

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148360.3 - phase: 0
(301 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
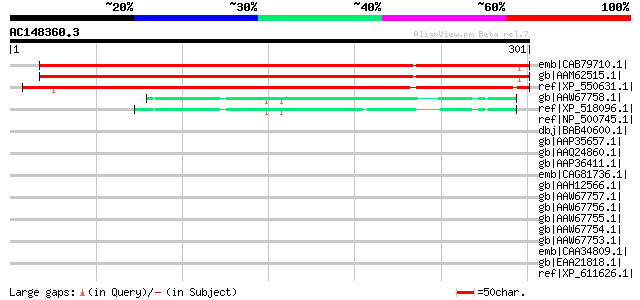
Score E
Sequences producing significant alignments: (bits) Value
emb|CAB79710.1| putative protein [Arabidopsis thaliana] gi|51235... 384 e-105
gb|AAM62515.1| unknown [Arabidopsis thaliana] 384 e-105
ref|XP_550631.1| unknown protein [Oryza sativa (japonica cultiva... 362 6e-99
gb|AAW67758.1| nucleophosmin [Homo sapiens] 47 5e-04
ref|XP_518096.1| PREDICTED: similar to nucleophosmin 1; nucleola... 47 7e-04
ref|NP_500745.1| putative protein, with a coiled coil domain (24... 45 0.002
dbj|BAB40600.1| nucleophosmin/B23.2 [Homo sapiens] 45 0.003
gb|AAP35657.1| nucleophosmin (nucleolar phosphoprotein B23, numa... 45 0.003
gb|AAQ24860.1| nucleophosmin [Homo sapiens] 45 0.003
gb|AAP36411.1| Homo sapiens nucleophosmin (nucleolar phosphoprot... 45 0.003
emb|CAG81736.1| unnamed protein product [Yarrowia lipolytica CLI... 45 0.003
gb|AAH12566.1| Nucleophosmin 1 [Homo sapiens] 45 0.003
gb|AAW67757.1| nucleophosmin [Homo sapiens] 45 0.003
gb|AAW67756.1| nucleophosmin [Homo sapiens] 45 0.003
gb|AAW67755.1| nucleophosmin [Homo sapiens] gi|58220457|gb|AAW67... 45 0.003
gb|AAW67754.1| nucleophosmin [Homo sapiens] 45 0.003
gb|AAW67753.1| nucleophosmin [Homo sapiens] 45 0.003
emb|CAA34809.1| B23 nucleophosmin (280 AA) [Homo sapiens] 45 0.003
gb|EAA21818.1| SNF2 family N-terminal domain, putative [Plasmodi... 45 0.003
ref|XP_611626.1| PREDICTED: hypothetical protein XP_611626, part... 45 0.003
>emb|CAB79710.1| putative protein [Arabidopsis thaliana] gi|5123546|emb|CAB45312.1|
putative protein [Arabidopsis thaliana]
gi|15233608|ref|NP_194681.1| expressed protein
[Arabidopsis thaliana] gi|7487103|pir||T09915
hypothetical protein T16L4.30 - Arabidopsis thaliana
Length = 306
Score = 384 bits (987), Expect = e-105
Identities = 185/286 (64%), Positives = 228/286 (79%), Gaps = 3/286 (1%)
Query: 18 IPLSHCAKKPVGIARKEDVPYIKCQVCEILAKQLYQQVQSKKAEISPKKISEYQIIEIAE 77
+P+S AKKP RKEDVPYIKCQVCE L+ +L+Q V+ K+ +ISPKKISEY+IIEIAE
Sbjct: 22 LPVSDAAKKPSSTPRKEDVPYIKCQVCEKLSSRLHQLVKEKQQQISPKKISEYEIIEIAE 81
Query: 78 NVCNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAE 137
NVCNLKK EADW+L+IDIVEK D L L E EG CNS+CKT+E ACQ+V+GYSDTDVAE
Sbjct: 82 NVCNLKKEEADWMLKIDIVEKGDNLVLVEQQEEGMCNSKCKTIENACQKVIGYSDTDVAE 141
Query: 138 YLYSSKPDIDSLTNYLCKDLSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEG 197
Y+Y SKPD+ SL N+LCKDL+ AC+ KPPPVPKDR PGEPFVAK SK+AEM+K+L+SM+G
Sbjct: 142 YIYKSKPDLVSLVNHLCKDLTDACSKKPPPVPKDRVPGEPFVAKPSKDAEMDKILRSMQG 201
Query: 198 MPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGDWKQ 257
MPGAPGMK+YSR+D+ K N G E++D D+DEDEEED+ FP LGK+LK KE++ + K+
Sbjct: 202 MPGAPGMKVYSREDIEKGNIGNEDDDGDDDEDEEEDD-KFPKNLGKVLKEKESKTEELKK 260
