

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148482.2 + phase: 0
(530 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
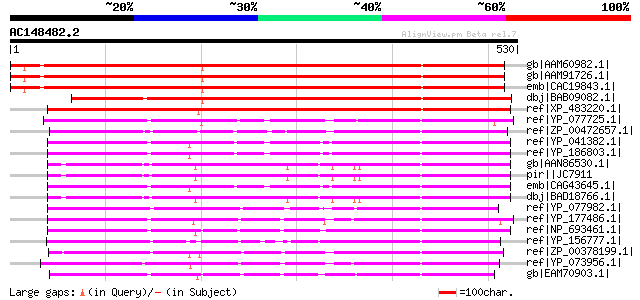
Score E
Sequences producing significant alignments: (bits) Value
gb|AAM60982.1| sodium-dicarboxylate cotransporter-like [Arabidop... 684 0.0
gb|AAM91726.1| putative sodium-dicarboxylate cotransporter prote... 683 0.0
emb|CAC19843.1| SDAT(At) [Arabidopsis thaliana] 674 0.0
dbj|BAB09082.1| sodium-dicarboxylate cotransporter-like [Arabido... 642 0.0
ref|XP_483220.1| putative sodium-dicarboxylate cotransporter [Or... 577 e-163
ref|YP_077725.1| Sodium/sulphate symporter [Bacillus licheniform... 308 3e-82
ref|ZP_00472657.1| Sodium/sulphate symporter [Chromohalobacter s... 299 1e-79
ref|YP_041382.1| putative sodium:sulfate symporter [Staphylococc... 298 4e-79
ref|YP_186803.1| sodium-dependent transporter [Staphylococcus au... 297 5e-79
gb|AAN86530.1| Na+-coupled citrate transporter protein [Homo sap... 297 5e-79
pir||JC7911 Na+-coupled citrate transporter NaCT - human 297 5e-79
emb|CAG43645.1| putative sodium:sulfate symporter [Staphylococcu... 297 6e-79
dbj|BAD18766.1| unnamed protein product [Homo sapiens] 292 2e-77
ref|YP_077982.1| Sodium/sulphate symporter [Bacillus licheniform... 285 2e-75
ref|YP_177486.1| sodium:dicarboxylate cotransporter [Bacillus cl... 284 6e-75
ref|NP_693461.1| sodium-dependent transporter [Oceanobacillus ih... 282 2e-74
ref|YP_156777.1| Na+/dicarboxylate symporter [Idiomarina loihien... 273 8e-72
ref|ZP_00378199.1| COG0471: Di- and tricarboxylate transporters ... 273 8e-72
ref|YP_073956.1| sodium-dependent transporter [Symbiobacterium t... 270 1e-70
gb|EAM70903.1| Sodium/sulphate symporter [Desulfuromonas acetoxi... 268 2e-70
>gb|AAM60982.1| sodium-dicarboxylate cotransporter-like [Arabidopsis thaliana]
Length = 540
Score = 684 bits (1765), Expect = 0.0
Identities = 338/527 (64%), Positives = 418/527 (79%), Gaps = 14/527 (2%)
Query: 1 MAGEDSLNATLLPL---QEPIQ-ETPNNNSITIFSILNLQNFYIILGPLLSIIICLLVKL 56
+AG D L + LLP+ EP + +T TIF+ +N YI LGPLL ++CL V L
Sbjct: 8 VAGSDDLKSPLLPVVHNDEPFERQTVGQQLRTIFTP---KNCYIALGPLLCAVVCLCVDL 64
Query: 57 --DAPTTSTKMLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFGIASADTVAHSYMDDVI 114
D TT+ MLGV+ W+F WW+T AVP+P+TSM PLFLFPLFGI++AD VA+SYMDDVI
Sbjct: 65 GGDETTTARNMLGVLVWMFAWWLTEAVPMPITSMTPLFLFPLFGISAADDVANSYMDDVI 124
Query: 115 TLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGLCATTFFVSMWLHNVAT 174
+LVLGSFILALAVE YN+HRRLALN+TLVFC +PLN +LLLG+CATT FVSMW+HNVA
Sbjct: 125 SLVLGSFILALAVEHYNIHRRLALNITLVFCVEPLNAPLLLLGICATTAFVSMWMHNVAA 184
Query: 175 AVMMMPVATGILHRLPPAEEQPELMN----KFSRAVILTVVYATPIGGISTLTGTGVNLI 230
AVMMMPVATGIL RLP + E+++ KFSRAV+L V+Y+ +GG+STLTGTGVNLI
Sbjct: 185 AVMMMPVATGILQRLPSSSSTTEVVHPAVGKFSRAVVLGVIYSAAVGGMSTLTGTGVNLI 244
Query: 231 IIGMWKSLAPEAKPISFNSWFFFGFPVAVLLLLCFWCILCLIYVRKGSSRALSDYLDRAL 290
++GMWKS PEA PISF+ WFFFGFP+A+ + + WC+LC++Y KG+ +ALS YL ++
Sbjct: 245 LVGMWKSYFPEADPISFSQWFFFGFPLALCIFVVLWCVLCVMYCPKGAGQALSPYLHKSH 304
Query: 291 LKRDLEALGPMSFAEKMVLCVFGLLIVLWMTRRISDDLPGWGVLFHGLVGDGSISVLAAV 350
L+R+LE LGPM+FAEKMVL VFG L+VLWMTR I+DD+PGWG +F G GDG++SV+ A
Sbjct: 305 LRRELELLGPMNFAEKMVLAVFGGLVVLWMTRNITDDIPGWGRIFAGRAGDGTVSVMMAT 364
Query: 351 LLFIIPNMKQNGEKLMDWNECKKLPWNLILLLGAGFALADGVQSSGLADVMSKALEFLKD 410
LLFIIP+ + GEKLMDWN+CKKLPWN++LLLGAGFA+ADGV++SGLA+V+SK L FL+
Sbjct: 365 LLFIIPSNIKKGEKLMDWNKCKKLPWNIVLLLGAGFAIADGVRTSGLAEVLSKGLVFLET 424
Query: 411 VPYLAIVPAVSLLCSIITEFITSNDATATLLVPLLYHIAINMHVHPLLLMIPGGIATEFA 470
PY AI P V L+ + ITEF TSN+AT TLLVPLL IA NM +HPLLLM+PG I +FA
Sbjct: 425 APYWAIAPTVCLIAATITEF-TSNNATTTLLVPLLIEIAKNMGIHPLLLMVPGAIGAQFA 483
Query: 471 FWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAGVSVLALLMPTLG 517
F LPT TPSNVVGF TGHIEIKDM+K G+PLK+AG L++LMPTLG
Sbjct: 484 FLLPTGTPSNVVGFTTGHIEIKDMIKTGLPLKIAGTIFLSILMPTLG 530
>gb|AAM91726.1| putative sodium-dicarboxylate cotransporter protein [Arabidopsis
thaliana] gi|18176009|gb|AAL59967.1| putative
sodium-dicarboxylate cotransporter protein [Arabidopsis
thaliana] gi|15238130|ref|NP_199567.1|
sodium/dicarboxylate cotransporter, putative
[Arabidopsis thaliana] gi|8051712|dbj|BAA96091.1| sodium
sulfate or dicarboxylate transporter [Arabidopsis
thaliana]
Length = 540
Score = 683 bits (1762), Expect = 0.0
Identities = 337/527 (63%), Positives = 418/527 (78%), Gaps = 14/527 (2%)
Query: 1 MAGEDSLNATLLPL---QEPIQ-ETPNNNSITIFSILNLQNFYIILGPLLSIIICLLVKL 56
+AG D L + LLP+ EP + +T TIF+ +N YI LGPLL ++CL V L
Sbjct: 8 VAGSDDLKSPLLPVVHNDEPFERQTVGQQLRTIFTP---KNCYIALGPLLCAVVCLCVDL 64
Query: 57 --DAPTTSTKMLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFGIASADTVAHSYMDDVI 114
D TT+ MLGV+ W+F WW+T AVP+P+TSM PLFLFPLFGI++AD VA+SYMDDVI
Sbjct: 65 GGDETTTARNMLGVLVWMFAWWLTEAVPMPITSMTPLFLFPLFGISAADDVANSYMDDVI 124
Query: 115 TLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGLCATTFFVSMWLHNVAT 174
+LVLGSFILALAVE YN+HRRLALN+TLVFC +PLN +LLLG+CATT FVSMW+HNVA
Sbjct: 125 SLVLGSFILALAVEHYNIHRRLALNITLVFCVEPLNAPLLLLGICATTAFVSMWMHNVAA 184
Query: 175 AVMMMPVATGILHRLPPAEEQPELMN----KFSRAVILTVVYATPIGGISTLTGTGVNLI 230
AVMMMPVATGIL RLP + E+++ KFSRAV+L V+Y+ +GG+STLTGTGVNLI
Sbjct: 185 AVMMMPVATGILQRLPSSSSTTEVVHPAVGKFSRAVVLGVIYSAAVGGMSTLTGTGVNLI 244
Query: 231 IIGMWKSLAPEAKPISFNSWFFFGFPVAVLLLLCFWCILCLIYVRKGSSRALSDYLDRAL 290
++GMWKS PEA PISF+ WFFFGFP+A+ + + WC+LC++Y KG+ +ALS YL ++
Sbjct: 245 LVGMWKSYFPEADPISFSQWFFFGFPLALCIFVVLWCVLCVMYCPKGAGQALSPYLHKSH 304
Query: 291 LKRDLEALGPMSFAEKMVLCVFGLLIVLWMTRRISDDLPGWGVLFHGLVGDGSISVLAAV 350
L+R+L+ LGPM+FAEKMVL VFG L+VLWMTR I+DD+PGWG +F G GDG++SV+ A
