

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148348.6 + phase: 0
(552 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
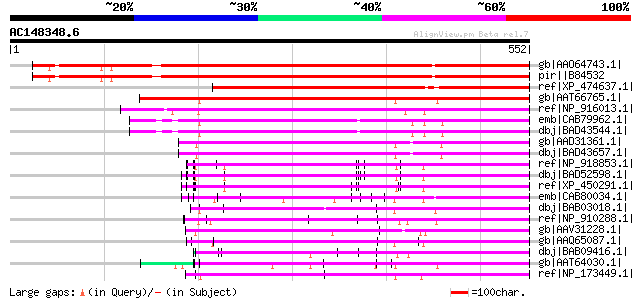
Score E
Sequences producing significant alignments: (bits) Value
gb|AAO64743.1| At2g15690/F9O13.24 [Arabidopsis thaliana] gi|2019... 540 e-152
pir||B84532 hypothetical protein At2g15690 [imported] - Arabidop... 540 e-152
ref|XP_474637.1| OSJNBa0039G19.7 [Oryza sativa (japonica cultiva... 460 e-128
gb|AAT66765.1| hypothetical protein PGEC160O2.3 [Solanum demissum] 357 5e-97
ref|NP_916013.1| putative selenium-binding protein [Oryza sativa... 326 1e-87
emb|CAB79962.1| putative protein [Arabidopsis thaliana] gi|40493... 285 2e-75
dbj|BAD43544.1| putative protein [Arabidopsis thaliana] 285 2e-75
gb|AAD31361.1| putative selenium-binding protein [Arabidopsis th... 276 2e-72
dbj|BAD43657.1| putative selenium-binding protein [Arabidopsis t... 275 2e-72
ref|NP_918853.1| P0458A05.10 [Oryza sativa (japonica cultivar-gr... 261 5e-68
dbj|BAD52598.1| pentatricopeptide (PPR) repeat-containing protei... 261 5e-68
ref|XP_450291.1| pentatricopeptide (PPR) repeat-containing prote... 259 1e-67
emb|CAB80034.1| putative protein [Arabidopsis thaliana] gi|44553... 258 3e-67
dbj|BAB03018.1| unnamed protein product [Arabidopsis thaliana] g... 254 4e-66
ref|NP_910288.1| pentatricopeptide (PPR) repeat-containing prote... 250 7e-65
gb|AAV31228.1| unknown protein [Oryza sativa (japonica cultivar-... 247 8e-64
gb|AAQ65087.1| At4g14850 [Arabidopsis thaliana] 245 3e-63
dbj|BAB09416.1| selenium-binding protein-like [Arabidopsis thali... 244 7e-63
gb|AAT64030.1| putative pentatricopeptide repeat protein [Gossyp... 244 7e-63
ref|NP_173449.1| pentatricopeptide (PPR) repeat-containing prote... 244 7e-63
>gb|AAO64743.1| At2g15690/F9O13.24 [Arabidopsis thaliana]
gi|20197709|gb|AAD17413.2| Expressed protein
[Arabidopsis thaliana] gi|14335136|gb|AAK59848.1|
At2g15690/F9O13.24 [Arabidopsis thaliana]
gi|18397896|ref|NP_565377.1| pentatricopeptide (PPR)
repeat-containing protein [Arabidopsis thaliana]
Length = 579
Score = 540 bits (1392), Expect = e-152
Identities = 280/547 (51%), Positives = 358/547 (65%), Gaps = 33/547 (6%)
Query: 25 KTLSTSAIPNEYGRLPP------HQVQQPPSDHNQHFNHHNTQNAPGRFPQQ--DWNPQN 76
K LSTSA N+Y + P HQ PP Q F+ N N R PQ W+ Q+
Sbjct: 47 KHLSTSAAANDYHQNPQSGSPSQHQRPYPP----QSFDSQNQTNTNQRVPQSPNQWSTQH 102
Query: 77 PNFRPPQPPQNPNYRQQPPP---QNPNFRPPPP------PQNPNFRPPPPPQNPNFRQPI 127
P QNP + Q PP QNP PQ+ RP Q P + P
Sbjct: 103 GGQIPQYGGQNPQHGGQRPPYGGQNPQQGGQMSQYGGHNPQHGGHRPQYGGQRPQYGGPG 162
Query: 128 R--QNPNFQPPNGPNRGINQNQWNPQNGNLNQFQNPNNQFQTPNVQEQAPPPPSIVDLTR 185
QN N Q N Q Q PQ P + Q+PN + PPPS+ ++ R
Sbjct: 163 NNYQNQNVQQSNQSQYYTPQQQQQPQP--------PRSSNQSPNQMNEVAPPPSVEEVMR 214
Query: 186 FCQEGKVKEALELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMH 245
CQ K+A+EL++KG D CF +LF+ C KS+E +KKVHD+FLQS FR D K++
Sbjct: 215 LCQRRLYKDAIELLDKGAMPDRECFVLLFESCANLKSLEHSKKVHDHFLQSKFRGDPKLN 274
Query: 246 NKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLE 305
N VI M+G C S+TDA+RVFDHM +++MDSWH+M+ Y+++ MGD+ L LFE+M + GL+
Sbjct: 275 NMVISMFGECSSITDAKRVFDHMVDKDMDSWHLMMCAYSDNGMGDDALHLFEEMTKHGLK 334
Query: 306 ITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEE 365
ET L V AC + +E+A+++ +SMK+++GI P EHY+G+L VLG+ G+L EAE+
Sbjct: 335 PNEETFLTVFLACATVGGIEEAFLHFDSMKNEHGISPKTEHYLGVLGVLGKCGHLVEAEQ 394
Query: 366 FIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSKAVANKIPTPPPKKYTA 425
+I LPFEPT +E ++NYAR+HGD+DLED++EEL+V +DPSKAV NKIPTPPPK +
Sbjct: 395 YIRDLPFEPTADFWEAMRNYARLHGDIDLEDYMEELMVDVDPSKAVINKIPTPPPKSFKE 454
Query: 426 ISMLDGKNRIIEYKNPTLYKDDEKLIAMNSMKDAGYVPDTRYVLHDIDQEAKEQALLYHS 485
+M+ K+RI+E++N T YKD+ K M + K YVPDTR+VLHDIDQEAKEQALLYHS
Sbjct: 455 TNMVTSKSRILEFRNLTFYKDEAK--EMAAKKGVVYVPDTRFVLHDIDQEAKEQALLYHS 512
Query: 486 ERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKIMSRIVGRELIVRDNKRFHHFKDGK 545
ERLAIAYG+I TPPR L IIKNLRVCGDCHN IKIMS+I+GR LIVRDNKRFHHFKDGK
Sbjct: 513 ERLAIAYGIICTPPRKTLTIIKNLRVCGDCHNFIKIMSKIIGRVLIVRDNKRFHHFKDGK 572
Query: 546 CSCGDYW 552
CSCGDYW
Sbjct: 573 CSCGDYW 579
>pir||B84532 hypothetical protein At2g15690 [imported] - Arabidopsis thaliana
Length = 989
Score = 540 bits (1392), Expect = e-152
Identities = 280/547 (51%), Positives = 358/547 (65%), Gaps = 33/547 (6%)
Query: 25 KTLSTSAIPNEYGRLPP------HQVQQPPSDHNQHFNHHNTQNAPGRFPQQ--DWNPQN 76
K LSTSA N+Y + P HQ PP Q F+ N N R PQ W+ Q+
Sbjct: 457 KHLSTSAAANDYHQNPQSGSPSQHQRPYPP----QSFDSQNQTNTNQRVPQSPNQWSTQH 512
Query: 77 PNFRPPQPPQNPNYRQQPPP---QNPNFRPPPP------PQNPNFRPPPPPQNPNFRQPI 127
P QNP + Q PP QNP PQ+ RP Q P + P
Sbjct: 513 GGQIPQYGGQNPQHGGQRPPYGGQNPQQGGQMSQYGGHNPQHGGHRPQYGGQRPQYGGPG 572
Query: 128 R--QNPNFQPPNGPNRGINQNQWNPQNGNLNQFQNPNNQFQTPNVQEQAPPPPSIVDLTR 185
QN N Q N Q Q PQ P + Q+PN + PPPS+ ++ R
Sbjct: 573 NNYQNQNVQQSNQSQYYTPQQQQQPQP--------PRSSNQSPNQMNEVAPPPSVEEVMR 624
Query: 186 FCQEGKVKEALELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMH 245
CQ K+A+EL++KG D CF +LF+ C KS+E +KKVHD+FLQS FR D K++
Sbjct: 625 LCQRRLYKDAIELLDKGAMPDRECFVLLFESCANLKSLEHSKKVHDHFLQSKFRGDPKLN 684
Query: 246 NKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLE 305
N VI M+G C S+TDA+RVFDHM +++MDSWH+M+ Y+++ MGD+ L LFE+M + GL+
Sbjct: 685 NMVISMFGECSSITDAKRVFDHMVDKDMDSWHLMMCAYSDNGMGDDALHLFEEMTKHGLK 744
Query: 306 ITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEE 365
ET L V AC + +E+A+++ +SMK+++GI P EHY+G+L VLG+ G+L EAE+
Sbjct: 745 PNEETFLTVFLACATVGGIEEAFLHFDSMKNEHGISPKTEHYLGVLGVLGKCGHLVEAEQ 804
Query: 366 FIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSKAVANKIPTPPPKKYTA 425
+I LPFEPT +E ++NYAR+HGD+DLED++EEL+V +DPSKAV NKIPTPPPK +
Sbjct: 805 YIRDLPFEPTADFWEAMRNYARLHGDIDLEDYMEELMVDVDPSKAVINKIPTPPPKSFKE 864
Query: 426 ISMLDGKNRIIEYKNPTLYKDDEKLIAMNSMKDAGYVPDTRYVLHDIDQEAKEQALLYHS 485
+M+ K+RI+E++N T YKD+ K M + K YVPDTR+VLHDIDQEAKEQALLYHS
Sbjct: 865 TNMVTSKSRILEFRNLTFYKDEAK--EMAAKKGVVYVPDTRFVLHDIDQEAKEQALLYHS 922
Query: 486 ERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKIMSRIVGRELIVRDNKRFHHFKDGK 545
ERLAIAYG+I TPPR L IIKNLRVCGDCHN IKIMS+I+GR LIVRDNKRFHHFKDGK
Sbjct: 923 ERLAIAYGIICTPPRKTLTIIKNLRVCGDCHNFIKIMSKIIGRVLIVRDNKRFHHFKDGK 982
Query: 546 CSCGDYW 552
CSCGDYW
Sbjct: 983 CSCGDYW 989
