

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148470.10 - phase: 0 /pseudo
(598 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
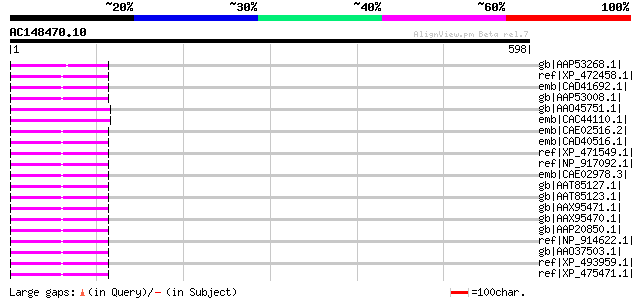
Score E
Sequences producing significant alignments: (bits) Value
gb|AAP53268.1| putative 22 kDa kafirin cluster; Ty3-Gypsy type [... 94 2e-17
ref|XP_472458.1| OSJNBb0108J11.19 [Oryza sativa (japonica cultiv... 92 5e-17
emb|CAD41692.1| OSJNBb0015D13.7 [Oryza sativa (japonica cultivar... 92 5e-17
gb|AAP53008.1| putative polyprotein [Oryza sativa (japonica cult... 92 6e-17
gb|AAO45751.1| gag-protease polyprotein [Cucumis melo] 91 8e-17
emb|CAC44110.1| gag polyprotein [Cicer arietinum] 91 1e-16
emb|CAE02516.2| OSJNBb0003A12.3 [Oryza sativa (japonica cultivar... 89 3e-16
emb|CAD40516.1| OSJNBa0023J03.1 [Oryza sativa (japonica cultivar... 89 3e-16
ref|XP_471549.1| OSJNBa0019J05.10 [Oryza sativa (japonica cultiv... 89 4e-16
ref|NP_917092.1| putative polyprotein [Oryza sativa (japonica cu... 89 4e-16
emb|CAE02978.3| OSJNBa0086B14.15 [Oryza sativa (japonica cultiva... 89 4e-16
gb|AAT85127.1| putative polyprotein [Oryza sativa (japonica cult... 89 4e-16
gb|AAT85123.1| putative polyprotein [Oryza sativa (japonica cult... 89 4e-16
gb|AAX95471.1| Retrotransposon gag protein, putative [Oryza sati... 89 4e-16
gb|AAX95470.1| Retrotransposon gag protein, putative [Oryza sati... 89 4e-16
gb|AAP20850.1| retrotransposon protein, putative, Ty3-gypsy sub-... 89 5e-16
ref|NP_914622.1| similar to polyprotein [Oryza sativa (japonica ... 89 5e-16
gb|AAO37503.1| retrotransposon protein, putative, Ty3-gypsy sub-... 88 7e-16
ref|XP_493959.1| Similar to Sorghum bicolor 22 kDa kafirin clust... 88 7e-16
ref|XP_475471.1| putative polyprotein [Oryza sativa (japonica cu... 88 7e-16
>gb|AAP53268.1| putative 22 kDa kafirin cluster; Ty3-Gypsy type [Oryza sativa
(japonica cultivar-group)] gi|37533358|ref|NP_920981.1|
putative 22 kDa kafirin cluster; Ty3-Gypsy type [Oryza
sativa (japonica cultivar-group)]
gi|21327374|gb|AAM48279.1| Putative 22 kDa kafirin
cluster; Ty3-Gypsy type [Oryza sativa (japonica
cultivar-group)] gi|18767374|gb|AAL79340.1| Putative 22
kDa kafirin cluster; Ty3-Gypsy type [Oryza sativa]
Length = 1230
Score = 93.6 bits (231), Expect = 2e-17
Identities = 45/114 (39%), Positives = 67/114 (58%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
F + PPTF G +P A++W+ +E F M C++ +K+ + T+ML A +WW +
Sbjct: 74 FQKLKPPTFSGTANPLEAEEWIVAMEKSFEAMGCTDKEKIIYATYMLQSSAFEWWDAHKK 133
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
+ +TW +F+ F +Y PE V+ KE EFLELKQG+ SV EY EFS+
Sbjct: 134 SYSER-IFITWELFKEAFYKKYFPESVKRMKEKEFLELKQGNKSVAEYEIEFSR 186
>ref|XP_472458.1| OSJNBb0108J11.19 [Oryza sativa (japonica cultivar-group)]
gi|32488367|emb|CAE02926.1| OSJNBb0108J11.19 [Oryza
sativa (japonica cultivar-group)]
gi|32489278|emb|CAE04619.1| OSJNBa0028I23.1 [Oryza
sativa (japonica cultivar-group)]
Length = 1516
Score = 92.0 bits (227), Expect = 5e-17