Sbjct: 305 LRRELDLLGPMNFAEKMVLAVFGGLVVLWMTRNITDDIPGWGRIFAGRAGDGTVSVMMAT 364
Query: 351 LLFIIPNMKQNGEKLMDWNECKKLPWNLILLLGAGFALADGVQSSGLADVMSKALEFLKD 410
LLFIIP+ + GEKLMDWN+CKKLPWN++LLLGAGFA+ADGV++SGLA+V+SK L FL+
Sbjct: 365 LLFIIPSNIKKGEKLMDWNKCKKLPWNIVLLLGAGFAIADGVRTSGLAEVLSKGLVFLET 424
Query: 411 VPYLAIVPAVSLLCSIITEFITSNDATATLLVPLLYHIAINMHVHPLLLMIPGGIATEFA 470
PY AI P V L+ + ITEF TSN+AT TLLVPLL IA NM +HPLLLM+PG I +FA
Sbjct: 425 APYWAIAPTVCLIAATITEF-TSNNATTTLLVPLLIEIAKNMGIHPLLLMVPGAIGAQFA 483
Query: 471 FWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAGVSVLALLMPTLG 517
F LPT TPSNVVGF TGHIEIKDM+K G+PLK+AG L++LMPTLG
Sbjct: 484 FLLPTGTPSNVVGFTTGHIEIKDMIKTGLPLKIAGTIFLSILMPTLG 530
>emb|CAC19843.1| SDAT(At) [Arabidopsis thaliana]
Length = 540
Score = 674 bits (1740), Expect = 0.0
Identities = 334/527 (63%), Positives = 415/527 (78%), Gaps = 14/527 (2%)
Query: 1 MAGEDSLNATLLPL---QEPIQ-ETPNNNSITIFSILNLQNFYIILGPLLSIIICLLVKL 56
+AG D L + LLP+ EP + +T TIF+ +N YI LGPLL ++CL V L
Sbjct: 8 VAGSDDLKSPLLPVVHNDEPFERQTVGQQLRTIFTP---KNCYIALGPLLCAVVCLCVDL 64
Query: 57 --DAPTTSTKMLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFGIASADTVAHSYMDDVI 114
D TT+ MLGV+ W+F WW+T AVP+P+TSM PLFLFPLFGI++AD VA+SYMDDVI
Sbjct: 65 GGDETTTARNMLGVLVWMFAWWLTEAVPMPITSMTPLFLFPLFGISAADDVANSYMDDVI 124
Query: 115 TLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGLCATTFFVSMWLHNVAT 174
+LVLGSFILALAVE YN+HRRLALN+TLVFC +PLN +LLLG+CATT FVSMW+HNVA
Sbjct: 125 SLVLGSFILALAVEHYNIHRRLALNITLVFCVEPLNAPLLLLGICATTAFVSMWMHNVAA 184
Query: 175 AVMMMPVATGILHRLPPAEEQPELMN----KFSRAVILTVVYATPIGGISTLTGTGVNLI 230
AVMMMPVATGIL RLP + E+++ KFSRAV+L V+Y+ +GG+STLTGTGVNLI
Sbjct: 185 AVMMMPVATGILQRLPSSSSTTEVVHPAVGKFSRAVVLGVIYSAAVGGMSTLTGTGVNLI 244
Query: 231 IIGMWKSLAPEAKPISFNSWFFFGFPVAVLLLLCFWCILCLIYVRKGSSRALSDYLDRAL 290
++GMWKS PEA PISF+ WFFFGFP+A+ + + WC+LC++Y KG+ +ALS YL ++
Sbjct: 245 LVGMWKSYFPEADPISFSQWFFFGFPLALCIFVVLWCVLCVMYCPKGAGQALSPYLHKSH 304
Query: 291 LKRDLEALGPMSFAEKMVLCVFGLLIVLWMTRRISDDLPGWGVLFHGLVGDGSISVLAAV 350
L+R+L+ LGPM+FAEKMVL VFG L+VLWMTR I+DD+PGWG +F G GDG++SV+ A
Sbjct: 305 LRRELDLLGPMNFAEKMVLAVFGGLVVLWMTRNITDDIPGWGRIFAGRAGDGTVSVMMAT 364
Query: 351 LLFIIPNMKQNGEKLMDWNECKKLPWNLILLLGAGFALADGVQSSGLADVMSKALEFLKD 410
LLFIIP+ + GEKLMDWN+CKKLPWN++LLLGAGFA+ADGV +SGLA+V+SK L FL+
Sbjct: 365 LLFIIPSNIKKGEKLMDWNKCKKLPWNIVLLLGAGFAIADGVPTSGLAEVLSKGLVFLET 424
Query: 411 VPYLAIVPAVSLLCSIITEFITSNDATATLLVPLLYHIAINMHVHPLLLMIPGGIATEFA 470
PY AI P V L+ + ITEF TSN+AT TLLVPLL IA NM +HPLLLM+ + +FA
Sbjct: 425 APYWAIAPTVCLIAATITEF-TSNNATTTLLVPLLIEIAKNMGIHPLLLMVQEPLGAQFA 483
Query: 471 FWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAGVSVLALLMPTLG 517
F LPT TPSNVVGF TGHIEIKDM+K G+PLK+AG L++LMPTLG
Sbjct: 484 FLLPTGTPSNVVGFTTGHIEIKDMIKTGLPLKIAGTIFLSILMPTLG 530
>dbj|BAB09082.1| sodium-dicarboxylate cotransporter-like [Arabidopsis thaliana]
Length = 462
Score = 642 bits (1655), Expect = 0.0
Identities = 309/464 (66%), Positives = 380/464 (81%), Gaps = 8/464 (1%)
Query: 65 MLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFGIASADTVAHSYMDDVITLVLGSFILA 124
MLGV+ W+F WW+T AVP+P+TSM PLFLFPLFGI++AD VA+SYMDDVI+LVLGSFILA
Sbjct: 1 MLGVLVWMFAWWLTEAVPMPITSMTPLFLFPLFGISAADDVANSYMDDVISLVLGSFILA 60
Query: 125 LAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGLCATTFFVSMWLHNVATAVMMMPVATG 184
LAVE YN+HRRLALNV C +PLN +LLLG+CATT FVSMW+HNVA AVMMMPVATG
Sbjct: 61 LAVEHYNIHRRLALNV---ICVEPLNAPLLLLGICATTAFVSMWMHNVAAAVMMMPVATG 117
Query: 185 ILHRLPPAEEQPELMN----KFSRAVILTVVYATPIGGISTLTGTGVNLIIIGMWKSLAP 240
IL RLP + E+++ KFSRAV+L V+Y+ +GG+STLTGTGVNLI++GMWKS P
Sbjct: 118 ILQRLPSSSSTTEVVHPAVGKFSRAVVLGVIYSAAVGGMSTLTGTGVNLILVGMWKSYFP 177
Query: 241 EAKPISFNSWFFFGFPVAVLLLLCFWCILCLIYVRKGSSRALSDYLDRALLKRDLEALGP 300
EA PISF+ WFFFGFP+A+ + + WC+LC++Y KG+ +ALS YL ++ L+R+L+ LGP
Sbjct: 178 EADPISFSQWFFFGFPLALCIFVVLWCVLCVMYCPKGAGQALSPYLHKSHLRRELDLLGP 237
Query: 301 MSFAEKMVLCVFGLLIVLWMTRRISDDLPGWGVLFHGLVGDGSISVLAAVLLFIIPNMKQ 360
M+FAEKMVL VFG L+VLWMTR I+DD+PGWG +F G GDG++SV+ A LLFIIP+ +
Sbjct: 238 MNFAEKMVLAVFGGLVVLWMTRNITDDIPGWGRIFAGRAGDGTVSVMMATLLFIIPSNIK 297
Query: 361 NGEKLMDWNECKKLPWNLILLLGAGFALADGVQSSGLADVMSKALEFLKDVPYLAIVPAV 420
GEKLMDWN+CKKLPWN++LLLGAGFA+ADGV++SGLA+V+SK L FL+ PY AI P V
Sbjct: 298 KGEKLMDWNKCKKLPWNIVLLLGAGFAIADGVRTSGLAEVLSKGLVFLETAPYWAIAPTV 357
Query: 421 SLLCSIITEFITSNDATATLLVPLLYHIAINMHVHPLLLMIPGGIATEFAFWLPTSTPSN 480
L+ + ITEF TSN+AT TLLVPLL IA NM +HPLLLM+PG I +FAF LPT TPSN
Sbjct: 358 CLIAATITEF-TSNNATTTLLVPLLIEIAKNMGIHPLLLMVPGAIGAQFAFLLPTGTPSN 416
Query: 481 VVGFATGHIEIKDMLKVGVPLKVAGVSVLALLMPTLGKLLLSLF 524
VVGF TGHIEIKDM+K G+PLK+AG L++LMPTLG LL + F
Sbjct: 417 VVGFTTGHIEIKDMIKTGLPLKIAGTIFLSILMPTLGNLLKTFF 460
>ref|XP_483220.1| putative sodium-dicarboxylate cotransporter [Oryza sativa (japonica
cultivar-group)] gi|51965002|ref|XP_507285.1| PREDICTED
OJ1506_F01.29 gene product [Oryza sativa (japonica
cultivar-group)] gi|42408139|dbj|BAD09278.1| putative
sodium-dicarboxylate cotransporter [Oryza sativa
(japonica cultivar-group)]
Length = 548
Score = 577 bits (1488), Expect = e-163
Identities = 276/488 (56%), Positives = 364/488 (74%), Gaps = 5/488 (1%)
Query: 40 IILGPLLSIIICLLVKL-DAPTTSTKMLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFG 98
I GP +C V L D + MLGV+AWVF WW+T AVPL V SM PLFLFP G
Sbjct: 58 IAAGPAACAAVCAAVDLGDGHGEARNMLGVLAWVFLWWVTGAVPLAVASMAPLFLFPALG 117
Query: 99 IASADTVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGL 158
I+S+D VA +YM DVI+LVLGSFILALAV+ + +HRRLALNV +FCGDP+ PS+LLLG+
Sbjct: 118 ISSSDDVARAYMGDVISLVLGSFILALAVDHHRIHRRLALNVLSLFCGDPVRPSLLLLGV 177
Query: 159 CATTFFVSMWLHNVATAVMMMPVATGILHRLPPAEEQP---ELMNKFSRAVILTVVYATP 215
