

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148483.6 + phase: 0
(125 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
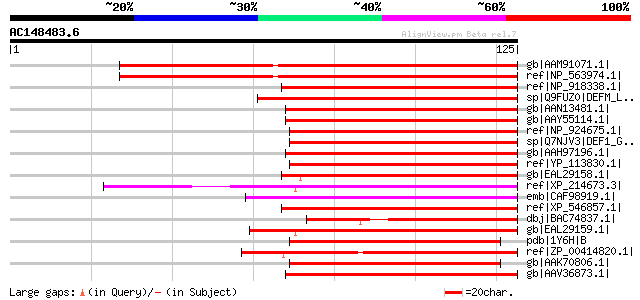
Score E
Sequences producing significant alignments: (bits) Value
gb|AAM91071.1| At1g15390/F9L1_34 [Arabidopsis thaliana] gi|13605... 90 1e-17
ref|NP_563974.1| peptide deformylase, mitochondrial / polypeptid... 90 1e-17
ref|NP_918338.1| peptide deformylase-like protein [Oryza sativa ... 84 8e-16
sp|Q9FUZ0|DEFM_LYCES Peptide deformylase, mitochondrial precurso... 84 8e-16
gb|AAN13481.1| CG31373-PA [Drosophila melanogaster] gi|24645728|... 58 6e-08
gb|AAY55114.1| IP07194p [Drosophila melanogaster] 58 6e-08
ref|NP_924675.1| polypeptide deformylase [Gloeobacter violaceus ... 57 8e-08
sp|Q7NJV3|DEF1_GLOVI Peptide deformylase 1 (PDF 1) (Polypeptide ... 57 8e-08
gb|AAH97196.1| Unknown (protein for MGC:114141) [Danio rerio] 56 2e-07
ref|YP_113830.1| polypeptide deformylase [Methylococcus capsulat... 55 5e-07
gb|EAL29158.1| GA16218-PA [Drosophila pseudoobscura] 54 9e-07
ref|XP_214673.3| PREDICTED: similar to Conserved oligomeric Golg... 52 2e-06
emb|CAF98919.1| unnamed protein product [Tetraodon nigroviridis] 52 3e-06
ref|XP_546857.1| PREDICTED: similar to component of oligomeric g... 50 1e-05
dbj|BAC74837.1| putative polypeptide deformylase [Streptomyces a... 50 1e-05
gb|EAL29159.1| GA16144-PA [Drosophila pseudoobscura] 50 1e-05
pdb|1Y6H|B Chain B, Crystal Structure Of Lipdf gi|61679585|pdb|1... 50 2e-05
ref|ZP_00414820.1| Peptide deformylase [Arthrobacter sp. FB24] g... 50 2e-05
gb|AAK70806.1| peptide deformylase [Leptospira interrogans] gi|4... 50 2e-05
gb|AAV36873.1| RE57173p [Drosophila melanogaster] gi|23170931|gb... 50 2e-05
>gb|AAM91071.1| At1g15390/F9L1_34 [Arabidopsis thaliana] gi|13605760|gb|AAK32873.1|
At1g15390/F9L1_34 [Arabidopsis thaliana]
gi|17433051|sp|Q9FV53|DEFM_ARATH Peptide deformylase,
mitochondrial precursor (PDF) (Polypeptide deformylase)
gi|5103837|gb|AAD39667.1| Simalar to gi|4377403
Polypeptide Deformylase from Chlamydia pneumoniae genome
gb|AE001687. [Arabidopsis thaliana]
Length = 259
Score = 89.7 bits (221), Expect = 1e-17
Identities = 53/98 (54%), Positives = 64/98 (65%), Gaps = 1/98 (1%)
Query: 28 SSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVNKLPYIVKAGDPVIHEPAREVDH 87
SS N K+ T S ST + +K V+ LP IV +GDPV+HE AREVD
Sbjct: 32 SSHLLNRKLYNLPTSSSSSLSTKAGWLLGLGEKKKKVD-LPEIVASGDPVLHEKAREVDP 90
Query: 88 SEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
EI S++IQ IIDDMI VMR APGVG+AAPQIG+PLR+
Sbjct: 91 GEIGSERIQKIIDDMIKVMRLAPGVGLAAPQIGVPLRI 128
>ref|NP_563974.1| peptide deformylase, mitochondrial / polypeptide deformylase 1A
(PDF1A) [Arabidopsis thaliana]
gi|11320952|gb|AAG33973.1| peptide deformylase-like
protein [Arabidopsis thaliana]
Length = 269
Score = 89.7 bits (221), Expect = 1e-17
Identities = 53/98 (54%), Positives = 64/98 (65%), Gaps = 1/98 (1%)
Query: 28 SSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVNKLPYIVKAGDPVIHEPAREVDH 87
SS N K+ T S ST + +K V+ LP IV +GDPV+HE AREVD
Sbjct: 42 SSHLLNRKLYNLPTSSSSSLSTKAGWLLGLGEKKKKVD-LPEIVASGDPVLHEKAREVDP 100
Query: 88 SEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
