

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148532.13 + phase: 0
(303 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
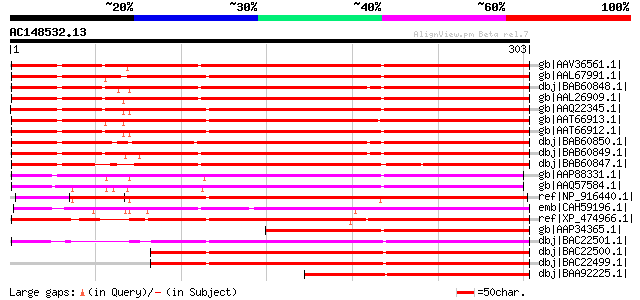
Score E
Sequences producing significant alignments: (bits) Value
gb|AAV36561.1| RD22-like protein [Vitis vinifera] 390 e-107
gb|AAL67991.1| dehydration-induced protein RD22-like protein [Go... 375 e-102
dbj|BAB60848.1| BURP domain-containing protein [Bruguiera gymnor... 375 e-102
gb|AAL26909.1| dehydration-responsive protein RD22 [Prunus persica] 371 e-101
gb|AAQ22345.1| BURP domain-containing protein [Gossypium hirsutum] 365 1e-99
gb|AAT66913.1| dehydration-induced protein RD22-like protein 2 [... 364 2e-99
gb|AAT66912.1| dehydration-induced protein RD22-like protein 1 [... 360 2e-98
dbj|BAB60850.1| BURP domain-containing protein [Bruguiera gymnor... 360 3e-98
dbj|BAB60849.1| BURP domain-containing protein [Bruguiera gymnor... 356 5e-97
dbj|BAB60847.1| BURP domain-containing protein [Bruguiera gymnor... 318 1e-85
gb|AAP88331.1| At5g25610/T14C9_150 [Arabidopsis thaliana] gi|391... 308 2e-82
gb|AAQ57584.1| BURP domain-containing protein [Brassica napus] 300 3e-80
ref|NP_916440.1| putative dehydration-responsive protein RD22 pr... 290 5e-77
emb|CAH59196.1| BURP-domain containing protein [Plantago major] 288 1e-76
ref|XP_474966.1| OSJNBa0036B17.11 [Oryza sativa (japonica cultiv... 241 2e-62
gb|AAP34365.1| putative dehydration-induced protein [Gossypium b... 237 4e-61
dbj|BAC22501.1| resistant specific protein-3 [Vigna radiata] 232 9e-60
dbj|BAC22500.1| resistant specific protein-2 [Vigna radiata] 226 5e-58
dbj|BAC22499.1| resistant specific protein-1(8) [Vigna radiata] ... 223 4e-57
dbj|BAA92225.1| similar to the BURP domain [Vigna unguiculata] 221 3e-56
>gb|AAV36561.1| RD22-like protein [Vitis vinifera]
Length = 364
Score = 390 bits (1003), Expect = e-107
Identities = 206/347 (59%), Positives = 240/347 (68%), Gaps = 50/347 (14%)
Query: 2 LPPQLYWKSMLPNSPMPKAITNLLHPAGYWSEEKGTWVDVGKGGVDVGVRKGYYEGGGTD 61
LP ++YW S+LPN+PMPKAI ++L P EEKGT V VGKGGV+V KG GGGT
Sbjct: 23 LPTKVYWNSVLPNTPMPKAIRDILRPD--LMEEKGTSVSVGKGGVNVHAGKGK-SGGGTT 79
Query: 62 VNVGVG---------------------------------------------RSPFIYNYA 76
V VG G +SPF Y YA
Sbjct: 80 VGVGKGTGVNVHAGKGKPGGGTTVGVGKGGVSVNAGHKGKHVYVGVGKGKSKSPFDYKYA 139
Query: 77 ASETQLHDKPNVALFFLEKDLHHGTKLNLQFSKTTSNAATFLPRQVANSIPFSSNKMEYI 136
A+E QLHD PNVALFF EK++ GTK+ L F + +N ATFLPRQVANSIPFSS K I
Sbjct: 140 ATEDQLHDDPNVALFFFEKNMQPGTKMELHFIRD-ANLATFLPRQVANSIPFSSKKFPEI 198
Query: 137 INKLNIKKGSKGVQIVKNTISECEEQGIKGEEKVCVTSLESMVDFTTSKLGKNVEAVSTE 196
+N+ +IK S+ + +KNTI ECEE GIKGEEK C TSLESMVDF+TSKLGK V+ +STE
Sbjct: 199 LNEFSIKPESEEAETIKNTIRECEEPGIKGEEKYCATSLESMVDFSTSKLGKGVQMISTE 258
Query: 197 VNKESNLQQYTIASGVKKLGEKNKAVVCHKENYPYAVFYCHKTDTTKAYSVPLEGADGSR 256
V KE+ QQYTI +GVKKL +KAVVCHK++YPYAVFYCHKT TT+AY VPL GADGS+
Sbjct: 259 VEKETPEQQYTITTGVKKLAG-DKAVVCHKQSYPYAVFYCHKTQTTRAYMVPLVGADGSK 317
Query: 257 VKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVWIRK 303
VKA+AVCHTDTS WNPKHLAFQVLKV+PGTVP+CH LPEDHVVW+ K
Sbjct: 318 VKAVAVCHTDTSAWNPKHLAFQVLKVKPGTVPICHFLPEDHVVWVPK 364
>gb|AAL67991.1| dehydration-induced protein RD22-like protein [Gossypium hirsutum]
Length = 335
Score = 375 bits (962), Expect = e-102
Identities = 187/321 (58%), Positives = 234/321 (72%), Gaps = 26/321 (8%)
Query: 2 LPPQLYWKSMLPNSPMPKAITNLLHPAGYWSEEKGTWVDVGKGGVDVGVRKGY------- 54
L P+ YW LPN+PMPKA+ +LHP EEK T V+VG GGV+V KG
Sbjct: 22 LSPEQYWSYKLPNTPMPKAVKEILHPE--LMEEKSTSVNVGGGGVNVNTGKGKPGGDTHV 79
Query: 55 ------------YEGGGTDVNVGVGRSPFIYNYAASETQLHDKPNVALFFLEKDLHHGTK 102