Identities = 45/114 (39%), Positives = 65/114 (56%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL+ +E ++QC++ +KV F TH L A+ WW + +
Sbjct: 84 FLRIQPPTFSSTTNPMEANDWLRAIEKKLNLLQCNDQEKVAFATHQLQGPASAWWDNYV- 142
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
D +TWA F + F +PE + Q++ EF L+QG MSVTEY EF++
Sbjct: 143 ATRLDATEVTWAEFCQSFRKAQIPEGIMAQQKREFRALQQGTMSVTEYLHEFNR 196
>emb|CAD41692.1| OSJNBb0015D13.7 [Oryza sativa (japonica cultivar-group)]
Length = 1516
Score = 92.0 bits (227), Expect = 5e-17
Identities = 45/114 (39%), Positives = 65/114 (56%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL+ +E ++QC++ +KV F TH L A+ WW + +
Sbjct: 84 FLRIQPPTFSSTTNPMEANDWLRAIEKKLNLLQCNDQEKVAFATHQLQGPASAWWDNYV- 142
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
D +TWA F + F +PE + Q++ EF L+QG MSVTEY EF++
Sbjct: 143 ATRLDATEVTWAEFCQSFRKAQIPEGIMAQQKREFRALQQGTMSVTEYLHEFNR 196
>gb|AAP53008.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|37532838|ref|NP_920721.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
gi|16905208|gb|AAL31078.1| putative polyprotein [Oryza
sativa]
Length = 1524
Score = 91.7 bits (226), Expect = 6e-17
Identities = 46/114 (40%), Positives = 63/114 (54%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL +E ++QC+E +KV F TH L A+ WW + +
Sbjct: 96 FLRVKPPTFSSTTNPIEAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASIWWDNYM- 154
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
+ A +TW FR F VPE + GQK+ EF L+QG +V EY EF++
Sbjct: 155 VTRPTGAEVTWTEFRHSFNKAQVPEGIVGQKKREFRSLQQGTKTVIEYLHEFNR 208
>gb|AAO45751.1| gag-protease polyprotein [Cucumis melo]
Length = 429
Score = 91.3 bits (225), Expect = 8e-17
Identities = 47/117 (40%), Positives = 65/117 (55%), Gaps = 1/117 (0%)
Query: 1 FLRNHPPTFKGRY-DPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLL 59
F + +P TF G DP AQ WL +E+IFR M+C E QKV+ ML + WW +
Sbjct: 63 FRKYNPTTFDGSLEDPTRAQMWLSSLETIFRYMKCPEDQKVQCAVFMLTDRGTAWWETTE 122
Query: 60 PILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSKSS 116
+L D + +TW F+ F ++ +R K EFL L+QGDM+V +Y AEF S
Sbjct: 123 RMLGGDVSQITWQQFKESFYAKFFSASLRDAKRQEFLNLEQGDMTVEQYDAEFDMLS 179
>emb|CAC44110.1| gag polyprotein [Cicer arietinum]
Length = 277
Score = 90.5 bits (223), Expect = 1e-16
Identities = 44/116 (37%), Positives = 67/116 (56%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
F R +PP FKG + A +W++EVE IF ++ C KV + T+ML +A WW S
Sbjct: 59 FRRYNPPKFKGDEGSEKADQWIQEVEKIFDMINCQAGVKVSYATYMLLGDAEYWWRSARL 118
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSKSS 116
++ + W F+R+FL++Y P R + +FL+L QG ++V EYAA+F S
Sbjct: 119 LMGAAHEEVNWESFKRKFLDKYFPMSTRTKLGDDFLKLHQGSLTVGEYAAKFESLS 174
>emb|CAE02516.2| OSJNBb0003A12.3 [Oryza sativa (japonica cultivar-group)]
gi|38347085|emb|CAE05109.2| OSJNBa0001M07.5 [Oryza
sativa (japonica cultivar-group)]
gi|50930335|ref|XP_474695.1| OSJNBa0001M07.5 [Oryza
sativa (japonica cultivar-group)]
Length = 1471
Score = 89.4 bits (220), Expect = 3e-16
