

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148360.12 + phase: 0 /pseudo
(102 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
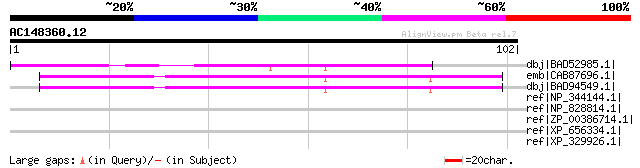
Score E
Sequences producing significant alignments: (bits) Value
dbj|BAD52985.1| unknown protein [Oryza sativa (japonica cultivar... 48 7e-05
emb|CAB87696.1| putative protein [Arabidopsis thaliana] gi|15239... 47 2e-04
dbj|BAD94549.1| putative protein [Arabidopsis thaliana] 47 2e-04
ref|NP_344144.1| Formate dehydrogenase Alpha subunit (fdhF-2) [S... 32 3.8
ref|NP_828814.1| putative IS116/IS110/IS902-family transposase [... 32 5.0
ref|ZP_00386714.1| COG1574: Predicted metal-dependent hydrolase ... 31 6.5
ref|XP_656334.1| hypothetical protein 14.t00038 [Entamoeba histo... 31 6.5
ref|XP_329926.1| hypothetical protein [Neurospora crassa] gi|289... 31 8.5
>dbj|BAD52985.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|53791397|dbj|BAD53434.1| unknown protein [Oryza
sativa (japonica cultivar-group)]
Length = 524
Score = 47.8 bits (112), Expect = 7e-05
Identities = 31/95 (32%), Positives = 51/95 (53%), Gaps = 20/95 (21%)
Query: 1 MQTKVDPPFDVIWVYSAIKFGCRKSLKGDILEQISAAKALFQLISACSASV-GGSKSIAL 59
+ T DPP + +W +SA+ F GD+ ++L L+SA +AS +K +AL
Sbjct: 56 LNTYPDPPLEAVWFFSALSF---HDNPGDL-------RSLLHLLSAFTASSRSAAKPLAL 105
Query: 60 LAP---------KAMKEVKSLVDMILGFMSICCSK 85
LAP K +E ++LV+ +L ++SIC S+
Sbjct: 106 LAPVVSELYHSAKPRREAEALVEAVLSYISICSSR 140
>emb|CAB87696.1| putative protein [Arabidopsis thaliana]
gi|15239784|ref|NP_196739.1| expressed protein
[Arabidopsis thaliana] gi|11281142|pir||T48537
hypothetical protein T22P22.170 - Arabidopsis thaliana
Length = 494
Score = 46.6 bits (109), Expect = 2e-04
Identities = 31/105 (29%), Positives = 53/105 (49%), Gaps = 14/105 (13%)
Query: 7 PPFDVIWVYSAIKFGCRKSLKGDILEQISAAKALFQLISACSASVGGSKSIALLAP---- 62
PP +++W YSAI+F K D + + FQLI + S S G K ++LL+P
Sbjct: 57 PPLELVWFYSAIRFYSSKLAFRD--DSVRLTSCFFQLIVSFSDSFSGVKKVSLLSPVVYQ 114
Query: 63 ------KAMKEVKSLVDMILGFMSICC--SKISEEKDLDLVLSFS 99
++ SL++ I+ ++S+ C +E+ D+ +V FS
Sbjct: 115 LSRLVISRRRDALSLLEGIVSYISMYCVDEPGNEDDDVLMVSGFS 159
>dbj|BAD94549.1| putative protein [Arabidopsis thaliana]
Length = 504
Score = 46.6 bits (109), Expect = 2e-04
Identities = 31/105 (29%), Positives = 53/105 (49%), Gaps = 14/105 (13%)
Query: 7 PPFDVIWVYSAIKFGCRKSLKGDILEQISAAKALFQLISACSASVGGSKSIALLAP---- 62
PP +++W YSAI+F K D + + FQLI + S S G K ++LL+P
Sbjct: 57 PPLELVWFYSAIRFYSSKLAFRD--DSVRLTSCFFQLIVSFSDSFSGVKKVSLLSPVVYQ 114
Query: 63 ------KAMKEVKSLVDMILGFMSICC--SKISEEKDLDLVLSFS 99
++ SL++ I+ ++S+ C +E+ D+ +V FS
Sbjct: 115 LSRLVISRRRDALSLLEGIVSYISMYCVDEPGNEDDDVLMVSGFS 159
>ref|NP_344144.1| Formate dehydrogenase Alpha subunit (fdhF-2) [Sulfolobus
solfataricus P2] gi|13816176|gb|AAK42934.1| Formate
dehydrogenase Alpha subunit (fdhF-2) [Sulfolobus
solfataricus P2] gi|25387635|pir||G90459 formate
dehydrogenase Alpha subunit (fdhF-2) [imported] -
Sulfolobus solfataricus