GGGT VNVG PF Y YAASETQ+H+ PNVALFFLEKD+H G
Sbjct: 80 NVGGKGVGVNTGKPGGGTHVNVG---DPFNYLYAASETQIHEDPNVALFFLEKDMHPGAT 136
Query: 103 LNLQFSKTTSNAATFLPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEEQ 162
++L F++ T +A FLP Q A IPFSS+K+ I NK ++K GS +++KNTI ECE+
Sbjct: 137 MSLHFTENTEKSA-FLPYQTAQKIPFSSDKLPEIFNKFSVKPGSLKAEMMKNTIKECEQP 195
Query: 163 GIKGEEKVCVTSLESMVDFTTSKLGKNVEAVSTEVNKESNLQQYTIASGVKKLGEKNKAV 222
I+GEEK C TSLESM+D++ SKLGK +AVSTEV K++ +Q+YTIA+GV+K+ + +KAV
Sbjct: 196 AIEGEEKYCATSLESMIDYSISKLGKVDQAVSTEVEKQTPMQKYTIAAGVQKMTD-DKAV 254
Query: 223 VCHKENYPYAVFYCHKTDTTKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKV 282
VCHK+NY YAVFYCHK++TT+AY VPLEGADG++ KA+AVCHTDTS WNPKHLAFQVLKV
Sbjct: 255 VCHKQNYAYAVFYCHKSETTRAYMVPLEGADGTKAKAVAVCHTDTSAWNPKHLAFQVLKV 314
Query: 283 QPGTVPVCHLLPEDHVVWIRK 303
+PGT+PVCH LP DH+VW+ K
Sbjct: 315 EPGTIPVCHFLPRDHIVWVPK 335
>dbj|BAB60848.1| BURP domain-containing protein [Bruguiera gymnorrhiza]
Length = 346
Score = 375 bits (962), Expect = e-102
Identities = 194/339 (57%), Positives = 241/339 (70%), Gaps = 43/339 (12%)
Query: 2 LPPQLYWKSMLPNSPMPKAITNLLHPAGYWSEEKGTWVDVGKGGVDVGVRKGYYEGGGTD 61
L P+ YW S+LPN+PMPKAI +LLHP E+KGT V VGKGGV+V KG GGGT
Sbjct: 14 LAPEAYWNSVLPNTPMPKAIKDLLHPD--MLEDKGTSVAVGKGGVNVDAGKGK-PGGGTH 70
Query: 62 V-------------------NVGVGR------------------SPFIYNYAASETQLHD 84
V NVGVG+ SPF+Y YAASE Q+HD
Sbjct: 71 VGVGGKGVGVDAGTPGGGGTNVGVGKGGVSVNTGHKGTPVNVNVSPFVYRYAASEDQIHD 130
Query: 85 KPNVALFFLEKDLHHGTKLNLQFSKTTSNAATFLPRQVANSIPFSSNKMEYIINKLNIKK 144
PNVALFFLEKD+ G +NL FS++ SN ATFLPR VA+SIPFSSNK++ + ++K
Sbjct: 131 NPNVALFFLEKDMKPGKSMNLDFSES-SNTATFLPRHVADSIPFSSNKLQDAFREFSVKP 189
Query: 145 GSKGVQIVKNTISECEEQGIKGEEKVCVTSLESMVDFTTSKLGKNVEAVSTEVNKESNLQ 204
GS +I++NT+ ECE GI+GEEK C TSLESMVDF+TSKLGK+++A+STEV K++ Q
Sbjct: 190 GSMEAEIMENTVKECENPGIEGEEKYCATSLESMVDFSTSKLGKDIQAISTEVEKQTGRQ 249
Query: 205 QYTIASGVKKLGEKNKAVVCHKENYPYAVFYCHKTDTTKAYSVPLEGADGSRVKAIAVCH 264
QYTIA GVKK +K+VVCHK+NYPYAVFYCH T +T+AY V L+GAD +R++A+A+CH
Sbjct: 250 QYTIA-GVKKTAG-DKSVVCHKQNYPYAVFYCHSTQSTRAYLVQLKGADRTRLQAVAICH 307
Query: 265 TDTSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVWIRK 303
T+TS WNPKHLAFQVL V+PGTVP+CH LP+ HVVW+ K
Sbjct: 308 TNTSAWNPKHLAFQVLAVKPGTVPICHFLPQSHVVWVPK 346
>gb|AAL26909.1| dehydration-responsive protein RD22 [Prunus persica]
Length = 349
Score = 371 bits (953), Expect = e-101
Identities = 194/344 (56%), Positives = 232/344 (67%), Gaps = 47/344 (13%)
Query: 2 LPPQLYWKSMLPNSPMPKAITNLLHPAGYWSEEKGTWVDVGKGGVDVGVRKGYYEGGGTD 61
LPPQLYW S+LPN+ MPK+I+ LL P ++EEKGT V VGKGGV+V KG GGGT+
Sbjct: 8 LPPQLYWNSVLPNTQMPKSISELLQPD--FAEEKGTTVGVGKGGVNVQAGKGK-PGGGTN 64
Query: 62 VNVG------------------------------------------VGRSPFIYNYAASE 79
VNVG G +PF Y YAA+E
Sbjct: 65 VNVGGKGGGVGVTTGKPGGGTNVGVGKGGVSVNTGHKGKPVYVGVKPGPNPFNYVYAATE 124
Query: 80 TQLHDKPNVALFFLEKDLHHGTKLNLQFSKTTSNAATFLPRQVANSIPFSSNKMEYIINK 139
TQLH N A+FFLEKD+ GT + L FS SN A F+PR+ A+SIPFSSNK+ I ++
Sbjct: 125 TQLHGNRNAAIFFLEKDIRPGTSMTLTFSGN-SNTAAFVPRKTADSIPFSSNKLPEIFSQ 183
Query: 140 LNIKKGSKGVQIVKNTISECEEQGIKGEEKVCVTSLESMVDFTTSKLGKNVEAVSTEVNK 199
++K S I+K TI ECE GI+GEEK C TSLESMVDF+TSKLG N++A+STE K
Sbjct: 184 FSVKPESVEADIIKGTIEECESSGIRGEEKYCATSLESMVDFSTSKLGGNIQAISTEAEK 243
Query: 200 ESNLQQYTIASGVKKLGEKNKAVVCHKENYPYAVFYCHKTDTTKAYSVPLEGADGSRVKA 259
+ LQ+YTI GVKKL K+VVCHK+ YPYAVFYCH T TT+AY VPL+GADG KA
Sbjct: 244 GATLQKYTITPGVKKLAA-GKSVVCHKQTYPYAVFYCHATKTTRAYVVPLKGADGLEAKA 302
Query: 260 IAVCHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVWIRK 303
+AVCHTDTSEWNPKHLAFQVLKV+PGTVPVCH LP+DH+VW+ K
Sbjct: 303 VAVCHTDTSEWNPKHLAFQVLKVKPGTVPVCHFLPKDHIVWVPK 346