TT VSMW+HN A VMMMPVATGIL R P + + + +FS+AV+L VVYA+
Sbjct: 178 TGTTALVSMWIHNTACTVMMMPVATGILQRFPRGDIDDGGGQEVRRFSKAVVLGVVYASA 237
Query: 216 IGGISTLTGTGVNLIIIGMWKSLAPEAKPISFNSWFFFGFPVAVLLLLCFWCILCLIYVR 275
IGG++TLTGTGVN+I++GMW S PE +PI+F+SW FG P+A++L L W LCL+Y
Sbjct: 238 IGGMATLTGTGVNIILVGMWSSYFPEQRPITFSSWMSFGLPMAIILFLALWLTLCLMYCS 297
Query: 276 KGSSRALSDYLDRALLKRDLEALGPMSFAEKMVLCVFGLLIVLWMTRRISDDLPGWGVLF 335
+++ALS YLDR+ L+R+L LGPM+FAEKMVL VFG L+VLWM+R I+D++PGWGVLF
Sbjct: 298 NNTAKALSAYLDRSHLRRELSLLGPMAFAEKMVLAVFGGLVVLWMSRNITDNIPGWGVLF 357
Query: 336 HGLVGDGSISVLAAVLLFIIPNMKQNGEKLMDWNECKKLPWNLILLLGAGFALADGVQSS 395
H VGDG+++++ A LLFIIP+ K+ GEKLMDWN+CKK+ WN+ILLLGAGFA+ADG ++S
Sbjct: 358 HNKVGDGTVTIMMATLLFIIPSGKREGEKLMDWNKCKKIQWNIILLLGAGFAIADGFKTS 417
Query: 396 GLADVMSKALEFLKDVPYLAIVPAVSLLCSIITEFITSNDATATLLVPLLYHIAINMHVH 455
GL D++S L FLK P L IVP + I+TEF TS+D+T TL++PL +A ++ VH
Sbjct: 418 GLTDILSNGLRFLKGAPTLVIVPVACVFSGIMTEF-TSDDSTTTLVLPLFAELAKSIEVH 476
Query: 456 PLLLMIPGGIATEFAFWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAGVSVLALLMPT 515
P LLM+ G I + ++ LPT +PSNVVGF+TG+I IKD++ G+PLK+ ++ L +L+PT
Sbjct: 477 PALLMVSGAIGAQLSYLLPTGSPSNVVGFSTGYITIKDLVATGLPLKIVAIAALTVLLPT 536
Query: 516 LGKLLLSL 523
LG + +
Sbjct: 537 LGSTIFGM 544
>ref|YP_077725.1| Sodium/sulphate symporter [Bacillus licheniformis ATCC 14580]
gi|52002145|gb|AAU22087.1| Sodium/sulphate symporter
[Bacillus licheniformis ATCC 14580]
gi|52784308|ref|YP_090137.1| hypothetical protein
BLi00488 [Bacillus licheniformis ATCC 14580]
gi|52346810|gb|AAU39444.1| putative protein [Bacillus
licheniformis DSM 13]
Length = 546
Score = 308 bits (789), Expect = 3e-82
Identities = 184/505 (36%), Positives = 281/505 (55%), Gaps = 34/505 (6%)
Query: 36 QNFYIILGPLLSIIICLLVKLDAPTTSTKM-LGVIAWVFTWWITNAVPLPVTSMCPLFLF 94
Q ++LGP L + L + + +M L WV WWIT AVP+P S+ P+ L
Sbjct: 57 QKIGLLLGPALFFAVLLFFFPEGLSYEGRMVLATTLWVAVWWITEAVPIPAASLLPIVLL 116
Query: 95 PLFGIASADTVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSML 154
PL G V SY D ++ L LG F++ALA+ER+N+H+R+ALN+ V + S +
Sbjct: 117 PLTGALEGAAVTSSYGDPIVFLFLGGFLIALAMERWNLHKRIALNIISVV---GTSTSRI 173
Query: 155 LLGLCATTFFVSMWLHNVATAVMMMPVATGILHRLPPA--EEQPELM---NKFSRAVILT 209
+LG A T F+SMW+ N A +MM+P+ T I+H++ E+ +L KFS+A+I +
Sbjct: 174 VLGFMAATGFLSMWVSNTAAVMMMLPIGTAIIHQVSAVIKSERKDLAAEEAKFSKALIFS 233
Query: 210 VVYATPIGGISTLTGTGVNLIIIGMWKSLAPEAKPISFNSWFFFGFPVAVLLLLCFWCIL 269
+ YA IGG+ TL GT N+I+ K L +SF W F PV V+LL+ W
Sbjct: 234 IGYAGTIGGLGTLIGTPPNIILAANIKKL--YGVEVSFGGWMAFAVPVVVILLVAVW--- 288
Query: 270 CLIYVRKGSSRALSDYL--DRALLKRDLEALGPMSFAEKMVLCVFGLLIVLWMTRRI--S 325
+Y+ K + L + L+ + LG MSF E MVL VFG +W+TR
Sbjct: 289 --LYLTKVAHPIKMKELPGGKELILEEKRKLGKMSFEETMVLLVFGFAAFMWVTRTFLWD 346
Query: 326 DDLPGWGVLFHGLVGDGSISVLAAVLLFIIPNMKQNGEKLMDWNECKKLPWNLILLLGAG 385
D +PG + D I++ AA LLF+IP++ + G +++DW+ K LPW ++LL G G
Sbjct: 347 DKIPG--------IDDTMIAIFAASLLFLIPSLNKGG-RVLDWSVSKDLPWGILLLFGGG 397
Query: 386 FALADGVQSSGLADVMSKALEFLKDVPYLAIVPAVSLLCSIITEFITSNDATATLLVPLL 445
ALA G + +GLA+ + L L ++ IV + L +TE ITSN ATAT+++P+L
Sbjct: 398 LALATGFKETGLAEWIGGRLTVLDGFNFVVIVIISTALVLFLTE-ITSNTATATMILPVL 456
Query: 446 YHIAINMHVHPLLLMIPGGIATEFAFWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAG 505
+A+ ++VHP LM+P +A AF LP TP N + FA+G ++I +M++ G + +
Sbjct: 457 ASLALALNVHPYALMVPAAMAANCAFMLPVGTPPNAIIFASGKLKISEMVRTGFVINIFT 516
Query: 506 ----VSVLALLMPTLGKLLLSLFSD 526
V + ++P L + L++F D
Sbjct: 517 LILIVGAVFYILPHLWGVDLTVFPD 541
>ref|ZP_00472657.1| Sodium/sulphate symporter [Chromohalobacter salexigens DSM 3043]
gi|67520172|gb|EAM24118.1| Sodium/sulphate symporter
[Chromohalobacter salexigens DSM 3043]
Length = 481
Score = 299 bits (766), Expect = 1e-79
Identities = 172/482 (35%), Positives = 277/482 (56%), Gaps = 25/482 (5%)
Query: 42 LGPLLSIIICLLVKLDAPTTSTKM--LGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFGI 99
LGPL +I+CLL+ A ++T +G + TWW T A+P+P TS+ P+ L P GI
Sbjct: 20 LGPLW-VILCLLLPAPAEMSATAWACVGFSLLMATWWATEAIPIPATSLLPMVLAPALGI 78
Query: 100 ASADTVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGLC 159
+ DTVA +Y D +I L LG F+L +A++R+N+HRR+AL V L G P + G
Sbjct: 79 SDLDTVAPAYADPIIYLFLGGFLLGIAMQRWNLHRRIALKV-LSIVGQ--KPRQQIAGFM 135
Query: 160 ATTFFVSMWLHNVATAVMMMPVATGILHRLPPAEEQPELMNKFSRAVILTVVYATPIGGI 219
T F+SMW+ N AT++MM+P+ ++ L + + + ++S A++L + YA IGG+
Sbjct: 136 IATGFISMWVSNTATSIMMLPIGMSVVSLLGGNDSRD--VARYSTALLLAIAYAASIGGV 193
Query: 220 STLTGTGVNLIIIGMWKSLAPEAKPISFNSWFFFGFPVAVLLLLCFWCILCLIYVRKGSS 279
+TL GT N ++ G LA E I F W G P+++ ++ W L RKG
Sbjct: 194 ATLIGTPPNALLAGY---LAGEGIDIGFAQWMTIGLPISIAMMAVTWWWL----TRKGFD 246
Query: 280 RALSDYLDRALLKRDLEALGPMSFAEKMVLCVFGLLIVLWMTRRISDDLP-GWGVLFHGL 338
S+ D+ +++R+L+ALG MS AE+ V +F + W+ R + + W
Sbjct: 247 LEQSEDGDQ-VIRRELDALGGMSRAERRVGLIFLVAAAAWILRPLMNQAGLDW------- 298
Query: 339 VGDGSISVLAAVLLFIIPNMKQNGEKLMDWNECKKLPWNLILLLGAGFALADGVQSSGLA 398
+ D +I++ A + LF++P+ + L+DW K LPW ++LL G G ALA ++ SGLA
Sbjct: 299 LSDTTIALAAGLALFLVPSGDKESGALLDWESAKDLPWGVLLLFGGGLALAGMIKRSGLA 358
Query: 399 DVMSKALEFLKDVPYLAIVPAVSLLCSIITEFITSNDATATLLVPLLYHIAINMHVHPLL 458
+ +++ L +P L +V + L+ +TE +TSN ATA +PLL +AI++ + PLL
Sbjct: 359 EWIAQQLSVFSALPLLMLVGIIVLVIIFLTE-VTSNTATAAAFLPLLGALAISLDIDPLL 417
Query: 459 LMIPGGIATEFAFWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAGVSVLALLMPTLGK 518
+ P +A AF +P +TP N + FATGH++I+ M++ G+ L + G ++ L+ L +
Sbjct: 418 ITAPAAVAASCAFMMPVATPPNAIVFATGHMKIQSMIRAGLMLNLIGTVLITLITVGLLR 477
Query: 519 LL 520
LL
Sbjct: 478 LL 479
>ref|YP_041382.1| putative sodium:sulfate symporter [Staphylococcus aureus subsp.