Identities = 45/114 (39%), Positives = 62/114 (53%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL +E ++QC+E +KV F TH L A+ WW + +
Sbjct: 70 FLRVKPPTFSSTTNPIEAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASVWWDNYM- 128
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
+ A +TW FR F VPE + QK+ EF L+QG +V EY EF++
Sbjct: 129 VTRPTGAEVTWTEFRHSFNKAQVPEGIVAQKKREFRSLQQGTRTVIEYLHEFNR 182
>emb|CAD40516.1| OSJNBa0023J03.1 [Oryza sativa (japonica cultivar-group)]
gi|50922727|ref|XP_471724.1| OSJNBa0023J03.1 [Oryza
sativa (japonica cultivar-group)]
Length = 1827
Score = 89.4 bits (220), Expect = 3e-16
Identities = 46/114 (40%), Positives = 63/114 (54%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL +E ++QC++ +KV F TH L A+ WW + +
Sbjct: 401 FLRVRPPTFSSTTNPMEANDWLHAIEKKLNLLQCNDQEKVAFATHQLQGPASAWWDNHM- 459
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
+TWA F R F VP+ V QK+ EF L QG+M+VTEY EF++
Sbjct: 460 ATRPPGTEVTWAEFCRSFRKAQVPDGVVAQKKREFRALHQGNMTVTEYLHEFNR 513
>ref|XP_471549.1| OSJNBa0019J05.10 [Oryza sativa (japonica cultivar-group)]
gi|38346425|emb|CAD40212.2| OSJNBa0019J05.10 [Oryza
sativa (japonica cultivar-group)]
Length = 1525
Score = 89.0 bits (219), Expect = 4e-16
Identities = 45/114 (39%), Positives = 62/114 (53%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL +E ++QC+E +KV F TH L A+ WW + +
Sbjct: 96 FLRVKPPTFSSTTNPIEAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASIWWDNYM- 154
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
+ A +TW FR F VPE + QK+ EF L+QG +V EY EF++
Sbjct: 155 VTRPAGAEVTWTEFRHSFNKAQVPEGIVAQKKREFRSLQQGTRTVIEYLHEFNR 208
>ref|NP_917092.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1524
Score = 89.0 bits (219), Expect = 4e-16
Identities = 45/114 (39%), Positives = 62/114 (53%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL +E ++QC+E +KV F TH L A+ WW + +
Sbjct: 96 FLRVKPPTFSSTTNPIEAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASIWWDNYM- 154
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
+ A +TW FR F VPE + QK+ EF L+QG +V EY EF++
Sbjct: 155 VTRPAGAEVTWTEFRHSFNKAQVPEGIVAQKKREFRSLQQGTRTVIEYLHEFNR 208
>emb|CAE02978.3| OSJNBa0086B14.15 [Oryza sativa (japonica cultivar-group)]
gi|50924632|ref|XP_472673.1| OSJNBa0086B14.15 [Oryza
sativa (japonica cultivar-group)]
Length = 1516
Score = 89.0 bits (219), Expect = 4e-16
Identities = 42/114 (36%), Positives = 64/114 (55%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL+ +E ++QC++ +KV F TH L A+ WW + +
Sbjct: 84 FLRIQPPTFSSTTNPMEANDWLRAIEKKLNLLQCNDQEKVAFATHQLQGPASAWWDNYV- 142
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
+ D +TW F + F +PE + Q++ EF L+QG +VTEY EF++
Sbjct: 143 VTRPDATEVTWVEFCQNFRKALIPEGIMAQQKREFRALQQGTRTVTEYLHEFNR 196
>gb|AAT85127.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1524
Score = 89.0 bits (219), Expect = 4e-16
Identities = 45/114 (39%), Positives = 62/114 (53%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL +E ++QC+E +KV F TH L A+ WW + +
