDLSKVTKLLSYKGTFCSIFSCYRLTYSK 174
I+RPYFHV+ L +L NW YLDF G +V L C+++ + + Y+K
Sbjct: 413 IKRPYFHVKALEKTQLNNWREYLDFELENGTPERVVVLFERCLIACALYEEFWIKYAK 470
>gb|EAL89886.1| conserved hypothetical protein [Aspergillus fumigatus Af293]
Length = 591
Score = 74.3 bits (181), Expect = 1e-12
Identities = 48/173 (27%), Positives = 81/173 (46%), Gaps = 22/173 (12%)
Query: 7 RLAMIYTRILENPNQQLDRYFNSFKELASNRPLSELRTADEAAAVAGVVSEGIDQGVEGE 66
++ I R++ P Q RYF +++LA RPLSEL A+ + + + G+
Sbjct: 164 KIFAILGRVIHIPMHQYARYFERYRQLAQTRPLSELAPAETLSQFRAEL-----EAAAGQ 218
Query: 67 VHPDGADNSPKPASAGLTEAEELEKYIAIRE-----EMYKKAKEFDSKIIGFETTIRRPY 121
+ P G E+E+ + +R E++ K + +K +E+ I+RPY
Sbjct: 219 IPP------------GAKAEAEIERDLRLRVDAYHLEIFSKTQTETTKRWTYESEIKRPY 266
Query: 122 FHVRPLNIGELENWHNYLDFIEREGDLSKVTKLLSYKGTFCSIFSCYRLTYSK 174
FHV L+ G+L NW YLDF E EG ++ L C+ + + Y++
Sbjct: 267 FHVTELDEGQLTNWKKYLDFEEAEGSYPRIQFLYERCLVTCAHYDEFWQRYAR 319
>gb|EAA68105.1| hypothetical protein FG01244.1 [Gibberella zeae PH-1]
gi|46108724|ref|XP_381420.1| hypothetical protein
FG01244.1 [Gibberella zeae PH-1]
Length = 587
Score = 72.8 bits (177), Expect = 4e-12
Identities = 49/174 (28%), Positives = 84/174 (48%), Gaps = 14/174 (8%)
Query: 2 QQDWARLAMIYTRILENPNQQLDRYFNSFKELASNRPLSELRTADEAAAV-AGVVSEGID 60
Q+ ++ I+ RI+ P Q RY+ F+ L+ ++P++E+ A++ A A V +E +
Sbjct: 159 QEAQDKIYAIHARIIRIPMHQYARYYERFRSLSHSQPITEVVPAEDLARFRAEVEAENVA 218
Query: 61 QGVEGEVHPDGADNSPKPASAGLTEAEELEKYIAIREEMYKKAKEFDSKIIGFETTIRRP 120
G +PKP E + K A+ +++ + SK +E+ I+RP
Sbjct: 219 FG-----------GAPKPELE--IERDVRAKIDAMFYDIFTTTQTEVSKRWTYESEIKRP 265
Query: 121 YFHVRPLNIGELENWHNYLDFIEREGDLSKVTKLLSYKGTFCSIFSCYRLTYSK 174
YFHV L +L NW YLDF E EGD +++ L C+ + Y++
Sbjct: 266 YFHVTALEHKDLANWRKYLDFEESEGDYARIVALYERCLVTCAFYDDLWFRYAR 319
>gb|EAA64755.1| hypothetical protein AN1635.2 [Aspergillus nidulans FGSC A4]
gi|67522356|ref|XP_659239.1| hypothetical protein
AN1635_2 [Aspergillus nidulans FGSC A4]
gi|49087740|ref|XP_405772.1| hypothetical protein
AN1635.2 [Aspergillus nidulans FGSC A4]
Length = 588
Score = 72.0 bits (175), Expect = 6e-12
Identities = 47/179 (26%), Positives = 81/179 (44%), Gaps = 26/179 (14%)
Query: 3 QDWARLAMIYTRILENPNQQLDRYFNSFKELASNRPLSELRTAD-------EAAAVAGVV 55
+ + ++ I R++E P Q RYF +++LA RP++EL + + A AG+V
Sbjct: 160 EGYDKIFAILARVIEIPMHQYARYFERYRQLAQTRPVAELAPPNVISQFRADLDAAAGIV 219
Query: 56 SEGIDQGVEGEVHPDGADNSPKPASAGLTEAEELEKYIAIREEMYKKAKEFDSKIIGFET 115
+ G E E + + E++ K + +K +E+
Sbjct: 220 APGAKADAE-------------------IERDLRLRLDGYHLEIFSKTQTETTKRWTYES 260
Query: 116 TIRRPYFHVRPLNIGELENWHNYLDFIEREGDLSKVTKLLSYKGTFCSIFSCYRLTYSK 174
I+RPYFHV L+ G+L NW YLDF E EG +++ L C+ + + Y++
Sbjct: 261 EIKRPYFHVTELDEGQLANWRKYLDFEEAEGSYARIQFLYERCLVTCAHYDEFWQRYAR 319