>gb|AAQ22345.1| BURP domain-containing protein [Gossypium hirsutum]
Length = 335
Score = 365 bits (936), Expect = 1e-99
Identities = 184/320 (57%), Positives = 232/320 (72%), Gaps = 22/320 (6%)
Query: 1 MLPPQLYWKSMLPNSPMPKAITNLLHPAGYWSEEKGTWVDVGKGGVDVGVRKGYYEGGGT 60
+L P+ YW LPN PMPKA+ +LHP EEK T V+VG GGV+V KG GG T
Sbjct: 21 VLSPEQYWSYKLPNIPMPKAVKEILHPE--LMEEKSTSVNVGGGGVNVNTGKGK-PGGDT 77
Query: 61 DVNVG-------VGR----------SPFIYNYAASETQLHDKPNVALFFLEKDLHHGTKL 103
VNVG G+ PF Y YAASETQ+H+ PNVALFFLEKD+H G +
Sbjct: 78 HVNVGGKGVGVNTGKPGGGTHVNDPDPFNYLYAASETQIHEDPNVALFFLEKDMHPGATM 137
Query: 104 NLQFSKTTSNAATFLPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEEQG 163
+L F + T +A FLP Q A IPFSS+K+ I NK ++K GS +++KNTI ECE+
Sbjct: 138 SLHFIENTEKSA-FLPYQTAQKIPFSSDKLPEIFNKFSVKPGSVKAEMMKNTIKECEQPA 196
Query: 164 IKGEEKVCVTSLESMVDFTTSKLGKNVEAVSTEVNKESNLQQYTIASGVKKLGEKNKAVV 223
I+GEEK C TSLESM+D++ SKLGK +AVSTEV K++ +Q+YTIA+GV+K+ + +KAVV
Sbjct: 197 IEGEEKYCATSLESMIDYSISKLGKVDQAVSTEVEKQTPMQKYTIAAGVQKMTD-DKAVV 255
Query: 224 CHKENYPYAVFYCHKTDTTKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQ 283
CHK+NY YAVFYCHK++TT+AY VPLEGA G++ +A+AVCHTDTS WNPKHLAFQ LKV+
Sbjct: 256 CHKQNYAYAVFYCHKSETTRAYMVPLEGAGGTKAQALAVCHTDTSAWNPKHLAFQFLKVE 315
Query: 284 PGTVPVCHLLPEDHVVWIRK 303
PGT+PVCH LP DH+VW+ K
Sbjct: 316 PGTIPVCHFLPRDHIVWVPK 335
>gb|AAT66913.1| dehydration-induced protein RD22-like protein 2 [Gossypium
arboreum]
Length = 376
Score = 364 bits (934), Expect = 2e-99
Identities = 189/359 (52%), Positives = 233/359 (64%), Gaps = 61/359 (16%)
Query: 2 LPPQLYWKSMLPNSPMPKAITNLLHPAGYWSEEKGTWVDVGKGGVDVGVRKGY------- 54
L P+ YW LPN+PMPKA+ +LHP EEK T V+VG GGV+V KG
Sbjct: 22 LSPEQYWSYKLPNTPMPKAVKEILHPE--LMEEKSTSVNVGGGGVNVNTGKGKPAGGTHV 79
Query: 55 --------------------------------YEGGGTDVNVG----------------- 65
GGGT VNVG
Sbjct: 80 NVGRKGVGVNTGKPGGGTHVNVGGKGVGVNTGKPGGGTHVNVGGKGGGVSVHTGHKGKPV 139
Query: 66 -VGRSPFIYNYAASETQLHDKPNVALFFLEKDLHHGTKLNLQFSKTTSNAATFLPRQVAN 124
V SPF+Y YAASETQ+HD PNVALFFLEKDLH G ++L F++ T +A FLP Q A
Sbjct: 140 NVNVSPFLYQYAASETQIHDDPNVALFFLEKDLHPGATMSLHFTENTEKSA-FLPYQTAQ 198
Query: 125 SIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEEQGIKGEEKVCVTSLESMVDFTTS 184
IPFSSN++ I NK ++K GS +++KNTI ECE+ I+GEEK C TSLESM+D++ S
Sbjct: 199 KIPFSSNELPEIFNKFSVKPGSVKAEMMKNTIKECEQPAIEGEEKYCATSLESMIDYSIS 258
Query: 185 KLGKNVEAVSTEVNKESNLQQYTIASGVKKLGEKNKAVVCHKENYPYAVFYCHKTDTTKA 244
KLGK +AVSTEV K++ +YTI +GV+K+ +KAVVCHK+NY YAVFYCHK++TT+A
Sbjct: 259 KLGKVDQAVSTEVEKQTPTHKYTITAGVQKM-TNDKAVVCHKQNYAYAVFYCHKSETTRA 317
Query: 245 YSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVWIRK 303
Y VPLEGADG++ KA+AVCHTDTS WNPKHLAFQVLKV+PGT+PVCH LP DH+VW+ K
Sbjct: 318 YMVPLEGADGTKAKAVAVCHTDTSAWNPKHLAFQVLKVEPGTIPVCHFLPRDHIVWVPK 376
>gb|AAT66912.1| dehydration-induced protein RD22-like protein 1 [Gossypium
arboreum]
Length = 335
Score = 360 bits (925), Expect = 2e-98
Identities = 183/319 (57%), Positives = 230/319 (71%), Gaps = 22/319 (6%)
Query: 2 LPPQLYWKSMLPNSPMPKAITNLLHPAGYWSEEKGTWVDVGKGGVDVGVRKGYYEGGGTD 61
L P+ YW LPN+PMPKA+ +LHP EEK T V+VG GGV+V KG GG T
Sbjct: 22 LSPEQYWSYKLPNTPMPKAVKEILHPE--LMEEKSTSVNVGGGGVNVNTGKGK-PGGDTH 78
Query: 62 VNVG-------VGR----------SPFIYNYAASETQLHDKPNVALFFLEKDLHHGTKLN 104
VNVG G+ PF Y YAASETQ+H+ PNVALFFLEKD+H G ++
Sbjct: 79 VNVGGKGVGVNTGKPGGGTHVNDPDPFNYLYAASETQIHEDPNVALFFLEKDMHPGATMS 138
Query: 105 LQFSKTTSNAATFLPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEEQGI 164
L F + T +A FLP Q A FSS+K+ I NK ++K GS +++KNTI ECE+ I
Sbjct: 139 LHFIENTEKSA-FLPYQTAPKNTFSSDKLPEIFNKFSVKPGSVKAEMMKNTIKECEQPAI 197