Query: 258 KIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSK--KNSKSEL 301
I K LK+HA KVSN ++RWWKG ++SSK K+ KSEL
Sbjct: 261 TITKEFKKKGEALKRHAQKVSNRVRRWWKGLGSSSSKKPKSGKSEL 306
>gb|AAM62515.1| unknown [Arabidopsis thaliana]
Length = 303
Score = 384 bits (987), Expect = e-105
Identities = 185/286 (64%), Positives = 228/286 (79%), Gaps = 3/286 (1%)
Query: 18 IPLSHCAKKPVGIARKEDVPYIKCQVCEILAKQLYQQVQSKKAEISPKKISEYQIIEIAE 77
+P+S AKKP RKEDVPYIKCQVCE L+ +L+Q V+ K+ +ISPKKISEY+IIEIAE
Sbjct: 19 LPVSDAAKKPSSTPRKEDVPYIKCQVCEKLSSRLHQLVKEKQQQISPKKISEYEIIEIAE 78
Query: 78 NVCNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAE 137
NVCNLKK EADW+L+IDIVEK D L L E EG CNS+CKT+E ACQ+V+GYSDTDVAE
Sbjct: 79 NVCNLKKEEADWMLKIDIVEKGDNLVLVEQQEEGMCNSKCKTIENACQKVIGYSDTDVAE 138
Query: 138 YLYSSKPDIDSLTNYLCKDLSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEG 197
Y+Y SKPD+ SL N+LCKDL+ AC+ KPPPVPKDR PGEPFVAK SK+AEM+K+L+SM+G
Sbjct: 139 YIYKSKPDLVSLVNHLCKDLTDACSKKPPPVPKDRVPGEPFVAKPSKDAEMDKILRSMQG 198
Query: 198 MPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKGDWKQ 257
MPGAPGMK+YSR+D+ K N G E++D D+DEDEEED+ FP LGK+LK KE++ + K+
Sbjct: 199 MPGAPGMKVYSREDIEKGNIGNEDDDGDDDEDEEEDD-KFPKNLGKVLKEKESKTEELKK 257
Query: 258 KIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSK--KNSKSEL 301
I K LK+HA KVSN ++RWWKG ++SSK K+ KSEL
Sbjct: 258 TITKEFKKKGEALKRHAQKVSNRVRRWWKGLGSSSSKKPKSGKSEL 303
>ref|XP_550631.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|55297262|dbj|BAD69047.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
gi|55296609|dbj|BAD69311.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 306
Score = 362 bits (930), Expect = 6e-99
Identities = 178/303 (58%), Positives = 225/303 (73%), Gaps = 14/303 (4%)
Query: 8 LVVSVVIASWIPLSHCA--------KKPVGIARKEDVPYIKCQVCEILAKQLYQQVQSKK 59
+ ++V +A+ + L H A KK ARKED+PYI+CQVCE +A+++ QV K+
Sbjct: 9 VAMAVAVAAVVLLLHPAASAAAAGPKKVATAARKEDIPYIRCQVCERIAREISAQVAKKQ 68
Query: 60 AEI-SPKKISEYQIIEIAENVCNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECK 118
+ + KK+ E +IIEIAENVCNLKK EADW+L+IDIVEK D+LEL E D EG CN+ECK
Sbjct: 69 QALPATKKVPEIEIIEIAENVCNLKKQEADWMLKIDIVEKGDKLELVEQDEEGHCNAECK 128
Query: 119 TVERACQEVMGYSDTDVAEYLYSSKPDIDSLTNYLCKDLSKACNTKPPPVPKDRTPGEPF 178
T+ERACQEVMGY+DTDVAE++Y KP D L +LCKDLS+AC PPPVPKDR PGEPF
Sbjct: 129 TIERACQEVMGYADTDVAEFVYKKKPSADQLVKFLCKDLSEACVVDPPPVPKDRVPGEPF 188
Query: 179 VAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFP 238
AK SK+AEM+++LKSMEG+PGAP MKMYSRDDLMK NFG + +DDD+DEDE++ DFP
Sbjct: 189 AAKPSKDAEMDRILKSMEGIPGAPSMKMYSRDDLMKNNFGVDGDDDDDDEDEDD---DFP 245
Query: 239 SKLGKILKSKENEKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSKKNSK 298
LGK+ K K + K D KQ++ K I DT LK H KVS +++WW+GKK S K+SK
Sbjct: 246 KNLGKVFKDKGSPKKDLKQQVVKQIKDTGKKLKGHVNKVSKVVKKWWQGKKKPS--KSSK 303
Query: 299 SEL 301
+EL
Sbjct: 304 TEL 306
>gb|AAW67758.1| nucleophosmin [Homo sapiens]