>emb|CAG09750.1| unnamed protein product [Tetraodon nigroviridis]
Length = 509
Score = 70.9 bits (172), Expect = 1e-11
Identities = 47/151 (31%), Positives = 77/151 (50%), Gaps = 14/151 (9%)
Query: 2 QQDWARLAMIYTRILENPNQQLDRYFNSFKE-LASNRP--------LSELRTADEAAAVA 52
Q + + + R+L+ P Q + +++ FKE L SN P ELR A + A
Sbjct: 66 QGNMRKATAVLDRVLKVPTQLYNTHYDKFKEHLNSNEPKEVLSTEEYEELRAACRQSQRA 125
Query: 53 GVVSEGIDQGVEGEVHPDGADNSPKPASAGLTEAEEL--EKYIAIREEMYKKAKEFDSKI 110
+ ++ EG P G + P P E ++ EK + R+++Y++ +E K
Sbjct: 126 KRAEQSQEETQEG---PPGEEKPPTPEGPDTEELKQKIREKVLLQRDKVYQENEEQVRKR 182
Query: 111 IGFETTIRRPYFHVRPLNIGELENWHNYLDF 141
FE I+RPYFHV+PL+ +L+ WH+YLD+
Sbjct: 183 WHFEDAIKRPYFHVKPLDHLQLQAWHSYLDW 213
>emb|CAA19289.1| SPBC4B4.09 [Schizosaccharomyces pombe] gi|19113218|ref|NP_596426.1|
hypothetical protein SPBC4B4.09 [Schizosaccharomyces
pombe 972h-] gi|7491683|pir||T40481 hypothetical protein
SPBC4B4.09 - fission yeast (Schizosaccharomyces pombe)
Length = 612
Score = 70.1 bits (170), Expect = 2e-11
Identities = 43/161 (26%), Positives = 77/161 (47%), Gaps = 5/161 (3%)
Query: 14 RILENPNQQLDRYFNSFKELASNRPLSELRTADEAAAVAGVVSEGIDQGVEGEVHPDGAD 73
R++ P Q RYF F +++ ++P+ +L D A++ V+ +V G+
Sbjct: 169 RLIHIPLHQYARYFERFVQVSQSQPIQQLLPPDVLASIRADVTRE-----PAKVVSAGSK 223
Query: 74 NSPKPASAGLTEAEELEKYIAIREEMYKKAKEFDSKIIGFETTIRRPYFHVRPLNIGELE 133
E E + I ++++K + +K FE+ I+RPYFHV+ L+ +L
Sbjct: 224 QITVERGELEIEREMRARIYNIHLQIFQKVQLETAKRWTFESEIKRPYFHVKELDEAQLV 283
Query: 134 NWHNYLDFIEREGDLSKVTKLLSYKGTFCSIFSCYRLTYSK 174
NW YLDF E EGD ++ L C+++ + Y++
Sbjct: 284 NWRKYLDFEEVEGDFQRICHLYERCLITCALYDEFWFRYAR 324
>ref|XP_581849.1| PREDICTED: similar to PRP39 pre-mRNA processing factor 39 homolog,
partial [Bos taurus]
Length = 295
Score = 67.4 bits (163), Expect = 2e-10
Identities = 46/150 (30%), Positives = 71/150 (46%), Gaps = 6/150 (4%)
Query: 30 FKELASNRPLSELRTADE----AAAVAGVVSEGIDQGVEGEVHPDGADNSPKPASAGLTE 85
FK+ N +L T ++ +A V D G G+ P G ++ PA +TE
Sbjct: 1 FKDHVQNNLPRDLLTGEQFIQLRRELASVNGHSGDDGPPGDDLPSGIEDITDPAKL-ITE 59
Query: 86 AEELE-KYIAIREEMYKKAKEFDSKIIGFETTIRRPYFHVRPLNIGELENWHNYLDFIER 144
E + + I I +EM+ + SK FE I+RPYFHV+PL +L+NW YL+F
Sbjct: 60 IENMRHRIIEIHQEMFNYNEHEVSKRWTFEEGIKRPYFHVKPLEKAQLKNWKEYLEFEIE 119
Query: 145 EGDLSKVTKLLSYKGTFCSIFSCYRLTYSK 174
G +V L C+++ + + SK
Sbjct: 120 NGTHERVVVLFERCVISCALYEEFWIKVSK 149
>gb|AAH88586.1| LOC496864 protein [Xenopus tropicalis]
Length = 656
Score = 66.2 bits (160), Expect = 4e-10
Identities = 42/173 (24%), Positives = 82/173 (47%), Gaps = 2/173 (1%)
Query: 2 QQDWARLAMIYTRILENPNQQLDRYFNSFKE-LASNRPLSELRTADEAAAVAGVVSEGID 60
Q ++ +Y ++L P Q ++ FK+ ++++ P LR + + + +EG +