Sbjct: 96 FLRVKPPTFSSTTNPIEAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASIWWDNYM- 154
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
+ A +TW FR F VPE + QK+ EF L+QG +V EY EF++
Sbjct: 155 VTRPAGAEVTWTEFRHSFNKAQVPEGIVAQKKREFRSLQQGTRTVIEYLHEFNR 208
>gb|AAT85123.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1495
Score = 89.0 bits (219), Expect = 4e-16
Identities = 42/114 (36%), Positives = 64/114 (55%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL+ +E ++QC++ +KV F TH L A+ WW + +
Sbjct: 84 FLRIQPPTFSSTTNPMEANDWLRAIEKKLNLLQCNDQEKVAFATHQLQGPASAWWDNYV- 142
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
+ D +TW F + F +PE + Q++ EF L+QG +VTEY EF++
Sbjct: 143 VTRPDATKVTWVEFCQNFRKAQIPEGIMAQQKREFRALQQGTRTVTEYLHEFNR 196
>gb|AAX95471.1| Retrotransposon gag protein, putative [Oryza sativa (japonica
cultivar-group)]
Length = 1441
Score = 89.0 bits (219), Expect = 4e-16
Identities = 45/114 (39%), Positives = 62/114 (53%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL +E ++QC+E +KV F TH L A+ WW + +
Sbjct: 96 FLRVKPPTFSSTTNPIEAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASIWWDNYM- 154
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
+ A +TW FR F VPE + QK+ EF L+QG +V EY EF++
Sbjct: 155 VTRPAGAEVTWTEFRHSFNKAQVPEGIVAQKKREFRSLQQGTRTVIEYLHEFNR 208
>gb|AAX95470.1| Retrotransposon gag protein, putative [Oryza sativa (japonica
cultivar-group)]
Length = 1441
Score = 89.0 bits (219), Expect = 4e-16
Identities = 45/114 (39%), Positives = 62/114 (53%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL +E ++QC+E +KV F TH L A+ WW + +
Sbjct: 96 FLRVKPPTFSSTTNPIEAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASIWWDNYM- 154
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
+ A +TW FR F VPE + QK+ EF L+QG +V EY EF++
Sbjct: 155 VTRPAGAEVTWTEFRHSFNKAQVPEGIVAQKKREFRSLQQGTRTVIEYLHEFNR 208
>gb|AAP20850.1| retrotransposon protein, putative, Ty3-gypsy sub-class [Oryza
sativa (japonica cultivar-group)]
gi|50916687|ref|XP_468740.1| putative retrotransposon
gag protein [Oryza sativa (japonica cultivar-group)]
Length = 486
Score = 88.6 bits (218), Expect = 5e-16
Identities = 42/114 (36%), Positives = 64/114 (55%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL+ +E ++QC++ +KV F TH L A+ WW + +
Sbjct: 84 FLRIQPPTFSSTTNPMEANDWLRAIEKKLNLLQCNDQEKVAFATHQLQGPASVWWDNYV- 142
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
+ D +TW F + F +PE + Q++ EF L+QG +VTEY EF++
Sbjct: 143 VTRPDATKVTWVEFCQNFRKAQIPEGIMAQQKREFRALQQGTRTVTEYLHEFNR 196
>ref|NP_914622.1| similar to polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1524
Score = 88.6 bits (218), Expect = 5e-16
Identities = 45/114 (39%), Positives = 62/114 (53%), Gaps = 1/114 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL +E ++QC+E +KV F TH L A+ WW + +
Sbjct: 96 FLRVKPPTFSSTTNPIEAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASVWWDNYM- 154