aureus MRSA252] gi|49242287|emb|CAG40994.1| putative
sodium:sulfate symporter [Staphylococcus aureus subsp.
aureus MRSA252]
Length = 520
Score = 298 bits (762), Expect = 4e-79
Identities = 174/491 (35%), Positives = 276/491 (55%), Gaps = 22/491 (4%)
Query: 40 IILGPLLSIIICLLVK-LDAPTTSTKMLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFG 98
+ILGPLL ++ L D P +L + W+ TWWIT A+P+ TS+ P+ L PL
Sbjct: 36 LILGPLLFLLTLLFFHPQDLPWKGVYVLAITLWIATWWITEAIPIAATSLLPIVLLPLGH 95
Query: 99 IASADTVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGL 158
I + + V+ Y +D+I L LG FILA+A+ER+N+H R+AL + + + S +LLG
Sbjct: 96 ILTPEQVSSEYGNDIIFLFLGGFILAIAMERWNLHTRVALTIINLI---GASTSKILLGF 152
Query: 159 CATTFFVSMWLHNVATAVMMMPVATGIL---HRLPPAEEQPELMNKFSRAVILTVVYATP 215
T F+SM++ N A ++M+P+ I+ H L A + KF ++++L + YA
Sbjct: 153 MVATGFLSMFVSNTAAVMIMIPIGLAIIKEAHDLQEANTNQTSIQKFEKSLVLAIGYAGT 212
Query: 216 IGGISTLTGTGVNLIIIGMWKSLAPEAKPISFNSWFFFGFPVAVLLLLCFWCILCLIYVR 275
IGG+ TL GT +I+ G + + ISF W G P ++LL W +Y+R
Sbjct: 213 IGGLGTLIGTPPLIILKGQY--MQHFGHEISFAKWMIVGIPTVIVLLGITW-----LYLR 265
Query: 276 KGSSRALSDYL--DRALLKRDLEALGPMSFAEKMVLCVFGLLIVLWMTRRISDDLPGWGV 333
+ R YL + L+K+ L+ LG M + EK+V +F L +LW+TR L W V
Sbjct: 266 YVAFRHDLKYLPGGQTLIKQKLDELGKMKYEEKVVQTIFVLASLLWITREFL--LKKWEV 323
Query: 334 LFHGLVGDGSISVLAAVLLFIIP-NMKQNGEKLMDWNECKKLPWNLILLLGAGFALADGV 392
V DG+I++ ++LLFIIP + +++DW K+LPW +++L G G ALA G+
Sbjct: 324 T--SSVADGTIAIFISILLFIIPAKNTEKHRRIIDWEVAKELPWGVLILFGGGLALAKGI 381
Query: 393 QSSGLADVMSKALEFLKDVPYLAIVPAVSLLCSIITEFITSNDATATLLVPLLYHIAINM 452
SGLA + + L+ L V + IV +++ +TE +TSN ATAT+++P+L +++ +
Sbjct: 382 SESGLAKWLGEQLKSLNGVSPILIVIVITIFVLFLTE-VTSNTATATMILPILATLSVAV 440
Query: 453 HVHPLLLMIPGGIATEFAFWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAGVSVLALL 512
VHPLLLM P +A A+ LP TP N + F +G I IK M VG + + ++ L+
Sbjct: 441 GVHPLLLMAPAAMAANCAYMLPVGTPPNAIIFGSGKISIKQMASVGFWVNLISAIIIILV 500
Query: 513 MPTLGKLLLSL 523
+ + ++L +
Sbjct: 501 VYYIMPIVLGI 511
>ref|YP_186803.1| sodium-dependent transporter [Staphylococcus aureus subsp. aureus
COL] gi|14247688|dbj|BAB58078.1| similar to
sodium-dependent transporter [Staphylococcus aureus
subsp. aureus Mu50] gi|15927490|ref|NP_375023.1|
hypothetical protein SA1732 [Staphylococcus aureus
subsp. aureus N315] gi|13701709|dbj|BAB43002.1| SA1732
[Staphylococcus aureus subsp. aureus N315]
gi|25506563|pir||C89980 hypothetical protein SA1732
[imported] - Staphylococcus aureus (strain N315)
gi|57284855|gb|AAW36949.1| sodium-dependent transporter
[Staphylococcus aureus subsp. aureus COL]
gi|15924906|ref|NP_372440.1| similar to sodium-dependent
transporter [Staphylococcus aureus subsp. aureus Mu50]
Length = 520
Score = 297 bits (761), Expect = 5e-79
Identities = 174/491 (35%), Positives = 276/491 (55%), Gaps = 22/491 (4%)
Query: 40 IILGPLLSIIICLLVK-LDAPTTSTKMLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFG 98
+ILGPLL ++ L D P +L + W+ TWWIT A+P+ TS+ P+ L PL
Sbjct: 36 LILGPLLFLLTLLFFHPQDLPWKGVYVLAITLWIATWWITEAIPIAATSLLPIVLLPLGH 95
Query: 99 IASADTVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGL 158
I + + V+ Y +D+I L LG FILA+A+ER+N+H R+AL + + + S +LLG
Sbjct: 96 ILTPEQVSSEYGNDIIFLFLGGFILAIAMERWNLHTRVALTIINLI---GASTSKILLGF 152
Query: 159 CATTFFVSMWLHNVATAVMMMPVATGIL---HRLPPAEEQPELMNKFSRAVILTVVYATP 215
T F+SM++ N A ++M+P+ I+ H L A + KF ++++L + YA
Sbjct: 153 MVATGFLSMFVSNTAAVMIMIPIGLAIIKEAHDLQEANTNQTSIQKFEKSLVLAIGYAGT 212
Query: 216 IGGISTLTGTGVNLIIIGMWKSLAPEAKPISFNSWFFFGFPVAVLLLLCFWCILCLIYVR 275
IGG+ TL GT +I+ G + + ISF W G P ++LL W +Y+R
Sbjct: 213 IGGLGTLIGTPPLIILKGQY--MQHFGHEISFAKWMIVGIPTVIVLLGITW-----LYLR 265
Query: 276 KGSSRALSDYL--DRALLKRDLEALGPMSFAEKMVLCVFGLLIVLWMTRRISDDLPGWGV 333
+ R YL + L+K+ L+ LG M + EK+V +F L +LW+TR L W V
Sbjct: 266 YVAFRHDLKYLPGGQTLIKQKLDELGKMKYEEKVVQTIFVLASLLWITREFL--LKKWEV 323
Query: 334 LFHGLVGDGSISVLAAVLLFIIP-NMKQNGEKLMDWNECKKLPWNLILLLGAGFALADGV 392
V DG+I++ ++LLFIIP + +++DW K+LPW +++L G G ALA G+
Sbjct: 324 T--SSVADGTIAIFISILLFIIPAKNTEKHRRIIDWEVAKELPWGVLILFGGGLALAKGI 381
Query: 393 QSSGLADVMSKALEFLKDVPYLAIVPAVSLLCSIITEFITSNDATATLLVPLLYHIAINM 452
SGLA + + L+ L V + IV +++ +TE +TSN ATAT+++P+L +++ +
Sbjct: 382 SESGLAKWLGEQLKSLNGVSPILIVIVITIFVLFLTE-VTSNTATATMILPILATLSVAV 440
Query: 453 HVHPLLLMIPGGIATEFAFWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAGVSVLALL 512
VHPLLLM P +A A+ LP TP N + F +G I IK M VG + + ++ L+
Sbjct: 441 GVHPLLLMAPAAMAANCAYMLPVGTPPNAIIFGSGKISIKQMASVGFWVNLISAIIIILV 500
Query: 513 MPTLGKLLLSL 523
+ + ++L +
Sbjct: 501 VYYVMPIVLGI 511
>gb|AAN86530.1| Na+-coupled citrate transporter protein [Homo sapiens]
gi|29171306|ref|NP_808218.1| solute carrier family 13
(sodium-dependent citrate transporter), member 5 [Homo
sapiens]
Length = 568
Score = 297 bits (761), Expect = 5e-79
Identities = 186/547 (34%), Positives = 292/547 (53%), Gaps = 72/547 (13%)
Query: 40 IILGPLLSIIICLLVKLDAPTTSTKMLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFGI 99
+ + PLL + + +L+ P + VI + +W T +PL VTS+ P+ LFPLF I
Sbjct: 17 LFVTPLLLLPLVILM----PAKFVRCAYVIILMAIYWCTEVIPLAVTSLMPVLLFPLFQI 72
Query: 100 ASADTVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGLC 159
+ V YM D L LG I+A+AVER+N+H+R+AL TL++ G P+ L+LG
Sbjct: 73 LDSRQVCVQYMKDTNMLFLGGLIVAVAVERWNLHKRIALR-TLLWVG--AKPARLMLGFM 129
Query: 160 ATTFFVSMWLHNVATAVMMMPVATGILHRLPPA--------------------------- 192
T +SMW+ N AT MM+P+ IL ++
Sbjct: 130 GVTALLSMWISNTATTAMMVPIVEAILQQMEATSAATEAGLELVDKGKAKELPGSQVIFE 189
Query: 193 -----EEQPELMNKFSRAVILTVVYATPIGGISTLTGTGVNLIIIGMWKSLAPEAKP-IS 246
+++ + + +A+ L + YA IGG +TLTGTG N++++G L P++K ++