Query: 165 KGEEKVCVTSLESMVDFTTSKLGKNVEAVSTEVNKESNLQQYTIASGVKKLGEKNKAVVC 224
+GEEK C TSLESM+D++ SKLGK +AVSTEV K++ +Q+YTIA+GV+K+ + +KAVVC
Sbjct: 198 EGEEKYCATSLESMIDYSISKLGKVDQAVSTEVEKQTPMQKYTIAAGVQKMTD-DKAVVC 256
Query: 225 HKENYPYAVFYCHKTDTTKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQP 284
HK+NY YAVFYCHK++TT+AY VPLEGA G++ KA+AVCHTDTS WNPKHLAFQ LKV+P
Sbjct: 257 HKQNYAYAVFYCHKSETTRAYMVPLEGAGGTKAKALAVCHTDTSAWNPKHLAFQFLKVEP 316
Query: 285 GTVPVCHLLPEDHVVWIRK 303
GT+PVCH LP DH+VW+ K
Sbjct: 317 GTIPVCHFLPRDHIVWVPK 335
>dbj|BAB60850.1| BURP domain-containing protein [Bruguiera gymnorrhiza]
Length = 307
Score = 360 bits (924), Expect = 3e-98
Identities = 183/303 (60%), Positives = 229/303 (75%), Gaps = 10/303 (3%)
Query: 2 LPPQLYWKSMLPNSPMPKAITNLLHPAGYWSEEKGTWVDVGKGGVDVGVRKGYYEGGGTD 61
L P+ YW S+LPN+PMPKAI +LLHP E+KGT V VGKGG +V KG GGGT
Sbjct: 14 LAPEAYWNSVLPNTPMPKAIKDLLHPD--MLEDKGTSVAVGKGGENVDAGKGKPGGGGT- 70
Query: 62 VNVGVGRSPFIYN-YAASETQLHDKPNVALFFLEKDLHHGTKLNLQFSKTTSNAATFLPR 120
N G+ R+ + N YAASE Q+HD PNVALFFL+KDL G +NL F +SN ATFLPR
Sbjct: 71 -NFGIDRA--VSNIYAASEDQIHDNPNVALFFLQKDLKPGKSMNLDFP-ASSNTATFLPR 126
Query: 121 QVANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEEQGIKGEEKVCVTSLESMVD 180
VA+SIPFSSNK++ + ++K GS +I++NT+ ECE GI+GEEK C TSLESMVD
Sbjct: 127 HVADSIPFSSNKLQDAFREFSVKPGSLEAEIMENTVKECENPGIEGEEKYCATSLESMVD 186
Query: 181 FTTSKLGKNVEAVSTEVNKESNLQQYTIASGVKKLGEKNKAVVCHKENYPYAVFYCHKTD 240
F+TSKLGK+V+A+STEV K++ Q+YTIA GVKK+ +K VVCHK+NYPYAVFYCH
Sbjct: 187 FSTSKLGKDVQAISTEVEKQTGRQKYTIA-GVKKIAG-DKCVVCHKQNYPYAVFYCHSIQ 244
Query: 241 TTKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVW 300
+T+AY V L+G D +R++A+A+CHT+TS WNPKH AFQVL V+PGTVP CH LP+ HVVW
Sbjct: 245 STRAYLVQLKGVDRTRLQAVAICHTNTSAWNPKHPAFQVLAVKPGTVPTCHFLPQSHVVW 304
Query: 301 IRK 303
+ K
Sbjct: 305 VPK 307
>dbj|BAB60849.1| BURP domain-containing protein [Bruguiera gymnorrhiza]
Length = 336
Score = 356 bits (913), Expect = 5e-97
Identities = 183/320 (57%), Positives = 233/320 (72%), Gaps = 24/320 (7%)
Query: 2 LPPQLYWKSMLPNSPMPKAITNLLHPAGYWSEEKGTWVDVGKGGVDVGVRKGYYEGGGTD 61
L P+ YW S+LPN+PMPKAI +LLHP E+KGT V VGKGGV+V KG GGGT
Sbjct: 23 LAPEAYWNSVLPNTPMPKAIKDLLHPD--MLEDKGTSVAVGKGGVNVDAGKGK-PGGGTH 79
Query: 62 VNVGV------------GRSPFIYN------YAASETQLHDKPNVALFFLEKDLHHGTKL 103
V VG G + F ++ YAASE Q+HD PN+ALFFL+KDL G +
Sbjct: 80 VGVGGKGVGVDAGKPGGGGTNFGFDRAVSNIYAASEDQIHDNPNLALFFLQKDLKPGKSM 139
Query: 104 NLQFSKTTSNAATFLPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEEQG 163
NL F ++ SN ATFLPR VA+SIPFSSNK++ + ++K GS +I++NT+ ECE G
Sbjct: 140 NLDFPES-SNTATFLPRHVADSIPFSSNKLQDAFREFSVKPGSLEAEIMENTVKECENPG 198
Query: 164 IKGEEKVCVTSLESMVDFTTSKLGKNVEAVSTEVNKESNLQQYTIASGVKKLGEKNKAVV 223
I+GEEK C TSLESMVDF+TSKLGK+++A+STEV K++ Q+YTIA GVKK+ +K VV
Sbjct: 199 IEGEEKYCATSLESMVDFSTSKLGKDIQAISTEVEKQTGRQKYTIA-GVKKIAG-DKCVV 256
Query: 224 CHKENYPYAVFYCHKTDTTKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQ 283
CHK+NYPYAVFYCH +T+AY V L+G D +R++A+A+CHT+TS WNPKH AFQVL V+
Sbjct: 257 CHKQNYPYAVFYCHSIQSTRAYLVQLKGVDRTRLQAVAICHTNTSAWNPKHPAFQVLAVK 316
Query: 284 PGTVPVCHLLPEDHVVWIRK 303
PGTVP+CH LP+ HVVW+ K
Sbjct: 317 PGTVPICHFLPQSHVVWVPK 336
>dbj|BAB60847.1| BURP domain-containing protein [Bruguiera gymnorrhiza]
Length = 292
Score = 318 bits (815), Expect = 1e-85
Identities = 166/302 (54%), Positives = 213/302 (69%), Gaps = 23/302 (7%)
Query: 2 LPPQLYWKSMLPNSPMPKAITNLLHPAGYWSEEKGTWVDVGKGGVDVGVRKGYYEGGGTD 61
L P+ YW S+LPN+PMPKAI LLHP E+KGT V VGKG V+V G+
Sbjct: 14 LAPEAYWNSVLPNTPMPKAIKVLLHPD--MLEDKGTSVAVGKGAVNVDA--------GSF 63
Query: 62 VNVGVGRSPFIYNYAASETQLHDKPNVALFFLEKDLHHGTKLNLQFSKTTSNAATFLPRQ 121