Length = 293
Score = 47.4 bits (111), Expect = 5e-04
Identities = 56/221 (25%), Positives = 88/221 (39%), Gaps = 25/221 (11%)
Query: 80 CNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYL 139
C LK + D+ ++D E +L L E VE E M Y + + L
Sbjct: 21 CELK-ADKDYHFKVDNDENEHQLSLRTVSLGAGAKDELHIVEA---EAMNYEGSPIKVTL 76
Query: 140 YSSKPDID---SLTNYLCKD---LSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLK 193
+ K + SL + L C + P + A+S E E + L
Sbjct: 77 ATLKMSVQPTVSLGGFEITPPVVLRLKCGSGPVHISGQHLVAVEEDAESEDEEEEDVKLL 136
Query: 194 SMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGKILKSKENEKG 253
S+ G APG +K +++DD+ED+DE++D+ DF ++E+
Sbjct: 137 SISGKRSAPGGGSKVPQKKVKLAADEDDDDDEEDDDEDDDDDDF-----------DDEEA 185
Query: 254 DWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSK 294
+ K ++K I DT K+A K SN + K T SK
Sbjct: 186 EEKAPVKKSIRDTPA---KNAQK-SNQNGKDSKPSSTPRSK 222
>ref|XP_518096.1| PREDICTED: similar to nucleophosmin 1; nucleolar phosphoprotein
B23; numatrin; nucleophosmin/nucleoplasmin family,
member 1 [Pan troglodytes]
Length = 376
Score = 47.0 bits (110), Expect = 7e-04
Identities = 58/228 (25%), Positives = 90/228 (39%), Gaps = 29/228 (12%)
Query: 73 IEIAENVCNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSD 132
++ N C LK + D+ ++D E +L L E VE E M Y
Sbjct: 101 VKYIPNRCELK-ADKDYHFKVDNDENEHQLSLRTVSLGAGAKDELHIVEA---EAMNYEG 156
Query: 133 TDVAEYLYSSKPDID---SLTNYLCKD---LSKACNTKPPPVPKDRTPGEPFVAKSSKEA 186
+ + L + K + SL + L C + P + A+S E
Sbjct: 157 SPIKVTLATLKMSVQPTVSLGGFEITPPVVLRLKCGSGPVHISGQHLVAVEEDAESEDEE 216
Query: 187 EMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSKLGKILK 246
E + L S+ G APG S+ K A+ +DDD+DED+++D+ D
Sbjct: 217 EEDVKLLSISGKRSAPGGG--SKVPQKKVKLAADEDDDDDDEDDDDDDFD---------- 264
Query: 247 SKENEKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSK 294
+E+ + K ++K I DT K+A K SN + K T SK
Sbjct: 265 ---DEEAEEKAPVKKSIRDTPA---KNAQK-SNQNGKDSKPSSTPRSK 305
>ref|NP_500745.1| putative protein, with a coiled coil domain (24.0 kD) (4F990)
[Caenorhabditis elegans] gi|7332136|gb|AAF60823.1|
Hypothetical protein Y59E9AL.6 [Caenorhabditis elegans]
Length = 214
Score = 45.4 bits (106), Expect = 0.002
Identities = 36/135 (26%), Positives = 55/135 (40%), Gaps = 3/135 (2%)
Query: 166 PPVPKDRTPGEPFVAKSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDD 225
PP D P P A + + E + K E G D + E +D D
Sbjct: 49 PPAAPDAAPAAPDAAAAPADGEKKDGDKKSEMKEGEEKKNEKKEGDEKDEKKDEEKKDGD 108
Query: 226 EDEDEEEDEADFPSKLGKILKSKENEKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWW 285
+ +DE++D+ K K K+ +K D K+K + D KK K + +
Sbjct: 109 DKKDEKKDD---DKKDEKKSDKKDEKKDDKKEKDEEKKDDKKEDEKKDDDKKDDKKEAKS 165
Query: 286 KGKKTTSSKKNSKSE 300
K KK+ SKK+ KS+
Sbjct: 166 KSKKSNKSKKSKKSK 180
>dbj|BAB40600.1| nucleophosmin/B23.2 [Homo sapiens]
Length = 259
Score = 45.1 bits (105), Expect = 0.003