EI S++IQ IIDDMI VMR APGVG+AAPQIG+PLR+
Sbjct: 101 GEIGSERIQKIIDDMIKVMRLAPGVGLAAPQIGVPLRI 138
>ref|NP_918338.1| peptide deformylase-like protein [Oryza sativa (japonica
cultivar-group)]
Length = 246
Score = 84.0 bits (206), Expect = 8e-16
Identities = 37/58 (63%), Positives = 49/58 (83%)
Query: 68 PYIVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
P VKAGDPV+HEPA++V +I S+K+Q +ID M+ VMRKAPGVG+AAPQIG+PL++
Sbjct: 72 PGTVKAGDPVLHEPAQDVAPGDIPSEKVQGVIDRMVAVMRKAPGVGLAAPQIGVPLKI 129
>sp|Q9FUZ0|DEFM_LYCES Peptide deformylase, mitochondrial precursor (PDF) (Polypeptide
deformylase) gi|11320968|gb|AAG33981.1| peptide
deformylase-like protein [Lycopersicon esculentum]
Length = 277
Score = 84.0 bits (206), Expect = 8e-16
Identities = 38/64 (59%), Positives = 52/64 (80%)
Query: 62 KTVNKLPYIVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGI 121
K +P IVKAGDPV+HEP++++ EI S++IQ II++M+ VMR APGVG+AAPQIGI
Sbjct: 83 KKKQAMPDIVKAGDPVLHEPSQDIPLEEIGSERIQKIIEEMVKVMRNAPGVGLAAPQIGI 142
Query: 122 PLRV 125
PL++
Sbjct: 143 PLKI 146
>gb|AAN13481.1| CG31373-PA [Drosophila melanogaster] gi|24645728|ref|NP_731495.1|
CG31373-PA [Drosophila melanogaster]
Length = 196
Score = 57.8 bits (138), Expect = 6e-08
Identities = 28/57 (49%), Positives = 38/57 (66%)
Query: 69 YIVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
+ + GDPV+ + A EV +I S +I IID M+ V+R VGVAAPQ+GIPLR+
Sbjct: 8 HFTQIGDPVLRQRAEEVPPEDIDSREINQIIDGMVKVLRHYDCVGVAAPQVGIPLRI 64
>gb|AAY55114.1| IP07194p [Drosophila melanogaster]
Length = 206
Score = 57.8 bits (138), Expect = 6e-08
Identities = 28/57 (49%), Positives = 38/57 (66%)
Query: 69 YIVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
+ + GDPV+ + A EV +I S +I IID M+ V+R VGVAAPQ+GIPLR+
Sbjct: 18 HFTQIGDPVLRQRAEEVPPEDIDSREINQIIDGMVKVLRHYDCVGVAAPQVGIPLRI 74
>ref|NP_924675.1| polypeptide deformylase [Gloeobacter violaceus PCC 7421]
gi|35212295|dbj|BAC89670.1| polypeptide deformylase
[Gloeobacter violaceus PCC 7421]
Length = 275
Score = 57.4 bits (137), Expect = 8e-08
Identities = 27/56 (48%), Positives = 41/56 (73%)
Query: 70 IVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
IVK GDPV+ A+ ++ EI+S+ IQ +I M MR+APGVG+AAPQ+G+ +++
Sbjct: 96 IVKTGDPVLRLTAKPLNSDEIQSEAIQQLIAAMAERMREAPGVGLAAPQVGVSVQL 151
>sp|Q7NJV3|DEF1_GLOVI Peptide deformylase 1 (PDF 1) (Polypeptide deformylase 1)
Length = 227
Score = 57.4 bits (137), Expect = 8e-08
Identities = 27/56 (48%), Positives = 41/56 (73%)
Query: 70 IVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
IVK GDPV+ A+ ++ EI+S+ IQ +I M MR+APGVG+AAPQ+G+ +++
Sbjct: 48 IVKTGDPVLRLTAKPLNSDEIQSEAIQQLIAAMAERMREAPGVGLAAPQVGVSVQL 103
>gb|AAH97196.1| Unknown (protein for MGC:114141) [Danio rerio]
Length = 247
Score = 56.2 bits (134), Expect = 2e-07
Identities = 24/57 (42%), Positives = 39/57 (68%)
Query: 69 YIVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
++ + GDPV+ A EV+ I+ ++Q +I ++ VMRK VG++APQIG+PLR+
Sbjct: 70 HVCQVGDPVLRSHAAEVEPGAIQGPEVQKVIKTLVKVMRKLECVGLSAPQIGVPLRI 126
>ref|YP_113830.1| polypeptide deformylase [Methylococcus capsulatus str. Bath]
gi|53758098|gb|AAU92389.1| polypeptide deformylase
[Methylococcus capsulatus str. Bath]