>ref|XP_474637.1| OSJNBa0039G19.7 [Oryza sativa (japonica cultivar-group)]
gi|38347147|emb|CAD39490.2| OSJNBa0039G19.7 [Oryza
sativa (japonica cultivar-group)]
Length = 557
Score = 460 bits (1184), Expect = e-128
Identities = 217/338 (64%), Positives = 277/338 (81%), Gaps = 8/338 (2%)
Query: 216 LCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDS 275
L K +E+ +K+HD+FL+S FR+D +++NK++EMY C +M ARR FDHMP+RNMDS
Sbjct: 227 LAPNPKLLEELRKIHDFFLRSPFRADLQVNNKMLEMYAKCAAMNHARRTFDHMPDRNMDS 286
Query: 276 WHMMIRGYANSTMGDEGLQLFEQMN-ELGLEITSETMLAVLSACGSAEAVEDAYIYLESM 334
WH+MI GYA + +GD LQLFE+M + G+ T+ T VL+AC ++EA+E+A++Y ++M
Sbjct: 287 WHIMIDGYAVNGLGDVALQLFEEMKTKYGIAPTAHTFTLVLNACANSEAIEEAFLYFDAM 346
Query: 335 KSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETLKNYARIHGDVDL 394
+GIEPGVEHY+G+++VLG+SG+L EA E+IE+LPFEPT TV+E+L N AR++GD+DL
Sbjct: 347 SRDHGIEPGVEHYVGIIEVLGKSGHLNEAVEYIEKLPFEPTDTVWESLLNLARMNGDIDL 406
Query: 395 EDHVEELIVSLDPSKAVANKIPTPPPKKYTAISMLDGKNRIIEYKNPTLYKDDEKLIAMN 454
ED EEL+VSLDP+K K+PTPPPK+ I+MLDG+N+++EY+ P K ++K++
Sbjct: 407 EDRAEELLVSLDPTKVNPKKLPTPPPKRRLGINMLDGRNKLVEYRLPP--KIEKKVV--- 461
Query: 455 SMKDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGD 514
+ YVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTP RTPLRIIKNLR+CGD
Sbjct: 462 --NEQRYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPARTPLRIIKNLRICGD 519
Query: 515 CHNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
CHNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW
Sbjct: 520 CHNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 557
>gb|AAT66765.1| hypothetical protein PGEC160O2.3 [Solanum demissum]
Length = 741
Score = 357 bits (916), Expect = 5e-97
Identities = 180/437 (41%), Positives = 277/437 (63%), Gaps = 23/437 (5%)
Query: 139 PNRGINQNQWNPQNGNLN--QFQNPNNQFQTPNVQEQAPPPPSIVDLTRFCQEGKVKEAL 196
P+ +Q + QNG + + ++ Q+ + + + S+ +L C+EGKVKEA+
Sbjct: 305 PSSAGHQASYQYQNGIVGHQEMRSSTPVEQSIDSDDSSSKKGSVDELDDLCKEGKVKEAV 364
Query: 197 ELME----KGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMY 252
E+++ + + D + + +L D+C + KS+EDAK +H++ ++S D KM+NK++EMY
Sbjct: 365 EVLQLLDQQHVTVDLSRYIMLMDVCSEDKSLEDAKSIHEHLVRSHPHLDIKMYNKILEMY 424
Query: 253 GNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETML 312
G C SM DA VF MP RN+ SW MI + +G++ ++LF + E G++ + L
Sbjct: 425 GKCGSMKDAFLVFRKMPQRNLTSWDTMITWLGKNGLGEDAIELFGEFKETGMKPDGQMFL 484
Query: 313 AVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPF 372
V AC + + ++ ESM Y I+ +E Y+G++D+LG +GYL EA EFIE++P
Sbjct: 485 GVFHACSVVGDIVEGMLHFESMSKDYDIDLSMEQYVGVVDMLGSTGYLDEAMEFIERMPI 544
Query: 373 EPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSK-----------AVANKIPTPPPK 421
EP++ V+ET+ N RIHG+++L D E++ LDPS+ A+ I K
Sbjct: 545 EPSIEVWETMMNLCRIHGNLELGDRCAEIVELLDPSRLDEQSKAGFLAVKASDIAKEKEK 604
Query: 422 KYTAISMLDGKNRIIEYK-NPTLYKDDEKLIAM-----NSMKDAGYVPDTRYVLHDIDQE 475
K +A S+L+ ++++ EY+ + D EK+ A+ MK+ GY+P+T++VLHD+DQE
Sbjct: 605 KRSAQSLLEARSKVHEYRAGDRSHPDHEKIYALLRGLKQLMKEDGYIPETKFVLHDVDQE 664
Query: 476 AKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKIMSRIVGRELIVRDN 535
KE AL+ HSERLA A GL+++ R+P+RIIKNLRVCGDCHNA+KI S++VGRE+I+RD
Sbjct: 665 TKEDALMAHSERLAFAQGLMNSSARSPIRIIKNLRVCGDCHNALKIASKLVGREIIMRDA 724
Query: 536 KRFHHFKDGKCSCGDYW 552
KRFHH KDG CSC DYW
Sbjct: 725 KRFHHLKDGLCSCRDYW 741
>ref|NP_916013.1| putative selenium-binding protein [Oryza sativa (japonica
cultivar-group)] gi|20160509|dbj|BAB89460.1|
selenium-binding protein-like [Oryza sativa (japonica
cultivar-group)]
Length = 796
Score = 326 bits (836), Expect = 1e-87
Identities = 174/464 (37%), Positives = 278/464 (59%), Gaps = 31/464 (6%)
Query: 118 PQNPNFRQPIRQNPNFQ-PPNGPNRGINQNQWNPQNGNLNQFQNPNNQFQTPNVQ----- 171
PQ+P I NF P NR + N P ++NP +P++
Sbjct: 335 PQSPYVSSKIDAQGNFPGQPMNVNRSVQHNTHAPALYQDGIYRNPLTD--SPSIDGLPSG 392
Query: 172 ----EQAPPPPSIVDLTRFCQEGKVKEALELM----EKGIKADANCFEILFDLCGKSKSV 223
++ ++ + C++GKVKEA+EL+ E+G A + L CG + S+
Sbjct: 393 ASDVTSGESKVTVEEMDKLCEDGKVKEAVELLALLQEEGTVVHAPQYFKLMQACGDATSL 452
Query: 224 EDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGY 283
+A+K+H+ QS D ++NK++EMY C SM DA+++F+ + RN+ SW+ +I G+
Sbjct: 453 AEARKIHNQISQSALAVDTDINNKILEMYAKCGSMEDAKKLFNTIAQRNLASWNTIISGF 512
Query: 284 ANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPG 343
+ +GDE F+Q + G + S V ACG +V++ ++ ESM+ +G+ P
Sbjct: 513 VYNGLGDEATDFFDQFKQTGNKPDSTMFTHVFLACGILGSVDEGMLHFESMQKDFGVTPT 572
Query: 344 VEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIV 403
+EHY+ ++++LGQSGY+ EA EF+EQ+P EP++ V+E+L N R++G ++L + +++
Sbjct: 573 MEHYVSIVNMLGQSGYIDEACEFVEQMPVEPSIDVWESLMNMCRLNGFLELGNRCAQIVE 632
Query: 404 SLDPSKA-VANKIPTPP--------PKKYTAISMLDGKNRIIEYK-----NPTLYKDDEK 449
LD S+ +KI P K+ + ++ ++++ EY+ +P K E+
Sbjct: 633 RLDSSRLNDQSKIGLFPVDASELAKEKERKKANAVEARSKVHEYRAGDRSHPEHLKIYEE 692
Query: 450 LIAMNS-MKDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKN 508
L + + MK+AGY+ DTR+VLHD+DQE KE ALL HSERLA++YGLI++ R+P+R+IKN
Sbjct: 693 LRYLAAHMKEAGYIADTRFVLHDVDQETKEDALLAHSERLAVSYGLITSAARSPIRVIKN 752
Query: 509 LRVCGDCHNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
LR CGDCH A+KI+S++VGR +I RD KRFHHF++G CSC DYW
Sbjct: 753 LRSCGDCHTALKIISKLVGRLIIARDAKRFHHFENGVCSCKDYW 796
>emb|CAB79962.1| putative protein [Arabidopsis thaliana] gi|4049347|emb|CAA22572.1|
putative protein [Arabidopsis thaliana]
gi|50897242|gb|AAT85760.1| At4g32450 [Arabidopsis
thaliana] gi|15236806|ref|NP_194971.1| pentatricopeptide
(PPR) repeat-containing protein [Arabidopsis thaliana]
gi|7486631|pir||T05355 hypothetical protein F8B4.150 -
Arabidopsis thaliana
Length = 537
Score = 285 bits (730), Expect = 2e-75
Identities = 161/442 (36%), Positives = 250/442 (56%), Gaps = 29/442 (6%)
Query: 128 RQNPNFQPPNGPNRGINQNQWNPQNGNLNQFQNPNNQFQTPNVQEQAPPPPSIVDLTRFC 187
++N N+Q +G + PQ N N FQ + S+ +L C
Sbjct: 108 QRNQNWQSSDGCSSYGTTGNGVPQENNTG-----GNHFQQDHSGHS-----SLDELDSIC 157
Query: 188 QEGKVKEALELME----KGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFK 243
+EGKVK+A+E+++ +G D + LCG ++++++AK VH++ S SD
Sbjct: 158 REGKVKKAVEIIKSWRNEGYVVDLPRLFWIAQLCGDAQALQEAKVVHEFITSSVGISDIS 217
Query: 244 MHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELG 303
+N +IEMY C S+ DA VF+ MP RN+++W +IR +A + G++ + F + + G