Length = 979
Score = 32.0 bits (71), Expect = 3.8
Identities = 19/55 (34%), Positives = 31/55 (55%), Gaps = 1/55 (1%)
Query: 42 QLISACSASVGGSKSIALLAPKAMKEVKSLVDMILGFMSICCSKISEEKDLDLVL 96
+L+ ACS V SI++ + +AM+ K+ + IL + + CS I E + D VL
Sbjct: 59 KLVRACSTRVEDGMSISVNSKRAMEARKTAISRILRYHKLYCS-ICENNNGDCVL 112
>ref|NP_828814.1| putative IS116/IS110/IS902-family transposase [Streptomyces
avermitilis MA-4680] gi|29611306|dbj|BAC75349.1|
putative IS116/IS110/IS902-family transposase
[Streptomyces avermitilis MA-4680]
Length = 397
Score = 31.6 bits (70), Expect = 5.0
Identities = 20/66 (30%), Positives = 30/66 (45%)
Query: 37 AKALFQLISACSASVGGSKSIALLAPKAMKEVKSLVDMILGFMSICCSKISEEKDLDLVL 96
AKA + +V G K IA + KEV +L + I G + + E + D+VL
Sbjct: 212 AKAAVEAAERQHTAVAGEKVIAQMVHTLAKEVMALNEKITGTEKLIEGRFREHELADIVL 271
Query: 97 SFSSLG 102
S +G
Sbjct: 272 SMPGMG 277
>ref|ZP_00386714.1| COG1574: Predicted metal-dependent hydrolase with the TIM-barrel
fold [Lactobacillus delbrueckii subsp. bulgaricus ATCC
BAA-365]
Length = 362
Score = 31.2 bits (69), Expect = 6.5
Identities = 15/47 (31%), Positives = 25/47 (52%)
Query: 13 WVYSAIKFGCRKSLKGDILEQISAAKALFQLISACSASVGGSKSIAL 59
W + K +S D L+Q+S + +F S C +SVG S+++ L
Sbjct: 22 WGFDESKLAEHRSPTRDDLDQVSTTQPIFLYRSDCHSSVGNSRALEL 68
>ref|XP_656334.1| hypothetical protein 14.t00038 [Entamoeba histolytica HM-1:IMSS]
gi|56473527|gb|EAL50949.1| hypothetical protein
14.t00038 [Entamoeba histolytica HM-1:IMSS]
Length = 1194
Score = 31.2 bits (69), Expect = 6.5
Identities = 20/77 (25%), Positives = 39/77 (49%), Gaps = 9/77 (11%)
Query: 22 CRKSLKGDILEQISAAKALFQLISACSASVGGSKSIALLAPKAMKEVKSLVDMILGFMSI 81
C+ +LK +LE IS A+ + +IS+ + ++ ++ L+ +I F++
Sbjct: 510 CKNNLKNGLLEDISIAERIGGMISSILGQIDNYEN--------KEQQHELITLIFIFINE 561
Query: 82 CC-SKISEEKDLDLVLS 97
C K EK LD++L+
Sbjct: 562 CIEDKKCREKQLDIILN 578
>ref|XP_329926.1| hypothetical protein [Neurospora crassa] gi|28920925|gb|EAA30256.1|
hypothetical protein [Neurospora crassa]
Length = 463
Score = 30.8 bits (68), Expect = 8.5
Identities = 13/50 (26%), Positives = 32/50 (64%), Gaps = 4/50 (8%)
Query: 25 SLKGDILEQISAAKALFQLISACSASVGGSKSIALLAPKAMKEVKSLVDM 74
+L+GD + A+ +F L+ +C+++ G ++++ P+ + E+ SL+D+
Sbjct: 114 ALRGDPV----LARQVFDLVRSCASTESGDEALSSALPQTIPEMLSLIDL 159
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.321 0.135 0.381
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 149,796,119
Number of Sequences: 2540612
Number of extensions: 4116328
Number of successful extensions: 9524
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 9520
Number of HSP's gapped (non-prelim): 8
length of query: 102
length of database: 863,360,394
effective HSP length: 78
effective length of query: 24
effective length of database: 665,192,658
effective search space: 15964623792
effective search space used: 15964623792
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 68 (30.8 bits)
Medicago: description of AC148360.12