Length = 191
Score = 54.7 bits (130), Expect = 5e-07
Identities = 28/56 (50%), Positives = 39/56 (69%)
Query: 70 IVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
IV+AG+PV+ + AR + EI+S +Q +I M MR APGVG+AAPQIG L++
Sbjct: 10 IVQAGEPVLRQRARPLSPEEIRSAAVQALIGHMRETMRDAPGVGLAAPQIGQGLQL 65
>gb|EAL29158.1| GA16218-PA [Drosophila pseudoobscura]
Length = 196
Score = 53.9 bits (128), Expect = 9e-07
Identities = 27/60 (45%), Positives = 39/60 (65%), Gaps = 2/60 (3%)
Query: 68 PYI--VKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
PY+ + GDPV+ + A EV I + +I I+D M+ V+R VGVAAPQ+G+PLR+
Sbjct: 5 PYLHFTQIGDPVLRQIAEEVPPESIGTVEIDQIVDRMVKVLRHYDCVGVAAPQVGVPLRI 64
>ref|XP_214673.3| PREDICTED: similar to Conserved oligomeric Golgi complex component
8 [Rattus norvegicus]
Length = 889
Score = 52.4 bits (124), Expect = 2e-06
Identities = 30/107 (28%), Positives = 55/107 (51%), Gaps = 14/107 (13%)
Query: 24 VSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVNKLPY-----IVKAGDPVI 78
V+LS SC++ L ++ S LR+ V P + + GDPV+
Sbjct: 688 VTLSRGQSCSSSASLEGAARTR---------SYWRYLRRLVRGAPQPPYTRVCQVGDPVL 738
Query: 79 HEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
A V+ ++ ++Q +++ ++ VMR+ VG++APQ+G+PL+V
Sbjct: 739 RTVAAPVEPKQLAGPELQRLVEQLVQVMRRRGCVGLSAPQLGVPLQV 785
>emb|CAF98919.1| unnamed protein product [Tetraodon nigroviridis]
Length = 198
Score = 52.0 bits (123), Expect = 3e-06
Identities = 23/67 (34%), Positives = 40/67 (59%)
Query: 59 LLRKTVNKLPYIVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQ 118
L+ V ++ + GDPV+ A VD + ++Q ++ ++ VMR+ VG++APQ
Sbjct: 11 LIPPPVPPYKHVCQVGDPVLRSRAAPVDPGAVGGAEVQKVVHTLVKVMRELDCVGLSAPQ 70
Query: 119 IGIPLRV 125
IG+PLR+
Sbjct: 71 IGVPLRI 77
>ref|XP_546857.1| PREDICTED: similar to component of oligomeric golgi complex 8
[Canis familiaris]
Length = 836
Score = 50.1 bits (118), Expect = 1e-05
Identities = 19/58 (32%), Positives = 39/58 (66%)
Query: 68 PYIVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
P++ + GDP + A V+ +++ ++Q ++ ++ VMR+ VG++APQ+G+PL+V
Sbjct: 658 PHVCQVGDPALRTVAAPVEPAQLAGPQLQRLVQRLVQVMRRRHCVGLSAPQLGVPLQV 715
>dbj|BAC74837.1| putative polypeptide deformylase [Streptomyces avermitilis MA-4680]
gi|39931074|sp|Q826Q0|DEF2_STRAW Peptide deformylase 2
(PDF 2) (Polypeptide deformylase 2)
gi|29833668|ref|NP_828302.1| putative polypeptide
deformylase [Streptomyces avermitilis MA-4680]
Length = 186
Score = 50.1 bits (118), Expect = 1e-05
Identities = 26/53 (49%), Positives = 34/53 (64%), Gaps = 5/53 (9%)
Query: 74 GDPVIHEPAREV-DHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
GDPV+H P EV DH ++ +++DM M A GVG+AA QIG+PLRV
Sbjct: 20 GDPVLHAPCEEVTDHGP----ELARLVEDMFATMYAANGVGLAANQIGVPLRV 68
>gb|EAL29159.1| GA16144-PA [Drosophila pseudoobscura]
Length = 238
Score = 50.1 bits (118), Expect = 1e-05
Identities = 26/68 (38%), Positives = 41/68 (60%), Gaps = 2/68 (2%)
Query: 60 LRKTVNKLPY--IVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAP 117
+ + N PY + GDPV+ + A V I+ +I+ I++ M+ V+RK VG+AAP
Sbjct: 39 ITERTNLPPYNHFTQIGDPVLRQQAAAVPLELIEGPEIEAIVEQMVKVLRKYNCVGIAAP 98
Query: 118 QIGIPLRV 125
QIG+ LR+
Sbjct: 99 QIGVSLRI 106
>pdb|1Y6H|B Chain B, Crystal Structure Of Lipdf gi|61679585|pdb|1Y6H|A Chain A,