Identities = 59/224 (26%), Positives = 89/224 (39%), Gaps = 30/224 (13%)
Query: 80 CNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYL 139
C LK + D+ ++D E +L L E VE E M Y + + L
Sbjct: 21 CELK-ADKDYHFKVDNDENEHQLSLRTVSLGAGAKDELHIVEA---EAMNYEGSPIKVTL 76
Query: 140 YSSKPDID---SLTNYLCKD---LSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLK 193
+ K + SL + L C + P + A+S E E + L
Sbjct: 77 ATLKMSVQPTVSLGGFEITPPVVLRLKCGSGPVHISGQHLVAVEEDAESEDEEEEDVKLL 136
Query: 194 SMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDE---DEEEDEADFPSKLGKILKSKEN 250
S+ G APG S+ K A+ +DDD+DE DE++D+ DF ++
Sbjct: 137 SISGKRSAPGGG--SKVPQKKVKLAADEDDDDDDEEDDDEDDDDDDF-----------DD 183
Query: 251 EKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSK 294
E+ + K ++K I DT K+A K SN + K T SK
Sbjct: 184 EEAEEKAPVKKSIRDTPA---KNAQK-SNQNGKDSKPSSTPRSK 223
>gb|AAP35657.1| nucleophosmin (nucleolar phosphoprotein B23, numatrin) [Homo
sapiens] gi|61361629|gb|AAX42078.1| nucleophosmin
[synthetic construct] gi|2745723|gb|AAB94739.1|
nucleophosmin phosphoprotein [Homo sapiens]
gi|47940120|gb|AAH71891.1| Unknown (protein for
MGC:88570) [Homo sapiens] gi|29792279|gb|AAH50628.1|
Nucleophosmin 1 [Homo sapiens]
gi|14250152|gb|AAH08495.1| Nucleophosmin 1 [Homo
sapiens] gi|12803185|gb|AAH02398.1| Nucleophosmin 1
[Homo sapiens] gi|16876872|gb|AAH16716.1| Nucleophosmin
1 [Homo sapiens] gi|15680058|gb|AAH14349.1|
Nucleophosmin 1 [Homo sapiens]
gi|18203876|gb|AAH21668.1| Nucleophosmin 1 [Homo
sapiens] gi|16877099|gb|AAH16824.1| Nucleophosmin 1
[Homo sapiens] gi|10835063|ref|NP_002511.1|
nucleophosmin 1 [Homo sapiens]
gi|114762|sp|P06748|NPM_HUMAN Nucleophosmin (NPM)
(Nucleolar phosphoprotein B23) (Numatrin) (Nucleolar
protein NO38) gi|107221|pir||A32915 nucleophosmin -
human gi|557546|gb|AAA58386.1| nucleolar phosphoprotein
B23 gi|189312|gb|AAA36385.1| nucleolar protein B23
gi|189272|gb|AAA36380.1| nucleophosmin
Length = 294
Score = 45.1 bits (105), Expect = 0.003
Identities = 59/224 (26%), Positives = 89/224 (39%), Gaps = 30/224 (13%)
Query: 80 CNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYL 139
C LK + D+ ++D E +L L E VE E M Y + + L
Sbjct: 21 CELK-ADKDYHFKVDNDENEHQLSLRTVSLGAGAKDELHIVEA---EAMNYEGSPIKVTL 76
Query: 140 YSSKPDID---SLTNYLCKD---LSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLK 193
+ K + SL + L C + P + A+S E E + L
Sbjct: 77 ATLKMSVQPTVSLGGFEITPPVVLRLKCGSGPVHISGQHLVAVEEDAESEDEEEEDVKLL 136
Query: 194 SMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDE---DEEEDEADFPSKLGKILKSKEN 250
S+ G APG S+ K A+ +DDD+DE DE++D+ DF ++
Sbjct: 137 SISGKRSAPGGG--SKVPQKKVKLAADEDDDDDDEEDDDEDDDDDDF-----------DD 183
Query: 251 EKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSK 294
E+ + K ++K I DT K+A K SN + K T SK
Sbjct: 184 EEAEEKAPVKKSIRDTPA---KNAQK-SNQNGKDSKPSSTPRSK 223
>gb|AAQ24860.1| nucleophosmin [Homo sapiens]
Length = 294
Score = 45.1 bits (105), Expect = 0.003