+N Y AASE Q+ D N+ +FFLEKDL G +NL F ++ SN ATFLPR
Sbjct: 64 IN---------YYNAASEDQIRDNSNITIFFLEKDLKPGKSMNLYFFES-SNTATFLPRH 113
Query: 122 VANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEEQGIKGEEKVCVTSLESMVDF 181
VA+S+PFSSNK++ + ++K GS + +++TI ECE I+GEEK CVTSLES+VDF
Sbjct: 114 VADSMPFSSNKLQEAFREFSVKPGSLEAETMEDTIKECENPRIEGEEKYCVTSLESIVDF 173
Query: 182 TTSKLGKNVEAVSTEVNKESNLQQYTIASGVKKLGEKNKAVVCHKENYPYAVFYCHKTDT 241
++SKLGK+++A+STEV K++ Q+YTIA K G+ K+VVCHK+NYPYAVFYCH T
Sbjct: 174 SSSKLGKDIQAISTEVEKQTGRQKYTIAVVKKMAGD--KSVVCHKQNYPYAVFYCHTTQ- 230
Query: 242 TKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVWI 301
++AY V L+GAD S +KA+A CHT+TS WNPKH+AFQVL V+PGTVPVCH L + HVVW
Sbjct: 231 SRAYLVQLKGADRSELKAVATCHTNTSAWNPKHMAFQVLAVKPGTVPVCHFLTQSHVVWA 290
Query: 302 RK 303
K
Sbjct: 291 PK 292
>gb|AAP88331.1| At5g25610/T14C9_150 [Arabidopsis thaliana]
gi|391608|dbj|BAA01546.1| rd22 [Arabidopsis thaliana]
gi|19699081|gb|AAL90908.1| AT5g25610/T14C9_150
[Arabidopsis thaliana] gi|15239478|ref|NP_197943.1|
dehydration-responsive protein (RD22) [Arabidopsis
thaliana] gi|16974546|gb|AAL31189.1| AT5g25610/T14C9_150
[Arabidopsis thaliana] gi|479589|pir||S34823
dehydration-induced protein RD22 - Arabidopsis thaliana
gi|1172874|sp|Q08298|RD22_ARATH Dehydration-responsive
protein RD22 precursor gi|447134|prf||1913421A rd22 gene
Length = 392
Score = 308 bits (788), Expect = 2e-82
Identities = 171/370 (46%), Positives = 220/370 (59%), Gaps = 74/370 (20%)
Query: 2 LPPQLYWKSMLPNSPMPKAITNLLHPAGYWSEEKGTWVDVGKGGVDVGVRKGYY------ 55
L P+ YW + LPN+P+P ++ NLL +++EK T V VGKGGV+V KG
Sbjct: 23 LTPERYWSTALPNTPIPNSLHNLL--TFDFTDEKSTNVQVGKGGVNVNTHKGKTGSGTAV 80
Query: 56 ---------------EGGGTDVNVGVGR-------------------------------- 68
GGGT V+VG G+
Sbjct: 81 NVGKGGVRVDTGKGKPGGGTHVSVGSGKGHGGGVAVHTGKPGKRTDVGVGKGGVTVHTRH 140
Query: 69 -------------SPFIYNYAASETQLHDKPNVALFFLEKDLHHGTKLNLQFSKTTS--N 113
+PF+YNYAA ETQLHD PN ALFFLEKDL G ++N++F+
Sbjct: 141 KGRPIYVGVKPGANPFVYNYAAKETQLHDDPNAALFFLEKDLVRGKEMNVRFNAEDGYGG 200
Query: 114 AATFLPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEEQGIKGEEKVCVT 173
FLPR A ++PF S K + + +++ GS+ +++K TI ECE + + GEEK C T
Sbjct: 201 KTAFLPRGEAETVPFGSEKFSETLKRFSVEAGSEEAEMMKKTIEECEARKVSGEEKYCAT 260
Query: 174 SLESMVDFTTSKLGK-NVEAVSTEV-NKESNLQQYTIAS-GVKKLGEKNKAVVCHKENYP 230
SLESMVDF+ SKLGK +V AVSTEV K + +Q+Y IA+ GVKKL + +K+VVCHK+ YP
Sbjct: 261 SLESMVDFSVSKLGKYHVRAVSTEVAKKNAPMQKYKIAAAGVKKLSD-DKSVVCHKQKYP 319
Query: 231 YAVFYCHKTDTTKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVC 290
+AVFYCHK T Y+VPLEG +G R KA+AVCH +TS WNP HLAF+VLKV+PGTVPVC
Sbjct: 320 FAVFYCHKAMMTTVYAVPLEGENGMRAKAVAVCHKNTSAWNPNHLAFKVLKVKPGTVPVC 379
Query: 291 HLLPEDHVVW 300
H LPE HVVW
Sbjct: 380 HFLPETHVVW 389
>gb|AAQ57584.1| BURP domain-containing protein [Brassica napus]
Length = 387
Score = 300 bits (769), Expect = 3e-80
Identities = 174/366 (47%), Positives = 217/366 (58%), Gaps = 69/366 (18%)
Query: 2 LPPQLYWKSMLPNSPMPKAITNLLHPAGYWSEEK---------------------GTWVD 40
L P+ YW S LPN+P+P ++ +L + + EE GT V+
Sbjct: 21 LTPERYWNSALPNTPIPNSLRHLF-TSDFSDEESTNVQVGKGGVNVYTGKGKPGGGTAVN 79
Query: 41 VGKGGVDVGVRKGYY----------EGGG-----------TDVNVGVG------------ 67
VGKGGV V KG GGG TDV VG G
Sbjct: 80 VGKGGVHVNTGKGKGTHVSVSGGKGHGGGVGVHTGKPGKRTDVGVGKGGVIVHTRHKGKP 139
Query: 68 --------RSPFIYNYAASETQLHDKPNVALFFLEKDLHHGTKLNLQFSKTT--SNAATF 117
+PF YNYAASETQLHD P ALFFLEKD+ G +NL+F+ + F
Sbjct: 140 VYVGVKPGHNPFAYNYAASETQLHDDPKAALFFLEKDMVPGKAMNLRFNAEDGYNGKTAF 199
Query: 118 LPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEEQGIKGEEKVCVTSLES 177
LPR A ++PF S K I+N ++K GS +++K TI ECE + + GEEK C TSLES
Sbjct: 200 LPRGEAETVPFGSEKSSEILNTFSVKPGSGEAEMMKKTIEECEAKRVGGEEKYCATSLES 259
Query: 178 MVDFTTSKLGKN-VEAVSTEV-NKESNLQQYTIAS-GVKKLGEKNKAVVCHKENYPYAVF 234
MVDF+ SKLGK+ V AVSTEV K + +Q+Y IA+ GVKKL + +K+VVCHK+ YP+AVF
Sbjct: 260 MVDFSVSKLGKDHVRAVSTEVAEKNAPMQKYRIAAAGVKKLSD-DKSVVCHKQKYPFAVF 318
Query: 235 YCHKTDTTKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLP 294
YCHK T Y+VPLEG +G R KA+AVCH +TS WNP HLAF+VLKV+PG+VPVCH LP
Sbjct: 319 YCHKAMMTSVYAVPLEGENGLRAKAVAVCHKNTSAWNPNHLAFKVLKVKPGSVPVCHFLP 378
Query: 295 EDHVVW 300
E HVVW
Sbjct: 379 ETHVVW 384
>ref|NP_916440.1| putative dehydration-responsive protein RD22 precursor [Oryza
sativa (japonica cultivar-group)]
gi|20161002|dbj|BAB89935.1| putative BURP
domain-containing protein [Oryza sativa (japonica
cultivar-group)] gi|15624018|dbj|BAB68072.1| putative
BURP domain-containing protein [Oryza sativa (japonica
cultivar-group)]
Length = 429
Score = 290 bits (741), Expect = 5e-77
Identities = 151/287 (52%), Positives = 191/287 (65%), Gaps = 21/287 (7%)
Query: 36 GTWVDVGKGGVDVGVRKGYYEGGGTDVNVGVGR----------------SPFIYNYAASE 79
GT V VGKGGV V V+ GY + GGT V VG G +PFIYNYAA+E
Sbjct: 142 GTTVGVGKGGVGVNVQPGYGKPGGTTVGVGKGGVGVNVKPRGKPVHVNVAPFIYNYAATE 201
Query: 80 TQLHDKPNVALFFLEKDLHHGTKLNLQFSKTTSNAATFLPRQVANSIPFSSNKMEYIINK 139
TQLHD PNVALFFLEKDLH G + + F+ TT+ FLPR A+++PFSS K+ I+++
Sbjct: 202 TQLHDDPNVALFFLEKDLHPGKTMAVHFTATTAGEK-FLPRSEADAMPFSSEKVPEILSR 260
Query: 140 LNIKKGSKGVQIVKNTISECEEQGIKGEEKVCVTSLESMVDFTTSKLG-KNVEAVSTEVN 198
++K GS + T+ +CE +GE K C TSLESMVDF TS LG +V A ST V
Sbjct: 261 FSVKPGSVEAAEMAQTLRDCEAPPAQGERKACATSLESMVDFATSSLGTSHVRAASTVVG 320
Query: 199 KESNLQQYTIASGVKKL---GEKNKAVVCHKENYPYAVFYCHKTDTTKAYSVPLEGADGS 255
KE + +Q + VK+ G++++ V CH E Y YAVF CH T T+AY+V + G DG+
Sbjct: 321 KEGSPEQEYTVTAVKRAAAGGDQDQLVACHAEPYAYAVFACHLTRATRAYAVSMAGRDGT 380
Query: 256 RVKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVWIR 302
V+A+AVCH DT+ WNPKH+AFQVLKV+PGTVPVCH LP+DHVVW R
Sbjct: 381 GVEAVAVCHADTAGWNPKHVAFQVLKVKPGTVPVCHFLPQDHVVWTR 427
Score = 65.1 bits (157), Expect = 2e-09
Identities = 36/80 (45%), Positives = 44/80 (55%), Gaps = 16/80 (20%)
Query: 4 PQLYWKSMLPNSPMPKAITNLLHPAGYWSEEK----------------GTWVDVGKGGVD 47
P+ YWKS LPNSP+P +++ LL AG + GT VDVGKGGV
Sbjct: 25 PEQYWKSALPNSPIPSSLSQLLSTAGGGTSVNVGGGGVHVDAGHGKPGGTTVDVGKGGVG 84
Query: 48 VGVRKGYYEGGGTDVNVGVG 67
V V+ GY + GGT V VG G
Sbjct: 85 VNVKPGYGKPGGTTVGVGKG 104
>emb|CAH59196.1| BURP-domain containing protein [Plantago major]
Length = 348
Score = 288 bits (738), Expect = 1e-76
Identities = 162/334 (48%), Positives = 202/334 (59%), Gaps = 42/334 (12%)
Query: 3 PPQLYWKSMLPNSPMPKAITNLLHPAGYWSEEKGTWVDVGKGGVD---------VGVRKG 53
P + YWK++LPNSPMP+A+ LL G VD G ++ G
Sbjct: 23 PSERYWKAVLPNSPMPQAVKVLL------PTPTGVGVDAANGRIERHAAGRTIYAAAANG 76
Query: 54 YYEGGGTDVNVGV----GR-----SPFIYNYAASE-----------TQLHDKPNVALFFL 93
E + GR +P I YAA+ TQLHD P +LFFL
Sbjct: 77 KIERHAAAYTIYAAAANGRIVRHAAPIILIYAAATNGRIERANVTGTQLHDDPTASLFFL 136
Query: 94 EKDLHHGTKLNLQFSKTTSNAATFLPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQIVK 153
EKD+H GTK+ LQF KT SN ATFLPRQVA+SIPFSS + I+ K K S I+K
Sbjct: 137 EKDMHAGTKMKLQFVKT-SNQATFLPRQVADSIPFSSKRFPEILRKF--KPTSDEADIMK 193
Query: 154 NTISECEEQGIKGEEKVCVTSLESMVDFTTSKLG-KNVEAVSTEVNKE---SNLQQYTIA 209
+TI +CE++GIKGEEK C TSLESM+DF TSKLG +NVEA+ST + S+ ++YT+
Sbjct: 194 DTIKDCEDEGIKGEEKFCATSLESMIDFCTSKLGSRNVEAISTNAQENAENSSPKEYTLV 253
Query: 210 SGVKKLGEKNKAVVCHKENYPYAVFYCHKTDTTKAYSVPLEGADGSRVKAIAVCHTDTSE 269
+K+ V CHK +Y YAVFYCHK T AY V L ADG +V+A+AVCH DTS+
Sbjct: 254 GAPRKMPGNKAVVACHKMDYAYAVFYCHKIAKTVAYEVSLAAADGCKVEAVAVCHHDTSQ 313