Sbjct: 218 AYNSIIEMYSGCGSVEDALTVFNSMPERNLETWCGVIRCFAKNGQGEDAIDTFSRFKQEG 277
Query: 304 LEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEA 363
+ E + ACG + + ++ ESM +YGI P +EHY+ L+ +L + GYL EA
Sbjct: 278 NKPDGEMFKEIFFACGVLGDMNEGLLHFESMYKEYGIIPCMEHYVSLVKMLAEPGYLDEA 337
Query: 364 EEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSKA-VANKIPTPPPKK 422
F+E + EP V ++ETL N +R+HGD+ L D ++++ LD S+ +K P K
Sbjct: 338 LRFVESM--EPNVDLWETLMNLSRVHGDLILGDRCQDMVEQLDASRLNKESKAGLVPVKS 395
Query: 423 YTAIS-----MLDGKNRIIEYK---NPTLYKDDEKLIAMNSMKD----AGYVPDTRYVLH 470
+ M G N I Y + + ++ E +A+ S+K+ GYVP ++ LH
Sbjct: 396 SDLVKEKLQRMAKGPNYGIRYMAAGDISRPENRELYMALKSLKEHMIEIGYVPLSKLALH 455
Query: 471 DIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKIMSRIVGREL 530
D+DQE+K++ L H+ER A + TP R+ +R++KNLRVC DCHNA+K+MS+IVGREL
Sbjct: 456 DVDQESKDENLFNHNERFAFISTFLDTPARSLIRVMKNLRVCADCHNALKLMSKIVGREL 515
Query: 531 IVRDNKRFHHFKDGKCSCGDYW 552
I RD KRFHH KDG CSC +YW
Sbjct: 516 ISRDAKRFHHMKDGVCSCREYW 537
>dbj|BAD43544.1| putative protein [Arabidopsis thaliana]
Length = 537
Score = 285 bits (730), Expect = 2e-75
Identities = 161/442 (36%), Positives = 250/442 (56%), Gaps = 29/442 (6%)
Query: 128 RQNPNFQPPNGPNRGINQNQWNPQNGNLNQFQNPNNQFQTPNVQEQAPPPPSIVDLTRFC 187
++N N+Q +G + PQ N N FQ + S+ +L C
Sbjct: 108 QRNQNWQSSDGCSSYGTTGNGVPQENNTG-----GNHFQQDHSGHS-----SLDELDSIC 157
Query: 188 QEGKVKEALELME----KGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFK 243
+EGKVK+A+E+++ +G D + LCG ++++++AK VH++ S SD
Sbjct: 158 REGKVKKAVEIIKSWRNEGYVVDLPRLFWMAQLCGDAQALQEAKVVHEFITSSVGISDIS 217
Query: 244 MHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELG 303
+N +IEMY C S+ DA VF+ MP RN+++W +IR +A + G++ + F + + G
Sbjct: 218 AYNSIIEMYSGCGSVEDALTVFNSMPERNLETWCGVIRCFAKNGQGEDAIDTFSRFKQEG 277
Query: 304 LEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEA 363
+ E + ACG + + ++ ESM +YGI P +EHY+ L+ +L + GYL EA
Sbjct: 278 NKPDGEMFKEIFFACGVLGDMNEGLLHFESMYKEYGIIPCMEHYVSLVKMLAEPGYLDEA 337
Query: 364 EEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSKA-VANKIPTPPPKK 422
F+E + EP V ++ETL N +R+HGD+ L D ++++ LD S+ +K P K
Sbjct: 338 LRFVESM--EPNVDLWETLMNLSRVHGDLILGDRCQDMVEQLDASRLNKESKAGLVPVKS 395
Query: 423 YTAIS-----MLDGKNRIIEYK---NPTLYKDDEKLIAMNSMKD----AGYVPDTRYVLH 470
+ M G N I Y + + ++ E +A+ S+K+ GYVP ++ LH
Sbjct: 396 SDLVKEKLQRMAKGPNYGIRYMAAGDISRPENRELYMALKSLKEHMIEIGYVPLSKLALH 455
Query: 471 DIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKIMSRIVGREL 530
D+DQE+K++ L H+ER A + TP R+ +R++KNLRVC DCHNA+K+MS+IVGREL
Sbjct: 456 DVDQESKDENLFNHNERFAFISTFLDTPARSLIRVMKNLRVCADCHNALKLMSKIVGREL 515
Query: 531 IVRDNKRFHHFKDGKCSCGDYW 552
I RD KRFHH KDG CSC +YW
Sbjct: 516 ISRDAKRFHHMKDGVCSCREYW 537
>gb|AAD31361.1| putative selenium-binding protein [Arabidopsis thaliana]
gi|25412333|pir||B84650 probable selenium-binding
protein [imported] - Arabidopsis thaliana
gi|15224758|ref|NP_180129.1| pentatricopeptide (PPR)
repeat-containing protein [Arabidopsis thaliana]
Length = 567
Score = 276 bits (705), Expect = 2e-72
Identities = 149/395 (37%), Positives = 231/395 (57%), Gaps = 24/395 (6%)
Query: 180 IVDLTRFCQEGKVKEALE----LMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQ 235
I + FC+ GKVK+AL L D + L +CG+++ +++AK VH
Sbjct: 175 IEEYDAFCKHGKVKKALYTIDILASMNYVVDLSRLLRLAKICGEAEGLQEAKTVHGKISA 234
Query: 236 STFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQL 295
S D ++ ++EMY NC +A VF+ M +N+++W ++IR +A + G++ + +
Sbjct: 235 SVSHLDLSSNHVLLEMYSNCGLANEAASVFEKMSEKNLETWCIIIRCFAKNGFGEDAIDM 294
Query: 296 FEQMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLG 355
F + E G + + ACG V++ ++ ESM YGI P +E Y+ L+++
Sbjct: 295 FSRFKEEGNIPDGQLFRGIFYACGMLGDVDEGLLHFESMSRDYGIAPSIEDYVSLVEMYA 354
Query: 356 QSGYLKEAEEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSK------ 409
G+L EA EF+E++P EP V V+ETL N +R+HG+++L D+ E++ LDP++
Sbjct: 355 LPGFLDEALEFVERMPMEPNVDVWETLMNLSRVHGNLELGDYCAEVVEFLDPTRLNKQSR 414
Query: 410 -----AVANKIPTPPPKKYTAISMLDG-KNRIIEYK--NPTLYKDDEKLIAMNSMK---- 457
A+ + KK + I L G K+ + E++ + L ++DE + ++K
Sbjct: 415 EGFIPVKASDVEKESLKKRSGI--LHGVKSSMQEFRAGDTNLPENDELFQLLRNLKMHMV 472
Query: 458 DAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHN 517
+ GYV +TR LHDIDQE+KE LL HSER+A A ++++ PR P +IKNLRVC DCHN
Sbjct: 473 EVGYVAETRMALHDIDQESKETLLLGHSERIAFARAVLNSAPRKPFTVIKNLRVCVDCHN 532
Query: 518 AIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
A+KIMS IVGRE+I RD KRFH K+G C+C DYW
Sbjct: 533 ALKIMSDIVGREVITRDIKRFHQMKNGACTCKDYW 567
>dbj|BAD43657.1| putative selenium-binding protein [Arabidopsis thaliana]
Length = 615
Score = 275 bits (704), Expect = 2e-72
Identities = 149/395 (37%), Positives = 231/395 (57%), Gaps = 24/395 (6%)
Query: 180 IVDLTRFCQEGKVKEALE----LMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQ 235
I + FC+ GKVK+AL L D + L +CG+++ +++AK VH
Sbjct: 223 IEEYDAFCKHGKVKKALYTIDILASMNYVVDLSRLLRLAKICGEAEGLQEAKTVHGKISA 282
Query: 236 STFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQL 295
S D ++ ++EMY NC +A VF+ M +N+++W ++IR +A + G++ + +
Sbjct: 283 SVSHLDLSSNHVLLEMYSNCGLANEAASVFEKMSEKNLETWCIIIRCFAKNGFGEDAIDM 342
Query: 296 FEQMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLG 355
F + E G + + ACG V++ ++ ESM YGI P +E Y+ L+++
Sbjct: 343 FSRFKEEGDIPDGQLFRGIFYACGMLGDVDEGLLHFESMSRDYGIAPSIEDYVSLVEMYA 402
Query: 356 QSGYLKEAEEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSK------ 409
G+L EA EF+E++P EP V V+ETL N +R+HG+++L D+ E++ LDP++
Sbjct: 403 LPGFLDEALEFVERMPMEPNVDVWETLMNLSRVHGNLELGDYCAEVVEFLDPTRLNKQSR 462
Query: 410 -----AVANKIPTPPPKKYTAISMLDG-KNRIIEYK--NPTLYKDDEKLIAMNSMK---- 457
A+ + KK + I L G K+ + E++ + L ++DE + ++K
Sbjct: 463 EGFIPVKASDVEKESLKKRSGI--LHGVKSSMQEFRAGDTNLPENDELFQLLRNLKMHMV 520
Query: 458 DAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHN 517
+ GYV +TR LHDIDQE+KE LL HSER+A A ++++ PR P +IKNLRVC DCHN
Sbjct: 521 EVGYVAETRMALHDIDQESKETLLLGHSERIAFARAVLNSAPRKPFTVIKNLRVCVDCHN 580
Query: 518 AIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
A+KIMS IVGRE+I RD KRFH K+G C+C DYW
Sbjct: 581 ALKIMSDIVGREVITRDIKRFHQMKNGACTCKDYW 615
>ref|NP_918853.1| P0458A05.10 [Oryza sativa (japonica cultivar-group)]
Length = 784
Score = 261 bits (666), Expect = 5e-68
Identities = 140/391 (35%), Positives = 212/391 (53%), Gaps = 27/391 (6%)