Crystal Structure Of Lipdf
Length = 177
Score = 49.7 bits (117), Expect = 2e-05
Identities = 23/52 (44%), Positives = 35/52 (67%)
Query: 70 IVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGI 121
I++ GDP++ + + V EI++ + + +I DM MR A GVG+AAPQIGI
Sbjct: 5 ILRMGDPILRKISEPVTEDEIQTKEFKKLIRDMFDTMRHAEGVGLAAPQIGI 56
>ref|ZP_00414820.1| Peptide deformylase [Arthrobacter sp. FB24]
gi|66866946|gb|EAL94340.1| Peptide deformylase
[Arthrobacter sp. FB24]
Length = 226
Score = 49.7 bits (117), Expect = 2e-05
Identities = 30/74 (40%), Positives = 46/74 (61%), Gaps = 7/74 (9%)
Query: 58 ALLRKTVNK------LPYIVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPG 111
A +R+TV + LP IV+AG PV+ + A + ++ ++ +I M VM APG
Sbjct: 13 ARIRETVQEILSAGVLPAIVQAGHPVLRQQAAPYE-GQLDGTELAALIALMREVMHDAPG 71
Query: 112 VGVAAPQIGIPLRV 125
VG+AAPQ+GIPL++
Sbjct: 72 VGLAAPQLGIPLQL 85
>gb|AAK70806.1| peptide deformylase [Leptospira interrogans]
gi|45657385|ref|YP_001471.1| putative polypeptide
deformylase protein [Leptospira interrogans serovar
Copenhageni str. Fiocruz L1-130]
gi|24215138|ref|NP_712619.1| Polypeptide deformylase
[Leptospira interrogans serovar Lai str. 56601]
gi|24196204|gb|AAN49637.1| Polypeptide deformylase
[Leptospira interrogans serovar lai str. 56601]
gi|23396549|sp|Q93LE9|DEF_LEPIN Peptide deformylase
(PDF) (Polypeptide deformylase)
gi|59797588|sp|Q72S74|DEF_LEPIC Peptide deformylase
(PDF) (Polypeptide deformylase)
gi|45600624|gb|AAS70108.1| putative polypeptide
deformylase protein [Leptospira interrogans serovar
Copenhageni str. Fiocruz L1-130]
Length = 178
Score = 49.7 bits (117), Expect = 2e-05
Identities = 23/52 (44%), Positives = 35/52 (67%)
Query: 70 IVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGI 121
I++ GDP++ + + V EI++ + + +I DM MR A GVG+AAPQIGI
Sbjct: 6 ILRMGDPILRKISEPVTEDEIQTKEFKKLIRDMFDTMRHAEGVGLAAPQIGI 57
>gb|AAV36873.1| RE57173p [Drosophila melanogaster] gi|23170931|gb|AAF54540.2|
CG31278-PA [Drosophila melanogaster]
gi|24645726|ref|NP_731494.1| CG31278-PA [Drosophila
melanogaster]
Length = 238
Score = 49.7 bits (117), Expect = 2e-05
Identities = 23/57 (40%), Positives = 37/57 (64%)
Query: 69 YIVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
+ + GDPV+ + A V + S +I+ I++ M+ V+RK VG+AAPQIG+ LR+
Sbjct: 50 HFTQIGDPVLRQQAALVPKEHMASPEIKAIVERMVKVLRKFDCVGIAAPQIGVSLRI 106
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.316 0.132 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 196,304,845
Number of Sequences: 2540612
Number of extensions: 6837958
Number of successful extensions: 23579
Number of sequences better than 10.0: 375
Number of HSP's better than 10.0 without gapping: 267
Number of HSP's successfully gapped in prelim test: 108
Number of HSP's that attempted gapping in prelim test: 23255
Number of HSP's gapped (non-prelim): 391
length of query: 125
length of database: 863,360,394
effective HSP length: 101
effective length of query: 24
effective length of database: 606,758,582
effective search space: 14562205968
effective search space used: 14562205968
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 68 (30.8 bits)
Medicago: description of AC148483.6