Query: 270 WNPKHLAFQVLKVQPGTVPVCHLLPEDHVVWIRK 303
WNPKH AFQVLKVQPG+VPVCH LP+ +VW+ K
Sbjct: 314 WNPKHFAFQVLKVQPGSVPVCHFLPQQQIVWVPK 347
>ref|XP_474966.1| OSJNBa0036B17.11 [Oryza sativa (japonica cultivar-group)]
gi|38344535|emb|CAD39857.2| OSJNBa0036B17.11 [Oryza
sativa (japonica cultivar-group)]
Length = 285
Score = 241 bits (615), Expect = 2e-62
Identities = 133/307 (43%), Positives = 187/307 (60%), Gaps = 28/307 (9%)
Query: 2 LPPQLYWKSMLPNS-PMPKAITNLLHPAGYWSEEKGTWVDVGKGGVDVGVRKGYYEGGGT 60
+ P+ YW+S+LP+S PMP +I+ LL +S G + K G V +R G
Sbjct: 1 MSPEQYWRSILPDSTPMPISISQLLGDGYPYSPAVG----LPKRGDRVQIRYG------- 49
Query: 61 DVNVGVGRSPFIYNYAASETQLHDKPNVALFFLEKDLHHGTKLNLQFSKTTSNAATFLPR 120
P IY AAS+ Q P + LFFLE +L + L F+ + FLPR
Sbjct: 50 ---------PNIYGLAASQ-QFFKDPTMGLFFLETNLQSSKSIKLHFANMMAGTK-FLPR 98
Query: 121 QVANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEEQGIKGEEKVCVTSLESMVD 180
A+++PFSS ++ I+ + ++ GS +VKNT+ ECE KGE+K C TSLESMVD
Sbjct: 99 GEADAVPFSSKDLQEILARFGVRPGSVDASVVKNTLLECELPANKGEKKACATSLESMVD 158
Query: 181 FTTSKLG-KNVEAVSTEV---NKESNLQQYTIASGVKKLGEKNKAVVCHKENYPYAVFYC 236
F S LG ++++A ST + + ++ Q+YT+ +G +++ E + + CH E+YPYAVF C
Sbjct: 159 FVASSLGTRDIKAASTFLVGKDGDTPAQEYTV-TGARRMAETGQLIACHPESYPYAVFMC 217
Query: 237 HKTDTTKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLPED 296
H T+ T+AY L G DG+ V+A+AVCHTDT+EWNPKH AFQVL V+PGTVPVCH + D
Sbjct: 218 HLTEATRAYKASLVGKDGAAVEAVAVCHTDTAEWNPKHAAFQVLGVKPGTVPVCHFVQPD 277
Query: 297 HVVWIRK 303
VVW R+
Sbjct: 278 VVVWTRR 284
>gb|AAP34365.1| putative dehydration-induced protein [Gossypium barbadense]
Length = 156
Score = 237 bits (604), Expect = 4e-61
Identities = 106/154 (68%), Positives = 134/154 (86%), Gaps = 1/154 (0%)
Query: 150 QIVKNTISECEEQGIKGEEKVCVTSLESMVDFTTSKLGKNVEAVSTEVNKESNLQQYTIA 209
+++KNTI ECE+ I+GEEK C TSLESM+D++ SKLGK +AVSTEV K++ +Q+YTIA
Sbjct: 4 EMMKNTIKECEQPAIEGEEKYCATSLESMIDYSISKLGKVDQAVSTEVEKQTPMQKYTIA 63
Query: 210 SGVKKLGEKNKAVVCHKENYPYAVFYCHKTDTTKAYSVPLEGADGSRVKAIAVCHTDTSE 269
+GV+K+ + +KAVVCHK+NY YAVFYCHK++TT+AY VPLEGADG++ KA+AVCHTDTS
Sbjct: 64 AGVQKMTD-DKAVVCHKQNYAYAVFYCHKSETTRAYMVPLEGADGTKAKAVAVCHTDTSA 122
Query: 270 WNPKHLAFQVLKVQPGTVPVCHLLPEDHVVWIRK 303
WNPKHLAFQVLK +PGTVPVCH LP DH+VW+ K
Sbjct: 123 WNPKHLAFQVLKAEPGTVPVCHFLPRDHIVWVPK 156
>dbj|BAC22501.1| resistant specific protein-3 [Vigna radiata]
Length = 275
Score = 232 bits (592), Expect = 9e-60
Identities = 124/302 (41%), Positives = 168/302 (55%), Gaps = 49/302 (16%)
Query: 2 LPPQLYWKSMLPNSPMPKAITNLLHPAGYWSEEKGTWVDVGKGGVDVGVRKGYYEGGGTD 61
LPP++YW+ MLPN+P+PK I E G
Sbjct: 23 LPPEVYWERMLPNTPIPKVIRQF--------SELG------------------------- 49
Query: 62 VNVGVGRSPFIYNYAASETQLHDKPNVALFFLEKDLHHGTKLNLQFSKTTSNAATFLPRQ 121
P + ++ HD + FF E+ L G KL++ F K + L R+
Sbjct: 50 --------PIVNHHH------HDHLKPSNFFSEERLRRGAKLDVLFRKRNFSTP-LLTRE 94
Query: 122 VANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEEQGIKGEEKVCVTSLESMVDF 181
+A +PFSS K+ I+ L +K SK + V+ T++ CE+ +KGEEK C TS+ESMVDF
Sbjct: 95 IAEHLPFSSEKINEILEILAVKPDSKDAKNVEETLNHCEKPALKGEEKQCATSVESMVDF 154
Query: 182 TTSKLGKNVEAVSTEVNKESNLQQYTIASGVKKLGEKNKAVVCHKENYPYAVFYCHKTDT 241
TSKLG N STE+ S Q++ + GVK L E+ K + CH +YPY VFYCHK
Sbjct: 155 VTSKLGNNARVTSTELEIGSKFQKFIVKDGVKILAEE-KIIACHPMSYPYVVFYCHKMAN 213
Query: 242 TKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVWI 301
+ A+ +PLEG DG+RVKA+A+CH DTS+W+P H+AFQVLKV+PGT CH PE H+VW
Sbjct: 214 STAHFLPLEGEDGTRVKAVAICHKDTSQWDPHHVAFQVLKVKPGTSSACHFFPEGHLVWY 273
Query: 302 RK 303
K
Sbjct: 274 AK 275
>dbj|BAC22500.1| resistant specific protein-2 [Vigna radiata]
Length = 440
Score = 226 bits (577), Expect = 5e-58