Query: 189 EGKVKEALELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKV 248
EG +K +E++ KG++ D L + C + E K+VH + ++ F SD N +
Sbjct: 394 EGAIKLFMEMLRKGLEPDPFVLSSLLNACASLSAYEQGKQVHAHLIKRQFMSDAFAGNAL 453
Query: 249 IEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITS 308
+ Y C S+ DA F +P R + SW MI G A G L+LF +M + G+
Sbjct: 454 VYTYAKCGSIEDAELAFSSLPERGVVSWSAMIGGLAQHGHGKRALELFGRMVDEGINPNH 513
Query: 309 ETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIE 368
TM +VL AC A V++A Y SMK +GI+ EHY ++D+LG++G L +A E +
Sbjct: 514 ITMTSVLCACNHAGLVDEAKRYFNSMKEMFGIDRTEEHYSCMIDLLGRAGKLDDAMELVN 573
Query: 369 QLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSKA------------------ 410
+PF+ +++ L +R+H D +L E + L+P K+
Sbjct: 574 SMPFQANASIWGALLGASRVHKDPELGKLAAEKLFILEPEKSGTHVLLANTYASAGMWNE 633
Query: 411 ---VANKIPTPPPKKYTAISMLDGKNRIIEY-----KNPTLYKDDEKLIAMNS-MKDAGY 461
V + KK A+S ++ K+++ + +P + KL+ + M AG+
Sbjct: 634 VAKVRKLMKDSNIKKEPAMSWIEVKDKVHTFIVGDKSHPMTKEIYAKLVELGDLMSKAGF 693
Query: 462 VPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKI 521
VP+ LHD+D+ KE L +HSERLA+A+ L+STPP P+R+ KNLR+C DCH A K
Sbjct: 694 VPNVDVDLHDLDRSEKELLLSHHSERLAVAFALLSTPPGAPIRVKKNLRICRDCHVAFKF 753
Query: 522 MSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
+S+IV RE+I+RD RFHHF+DG CSCGDYW
Sbjct: 754 ISKIVSREIIIRDINRFHHFRDGTCSCGDYW 784
Score = 64.3 bits (155), Expect = 1e-08
Identities = 47/221 (21%), Positives = 99/221 (44%), Gaps = 2/221 (0%)
Query: 196 LELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNC 255
L++ G+ + + C + + + +++H + +++ SD + +++MY
Sbjct: 199 LQMKYSGLVPNVFTLSSILKACSGAGAFDLGRQIHGFMIKANADSDDYIGVGLVDMYAKN 258
Query: 256 KSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVL 315
+ DAR+VFD M +R++ + +I G ++ DE L LF ++ + GL + T+ AVL
Sbjct: 259 HFLDDARKVFDWMFHRDLILCNALISGCSHGGRHDEALSLFYELRKEGLGVNRTTLAAVL 318
Query: 316 SACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPT 375
+ S EA + ++ K G GL+D + L +A E+
Sbjct: 319 KSTASLEAASTTR-QVHALAVKIGFIFDAHVVNGLIDSYWKCSCLSDANRVFEECSSGDI 377
Query: 376 VTVFETLKNYARI-HGDVDLEDHVEELIVSLDPSKAVANKI 415
+ + ++ HG+ ++ +E L L+P V + +
Sbjct: 378 IACTSMITALSQCDHGEGAIKLFMEMLRKGLEPDPFVLSSL 418
Score = 61.2 bits (147), Expect = 8e-08
Identities = 45/161 (27%), Positives = 79/161 (48%), Gaps = 5/161 (3%)
Query: 221 KSVEDAK---KVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPN-RNMDSW 276
K V DA+ +VH + + F SD + N ++ MYG M DARRVF+ + RN SW
Sbjct: 18 KCVPDARLGAQVHAMAMATGFGSDVFVANALVAMYGGFGFMDDARRVFNEADSERNAVSW 77
Query: 277 HMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKS 336
+ ++ Y + + +Q+F +M G++ T V++AC + +E A + +M
Sbjct: 78 NGLMSAYVKNDQCGDAIQVFGEMVWSGIQPTEFGFSCVVNACTGSRNIE-AGRQVHAMVV 136
Query: 337 KYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVT 377
+ G + V L+D+ + G + A E++P V+
Sbjct: 137 RMGYDKDVFTANALVDMYMKMGRVDIASVIFEKMPDSDVVS 177
Score = 58.5 bits (140), Expect = 5e-07
Identities = 39/172 (22%), Positives = 80/172 (45%), Gaps = 1/172 (0%)
Query: 197 ELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCK 256
E++ GI+ F + + C S+++E ++VH ++ + D N +++MY
Sbjct: 99 EMVWSGIQPTEFGFSCVVNACTGSRNIEAGRQVHAMVVRMGYDKDVFTANALVDMYMKMG 158
Query: 257 SMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLS 316
+ A +F+ MP+ ++ SW+ +I G + ++L QM GL T+ ++L
Sbjct: 159 RVDIASVIFEKMPDSDVVSWNALISGCVLNGHDHRAIELLLQMKYSGLVPNVFTLSSILK 218
Query: 317 ACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIE 368
AC A A D + K + +GL+D+ ++ +L +A + +
Sbjct: 219 ACSGAGAF-DLGRQIHGFMIKANADSDDYIGVGLVDMYAKNHFLDDARKVFD 269
Score = 54.3 bits (129), Expect = 1e-05
Identities = 42/187 (22%), Positives = 85/187 (44%), Gaps = 7/187 (3%)
Query: 190 GKVKEAL----ELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMH 245
G+ EAL EL ++G+ + + ++ ++VH ++ F D +
Sbjct: 290 GRHDEALSLFYELRKEGLGVNRTTLAAVLKSTASLEAASTTRQVHALAVKIGFIFDAHVV 349
Query: 246 NKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLE 305
N +I+ Y C ++DA RVF+ + ++ + MI + G+ ++LF +M GLE
Sbjct: 350 NGLIDSYWKCSCLSDANRVFEECSSGDIIACTSMITALSQCDHGEGAIKLFMEMLRKGLE 409
Query: 306 ITSETMLAVLSACGSAEAVEDA-YIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAE 364
+ ++L+AC S A E ++ +K ++ + + L+ + G +++AE
Sbjct: 410 PDPFVLSSLLNACASLSAYEQGKQVHAHLIKRQFMSDAFAGN--ALVYTYAKCGSIEDAE 467
Query: 365 EFIEQLP 371
LP
Sbjct: 468 LAFSSLP 474
>dbj|BAD52598.1| pentatricopeptide (PPR) repeat-containing protein-like [Oryza
sativa (japonica cultivar-group)]
Length = 877
Score = 261 bits (666), Expect = 5e-68
Identities = 140/391 (35%), Positives = 212/391 (53%), Gaps = 27/391 (6%)
Query: 189 EGKVKEALELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKV 248
EG +K +E++ KG++ D L + C + E K+VH + ++ F SD N +
Sbjct: 487 EGAIKLFMEMLRKGLEPDPFVLSSLLNACASLSAYEQGKQVHAHLIKRQFMSDAFAGNAL 546
Query: 249 IEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITS 308
+ Y C S+ DA F +P R + SW MI G A G L+LF +M + G+
Sbjct: 547 VYTYAKCGSIEDAELAFSSLPERGVVSWSAMIGGLAQHGHGKRALELFGRMVDEGINPNH 606
Query: 309 ETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIE 368
TM +VL AC A V++A Y SMK +GI+ EHY ++D+LG++G L +A E +
Sbjct: 607 ITMTSVLCACNHAGLVDEAKRYFNSMKEMFGIDRTEEHYSCMIDLLGRAGKLDDAMELVN 666
Query: 369 QLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSKA------------------ 410
+PF+ +++ L +R+H D +L E + L+P K+
Sbjct: 667 SMPFQANASIWGALLGASRVHKDPELGKLAAEKLFILEPEKSGTHVLLANTYASAGMWNE 726
Query: 411 ---VANKIPTPPPKKYTAISMLDGKNRIIEY-----KNPTLYKDDEKLIAMNS-MKDAGY 461
V + KK A+S ++ K+++ + +P + KL+ + M AG+
Sbjct: 727 VAKVRKLMKDSNIKKEPAMSWIEVKDKVHTFIVGDKSHPMTKEIYAKLVELGDLMSKAGF 786
Query: 462 VPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKI 521
VP+ LHD+D+ KE L +HSERLA+A+ L+STPP P+R+ KNLR+C DCH A K
Sbjct: 787 VPNVDVDLHDLDRSEKELLLSHHSERLAVAFALLSTPPGAPIRVKKNLRICRDCHVAFKF 846
Query: 522 MSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
+S+IV RE+I+RD RFHHF+DG CSCGDYW
Sbjct: 847 ISKIVSREIIIRDINRFHHFRDGTCSCGDYW 877
Score = 64.3 bits (155), Expect = 1e-08
Identities = 53/199 (26%), Positives = 95/199 (47%), Gaps = 7/199 (3%)
Query: 183 LTRFCQEGKVKEALELMEKGIKADANCFEILFDLCGKSKSVEDAK---KVHDYFLQSTFR 239
+T + G + A++ G++A+ C F L K V DA+ +VH + + F
Sbjct: 75 VTAYSNNGLPRSAIQAFH-GMRAEGVCCNE-FALPVVLKCVPDARLGAQVHAMAMATGFG 132
Query: 240 SDFKMHNKVIEMYGNCKSMTDARRVFDHMPN-RNMDSWHMMIRGYANSTMGDEGLQLFEQ 298