Sbjct: 177 QGNFREATAVYDQVLSIPTQLYRQHHERFKQHISAHAPHELLREEEFKWICSKIKAEGEN 236
Query: 61 QGVEGEVHPDGADN-SPKPASAGLTEAEELEKYIAIREEMYKKAKEFDSKIIGFETTIRR 119
+ E P G D +P + +++ + + IRE+++ + K FE I R
Sbjct: 237 DQIAAEDSPSGDDQENPVDVTDPELQSKVKAQVLIIREQLFLLNEAEVRKRWSFEEAITR 296
Query: 120 PYFHVRPLNIGELENWHNYLDFIEREGDLSKVTKLLSYKGTFCSIFSCYRLTY 172
PYFH PL+ +L+NW YLD +G ++ L C+++ + L+Y
Sbjct: 297 PYFHATPLDRTQLQNWRKYLDLEISQGRHERIVTLYERCLVACALYEEFWLSY 349
>dbj|BAA91318.1| unnamed protein product [Homo sapiens]
Length = 241
Score = 63.9 bits (154), Expect = 2e-09
Identities = 46/143 (32%), Positives = 66/143 (45%), Gaps = 6/143 (4%)
Query: 2 QQDWARLAMIYTRILENPNQQLDRYFNSFKELASNRPLSELRTADE----AAAVAGVVSE 57
Q + + IY RIL P Q +F FKE N +L T ++ +A V
Sbjct: 98 QGNLREVTAIYDRILGIPTQLCSHHFQRFKEHVQNNLPRDLLTGEQFIQLRRELASVNGH 157
Query: 58 GIDQGVEGEVHPDGADNSPKPASAGLTEAEELE-KYIAIREEMYKKAKEFDSKIIGFETT 116
D G G+ P G ++ PA +TE E + + I I +EM+ + SK FE
Sbjct: 158 SGDDGPPGDDLPSGIEDITDPAKL-ITEIENMRHRIIEIHQEMFNYNEHEVSKRWTFEEG 216
Query: 117 IRRPYFHVRPLNIGELENWHNYL 139
I+RPYFHV+PL +L+ L
Sbjct: 217 IKRPYFHVKPLEKAQLKKLERIL 239
>ref|XP_392380.2| PREDICTED: similar to novel protein similar to pre-mRNA processing
proteins [Apis mellifera]
Length = 946
Score = 63.2 bits (152), Expect = 3e-09
Identities = 44/168 (26%), Positives = 79/168 (46%), Gaps = 10/168 (5%)
Query: 6 ARLAMIYTRILENPNQQLDRYFNSFKELASNRPLSELRTADEAAAVAGVVSEGIDQGVEG 65
+R+ +Y R+L P +F++F+E S+ + + D+ A+ V + +
Sbjct: 452 SRVTALYDRLLCTPTLGYISHFDAFQEFVSSNLPNRILNVDDFLALRAEVKALLKS--DD 509
Query: 66 EVHPDGADNSP---KPASAGLTEAEE-----LEKYIAIREEMYKKAKEFDSKIIGFETTI 117
AD++P +P L +E EK I+ R +M+K + +E I
Sbjct: 510 TTSTSAADDAPPGEEPPPHELPPTDEETRAIREKIISSRRKMHKANINAVAARWSYEEGI 569
Query: 118 RRPYFHVRPLNIGELENWHNYLDFIEREGDLSKVTKLLSYKGTFCSIF 165
+RPYFHV+PL +L+NW YLDF + D +++ L C+++
Sbjct: 570 KRPYFHVKPLERCQLKNWKEYLDFEIEQKDQNRIIILFERCLIACALY 617
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.316 0.135 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 297,649,664
Number of Sequences: 2540612
Number of extensions: 12427263
Number of successful extensions: 39317
Number of sequences better than 10.0: 63
Number of HSP's better than 10.0 without gapping: 36
Number of HSP's successfully gapped in prelim test: 27
Number of HSP's that attempted gapping in prelim test: 39243
Number of HSP's gapped (non-prelim): 79
length of query: 174
length of database: 863,360,394
effective HSP length: 119
effective length of query: 55
effective length of database: 561,027,566
effective search space: 30856516130
effective search space used: 30856516130
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 70 (31.6 bits)
Medicago: description of AC148607.15