Identities = 109/221 (49%), Positives = 144/221 (64%), Gaps = 2/221 (0%)
Query: 83 HDKPNVALFFLEKDLHHGTKLNLQFSKTTSNAATFLPRQVANSIPFSSNKMEYIINKLNI 142
HD + +F E+ L G KL + F K + L R++A +PFSS K+ I+ L +
Sbjct: 222 HDHLKPSSYFSEEGLRRGAKLVMLFHKRKFSTP-LLTREIAEHLPFSSEKINEILEILAV 280
Query: 143 KKGSKGVQIVKNTISECEEQGIKGEEKVCVTSLESMVDFTTSKLGKNVEAVSTEVNKESN 202
K SK + V+ T++ CEE +KGEEK C TS+ESMVDF TSKLG N STE+ ES
Sbjct: 281 KPDSKNAKNVEKTLNNCEEPALKGEEKHCATSVESMVDFVTSKLGNNARVTSTELEIESK 340
Query: 203 LQQYTIASGVKKLGEKNKAVVCHKENYPYAVFYCHKTDTTKAYSVPLEGADGSRVKAIAV 262
Q++ + GVK L E+ + + CH +YPY VFYCHK + A+ VPLEG DG+RVKAI +
Sbjct: 341 FQKFIVKDGVKILAEE-EIIACHPMSYPYVVFYCHKMSNSTAHVVPLEGEDGTRVKAIVI 399
Query: 263 CHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVWIRK 303
CH DTS+W+P H+AFQVLKV+PGT PVCH P H++W K
Sbjct: 400 CHKDTSQWDPDHVAFQVLKVKPGTSPVCHFFPNGHLLWYAK 440
>dbj|BAC22499.1| resistant specific protein-1(8) [Vigna radiata]
gi|24416614|dbj|BAC22498.1| resistant specific
protein-1(4) [Vigna radiata]
Length = 402
Score = 223 bits (569), Expect = 4e-57
Identities = 108/221 (48%), Positives = 144/221 (64%), Gaps = 2/221 (0%)
Query: 83 HDKPNVALFFLEKDLHHGTKLNLQFSKTTSNAATFLPRQVANSIPFSSNKMEYIINKLNI 142
HD + FF E+ L G KL++ F K + L R++A +PFSS K+ I+ L +
Sbjct: 184 HDHLKPSNFFSEERLRRGAKLDVLFRKRNFSTP-LLTREIAEHLPFSSEKINEILEILAV 242
Query: 143 KKGSKGVQIVKNTISECEEQGIKGEEKVCVTSLESMVDFTTSKLGKNVEAVSTEVNKESN 202
K SK + V+ T++ CE+ +KGEEK C TS+ESMVDF TSKLG N STE+ S
Sbjct: 243 KPDSKDAKNVEETLNHCEKPALKGEEKQCATSVESMVDFVTSKLGNNARVTSTELEIGSK 302
Query: 203 LQQYTIASGVKKLGEKNKAVVCHKENYPYAVFYCHKTDTTKAYSVPLEGADGSRVKAIAV 262
Q++ + GVK L E+ K + CH +YPY VFYCHK + A+ +PLEG DG+RVKA+A+
Sbjct: 303 FQKFIVKDGVKILAEE-KIIACHPMSYPYVVFYCHKMANSTAHFLPLEGEDGTRVKAVAI 361
Query: 263 CHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVWIRK 303
CH DTS+W+P H+AFQVLKV+PGT CH PE H+VW K
Sbjct: 362 CHKDTSQWDPHHVAFQVLKVKPGTSSACHFFPEGHLVWYAK 402
Score = 37.7 bits (86), Expect = 0.42
Identities = 13/20 (65%), Positives = 18/20 (90%)
Query: 2 LPPQLYWKSMLPNSPMPKAI 21
LPP++YW+ MLPN+P+PK I
Sbjct: 23 LPPEVYWERMLPNTPIPKVI 42
>dbj|BAA92225.1| similar to the BURP domain [Vigna unguiculata]
Length = 132
Score = 221 bits (562), Expect = 3e-56
Identities = 102/131 (77%), Positives = 119/131 (89%), Gaps = 1/131 (0%)
Query: 173 TSLESMVDFTTSKLGKNVEAVSTEVNKESNLQQYTIASGVKKLGEKNKAVVCHKENYPYA 232
TSLESMVDF+TSKLGKNV +STEV++E+ LQQYTIA GVKK+ N AVVCHK++YPYA
Sbjct: 3 TSLESMVDFSTSKLGKNVAVLSTEVDQETGLQQYTIAPGVKKVSGDN-AVVCHKQSYPYA 61
Query: 233 VFYCHKTDTTKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVCHL 292
VFYCHKT+TT+ Y VPLEGA+G RVKA+AVCHTDTS+WNPKHLAF+VLKV+PGTVPVCH
Sbjct: 62 VFYCHKTETTRTYPVPLEGANGIRVKAVAVCHTDTSQWNPKHLAFEVLKVKPGTVPVCHF 121
Query: 293 LPEDHVVWIRK 303
LPEDHVVW++K
Sbjct: 122 LPEDHVVWVQK 132
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.314 0.132 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 546,657,357
Number of Sequences: 2540612
Number of extensions: 23598205
Number of successful extensions: 49958
Number of sequences better than 10.0: 100
Number of HSP's better than 10.0 without gapping: 72
Number of HSP's successfully gapped in prelim test: 28
Number of HSP's that attempted gapping in prelim test: 49650
Number of HSP's gapped (non-prelim): 146
length of query: 303
length of database: 863,360,394
effective HSP length: 127
effective length of query: 176
effective length of database: 540,702,670
effective search space: 95163669920
effective search space used: 95163669920
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 75 (33.5 bits)
Medicago: description of AC148532.13