Sbjct: 190 GPTLGQQEDQERKRLCKAMTLCICYAASIGGTATLTGTGPNVVLLGQMNELFPDSKDLVN 249
Query: 247 FNSWFFFGFPVAVLLLLCFWCILCLIYVRKGSSRALSDYLDRA--------LLKRDLEAL 298
F SWF F FP +++LL W L +Y+R ++ L+ +L+ + L
Sbjct: 250 FASWFAFAFPNMLVMLLFAWLWLQFVYMRFNFKKSWGCGLESKKNEKAALKVLQEEYRKL 309
Query: 299 GPMSFAEKMVLCVFGLLIVLWMTRRISDDLPGWGVLFH-----GLVGDGSISVLAAVLLF 353
GP+SFAE VL F LL++LW +R +PGW + V D ++++ A LLF
Sbjct: 310 GPLSFAEINVLICFFLLVILWFSRD-PGFMPGWLTVAWVEGETKYVSDATVAIFVATLLF 368
Query: 354 IIPNMK-------QNGEK---------LMDWNECK-KLPWNLILLLGAGFALADGVQSSG 396
I+P+ K Q E+ L+DW + K+PW ++LLLG GFALA G ++SG
Sbjct: 369 IVPSQKPKFNFRSQTEEERKTPFYPPPLLDWKVTQEKVPWGIVLLLGGGFALAKGSEASG 428
Query: 397 LADVMSKALEFLKDVPYLAIVPAVSLLCSIITEFITSNDATATLLVPLLYHIAINMHVHP 456
L+ M K +E L VP AI +SLL ++ TE TSN AT TL +P+ ++ ++ ++P
Sbjct: 429 LSVWMGKQMEPLHAVPPAAITLILSLLVAVFTE-CTSNVATTTLFLPIFASMSRSIGLNP 487
Query: 457 LLLMIPGGIATEFAFWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAGVSVLALLMPTL 516
L +M+P ++ FAF LP +TP N + F GH+++ DM+K GV + + GV + L + T
Sbjct: 488 LYIMLPCTLSASFAFMLPVATPPNAIVFTYGHLKVADMVKTGVIMNIIGVFCVFLAVNTW 547
Query: 517 GKLLLSL 523
G+ + L
Sbjct: 548 GRAIFDL 554
>pir||JC7911 Na+-coupled citrate transporter NaCT - human
Length = 568
Score = 297 bits (761), Expect = 5e-79
Identities = 186/547 (34%), Positives = 292/547 (53%), Gaps = 72/547 (13%)
Query: 40 IILGPLLSIIICLLVKLDAPTTSTKMLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFGI 99
+ + PLL + + +L+ P + VI + +W T +PL VTS+ P+ LFPLF I
Sbjct: 17 LFVTPLLLLPLVILM----PAKFVRCAYVIILMAIYWCTEVIPLAVTSLMPVLLFPLFQI 72
Query: 100 ASADTVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGLC 159
+ V YM D L LG I+A+AVER+N+H+R+AL TL++ G P+ L+LG
Sbjct: 73 LDSRQVCVQYMKDTNMLFLGGLIVAVAVERWNLHKRIALR-TLLWVG--AKPARLMLGFM 129
Query: 160 ATTFFVSMWLHNVATAVMMMPVATGILHRLPPA--------------------------- 192
T +SMW+ N AT MM+P+ IL ++
Sbjct: 130 GVTALLSMWISNTATTAMMVPIVEAILQQMEATSAATEAGLELVDKGKAKELPGSQVIFE 189
Query: 193 -----EEQPELMNKFSRAVILTVVYATPIGGISTLTGTGVNLIIIGMWKSLAPEAKP-IS 246
+++ + + +A+ L + YA IGG +TLTGTG N++++G L P++K ++
Sbjct: 190 GPILGQQEDQERKRLCKAMTLCICYAASIGGTATLTGTGPNVVLLGQMNELFPDSKDLVN 249
Query: 247 FNSWFFFGFPVAVLLLLCFWCILCLIYVRKGSSRALSDYLDRA--------LLKRDLEAL 298
F SWF F FP +++LL W L +Y+R ++ L+ +L+ + L
Sbjct: 250 FASWFAFAFPNMLVMLLFAWLWLQFVYMRFNFKKSWGCGLESKKNEKAALKVLQEEYRKL 309
Query: 299 GPMSFAEKMVLCVFGLLIVLWMTRRISDDLPGWGVLFH-----GLVGDGSISVLAAVLLF 353
GP+SFAE VL F LL++LW +R +PGW + V D ++++ A LLF
Sbjct: 310 GPLSFAEINVLICFFLLVILWFSRD-PGFMPGWLTVAWVEGETKYVSDATVAIFVATLLF 368
Query: 354 IIPNMK-------QNGEK---------LMDWNECK-KLPWNLILLLGAGFALADGVQSSG 396
I+P+ K Q E+ L+DW + K+PW ++LLLG GFALA G ++SG
Sbjct: 369 IVPSQKPKFNFRSQTEEERKTPFYPPPLLDWKVTQEKVPWGIVLLLGGGFALAKGSEASG 428
Query: 397 LADVMSKALEFLKDVPYLAIVPAVSLLCSIITEFITSNDATATLLVPLLYHIAINMHVHP 456
L+ M K +E L VP AI +SLL ++ TE TSN AT TL +P+ ++ ++ ++P
Sbjct: 429 LSVWMGKQMEPLHAVPPAAITLILSLLVAVFTE-CTSNVATTTLFLPIFASMSRSIGLNP 487
Query: 457 LLLMIPGGIATEFAFWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAGVSVLALLMPTL 516
L +M+P ++ FAF LP +TP N + F GH+++ DM+K GV + + GV + L + T
Sbjct: 488 LYIMLPCTLSASFAFMLPVATPPNAIVFTYGHLKVADMVKTGVIMNIIGVFCVFLAVNTW 547
Query: 517 GKLLLSL 523
G+ + L
Sbjct: 548 GRAIFDL 554
>emb|CAG43645.1| putative sodium:sulfate symporter [Staphylococcus aureus subsp.
aureus MSSA476] gi|21205027|dbj|BAB95722.1| MW1857
[Staphylococcus aureus subsp. aureus MW2]
gi|49486736|ref|YP_043957.1| putative sodium:sulfate
symporter [Staphylococcus aureus subsp. aureus MSSA476]
gi|21283586|ref|NP_646674.1| hypothetical protein MW1857
[Staphylococcus aureus subsp. aureus MW2]
Length = 520
Score = 297 bits (760), Expect = 6e-79
Identities = 173/491 (35%), Positives = 276/491 (55%), Gaps = 22/491 (4%)
Query: 40 IILGPLLSIIICLLVK-LDAPTTSTKMLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFG 98
+ILGPLL ++ L D P +L + W+ TWWIT A+P+ TS+ P+ L PL
Sbjct: 36 LILGPLLFLLTLLFFHPQDLPWKGVYVLAITLWIATWWITEAIPIAATSLLPIVLLPLGH 95
Query: 99 IASADTVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGL 158
I + + V+ Y +D+I L LG FILA+A+ER+N+H R+AL + + + S +LLG
Sbjct: 96 ILTPEQVSSEYGNDIIFLFLGGFILAIAMERWNLHTRVALTIINLI---GASTSKILLGF 152
Query: 159 CATTFFVSMWLHNVATAVMMMPVATGIL---HRLPPAEEQPELMNKFSRAVILTVVYATP 215
T F+SM++ N A ++M+P+ I+ H L A + KF ++++L + YA
Sbjct: 153 MVATGFLSMFVSNTAAVMIMIPIGLAIIKEAHDLQEANTNQTSIQKFEKSLVLAIGYAGT 212
Query: 216 IGGISTLTGTGVNLIIIGMWKSLAPEAKPISFNSWFFFGFPVAVLLLLCFWCILCLIYVR 275
IGG+ TL GT +I+ G + + ISF W G P ++LL W +Y+R
Sbjct: 213 IGGLGTLIGTPPLIILKGQY--MQHFGHEISFAKWMIVGIPTVIVLLGITW-----LYLR 265
Query: 276 KGSSRALSDYL--DRALLKRDLEALGPMSFAEKMVLCVFGLLIVLWMTRRISDDLPGWGV 333
+ R YL + L+K+ L+ LG M + EK+V +F L +LW+TR L W V
Sbjct: 266 YVAFRHDLKYLPGGQTLIKQKLDELGKMKYEEKVVQTIFVLASLLWITREFL--LKKWEV 323
Query: 334 LFHGLVGDGSISVLAAVLLFIIP-NMKQNGEKLMDWNECKKLPWNLILLLGAGFALADGV 392
V DG+I++ ++LLF+IP + +++DW K+LPW +++L G G ALA G+
Sbjct: 324 T--SSVADGTIAIFISILLFVIPAKNTEKHRRIIDWEVAKELPWGVLILFGGGLALAKGI 381
Query: 393 QSSGLADVMSKALEFLKDVPYLAIVPAVSLLCSIITEFITSNDATATLLVPLLYHIAINM 452
SGLA + + L+ L V + IV +++ +TE +TSN ATAT+++P+L +++ +
Sbjct: 382 SESGLAKWLGEQLKSLNGVSPILIVIVITIFVLFLTE-VTSNTATATMILPILATLSVAV 440
Query: 453 HVHPLLLMIPGGIATEFAFWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAGVSVLALL 512
VHPLLLM P +A A+ LP TP N + F +G I IK M VG + + ++ L+
Sbjct: 441 GVHPLLLMAPAAMAANCAYMLPVGTPPNAIIFGSGKISIKQMASVGFWVNLISAIIIILV 500
Query: 513 MPTLGKLLLSL 523
+ + ++L +
Sbjct: 501 VYYVMPIVLGI 511
>dbj|BAD18766.1| unnamed protein product [Homo sapiens]
Length = 568
Score = 292 bits (747), Expect = 2e-77