Identities = 59/224 (26%), Positives = 89/224 (39%), Gaps = 30/224 (13%)
Query: 80 CNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYL 139
C LK + D+ ++D E +L L E VE E M Y + + L
Sbjct: 21 CELK-ADKDYHFKVDNDENEHQLSLRTVSLGAGAKDELHIVEA---EAMNYEGSPIKVTL 76
Query: 140 YSSKPDID---SLTNYLCKD---LSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLK 193
+ K + SL + L C + P + A+S E E + L
Sbjct: 77 ATLKMSVQPTVSLGGFEITPPVVLRLKCGSGPVHISGQHLVAVEEDAESEDEEEEDVKLL 136
Query: 194 SMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDE---DEEEDEADFPSKLGKILKSKEN 250
S+ G APG S+ K A+ +DDD+DE DE++D+ DF ++
Sbjct: 137 SISGKRSAPGGG--SKVPQKKVKLAADEDDDDDDEEDDDEDDDDDDF-----------DD 183
Query: 251 EKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSK 294
E+ + K ++K I DT K+A K SN + K T SK
Sbjct: 184 EEAEEKAPVKKSIRDTPA---KNAQK-SNQNGKDSKPSSTPRSK 223
>gb|AAP36411.1| Homo sapiens nucleophosmin (nucleolar phosphoprotein B23, numatrin)
[synthetic construct] gi|60653673|gb|AAX29530.1|
nucleophosmin [synthetic construct]
Length = 295
Score = 45.1 bits (105), Expect = 0.003
Identities = 59/224 (26%), Positives = 89/224 (39%), Gaps = 30/224 (13%)
Query: 80 CNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYL 139
C LK + D+ ++D E +L L E VE E M Y + + L
Sbjct: 21 CELK-ADKDYHFKVDNDENEHQLSLRTVSLGAGAKDELHIVEA---EAMNYEGSPIKVTL 76
Query: 140 YSSKPDID---SLTNYLCKD---LSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLK 193
+ K + SL + L C + P + A+S E E + L
Sbjct: 77 ATLKMSVQPTVSLGGFEITPPVVLRLKCGSGPVHISGQHLVAVEEDAESEDEEEEDVKLL 136
Query: 194 SMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDE---DEEEDEADFPSKLGKILKSKEN 250
S+ G APG S+ K A+ +DDD+DE DE++D+ DF ++
Sbjct: 137 SISGKRSAPGGG--SKVPQKKVKLAADEDDDDDDEEDDDEDDDDDDF-----------DD 183
Query: 251 EKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSK 294
E+ + K ++K I DT K+A K SN + K T SK
Sbjct: 184 EEAEEKAPVKKSIRDTPA---KNAQK-SNQNGKDSKPSSTPRSK 223
>emb|CAG81736.1| unnamed protein product [Yarrowia lipolytica CLIB99]
gi|50547935|ref|XP_501437.1| hypothetical protein
[Yarrowia lipolytica]
Length = 262
Score = 45.1 bits (105), Expect = 0.003
Identities = 35/111 (31%), Positives = 55/111 (49%), Gaps = 14/111 (12%)
Query: 181 KSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDEDEEEDEADFPSK 240
K +E E ++ K+M+ + A +S DD +K+ + +E +DE+EDEEEDE K
Sbjct: 78 KGDQEKERARMQKAMKDIERARS-GYFSTDD--EKDSDSGDESEDEEEDEEEDEEKTLKK 134
Query: 241 LGKILKSKENEKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTT 291
K + E++ DW+ +D LKK K + +GKKTT
Sbjct: 135 SKKSSSTDEDDFSDWEG------IDKPGVLKKQKKKYVD-----GQGKKTT 174
>gb|AAH12566.1| Nucleophosmin 1 [Homo sapiens]
Length = 294
Score = 45.1 bits (105), Expect = 0.003
Identities = 59/224 (26%), Positives = 89/224 (39%), Gaps = 30/224 (13%)
Query: 80 CNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYL 139
C LK + D+ ++D E +L L E VE E M Y + + L