SD + N ++ MYG M DARRVF+ + RN SW+ ++ Y + + +Q+F +
Sbjct: 133 SDVFVANALVAMYGGFGFMDDARRVFNEADSERNAVSWNGLMSAYVKNDQCGDAIQVFGE 192
Query: 299 MNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSG 358
M G++ T V++AC + +E A + +M + G + V L+D+ + G
Sbjct: 193 MVWSGIQPTEFGFSCVVNACTGSRNIE-AGRQVHAMVVRMGYDKDVFTANALVDMYMKMG 251
Query: 359 YLKEAEEFIEQLPFEPTVT 377
+ A E++P V+
Sbjct: 252 RVDIASVIFEKMPDSDVVS 270
Score = 64.3 bits (155), Expect = 1e-08
Identities = 47/221 (21%), Positives = 99/221 (44%), Gaps = 2/221 (0%)
Query: 196 LELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNC 255
L++ G+ + + C + + + +++H + +++ SD + +++MY
Sbjct: 292 LQMKYSGLVPNVFTLSSILKACSGAGAFDLGRQIHGFMIKANADSDDYIGVGLVDMYAKN 351
Query: 256 KSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVL 315
+ DAR+VFD M +R++ + +I G ++ DE L LF ++ + GL + T+ AVL
Sbjct: 352 HFLDDARKVFDWMFHRDLILCNALISGCSHGGRHDEALSLFYELRKEGLGVNRTTLAAVL 411
Query: 316 SACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPT 375
+ S EA + ++ K G GL+D + L +A E+
Sbjct: 412 KSTASLEAASTTR-QVHALAVKIGFIFDAHVVNGLIDSYWKCSCLSDANRVFEECSSGDI 470
Query: 376 VTVFETLKNYARI-HGDVDLEDHVEELIVSLDPSKAVANKI 415
+ + ++ HG+ ++ +E L L+P V + +
Sbjct: 471 IACTSMITALSQCDHGEGAIKLFMEMLRKGLEPDPFVLSSL 511
Score = 58.5 bits (140), Expect = 5e-07
Identities = 39/172 (22%), Positives = 80/172 (45%), Gaps = 1/172 (0%)
Query: 197 ELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCK 256
E++ GI+ F + + C S+++E ++VH ++ + D N +++MY
Sbjct: 192 EMVWSGIQPTEFGFSCVVNACTGSRNIEAGRQVHAMVVRMGYDKDVFTANALVDMYMKMG 251
Query: 257 SMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLS 316
+ A +F+ MP+ ++ SW+ +I G + ++L QM GL T+ ++L
Sbjct: 252 RVDIASVIFEKMPDSDVVSWNALISGCVLNGHDHRAIELLLQMKYSGLVPNVFTLSSILK 311
Query: 317 ACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIE 368
AC A A D + K + +GL+D+ ++ +L +A + +
Sbjct: 312 ACSGAGAF-DLGRQIHGFMIKANADSDDYIGVGLVDMYAKNHFLDDARKVFD 362
Score = 54.3 bits (129), Expect = 1e-05
Identities = 42/187 (22%), Positives = 85/187 (44%), Gaps = 7/187 (3%)
Query: 190 GKVKEAL----ELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMH 245
G+ EAL EL ++G+ + + ++ ++VH ++ F D +
Sbjct: 383 GRHDEALSLFYELRKEGLGVNRTTLAAVLKSTASLEAASTTRQVHALAVKIGFIFDAHVV 442
Query: 246 NKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLE 305
N +I+ Y C ++DA RVF+ + ++ + MI + G+ ++LF +M GLE
Sbjct: 443 NGLIDSYWKCSCLSDANRVFEECSSGDIIACTSMITALSQCDHGEGAIKLFMEMLRKGLE 502
Query: 306 ITSETMLAVLSACGSAEAVEDA-YIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAE 364
+ ++L+AC S A E ++ +K ++ + + L+ + G +++AE
Sbjct: 503 PDPFVLSSLLNACASLSAYEQGKQVHAHLIKRQFMSDAFAGN--ALVYTYAKCGSIEDAE 560
Query: 365 EFIEQLP 371
LP
Sbjct: 561 LAFSSLP 567
Score = 48.9 bits (115), Expect = 4e-04
Identities = 37/145 (25%), Positives = 68/145 (46%), Gaps = 6/145 (4%)
Query: 229 VHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTM 288
+H L+S + F+ H +I Y C+ ARRVFD +P+ SW ++ Y+N+ +
Sbjct: 26 LHASLLKSGSLASFRNH--LISFYSKCRRPCCARRVFDEIPDPCHVSWSSLVTAYSNNGL 83
Query: 289 GDEGLQLFEQMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYM 348
+Q F M G+ +E L V+ C +A A ++ +M + +G + V +
Sbjct: 84 PRSAIQAFHGMRAEGV-CCNEFALPVVLKC-VPDARLGAQVHAMAMATGFGSDVFVAN-- 139
Query: 349 GLLDVLGQSGYLKEAEEFIEQLPFE 373
L+ + G G++ +A + E
Sbjct: 140 ALVAMYGGFGFMDDARRVFNEADSE 164
>ref|XP_450291.1| pentatricopeptide (PPR) repeat-containing protein-like [Oryza
sativa (japonica cultivar-group)]
gi|47848643|dbj|BAD22491.1| pentatricopeptide (PPR)
repeat-containing protein-like [Oryza sativa (japonica
cultivar-group)] gi|47848472|dbj|BAD22327.1|
pentatricopeptide (PPR) repeat-containing protein-like
[Oryza sativa (japonica cultivar-group)]
Length = 877
Score = 259 bits (663), Expect = 1e-67
Identities = 141/391 (36%), Positives = 210/391 (53%), Gaps = 27/391 (6%)
Query: 189 EGKVKEALELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKV 248
EG +K +E++ KG++ D L + C + E K+VH + ++ F SD N +
Sbjct: 487 EGAIKLFMEMLRKGLEPDPFVLSSLLNACASLSAYEQGKQVHAHLIKRQFMSDAFAGNAL 546
Query: 249 IEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITS 308
+ Y C S+ DA F +P R + SW MI G A G L+LF +M + G+
Sbjct: 547 VYTYAKCGSIEDAELAFSSLPERGVVSWSAMIGGLAQHGHGKRALELFGRMVDEGINPNH 606
Query: 309 ETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIE 368
TM +VL AC A V++A Y SMK +GI+ EHY ++D+LG++G L +A E +
Sbjct: 607 ITMTSVLCACNHAGLVDEAKRYFNSMKEMFGIDRTEEHYSCMIDLLGRAGKLDDAMELVN 666
Query: 369 QLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSKA------------------ 410
+PF+ +V+ L +R+H D +L E + L+P K+
Sbjct: 667 SMPFQANASVWGALLGASRVHKDPELGKLAAEKLFILEPEKSGTHVLLANTYASSGMWNE 726
Query: 411 ---VANKIPTPPPKKYTAISMLDGKNRIIEY-----KNPTLYKDDEKLIAMNS-MKDAGY 461
V + KK A+S ++ K+++ + +P + KL + M AGY
Sbjct: 727 VAKVRKLMKDSNIKKEPAMSWVEVKDKVHTFIVGDKSHPMTKEIYSKLDELGDLMSKAGY 786
Query: 462 VPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKI 521
+P+ LHD+D+ KE L +HSERLA+A+ L+STPP P+R+ KNLR+C DCH A K
Sbjct: 787 IPNVDVDLHDLDRSEKELLLSHHSERLAVAFALLSTPPGAPIRVKKNLRICRDCHMAFKF 846
Query: 522 MSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
+S IV RE+I+RD RFHHF+DG CSCGDYW
Sbjct: 847 ISNIVSREIIIRDINRFHHFRDGTCSCGDYW 877
Score = 67.4 bits (163), Expect = 1e-09
Identities = 64/236 (27%), Positives = 110/236 (46%), Gaps = 13/236 (5%)
Query: 183 LTRFCQEGKVKEALELMEKGIKADANCFEILFDLCGKSKSVEDAK---KVHDYFLQSTFR 239
+T + G + A++ G++A+ C F L K V DA+ +VH + + F
Sbjct: 75 VTAYSNNGLPRSAIQAFH-GMRAEGVCCNE-FALPVVLKCVPDAQLGAQVHAMAMATGFG 132
Query: 240 SDFKMHNKVIEMYGNCKSMTDARRVFDHM-PNRNMDSWHMMIRGYANSTMGDEGLQLFEQ 298
SD + N ++ MYG M DARRVFD RN SW+ ++ Y + + +Q+F +
Sbjct: 133 SDVFVANALVAMYGGFGFMDDARRVFDEAGSERNAVSWNGLMSAYVKNDQCGDAIQVFGE 192
Query: 299 MNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSG 358
M G++ T V++AC + + DA + +M + G E V L+D+ + G
Sbjct: 193 MVWSGIQPTEFGFSCVVNACTGSRNI-DAGRQVHAMVVRMGYEKDVFTANALVDMYVKMG 251
Query: 359 YLKEAEEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDH-VEELIVSLDPSKAVAN 413
+ A E++P + V + L + ++G DH EL++ + S V N
Sbjct: 252 RVDIASVIFEKMP-DSDVVSWNALISGCVLNG----HDHRAIELLLQMKSSGLVPN 302
Score = 67.4 bits (163), Expect = 1e-09
Identities = 47/221 (21%), Positives = 99/221 (44%), Gaps = 2/221 (0%)
Query: 196 LELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNC 255