Identities = 185/547 (33%), Positives = 291/547 (52%), Gaps = 72/547 (13%)
Query: 40 IILGPLLSIIICLLVKLDAPTTSTKMLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFGI 99
+ + PLL + + +L+ P + VI + +W T +PL VTS+ P+ LFPLF I
Sbjct: 17 LFVTPLLLLPLVILM----PAKFVRCAYVIILMAIYWCTEVIPLAVTSLMPVLLFPLFQI 72
Query: 100 ASADTVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGLC 159
+ V YM D L LG I+A+AVER+N+H+R+AL TL++ G P+ L+LG
Sbjct: 73 LDSRQVCVQYMKDTNMLFLGGLIVAVAVERWNLHKRIALR-TLLWVG--AKPARLMLGFM 129
Query: 160 ATTFFVSMWLHNVATAVMMMPVATGILHRLPPA--------------------------- 192
T +SMW+ N AT MM+P+ IL ++
Sbjct: 130 GVTALLSMWISNTATTAMMVPIVEAILQQMEATSAATEAGLELVDKGKAKELPGSQVIFE 189
Query: 193 -----EEQPELMNKFSRAVILTVVYATPIGGISTLTGTGVNLIIIGMWKSLAPEAKP-IS 246
+++ + + +A+ L + YA IGG +TLTGTG N++++G L P++K ++
Sbjct: 190 GPTLGQQEDQERKRLCKAMTLCICYAASIGGTATLTGTGPNVVLLGQMNELFPDSKDLVN 249
Query: 247 FNSWFFFGFPVAVLLLLCFWCILCLIYVRKGSSRALSDYLDRA--------LLKRDLEAL 298
F SWF F FP +++LL L +Y+R ++ L+ +L+ + L
Sbjct: 250 FASWFAFAFPNMLVMLLFARLWLQFVYMRFNFKKSWGCGLESKKNEKAALKVLQEEYRKL 309
Query: 299 GPMSFAEKMVLCVFGLLIVLWMTRRISDDLPGWGVLFH-----GLVGDGSISVLAAVLLF 353
GP+SFAE VL F LL++LW +R +PGW + V D ++++ A LLF
Sbjct: 310 GPLSFAEINVLICFFLLVILWFSRD-PGFMPGWLTVAWVEGETKYVSDATVAIFVATLLF 368
Query: 354 IIPNMK-------QNGEK---------LMDWNECK-KLPWNLILLLGAGFALADGVQSSG 396
I+P+ K Q E+ L+DW + K+PW ++LLLG GFALA G ++SG
Sbjct: 369 IVPSQKPRFNFRSQTEEERKTPFYPPPLLDWKVTQEKVPWGIVLLLGGGFALAKGSEASG 428
Query: 397 LADVMSKALEFLKDVPYLAIVPAVSLLCSIITEFITSNDATATLLVPLLYHIAINMHVHP 456
L+ M K +E L VP AI +SLL ++ TE TSN AT TL +P+ ++ ++ ++P
Sbjct: 429 LSVWMGKQMEPLHAVPPAAITLILSLLVAVFTE-CTSNVATTTLFLPIFASMSRSIGLNP 487
Query: 457 LLLMIPGGIATEFAFWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAGVSVLALLMPTL 516
L +M+P ++ FAF LP +TP N + F GH+++ DM+K GV + + GV + L + T
Sbjct: 488 LYIMLPCTLSASFAFMLPVATPPNAIVFTYGHLKVADMVKTGVIMNIIGVFCVFLAVNTW 547
Query: 517 GKLLLSL 523
G+ + L
Sbjct: 548 GRAIFDL 554
>ref|YP_077982.1| Sodium/sulphate symporter [Bacillus licheniformis ATCC 14580]
gi|52002402|gb|AAU22344.1| Sodium/sulphate symporter
[Bacillus licheniformis ATCC 14580]
gi|52784558|ref|YP_090387.1| hypothetical protein
BLi00759 [Bacillus licheniformis ATCC 14580]
gi|52347060|gb|AAU39694.1| putative protein [Bacillus
licheniformis DSM 13]
Length = 474
Score = 285 bits (730), Expect = 2e-75
Identities = 166/475 (34%), Positives = 267/475 (55%), Gaps = 26/475 (5%)
Query: 40 IILGPLLSIIIC-LLVKLDAPTTSTKMLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFG 98
++LGP+ ++I L+ + + +L V AW WWIT AVP+P S+ P+ L P G
Sbjct: 12 LLLGPISFLLILGLIPEAELAYAPRIVLAVTAWTAVWWITEAVPIPAASLLPVILLPTLG 71
Query: 99 IASADTVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGL 158
+T A +Y D ++ + +G FI+A+A+E++N+H+R+AL++ + ++LG+
Sbjct: 72 GLDMETTAKAYGDPIVFMYMGGFIIAIAIEKWNLHKRMALHILKRIGAE---SHRIVLGV 128
Query: 159 CATTFFVSMWLHNVATAVMMMPVATGILHRLPPAE-EQPELMNKFSRAVILTVVYATPIG 217
A T F+SMW+ N ATA+MM+PVA ++ + E + E + KFS++++L V YA IG
Sbjct: 129 MAATAFLSMWISNAATALMMLPVALAVIKEVQEKEILKGESLQKFSKSILLAVAYAASIG 188
Query: 218 GISTLTGTGVNLIIIGMWKSLAPEAKPISFNSWFFFGFPVAVLLLLCFWCILCLIYVRKG 277
G++TL G+ N + M K + I F WF FGFPV++L L + L I +
Sbjct: 189 GLATLVGSVPNAVFAAMSKQMLNH--EIMFFEWFLFGFPVSILFLAILYVYLTKIKFKVS 246
Query: 278 SSRALS-DYLDRALLKRDLEALGPMSFAEKMVLCVFGLLIVLWMTRRISDDLPGWGVLFH 336
+ + D++D DL+ALG M+ EK V VF L LWM + I LP
Sbjct: 247 AFQGKKPDFID-----HDLKALGAMTREEKSVFTVFLLTACLWMFKFI---LP------- 291
Query: 337 GLVGDGSISVLAAVLLFIIPNMKQNGEKLMDWNECKKLPWNLILLLGAGFALADGVQSSG 396
+ D SI++ AVLLF+IP+ K E++++W + + LPW L+LL G G +LA G ++
Sbjct: 292 FQISDTSIAIFGAVLLFLIPSTKD--ERILEWKDMQSLPWGLLLLFGGGLSLAAGFSATN 349
Query: 397 LADVMSKALEFLKDVPYLAIVPAVSLLCSIITEFITSNDATATLLVPLLYHIAINMHVHP 456
L + L+ L+ PYL ++ ++ +TE I SN A A +++P+ + + V P
Sbjct: 350 LTAWIGGKLKLLEGHPYLFVLIILTAAILFMTE-IMSNTAVANMVIPMTIGLGAALQVEP 408
Query: 457 LLLMIPGGIATEFAFWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAGVSVLAL 511
LM +A+ AF LP STP N F++ + IKDM++ G L +AGV ++ +
Sbjct: 409 YGLMAAAALASSCAFMLPISTPPNAAVFSSNLLTIKDMVRAGFWLNIAGVFLIVI 463
>ref|YP_177486.1| sodium:dicarboxylate cotransporter [Bacillus clausii KSM-K16]
gi|56911998|dbj|BAD66525.1| sodium:dicarboxylate
cotransporter [Bacillus clausii KSM-K16]
Length = 556
Score = 284 bits (726), Expect = 6e-75
Identities = 170/503 (33%), Positives = 274/503 (53%), Gaps = 35/503 (6%)
Query: 40 IILGPLLSIIICLLVKLDA-PTTSTKMLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFG 98
++LGPLL + + + PT +L V W+ TWW+T A+P+P TS+ PL L P+ G
Sbjct: 70 LLLGPLLFFLTLMFFHPEGLPTEGRHVLAVTLWIATWWMTEAIPIPATSLLPLILLPITG 129
Query: 99 IASADTVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGL 158
VA SY +D+I L LG F +A A+E++N+H+R+AL + V + +LLG
Sbjct: 130 AMEGSAVASSYGNDIIFLFLGGFFIATAMEKWNLHKRIALFIIAVI---GTSTQRILLGF 186
Query: 159 CATTFFVSMWLHNVATAVMMMPVATGILHRLP---PAEEQPELMNKFSRAVILTVVYATP 215
A T F+SMW+ N A +MM+P+A I ++ +++ + KF +A++ V YA
Sbjct: 187 MAATAFLSMWVSNTAAVMMMVPMALAITAQVAETLKGQKEGRELPKFEKAMLFGVGYAGT 246
Query: 216 IGGISTLTGTGVNLIIIGMWKSLAPEAKPISFNSWFFFGFPVAVLLLLCFWCILCLIYVR 275
IGG +TL GT +I G + L +SF SW F P+ +++LL W L I +
Sbjct: 247 IGGFATLIGTPPTIIFAGQVRELF--GIEVSFASWMLFATPLMLVVLLFTWFYLGRIAFK 304
Query: 276 KGSSRALSDYLDRALLKRDLEALGPMSFAEKMVLCVFGLLIVLWMTRRI--SDD------ 327
+ +++ + LG +S+ E +V VF +W+T+ S D
Sbjct: 305 TKIKELPG---GKEVIQSERSKLGRISYEEGIVAGVFAFAAFMWITKDFFWSGDNAMIFQ 361
Query: 328 LPGWGVLFHGLVGDGSISVLAAVLLFIIPNMKQNGEKLMDWNECKKLPWNLILLLGAGFA 387
LPG + DG ++++A + LF+IP + +++DW + + +PW ++LL G G A
Sbjct: 362 LPG--------ISDGMVAIMATMFLFLIPG--KASARILDWADSRDIPWGVLLLFGGGLA 411