Sbjct: 21 CELK-ADKDYHFKVDNDENEHQLSLRTVSLGAGAKDELHIVEA---EAMNYEGSPIKVTL 76
Query: 140 YSSKPDID---SLTNYLCKD---LSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLK 193
+ K + SL + L C + P + A+S E E + L
Sbjct: 77 ATLKMSVQPTVSLGGFEITPPVVLRLKCGSGPVHISGQHLVAVEEDAESEDEEEEDVKLL 136
Query: 194 SMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDE---DEEEDEADFPSKLGKILKSKEN 250
S+ G APG S+ K A+ +DDD+DE DE++D+ DF ++
Sbjct: 137 SISGKRSAPGGG--SKVPQKKVKLAADEDDDDDDEEDDDEDDDDDDF-----------DD 183
Query: 251 EKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSK 294
E+ + K ++K I DT K+A K SN + K T SK
Sbjct: 184 EEAEEKAPVKKSIRDTPA---KNAQK-SNQNGKDSKPSSTPRSK 223
>gb|AAW67757.1| nucleophosmin [Homo sapiens]
Length = 298
Score = 45.1 bits (105), Expect = 0.003
Identities = 59/224 (26%), Positives = 89/224 (39%), Gaps = 30/224 (13%)
Query: 80 CNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYL 139
C LK + D+ ++D E +L L E VE E M Y + + L
Sbjct: 21 CELK-ADKDYHFKVDNDENEHQLSLRTVSLGAGAKDELHIVEA---EAMNYEGSPIKVTL 76
Query: 140 YSSKPDID---SLTNYLCKD---LSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLK 193
+ K + SL + L C + P + A+S E E + L
Sbjct: 77 ATLKMSVQPTVSLGGFEITPPVVLRLKCGSGPVHISGQHLVAVEEDAESEDEEEEDVKLL 136
Query: 194 SMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDE---DEEEDEADFPSKLGKILKSKEN 250
S+ G APG S+ K A+ +DDD+DE DE++D+ DF ++
Sbjct: 137 SISGKRSAPGGG--SKVPQKKVKLAADEDDDDDDEEDDDEDDDDDDF-----------DD 183
Query: 251 EKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSK 294
E+ + K ++K I DT K+A K SN + K T SK
Sbjct: 184 EEAEEKAPVKKSIRDTPA---KNAQK-SNQNGKDSKPSSTPRSK 223
>gb|AAW67756.1| nucleophosmin [Homo sapiens]
Length = 298
Score = 45.1 bits (105), Expect = 0.003
Identities = 59/224 (26%), Positives = 89/224 (39%), Gaps = 30/224 (13%)
Query: 80 CNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYL 139
C LK + D+ ++D E +L L E VE E M Y + + L
Sbjct: 21 CELK-ADKDYHFKVDNDENEHQLSLRTVSLGAGAKDELHIVEA---EAMNYEGSPIKVTL 76
Query: 140 YSSKPDID---SLTNYLCKD---LSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLK 193
+ K + SL + L C + P + A+S E E + L
Sbjct: 77 ATLKMSVQPTVSLGGFEITPPVVLRLKCGSGPVHISGQHLVAVEEDAESEDEEEEDVKLL 136
Query: 194 SMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDE---DEEEDEADFPSKLGKILKSKEN 250
S+ G APG S+ K A+ +DDD+DE DE++D+ DF ++
Sbjct: 137 SISGKRSAPGGG--SKVPQKKVKLAADEDDDDDDEEDDDEDDDDDDF-----------DD 183
Query: 251 EKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSK 294
E+ + K ++K I DT K+A K SN + K T SK
Sbjct: 184 EEAEEKAPVKKSIRDTPA---KNAQK-SNQNGKDSKPSSTPRSK 223
>gb|AAW67755.1| nucleophosmin [Homo sapiens] gi|58220457|gb|AAW67752.1|
nucleophosmin [Homo sapiens]
Length = 298
Score = 45.1 bits (105), Expect = 0.003
Identities = 59/224 (26%), Positives = 89/224 (39%), Gaps = 30/224 (13%)
Query: 80 CNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYL 139