L++ G+ + + C + + + +++H + +++ SD + +++MY
Sbjct: 292 LQMKSSGLVPNVFMLSSILKACAGAGAFDLGRQIHGFMIKANADSDDYIGVGLVDMYAKN 351
Query: 256 KSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVL 315
+ DA +VFD M +R++ W+ +I G ++ DE +F + + GL + T+ AVL
Sbjct: 352 HFLDDAMKVFDWMSHRDLILWNALISGCSHGGRHDEAFSIFYGLRKEGLGVNRTTLAAVL 411
Query: 316 SACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPT 375
+ S EA A + ++ K G GL+D + L +A E+
Sbjct: 412 KSTASLEAA-SATRQVHALAEKIGFIFDAHVVNGLIDSYWKCSCLSDAIRVFEECSSGDI 470
Query: 376 VTVFETLKNYARI-HGDVDLEDHVEELIVSLDPSKAVANKI 415
+ V + ++ HG+ ++ +E L L+P V + +
Sbjct: 471 IAVTSMITALSQCDHGEGAIKLFMEMLRKGLEPDPFVLSSL 511
Score = 55.8 bits (133), Expect = 3e-06
Identities = 37/167 (22%), Positives = 77/167 (45%), Gaps = 1/167 (0%)
Query: 197 ELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCK 256
E++ GI+ F + + C S++++ ++VH ++ + D N +++MY
Sbjct: 192 EMVWSGIQPTEFGFSCVVNACTGSRNIDAGRQVHAMVVRMGYEKDVFTANALVDMYVKMG 251
Query: 257 SMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLS 316
+ A +F+ MP+ ++ SW+ +I G + ++L QM GL + ++L
Sbjct: 252 RVDIASVIFEKMPDSDVVSWNALISGCVLNGHDHRAIELLLQMKSSGLVPNVFMLSSILK 311
Query: 317 ACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEA 363
AC A A D + K + +GL+D+ ++ +L +A
Sbjct: 312 ACAGAGAF-DLGRQIHGFMIKANADSDDYIGVGLVDMYAKNHFLDDA 357
Score = 50.4 bits (119), Expect = 1e-04
Identities = 36/141 (25%), Positives = 68/141 (47%), Gaps = 6/141 (4%)
Query: 229 VHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTM 288
+H L+S F + + H +I Y C+ ARRVFD +P+ SW ++ Y+N+ +
Sbjct: 26 LHANLLKSGFLASLRNH--LISFYSKCRRPCCARRVFDEIPDPCHVSWSSLVTAYSNNGL 83
Query: 289 GDEGLQLFEQMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYM 348
+Q F M G+ +E L V+ C +A A ++ +M + +G + V +
Sbjct: 84 PRSAIQAFHGMRAEGV-CCNEFALPVVLKC-VPDAQLGAQVHAMAMATGFGSDVFVAN-- 139
Query: 349 GLLDVLGQSGYLKEAEEFIEQ 369
L+ + G G++ +A ++
Sbjct: 140 ALVAMYGGFGFMDDARRVFDE 160
Score = 48.1 bits (113), Expect = 7e-04
Identities = 37/175 (21%), Positives = 78/175 (44%), Gaps = 3/175 (1%)
Query: 198 LMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKS 257
L ++G+ + + ++ ++VH + F D + N +I+ Y C
Sbjct: 395 LRKEGLGVNRTTLAAVLKSTASLEAASATRQVHALAEKIGFIFDAHVVNGLIDSYWKCSC 454
Query: 258 MTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSA 317
++DA RVF+ + ++ + MI + G+ ++LF +M GLE + ++L+A
Sbjct: 455 LSDAIRVFEECSSGDIIAVTSMITALSQCDHGEGAIKLFMEMLRKGLEPDPFVLSSLLNA 514
Query: 318 CGSAEAVEDA-YIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLP 371
C S A E ++ +K ++ + + L+ + G +++AE LP
Sbjct: 515 CASLSAYEQGKQVHAHLIKRQFMSDAFAGN--ALVYTYAKCGSIEDAELAFSSLP 567
>emb|CAB80034.1| putative protein [Arabidopsis thaliana] gi|4455331|emb|CAB36791.1|
putative protein [Arabidopsis thaliana]
gi|15234095|ref|NP_195043.1| pentatricopeptide (PPR)
repeat-containing protein [Arabidopsis thaliana]
gi|7486443|pir||T05197 hypothetical protein F4I10.100 -
Arabidopsis thaliana
Length = 990
Score = 258 bits (660), Expect = 3e-67
Identities = 133/360 (36%), Positives = 206/360 (56%), Gaps = 31/360 (8%)
Query: 222 SVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIR 281
++E +++H L+ +D + +++MY C S+ DA +F + N+ +W+ M+
Sbjct: 633 ALEQGRQIHANALKLNCTNDPFVGTSLVDMYAKCGSIDDAYCLFKRIEMMNITAWNAMLV 692
Query: 282 GYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIE 341
G A G E LQLF+QM LG++ T + VLSAC + V +AY ++ SM YGI+
Sbjct: 693 GLAQHGEGKETLQLFKQMKSLGIKPDKVTFIGVLSACSHSGLVSEAYKHMRSMHGDYGIK 752
Query: 342 PGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEEL 401
P +EHY L D LG++G +K+AE IE + E + +++ TL R+ GD + V
Sbjct: 753 PEIEHYSCLADALGRAGLVKQAENLIESMSMEASASMYRTLLAACRVQGDTETGKRVATK 812
Query: 402 IVSLDP---------------------SKAVANKIPTPPPKKYTAISMLDGKNRIIEY-- 438
++ L+P K + KK S ++ KN+I +
Sbjct: 813 LLELEPLDSSAYVLLSNMYAAASKWDEMKLARTMMKGHKVKKDPGFSWIEVKNKIHIFVV 872
Query: 439 ------KNPTLYKDDEKLIAMNSMKDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAY 492
+ +Y+ + +I +K GYVP+T + L D+++E KE+AL YHSE+LA+A+
Sbjct: 873 DDRSNRQTELIYRKVKDMI--RDIKQEGYVPETDFTLVDVEEEEKERALYYHSEKLAVAF 930
Query: 493 GLISTPPRTPLRIIKNLRVCGDCHNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
GL+STPP TP+R+IKNLRVCGDCHNA+K ++++ RE+++RD RFH FKDG CSCGDYW
Sbjct: 931 GLLSTPPSTPIRVIKNLRVCGDCHNAMKYIAKVYNREIVLRDANRFHRFKDGICSCGDYW 990
Score = 62.0 bits (149), Expect = 5e-08
Identities = 51/197 (25%), Positives = 91/197 (45%), Gaps = 8/197 (4%)
Query: 183 LTRFCQEGKVKEALELMEKGIKADANCFEILFDL----CGKSKSVEDAKKVHDYFLQSTF 238
L+ + G+ L+ +++D C ++ F L K S+ ++VH L+
Sbjct: 287 LSEYLHSGQYSALLKCFADMVESDVECDQVTFILMLATAVKVDSLALGQQVHCMALKLGL 346
Query: 239 RSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQ 298
+ N +I MY + AR VFD+M R++ SW+ +I G A + + E + LF Q
Sbjct: 347 DLMLTVSNSLINMYCKLRKFGFARTVFDNMSERDLISWNSVIAGIAQNGLEVEAVCLFMQ 406
Query: 299 MNELGLEITSETMLAVLSACGS-AEAVE-DAYIYLESMKSKYGIEPGVEHYMGLLDVLGQ 356
+ GL+ TM +VL A S E + +++ ++K + V L+D +
Sbjct: 407 LLRCGLKPDQYTMTSVLKAASSLPEGLSLSKQVHVHAIKINNVSDSFVS--TALIDAYSR 464
Query: 357 SGYLKEAEEFIEQLPFE 373
+ +KEAE E+ F+
Sbjct: 465 NRCMKEAEILFERHNFD 481
Score = 58.9 bits (141), Expect = 4e-07
Identities = 43/175 (24%), Positives = 80/175 (45%), Gaps = 4/175 (2%)
Query: 191 KVKEALELMEK-GIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVI 249
K + LM K G ++D +F CG ++ K+VH Y ++S + D + + ++
Sbjct: 500 KTLKLFALMHKQGERSDDFTLATVFKTCGFLFAINQGKQVHAYAIKSGYDLDLWVSSGIL 559
Query: 250 EMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSE 309
+MY C M+ A+ FD +P + +W MI G + + +F QM +G+
Sbjct: 560 DMYVKCGDMSAAQFAFDSIPVPDDVAWTTMISGCIENGEEERAFHVFSQMRLMGVLPDEF 619
Query: 310 TMLAVLSACGSAEAVEDA-YIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEA 363
T+ + A A+E I+ ++K +P V L+D+ + G + +A
Sbjct: 620 TIATLAKASSCLTALEQGRQIHANALKLNCTNDPFVG--TSLVDMYAKCGSIDDA 672
Score = 57.8 bits (138), Expect = 9e-07
Identities = 41/149 (27%), Positives = 71/149 (47%), Gaps = 14/149 (9%)
Query: 246 NKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMG-----DEGLQLFEQMN 300
N +I MY C S+T ARRVFD MP+R++ SW+ ++ YA S+ + LF +
Sbjct: 78 NNLISMYSKCGSLTYARRVFDKMPDRDLVSWNSILAAYAQSSECVVENIQQAFLLFRILR 137
Query: 301 ELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGV---EHYMG-LLDVLGQ 356
+ + + T+ +L C + Y++ Y + G+ E G L+++ +
Sbjct: 138 QDVVYTSRMTLSPMLKLC-----LHSGYVWASESFHGYACKIGLDGDEFVAGALVNIYLK 192