Query: 388 LADGVQSSGLADVMSKALEFLKDVPYLAIVPAVSLLCSIITEFITSNDATATLLVPLLYH 447
+A G QSSGL+ + + L L + L I+ +LL ++TE ITSN ATAT+++P++
Sbjct: 412 IAAGFQSSGLSVWLGEQLTVLDGLHMLLIIGGATLLIMMLTE-ITSNTATATMIMPIVAS 470
Query: 448 IAINMHVHPLLLMIPGGIATEFAFWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAGVS 507
+A+ ++VHP LM P IA AF LP TP N + F TG ++I +M++ G L V
Sbjct: 471 LAVAINVHPFALMAPCAIAANCAFMLPVGTPPNAIIFGTGKVKIIEMVRAGFWLNVFSTI 530
Query: 508 VLAL----LMPTLGKLLLSLFSD 526
++ L +P + + LS+F D
Sbjct: 531 LIVLAVYAFLPIVFDIDLSVFPD 553
>ref|NP_693461.1| sodium-dependent transporter [Oceanobacillus iheyensis HTE831]
gi|22778226|dbj|BAC14496.1| sodium-dependent transporter
[Oceanobacillus iheyensis HTE831]
Length = 552
Score = 282 bits (722), Expect = 2e-74
Identities = 162/492 (32%), Positives = 268/492 (53%), Gaps = 23/492 (4%)
Query: 40 IILGPLLSIIICLLVKLDAPTTSTK-MLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFG 98
+ LGP+ I+ L + ++ + +L W+ WW+T A+P+P T++ P+ LFPL
Sbjct: 66 LFLGPISFILTLLFFSPEGLSSEGQAVLASTIWIAIWWMTEAIPIPATALLPIILFPLTN 125
Query: 99 IASADTVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGL 158
+Y D I L +G F++ALA+ER+N+HRR+A+++ V N ++LG
Sbjct: 126 GLDIGATTSAYGSDTIFLFMGGFVIALAMERWNLHRRIAISIISVI---GTNTDRIILGF 182
Query: 159 CATTFFVSMWLHNVATAVMMMPVATGILHRLPPA-------EEQPELMNKFSRAVILTVV 211
T F+SM++ N ATA+MM+P+ I++++ + PE + F ++++L +
Sbjct: 183 MVATGFLSMFISNTATAMMMVPIGLAIIYQVSDTLKEQGGFDTSPENFS-FGKSLMLGIA 241
Query: 212 YATPIGGISTLTGTGVNLIIIGMWKSLAPEAKPISFNSWFFFGFPVAVLLLLCFWCILCL 271
Y+ IGG++TL GT N+I G ++ +SF SW FG P A + + W L
Sbjct: 242 YSASIGGMATLIGTPPNVIFAGTINTM--YGIEVSFASWMMFGVPFAWIFIFLTWFYLSK 299
Query: 272 IYVRKGSSRALSDYLDRALLKRDLEALGPMSFAEKMVLCVFGLLIVLWMTRRISDDLPGW 331
I + S ++K + + LG S+ EK V VF L + W++R
Sbjct: 300 IAFKLEVSHIPG---GAKIIKEEKKKLGNASYEEKAVFVVFILAAIAWISREFL-----L 351
Query: 332 GVLFHGLVGDGSISVLAAVLLFIIPNMKQNGEKLMDWNECKKLPWNLILLLGAGFALADG 391
G + D I++ AA++LF+IP + G+ LM+W KLPW ++LL G G A+A G
Sbjct: 352 GPYVSENINDAIIAITAALVLFLIPAKNKKGDFLMNWESAIKLPWGILLLFGGGLAIAAG 411
Query: 392 VQSSGLADVMSKALEFLKDVPYLAIVPAVSLLCSIITEFITSNDATATLLVPLLYHIAIN 451
SGL+D + + L L++V + I+ +++ L +TE ITSN ATAT++ P++ +A+
Sbjct: 412 FTDSGLSDWIGEQLTGLENVAPIIIILSIAALVIFLTE-ITSNTATATMMFPIMAALAVA 470
Query: 452 MHVHPLLLMIPGGIATEFAFWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAGVSVLAL 511
+ VHP +LMI +A AF LP +TP N V F +G++ I DM K G L + G+ ++ L
Sbjct: 471 VGVHPFILMIAAALAASCAFMLPVATPPNAVVFGSGYLRIPDMAKAGFFLNIVGIILVTL 530
Query: 512 LMPTLGKLLLSL 523
+ L + L +
Sbjct: 531 TVYFLVPMALGI 542
>ref|YP_156777.1| Na+/dicarboxylate symporter [Idiomarina loihiensis L2TR]
gi|56180506|gb|AAV83228.1| Na+/dicarboxylate symporter
[Idiomarina loihiensis L2TR]
Length = 454
Score = 273 bits (699), Expect = 8e-72
Identities = 167/477 (35%), Positives = 260/477 (54%), Gaps = 31/477 (6%)
Query: 39 YIILGPLLSIIICLLVKL-DAPTTSTKMLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLF 97
++ LGP+ ++I+ L +L + P + L + WV WW+T +PLPVTS+ P L P F
Sbjct: 3 HLALGPITALIVLLTTQLANMPLPAALTLSTVVWVAWWWVTECIPLPVTSLLPFVLLPTF 62
Query: 98 GIASADTVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLG 157
G+ T A + D++I L +G+F+LA AVE VH+R+AL++ D N +++
Sbjct: 63 GVIDTQTAAGALGDNIILLFMGAFMLAKAVEASGVHKRMALSLIARIGAD--NGRKVVIS 120
Query: 158 LCATTFFVSMWLHNVATAVMMMPVATGILHRLPPAEEQPELMNKFSRAVILTVVYATPIG 217
A T F+SMW+ N A+ + ++PVA L A + F RA++L + YA +G
Sbjct: 121 FMAATAFLSMWISNTASVLALLPVALA----LADASDD----ENFQRALLLGLAYAGSLG 172
Query: 218 GISTLTGTGVNLIIIGMWKSLAPEAKPISFNSWFFFGFPVAVLLLLCFWCILCLIYVRKG 277
G++TL GT NLI +++++ SF W G P+ +L L I+ L R
Sbjct: 173 GVATLVGTPPNLIFASIYQNIT--GVEFSFTRWLGIGLPMVLLGL----PIIALWLTRNV 226
Query: 278 SSRALSDYLDRALLKRDLEALGPMSFAEKMVLCVFGLLIVLWMTRRISDDLPGW-GVLFH 336
L + L +L M+ EK VL VFG ++VLW+TR + GW G+L
Sbjct: 227 K-------LSKPL---ELPTTSEMTTYEKRVLMVFGTVVVLWITR--TAPFGGWTGLLGV 274
Query: 337 GLVGDGSISVLAAVLLFIIPNMKQNGEKLMDWNECKKLPWNLILLLGAGFALADGVQSSG 396
G + D ++++ V +F+IP EKL++W + +PW ++LL G LA GV SG
Sbjct: 275 GSLNDATVALAGVVSMFVIPTGNAKNEKLLNWPLARDIPWGILLLFAGGICLARGVMESG 334
Query: 397 LADVMSKALEFLKDVPYLAIVPAVSLLCSIITEFITSNDATATLLVPLLYHIAINMHVHP 456
L +++ ++L L +P ++ +SL S +TE +TSN ATATLL+P+L A + +
Sbjct: 335 LGELIGQSLTALDALPLWLLIFVLSLTVSFVTE-VTSNTATATLLMPILAATATALELPI 393
Query: 457 LLLMIPGGIATEFAFWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAGVSVLALLM 513
LLMIP IA AF +P +TP N + FA+ + IKDM K G+ L + V L++
Sbjct: 394 ELLMIPAVIACSCAFCMPVATPPNSIVFASNRVTIKDMAKEGLVLNILLAIVTTLVV 450
>ref|ZP_00378199.1| COG0471: Di- and tricarboxylate transporters [Brevibacterium linens
BL2]
Length = 539
Score = 273 bits (699), Expect = 8e-72
Identities = 167/509 (32%), Positives = 270/509 (52%), Gaps = 51/509 (10%)
Query: 41 ILGPLLSIIICLLVKLDAPTTSTKMLGVIAWVFT-WWITNAVPLPVTSMCPLFLFPLFGI 99
+LG +LS ++ ++ D K+ +A + WW+T A+P+P TS+ PL FP+FG
Sbjct: 40 LLGLILSTLVYAIMP-DGVEHGAKLTAAVAVLMAVWWMTEALPIPATSLLPLVAFPVFGT 98
Query: 100 -ASADTVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGL 158
DTV SY + +I L LG F++ALA++R+N+HRR+AL +TL G+ P ++ G
Sbjct: 99 EVEMDTVGASYGNPIIFLFLGGFLIALAMQRWNLHRRIAL-ITLSIMGN--KPGPMIAGF 155
Query: 159 CATTFFVSMWLHNVATAVMMMPVATGIL--------------------HRLPPAEEQPE- 197
T FVSMW+ N ATAVMM+P+ +L A E PE
Sbjct: 156 MIATGFVSMWVSNTATAVMMLPIGISVLMIVSKVVGGAITDAAETADSDTADSAAEAPET 215
Query: 198 ----------LMNKFSRAVILTVVYATPIGGISTLTGTGVNLIIIGMWKSLAPEAKPISF 247
+ + F A++L + Y+ IG + T+ GT NL ++G K + I F
Sbjct: 216 GEDDSGLGAVVKSNFGTALMLGIAYSASIGSLGTIIGTPPNLFLVGYLKDNHDIS--IGF 273
Query: 248 NSWFFFGFPVAVLLLLCFWCILCLIYVRKGSSRALSDYLDRALLKRDLEALGPMSFAEKM 307