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFSK 114
+ A +TW FR F VPE + QK+ EF L+QG +V EY EF++
Sbjct: 155 VTRPAGAEVTWTEFRHSFNKAQVPEGIVAQKKREFHSLQQGTRTVIEYLHEFNR 208
>gb|AAO37503.1| retrotransposon protein, putative, Ty3-gypsy sub-class [Oryza
sativa (japonica cultivar-group)]
gi|50916491|ref|XP_468642.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1079
Score = 88.2 bits (217), Expect = 7e-16
Identities = 45/113 (39%), Positives = 61/113 (53%), Gaps = 1/113 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL +E ++QC+E +KV F TH L A+ WW + +
Sbjct: 96 FLRVKPPTFSSTTNPIEAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASIWWDNYM- 154
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFS 113
+ A +TW FR F VPE + QK+ EF L+QG +V EY EF+
Sbjct: 155 VTRPAGAEVTWTEFRHSFNKAQVPEGIVAQKKREFRSLQQGTRTVIEYLHEFN 207
>ref|XP_493959.1| Similar to Sorghum bicolor 22 kDa kafirin cluster; polyprotein.
(AF061282) [Oryza sativa (japonica cultivar-group)]
Length = 1524
Score = 88.2 bits (217), Expect = 7e-16
Identities = 45/113 (39%), Positives = 61/113 (53%), Gaps = 1/113 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL +E ++QC+E +KV F TH L A+ WW + +
Sbjct: 96 FLRVKPPTFSSTTNPIEAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASVWWDNYM- 154
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFS 113
+ A +TW FR F VPE + QK+ EF L+QG +V EY EF+
Sbjct: 155 VTRPAGAEVTWTEFRHSFNKAQVPEGIVAQKKREFRSLQQGTRTVIEYLHEFN 207
>ref|XP_475471.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
gi|50080316|gb|AAT69650.1| putative polyprotein [Oryza
sativa (japonica cultivar-group)]
Length = 1458
Score = 88.2 bits (217), Expect = 7e-16
Identities = 45/113 (39%), Positives = 61/113 (53%), Gaps = 1/113 (0%)
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
FLR PPTF +P A WL +E ++QC+E +KV F TH L A+ WW + +
Sbjct: 96 FLRVKPPTFSSTTNPIEAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASIWWDNYM- 154
Query: 61 ILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEIEFLELKQGDMSVTEYAAEFS 113
+ A +TW FR F VPE + QK+ EF L+QG +V EY EF+
Sbjct: 155 VTRPAGAEVTWTEFRHSFNKAQVPEGIVAQKKREFRSLQQGTRTVIEYLHEFN 207
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.370 0.166 0.609
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 735,975,829
Number of Sequences: 2540612
Number of extensions: 23539989
Number of successful extensions: 127891
Number of sequences better than 10.0: 581
Number of HSP's better than 10.0 without gapping: 534
Number of HSP's successfully gapped in prelim test: 47
Number of HSP's that attempted gapping in prelim test: 126262
Number of HSP's gapped (non-prelim): 1093
length of query: 598
length of database: 863,360,394
effective HSP length: 134
effective length of query: 464
effective length of database: 522,918,386
effective search space: 242634131104
effective search space used: 242634131104
T: 11
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.8 bits)
S2: 78 (34.7 bits)
Medicago: description of AC148470.10