Query: 357 SGYLKEAEEFIEQLPFEPTVTVFETLKNY 385
G +KE + E++P+ V LK Y
Sbjct: 193 FGKVKEGKVLFEEMPYRDVVLWNLMLKAY 221
Score = 55.8 bits (133), Expect = 3e-06
Identities = 47/205 (22%), Positives = 97/205 (46%), Gaps = 6/205 (2%)
Query: 196 LELMEKGIKADANCF-EILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGN 254
++L+ G+K D +L + + +K+VH + ++ SD + +I+ Y
Sbjct: 405 MQLLRCGLKPDQYTMTSVLKAASSLPEGLSLSKQVHVHAIKINNVSDSFVSTALIDAYSR 464
Query: 255 CKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAV 314
+ M +A +F+ N ++ +W+ M+ GY S G + L+LF M++ G T+ V
Sbjct: 465 NRCMKEAEILFERH-NFDLVAWNAMMAGYTQSHDGHKTLKLFALMHKQGERSDDFTLATV 523
Query: 315 LSACGSAEAV-EDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFE 373
CG A+ + ++ ++KS Y ++ V G+LD+ + G + A+ + +P
Sbjct: 524 FKTCGFLFAINQGKQVHAYAIKSGYDLDLWVS--SGILDMYVKCGDMSAAQFAFDSIPV- 580
Query: 374 PTVTVFETLKNYARIHGDVDLEDHV 398
P + T+ + +G+ + HV
Sbjct: 581 PDDVAWTTMISGCIENGEEERAFHV 605
Score = 36.6 bits (83), Expect = 2.1
Identities = 20/99 (20%), Positives = 43/99 (43%)
Query: 213 LFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRN 272
+ LC S V ++ H Y + D + ++ +Y + + + +F+ MP R+
Sbjct: 151 MLKLCLHSGYVWASESFHGYACKIGLDGDEFVAGALVNIYLKFGKVKEGKVLFEEMPYRD 210
Query: 273 MDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETM 311
+ W++M++ Y +E + L + GL T+
Sbjct: 211 VVLWNLMLKAYLEMGFKEEAIDLSSAFHSSGLNPNEITL 249
>dbj|BAB03018.1| unnamed protein product [Arabidopsis thaliana]
gi|15229561|ref|NP_189042.1| pentatricopeptide (PPR)
repeat-containing protein [Arabidopsis thaliana]
Length = 633
Score = 254 bits (650), Expect = 4e-66
Identities = 139/391 (35%), Positives = 219/391 (55%), Gaps = 32/391 (8%)
Query: 193 KEALELME----KGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKV 248
++ALEL + G + + LF C + +E K VH Y ++S + N +
Sbjct: 244 EKALELFQGMLRDGFRPSHFSYASLFGACSSTGFLEQGKWVHAYMIKSGEKLVAFAGNTL 303
Query: 249 IEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITS 308
++MY S+ DAR++FD + R++ SW+ ++ YA G E + FE+M +G+
Sbjct: 304 LDMYAKSGSIHDARKIFDRLAKRDVVSWNSLLTAYAQHGFGKEAVWWFEEMRRVGIRPNE 363
Query: 309 ETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIE 368
+ L+VL+AC + +++ + Y E MK K GI P HY+ ++D+LG++G L A FIE
Sbjct: 364 ISFLSVLTACSHSGLLDEGWHYYELMK-KDGIVPEAWHYVTVVDLLGRAGDLNRALRFIE 422
Query: 369 QLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDP--------------------- 407
++P EPT +++ L N R+H + +L + E + LDP
Sbjct: 423 EMPIEPTAAIWKALLNACRMHKNTELGAYAAEHVFELDPDDPGPHVILYNIYASGGRWND 482
Query: 408 SKAVANKIPTPPPKKYTAISMLDGKNRIIEY-KNPTLYKDDEKLI-----AMNSMKDAGY 461
+ V K+ KK A S ++ +N I + N + E++ + +K+ GY
Sbjct: 483 AARVRKKMKESGVKKEPACSWVEIENAIHMFVANDERHPQREEIARKWEEVLAKIKELGY 542
Query: 462 VPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKI 521
VPDT +V+ +DQ+ +E L YHSE++A+A+ L++TPP + + I KN+RVCGDCH AIK+
Sbjct: 543 VPDTSHVIVHVDQQEREVNLQYHSEKIALAFALLNTPPGSTIHIKKNIRVCGDCHTAIKL 602
Query: 522 MSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
S++VGRE+IVRD RFHHFKDG CSC DYW
Sbjct: 603 ASKVVGREIIVRDTNRFHHFKDGNCSCKDYW 633
Score = 81.3 bits (199), Expect = 8e-14
Identities = 55/188 (29%), Positives = 84/188 (44%), Gaps = 1/188 (0%)
Query: 203 IKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDAR 262
I AD + L C K + + VH + LQS FR D M N ++ MY C S+ +AR
Sbjct: 56 IPADRRFYNTLLKKCTVFKLLIQGRIVHAHILQSIFRHDIVMGNTLLNMYAKCGSLEEAR 115
Query: 263 RVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAE 322
+VF+ MP R+ +W +I GY+ + L F QM G T+ +V+ A +AE
Sbjct: 116 KVFEKMPQRDFVTWTTLISGYSQHDRPCDALLFFNQMLRFGYSPNEFTLSSVIKA-AAAE 174
Query: 323 AVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETL 382
L K G + V LLD+ + G + +A+ + L V+ +
Sbjct: 175 RRGCCGHQLHGFCVKCGFDSNVHVGSALLDLYTRYGLMDDAQLVFDALESRNDVSWNALI 234
Query: 383 KNYARIHG 390
+AR G
Sbjct: 235 AGHARRSG 242
Score = 58.5 bits (140), Expect = 5e-07
Identities = 41/163 (25%), Positives = 82/163 (50%), Gaps = 2/163 (1%)
Query: 228 KVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANST 287
++H + ++ F S+ + + ++++Y M DA+ VFD + +RN SW+ +I G+A +
Sbjct: 182 QLHGFCVKCGFDSNVHVGSALLDLYTRYGLMDDAQLVFDALESRNDVSWNALIAGHARRS 241
Query: 288 MGDEGLQLFEQMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHY 347
++ L+LF+ M G + + ++ AC S +E ++ + K G +
Sbjct: 242 GTEKALELFQGMLRDGFRPSHFSYASLFGACSSTGFLEQGK-WVHAYMIKSGEKLVAFAG 300
Query: 348 MGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETLKNYARIHG 390
LLD+ +SG + +A + ++L V+ L YA+ HG
Sbjct: 301 NTLLDMYAKSGSIHDARKIFDRLAKRDVVSWNSLLTAYAQ-HG 342
>ref|NP_910288.1| pentatricopeptide (PPR) repeat-containing protein-like [Oryza
sativa (japonica cultivar-group)]
gi|7363286|dbj|BAA93030.1| pentatricopeptide (PPR)
repeat-containing protein-like [Oryza sativa (japonica
cultivar-group)]
Length = 734
Score = 250 bits (639), Expect = 7e-65
Identities = 138/393 (35%), Positives = 216/393 (54%), Gaps = 28/393 (7%)
Query: 187 CQEGKVKEALELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHN 246
C E V+ + + +K D+ + A+ +H Y ++ D +
Sbjct: 343 CSEDAVRLFTRMQLENVKPDSFTLVSVIPALADISDPLQARWIHGYSIRLHLDQDVYVLT 402
Query: 247 KVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEI 306
+I+MY C + AR +F+ R++ +W+ MI GY + G ++LFE+M +G+
Sbjct: 403 ALIDMYAKCGRVNIARILFNSARERHVITWNAMIHGYGSHGFGKAAVELFEEMKSIGIVP 462
Query: 307 TSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEF 366
T L+VLSAC A V++ Y SMK YG+EPG+EHY ++D+LG++G L EA F
Sbjct: 463 NETTFLSVLSACSHAGLVDEGREYFTSMKEDYGLEPGMEHYGTMVDLLGRAGKLDEAWAF 522
Query: 367 IEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSKAVAN------------- 413
I+++P +P ++V+ + ++H +V+L + + I L P + V +
Sbjct: 523 IQKMPMDPGLSVYGAMLGACKLHKNVELAEESAQKIFELGPQEGVYHVLLANIYANASMW 582
Query: 414 ----KIPTPPPK----KYTAISMLDGKNRI-IEYKNPTLYKDDEKLIA-----MNSMKDA 459
++ T K K S++ KN I Y T ++ +++ + + +K
Sbjct: 583 KDVARVRTAMEKNGLQKTPGWSIIQLKNEIHTFYSGSTNHQQAKEIYSRLAKLIEEIKAV 642
Query: 460 GYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAI 519
GYVPDT + HD++ + K Q L HSE+LAIA+GLI T P T ++I KNLRVC DCHNA
Sbjct: 643 GYVPDTDSI-HDVEDDVKAQLLNTHSEKLAIAFGLIRTAPGTTIQIKKNLRVCNDCHNAT 701
Query: 520 KIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
K++S + GRE+I+RD +RFHHFKDGKCSCGDYW
Sbjct: 702 KLISLVTGREIIMRDIQRFHHFKDGKCSCGDYW 734
Score = 79.0 bits (193), Expect = 4e-13