W G P+A+++++ W +L + + R L++ +LE LGPMS EK+
Sbjct: 274 GQWMLVGVPLAIVMMIIAWFLLVKVLFKPEIDEIPGG---RELIRDELEKLGPMSTGEKL 330
Query: 308 VLCVFGLLIVLWMTRRISDDLPGWGVLFHGLVGDGSISVLAAVLLFIIPNMKQNGEKLMD 367
VL +F L V W++ + + P + D I+++ +LLF+IP G +L+D
Sbjct: 331 VLGMFILAAVSWISLPLIFEEPP--------ISDEGIAMVIGLLLFLIPGGANRGVRLLD 382
Query: 368 WNECKKLPWNLILLLGAGFALADGVQSSGLADVMSKALEFLKDVPYLAIVPAVSLLCSII 427
W+ +KLPW ++LL G G AL+ SSGL + + K+ L +P + +V + + +
Sbjct: 383 WDTAEKLPWGVLLLFGGGLALSAQFSSSGLTEWIGKSTSGLGVLPTVFVVAIFATIILFL 442
Query: 428 TEFITSNDATATLLVPLLYHIAINMHVHPLLLMIPGGIATEFAFWLPTSTPSNVVGFATG 487
TE +TSN ATA VP++ +A+ + + PLLL IP +A AF LP +TP N V + +G
Sbjct: 443 TE-LTSNTATAATFVPVVGGVAMGLDLDPLLLTIPVALAATCAFMLPVATPPNAVAYGSG 501
Query: 488 HIEIKDMLKVGVPLKVAGVSVLALLMPTL 516
++ + M+K G+ L V G+ ++ + + L
Sbjct: 502 YVTVAQMVKGGLWLNVIGIVLITITVYVL 530
>ref|YP_073956.1| sodium-dependent transporter [Symbiobacterium thermophilum IAM
14863] gi|51854954|dbj|BAD39112.1| sodium-dependent
transporter [Symbiobacterium thermophilum IAM 14863]
Length = 495
Score = 270 bits (689), Expect = 1e-70
Identities = 166/489 (33%), Positives = 265/489 (53%), Gaps = 29/489 (5%)
Query: 33 LNL-QNFYIILGPLLSIIICLLVK-LDAPTTSTKMLGVIAWVFTWWITNAVPLPVTSMCP 90
LNL Q ++LGP L ++I L + L AW+ WW+T A+P+P+TS+ P
Sbjct: 16 LNLAQRIGLVLGPTLCLLIYLFFRPAGLGPEGVAALAGTAWIAAWWVTEAIPIPITSLLP 75
Query: 91 LFLFPLFGIASADTVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLN 150
+ LFP+ G+ S V +Y D + L LG FI+A+A+ER+N+HRR+AL V +
Sbjct: 76 VILFPVLGLGSVGEVVSAYADPTLFLFLGGFIIAVAIERWNLHRRIAL---AVISAVGTS 132
Query: 151 PSMLLLGLCATTFFVSMWLHNVATAVMMMPVATGILH-----RLPPAEEQPELMNKFSRA 205
PS L+LG T +S ++ N ATA+MM+P+ ++ R+ E+ FS
Sbjct: 133 PSRLVLGFMLATGVLSAFISNSATAMMMLPIGIAVVKQITELRVRNGEKVYPDQEPFSAL 192
Query: 206 VILTVVYATPIGGISTLTGTGVNLIIIGMWKSLAPEAKPISFNSWFFFGFPVAVLLLLCF 265
++L + YA IGG+ ++ G+ N I++G+ S ISF + G P+ ++ L
Sbjct: 193 ILLAIAYAASIGGVGSIIGSPPNAIMVGVAASTL--GVEISFFDYMLVGVPMFLVFTLVA 250
Query: 266 WCILCLIYVRKGSSRALSDYLDRALLKRDLEALGPMSFAEKMVLCVFGLLIVLWMTR--R 323
W IL + + A S R ++R + LGPMS E+ VL VF +LW+T
Sbjct: 251 WRILLWVMPPGFAEVAGS----REWVRRQMAELGPMSGEERKVLGVFCATALLWITYPFL 306
Query: 324 ISDDLPGWGVLFHGLVGDGSISVLAAVLLFIIPNMKQNGEKLMDWNECKKLPWNLILLLG 383
+ +PG + D +++ A LLF+IP G L+D + K+LPW ++LL G
Sbjct: 307 LKGVVPG--------ISDAMVAIFGASLLFLIPG--GEGGFLLDESALKELPWGVLLLFG 356
Query: 384 AGFALADGVQSSGLADVMSKALEFLKDVPYLAIVPAVSLLCSIITEFITSNDATATLLVP 443
AGF +A+ QS+G+A ++ L+ L PY IV AV+ + + +TE ITSN A A + +P
Sbjct: 357 AGFCIAEVFQSTGVAVWLAGFLQVLAGAPYFLIVIAVAAMVTFLTE-ITSNTAIAAIFMP 415
Query: 444 LLYHIAINMHVHPLLLMIPGGIATEFAFWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKV 503
++ +A + P LM+ +A AF LP TP N + F++G + I+ M++VG L +
Sbjct: 416 IMASLATALGAAPTGLMVVAALAGSLAFMLPVGTPPNAIVFSSGRVTIRQMIRVGFFLNL 475
Query: 504 AGVSVLALL 512
A ++ + L+
Sbjct: 476 AAIATITLI 484
>gb|EAM70903.1| Sodium/sulphate symporter [Desulfuromonas acetoxidans DSM 684]
Length = 573
Score = 268 bits (686), Expect = 2e-70
Identities = 172/480 (35%), Positives = 268/480 (55%), Gaps = 33/480 (6%)
Query: 42 LGPLLSIIICLL-VKLDAPTTSTKMLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFGIA 100
LGPLL I + L+ ++ KM V + TWW+ ++P+P TS+ PL LFPL GI
Sbjct: 85 LGPLLFITMLLIPTPAGMEPSAQKMAAVALLMATWWMCESIPIPATSLLPLMLFPLLGIM 144
Query: 101 SADTVAHSYMDDVITLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGLCA 160
+ A Y +I L +G FI+AL+++R+++HRR+A+ + + +PS L+ G A
Sbjct: 145 HTKSAAAPYASHLIFLFMGGFIIALSMQRWDLHRRIAMTIVKLV---GFSPSRLIFGFMA 201
Query: 161 TTFFVSMWLHNVATAVMMMPVATGIL-HRLPPAEEQ---------PELMNKFSRAVILTV 210
T +S ++ N ATAVMMMP+ I+ H + +++ PE + F ++L +
Sbjct: 202 ATATLSAFVSNTATAVMMMPIGLAIISHVIEEGKKEGLDKEIDFSPEHFS-FGLNLMLGI 260
Query: 211 VYATPIGGISTLTGTGVNLIIIG-MWKSLAPEAKPISFNSWFFFGFPVAVLLLLCFWCIL 269
YA IGG++TL GT N ++ G + K+ E IS+ W G P+ V++L W L
Sbjct: 261 AYAASIGGMATLIGTPPNTVLAGYLNKTYGFE---ISYVDWLKVGVPLVVVMLPLCWLWL 317
Query: 270 CLIYVRKGSSRALSDYLDRALLKRDLEALGPMSFAEKMVLCVFGLLIVLWMTRRISDDLP 329
I R R ++ +L+ +G MS EK VFGL + W+ R+
Sbjct: 318 TRIANPMKLKRVPG---GREMILDELKQMGRMSSGEKWTAVVFGLTALGWIFRK------ 368
Query: 330 GWGVLFHG--LVGDGSISVLAAVLLFIIP-NMKQNGEKLMDWNECKKLPWNLILLLGAGF 386
G LF LV D +I++ A++LF+IP NMK+N +MDW+ K+PW ++LL G G
Sbjct: 369 QLGFLFPDPTLVTDAAIAMTGALVLFMIPVNMKKN-IFVMDWHWASKMPWGVLLLFGGGL 427
Query: 387 ALADGVQSSGLADVMSKALEFLKDVPYLAIVPAVSLLCSIITEFITSNDATATLLVPLLY 446
A+A G + +GLA + + L + P L +V AV+ L +TE +TSN ATA +++P+L
Sbjct: 428 AMAAGFKQTGLATWIGSQVSLLNNAPILVLVVAVTALIIFLTE-LTSNTATAAMVMPILS 486
Query: 447 HIAINMHVHPLLLMIPGGIATEFAFWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAGV 506
+AI ++ +PLLL+IP IA AF LP +TP N + F +G++ I M++ G L + GV
Sbjct: 487 AVAIGINQNPLLLVIPAAIAASCAFMLPVATPPNAIVFGSGYVTIPQMVRSGFGLNIVGV 546
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.327 0.142 0.440
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 893,699,281
Number of Sequences: 2540612
Number of extensions: 37867007
Number of successful extensions: 118000
Number of sequences better than 10.0: 647
Number of HSP's better than 10.0 without gapping: 344
Number of HSP's successfully gapped in prelim test: 303
Number of HSP's that attempted gapping in prelim test: 115811
Number of HSP's gapped (non-prelim): 1086
length of query: 530
length of database: 863,360,394
effective HSP length: 133
effective length of query: 397
effective length of database: 525,458,998
effective search space: 208607222206
effective search space used: 208607222206
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 78 (34.7 bits)
Medicago: description of AC148482.2