C LK + D+ ++D E +L L E VE E M Y + + L
Sbjct: 21 CELK-ADKDYHFKVDNDENEHQLSLRTVSLGAGAKDELHIVEA---EAMNYEGSPIKVTL 76
Query: 140 YSSKPDID---SLTNYLCKD---LSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLK 193
+ K + SL + L C + P + A+S E E + L
Sbjct: 77 ATLKMSVQPTVSLGGFEITPPVVLRLKCGSGPVHISGQHLVAVEEDAESEDEEEEDVKLL 136
Query: 194 SMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDE---DEEEDEADFPSKLGKILKSKEN 250
S+ G APG S+ K A+ +DDD+DE DE++D+ DF ++
Sbjct: 137 SISGKRSAPGGG--SKVPQKKVKLAADEDDDDDDEEDDDEDDDDDDF-----------DD 183
Query: 251 EKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSK 294
E+ + K ++K I DT K+A K SN + K T SK
Sbjct: 184 EEAEEKAPVKKSIRDTPA---KNAQK-SNQNGKDSKPSSTPRSK 223
>gb|AAW67754.1| nucleophosmin [Homo sapiens]
Length = 298
Score = 45.1 bits (105), Expect = 0.003
Identities = 59/224 (26%), Positives = 89/224 (39%), Gaps = 30/224 (13%)
Query: 80 CNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYL 139
C LK + D+ ++D E +L L E VE E M Y + + L
Sbjct: 21 CELK-ADKDYHFKVDNDENEHQLSLRTVSLGAGAKDELHIVEA---EAMNYEGSPIKVTL 76
Query: 140 YSSKPDID---SLTNYLCKD---LSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLK 193
+ K + SL + L C + P + A+S E E + L
Sbjct: 77 ATLKMSVQPTVSLGGFEITPPVVLRLKCGSGPVHISGQHLVAVEEDAESEDEEEEDVKLL 136
Query: 194 SMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDE---DEEEDEADFPSKLGKILKSKEN 250
S+ G APG S+ K A+ +DDD+DE DE++D+ DF ++
Sbjct: 137 SISGKRSAPGGG--SKVPQKKVKLAADEDDDDDDEEDDDEDDDDDDF-----------DD 183
Query: 251 EKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSK 294
E+ + K ++K I DT K+A K SN + K T SK
Sbjct: 184 EEAEEKAPVKKSIRDTPA---KNAQK-SNQNGKDSKPSSTPRSK 223
>gb|AAW67753.1| nucleophosmin [Homo sapiens]
Length = 298
Score = 45.1 bits (105), Expect = 0.003
Identities = 59/224 (26%), Positives = 89/224 (39%), Gaps = 30/224 (13%)
Query: 80 CNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYL 139
C LK + D+ ++D E +L L E VE E M Y + + L
Sbjct: 21 CELK-ADKDYHFKVDNDENEHQLSLRTVSLGAGAKDELHIVEA---EAMNYEGSPIKVTL 76
Query: 140 YSSKPDID---SLTNYLCKD---LSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLK 193
+ K + SL + L C + P + A+S E E + L
Sbjct: 77 ATLKMSVQPTVSLGGFEITPPVVLRLKCGSGPVHISGQHLVAVEEDAESEDEEEEDVKLL 136
Query: 194 SMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDE---DEEEDEADFPSKLGKILKSKEN 250
S+ G APG S+ K A+ +DDD+DE DE++D+ DF ++
Sbjct: 137 SISGKRSAPGGG--SKVPQKKVKLAADEDDDDDDEEDDDEDDDDDDF-----------DD 183
Query: 251 EKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSK 294
E+ + K ++K I DT K+A K SN + K T SK
Sbjct: 184 EEAEEKAPVKKSIRDTPA---KNAQK-SNQNGKDSKPSSTPRSK 223
>emb|CAA34809.1| B23 nucleophosmin (280 AA) [Homo sapiens]
Length = 280
Score = 45.1 bits (105), Expect = 0.003
Identities = 59/224 (26%), Positives = 89/224 (39%), Gaps = 30/224 (13%)
Query: 80 CNLKKVEADWILRIDIVEKADRLELEEHDSEGQCNSECKTVERACQEVMGYSDTDVAEYL 139
C LK + D+ ++D E +L L E VE E M Y + + L
Sbjct: 7 CELK-ADKDYHFKVDNDENEHQLSLRTVSLGAGAKDELHIVEA---EAMNYEGSPIKVTL 62
Query: 140 YSSKPDID---SLTNYLCKD---LSKACNTKPPPVPKDRTPGEPFVAKSSKEAEMEKLLK 193
+ K + SL + L C + P + A+S E E + L
Sbjct: 63 ATLKMSVQPTVSLGGFEITPPVVLRLKCGSGPVHISGQHLVAVEEDAESEDEEEEDVKLL 122
Query: 194 SMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDE---DEEEDEADFPSKLGKILKSKEN 250
S+ G APG S+ K A+ +DDD+DE DE++D+ DF ++
Sbjct: 123 SISGKRSAPGGG--SKVPQKKVKLAADEDDDDDDEEDDDEDDDDDDF-----------DD 169
Query: 251 EKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKKTTSSK 294
E+ + K ++K I DT K+A K SN + K T SK
Sbjct: 170 EEAEEKAPVKKSIRDTPA---KNAQK-SNQNGKDSKPSSTPRSK 209
>gb|EAA21818.1| SNF2 family N-terminal domain, putative [Plasmodium yoelii yoelii]
Length = 1350
Score = 45.1 bits (105), Expect = 0.003
Identities = 38/142 (26%), Positives = 65/142 (45%), Gaps = 27/142 (19%)
Query: 187 EMEKLLKSMEGMPGAP------GMKMYSRDDLMKKNFGAE----NEDDDEDEDEEEDEAD 236
E+EK LK++E + G+ MY+ + + + ++ +EDDDED+D+EEDE +
Sbjct: 752 EIEKKLKNLENIFDLTNISLDGGLNMYNNLEKEESDNSSDEDGSDEDDDEDDDDEEDENE 811
Query: 237 FPSKLGKILKSKENEKG---------DWKQKIRKGIVDTSTTLKKHATKVSNHI------ 281
G L ++ E G K+K +K ++ S+T+KK H+
Sbjct: 812 SEGGDGNNLDNQIGEDGIVDENXKKKKKKKKKKKKKLNKSSTIKKFLKNNKKHMIFLDLG 871
Query: 282 --QRWWKGKKTTSSKKNSKSEL 301
+ WK T +K N K +
Sbjct: 872 ERKSKWKVMNTACTKTNKKKAI 893
>ref|XP_611626.1| PREDICTED: hypothetical protein XP_611626, partial [Bos taurus]
Length = 177
Score = 44.7 bits (104), Expect = 0.003
Identities = 38/132 (28%), Positives = 65/132 (48%), Gaps = 17/132 (12%)
Query: 171 DRTPGEPFVA-KSSKEAEMEKLLKSMEGMPGAPGMKMYSRDDLMKKNFGAENEDDDEDED 229
D T P VA S+K+AE+E+ +S + +K+ K+ E++++DE+ED
Sbjct: 60 DETGACPLVAGASAKKAELEENQRSYKQKKKRRRIKVQEDSSSENKSNSEEDKEEDEEED 119
Query: 230 EEEDEADFPSKLGKILKSKENEKGDWKQKIRKGIVDTSTTLKKHATKVSNHIQRWWKGKK 289
EEEDE D SK KG ++KIRK + D K T+ N ++ + +K
Sbjct: 120 EEEDEND---------DSKSPGKG--RKKIRKILKD-----DKLRTETQNALKEEEERRK 163
Query: 290 TTSSKKNSKSEL 301
+ ++ + +L
Sbjct: 164 RIAEREREREKL 175
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.311 0.129 0.376
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 528,329,621
Number of Sequences: 2540612
Number of extensions: 23285873
Number of successful extensions: 262941
Number of sequences better than 10.0: 1462
Number of HSP's better than 10.0 without gapping: 437
Number of HSP's successfully gapped in prelim test: 1143
Number of HSP's that attempted gapping in prelim test: 241180
Number of HSP's gapped (non-prelim): 7292
length of query: 301
length of database: 863,360,394
effective HSP length: 127
effective length of query: 174
effective length of database: 540,702,670
effective search space: 94082264580
effective search space used: 94082264580
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 75 (33.5 bits)
Medicago: description of AC148360.3