Identities = 45/190 (23%), Positives = 93/190 (48%), Gaps = 6/190 (3%)
Query: 186 FCQEGKVKEALELM-----EKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRS 240
+ + G + A+E++ E+G + D+ + C ++++ ++ H + ++S
Sbjct: 135 YARNGLARMAMEMVVRMQEEEGERPDSITLVSVLPACANARALAACREAHAFAIRSGLEE 194
Query: 241 DFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMN 300
+ +++ Y C + AR VFD MP +N SW+ MI GYA + E L LF +M
Sbjct: 195 LVNVATAILDAYCKCGDIRAARVVFDWMPTKNSVSWNAMIDGYAQNGDSREALALFNRMV 254
Query: 301 ELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYL 360
E G+++T ++LA L ACG +++ + + + + G++ V L+ + + +
Sbjct: 255 EEGVDVTDVSVLAALQACGELGCLDEG-MRVHELLVRIGLDSNVSVMNALITMYSKCKRV 313
Query: 361 KEAEEFIEQL 370
A ++L
Sbjct: 314 DLASHVFDEL 323
Score = 70.1 bits (170), Expect = 2e-10
Identities = 49/183 (26%), Positives = 85/183 (45%), Gaps = 3/183 (1%)
Query: 210 FEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMP 269
F L LC + + VH S+ + MY C+ DARRVFD MP
Sbjct: 62 FTSLLKLCAARGDLATGRAVHAQLAARGIDSEALAATALANMYAKCRRPADARRVFDRMP 121
Query: 270 NRNMDSWHMMIRGYANSTMGDEGLQLFEQM-NELGLEITSETMLAVLSACGSAEAVEDAY 328
R+ +W+ ++ GYA + + +++ +M E G S T+++VL AC +A A+ A
Sbjct: 122 VRDRVAWNALVAGYARNGLARMAMEMVVRMQEEEGERPDSITLVSVLPACANARALA-AC 180
Query: 329 IYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETLKNYARI 388
+ + G+E V +LD + G ++ A + +P + +V+ + YA+
Sbjct: 181 REAHAFAIRSGLEELVNVATAILDAYCKCGDIRAARVVFDWMPTKNSVSWNAMIDGYAQ- 239
Query: 389 HGD 391
+GD
Sbjct: 240 NGD 242
Score = 65.1 bits (157), Expect = 6e-09
Identities = 35/136 (25%), Positives = 69/136 (50%), Gaps = 4/136 (2%)
Query: 186 FCQEGKVKEALELMEKGIKADANCFEI----LFDLCGKSKSVEDAKKVHDYFLQSTFRSD 241
+ Q G +EAL L + ++ + ++ CG+ +++ +VH+ ++ S+
Sbjct: 237 YAQNGDSREALALFNRMVEEGVDVTDVSVLAALQACGELGCLDEGMRVHELLVRIGLDSN 296
Query: 242 FKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNE 301
+ N +I MY CK + A VFD + R SW+ MI G A + ++ ++LF +M
Sbjct: 297 VSVMNALITMYSKCKRVDLASHVFDELDRRTQVSWNAMILGCAQNGCSEDAVRLFTRMQL 356
Query: 302 LGLEITSETMLAVLSA 317
++ S T+++V+ A
Sbjct: 357 ENVKPDSFTLVSVIPA 372
>gb|AAV31228.1| unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 664
Score = 247 bits (630), Expect = 8e-64
Identities = 141/399 (35%), Positives = 209/399 (52%), Gaps = 37/399 (9%)
Query: 188 QEGKVKEAL----ELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFK 243
Q G+ EA+ E+ +GI+ ++ + ++ + H + L+ F D
Sbjct: 269 QNGRDLEAVDLFREMQSEGIEPNSVTIPCVLPAFANIAALMHGRSAHCFSLRKGFHHDIY 328
Query: 244 MHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELG 303
+ + +++MY C + DAR +F+ MP RN+ SW+ MI GYA + ++LF M
Sbjct: 329 VGSALVDMYAKCGRVRDARMIFEAMPYRNVVSWNAMIGGYAMHGEAENAVRLFRSMQSSK 388
Query: 304 LEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEA 363
+ T VL AC A E+ Y M+ K+GI P +EHY ++ +LG++G L +A
Sbjct: 389 EKPDLVTFTCVLGACSQAGWTEEGRSYFNEMQHKHGISPRMEHYACMVTLLGRAGKLDDA 448
Query: 364 EEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSKAVANKIPTPPPKKY 423
+ I Q+PFEP ++ +L R+HG+V L + E + L+P A + + Y
Sbjct: 449 YDIINQMPFEPDGCIWGSLLGSCRVHGNVVLAEVAAENLFQLEPENAGNYVLLS---NIY 505
Query: 424 TAISMLDGKNRI-----------------IEYKN------------PTLYKDDEKLIAMN 454
+ M DG NR+ IE KN P + EKL +
Sbjct: 506 ASKKMWDGVNRLRDMMKTVGLKKEKGCSWIEIKNKVHMLLAGDSSHPMMAAITEKLKHLT 565
Query: 455 -SMKDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCG 513
M+ G+ P T YVLHD++++ K+ L HSE+LA+A GLIST TPL++IKNLR+CG
Sbjct: 566 MEMRRLGFAPSTDYVLHDVEEQEKDDILSVHSEKLAVALGLISTSHGTPLQVIKNLRICG 625
Query: 514 DCHNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
DCH A+K +S RE+ VRD RFHHFKDGKCSC DYW
Sbjct: 626 DCHEAMKFISSFERREIYVRDTNRFHHFKDGKCSCADYW 664
Score = 58.2 bits (139), Expect = 7e-07
Identities = 51/245 (20%), Positives = 102/245 (40%), Gaps = 41/245 (16%)
Query: 188 QEGKVKEAL----ELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFK 243
+ G+ ++A+ + +G DA G V +++H Y +++ R D
Sbjct: 133 RSGRARDAVLALVRMHGEGFLPDATGVSCALSAVGDVGDVAVGEQLHGYVVKAGCRLDAC 192
Query: 244 MHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELG 303
+ +I+MYG C + RVFD + ++ S + ++ G + + E L+LF + G
Sbjct: 193 VATALIDMYGKCGRADEIVRVFDESSHMDVASCNALVAGLSRNAQVSEALRLFREFVGRG 252
Query: 304 LEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEP--------------------- 342
+E+ + ++++ C +A M+S+ GIEP
Sbjct: 253 IELNVVSWTSIVACCVQNGRDLEAVDLFREMQSE-GIEPNSVTIPCVLPAFANIAALMHG 311
Query: 343 ----------GVEH--YMG--LLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETLKNYARI 388
G H Y+G L+D+ + G +++A E +P+ V+ + YA +
Sbjct: 312 RSAHCFSLRKGFHHDIYVGSALVDMYAKCGRVRDARMIFEAMPYRNVVSWNAMIGGYA-M 370
Query: 389 HGDVD 393
HG+ +
Sbjct: 371 HGEAE 375
Score = 39.3 bits (90), Expect = 0.33
Identities = 32/140 (22%), Positives = 56/140 (39%), Gaps = 1/140 (0%)
Query: 226 AKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYAN 285
A+ +H + D + + ++ Y + DAR V D MP+R + W +I +A+
Sbjct: 39 ARALHAAAAVAGVSRDAFVASSLLHAYLRFGATADARSVLDGMPHRTVVGWSALIAAHAS 98
Query: 286 STMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVE 345
+ L E+M G+E T ++S + DA + L M + G P
Sbjct: 99 HGDAEGAWGLLERMRSDGVEPNVITWNGLVSGLNRSGRARDAVLALVRMHGE-GFLPDAT 157
Query: 346 HYMGLLDVLGQSGYLKEAEE 365
L +G G + E+
Sbjct: 158 GVSCALSAVGDVGDVAVGEQ 177
>gb|AAQ65087.1| At4g14850 [Arabidopsis thaliana]
Length = 634
Score = 245 bits (625), Expect = 3e-63
Identities = 133/365 (36%), Positives = 207/365 (56%), Gaps = 29/365 (7%)
Query: 217 CGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSW 276
C +E + +H + +++ + + +++MYG C + D+ + FD MP +N+ +
Sbjct: 270 CAGMAGLELGRSIHAHAVKACVERTIFVGSALVDMYGKCGCIEDSEQAFDEMPEKNLVTR 329
Query: 277 HMMIRGYANSTMGDEGLQLFEQMNELGLEITSE--TMLAVLSACGSAEAVEDAYIYLESM 334
+ +I GYA+ D L LFE+M G T T +++LSAC A AVE+ +SM
Sbjct: 330 NSLIGGYAHQGQVDMALALFEEMAPRGCGPTPNYMTFVSLLSACSRAGAVENGMKIFDSM 389
Query: 335 KSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETLKNYARIHGDVDL 394
+S YGIEPG EHY ++D+LG++G ++ A EFI+++P +PT++V+ L+N R+HG L
Sbjct: 390 RSTYGIEPGAEHYSCIVDMLGRAGMVERAYEFIKKMPIQPTISVWGALQNACRMHGKPQL 449
Query: 395 EDHVEELIVSLDP---------------------SKAVANKIPTPPPKKYTAISMLDGKN 433
E + LDP + V ++ KK S + KN
Sbjct: 450 GLLAAENLFKLDPKDSGNHVLLSNTFAAAGRWAEANTVREELKGVGIKKGAGYSWITVKN 509
Query: 434 RIIEY----KNPTLYKDDEKLIAM--NSMKDAGYVPDTRYVLHDIDQEAKEQALLYHSER 487
++ + ++ L K+ + +A N M+ AGY PD + L+D+++E K + +HSE+
Sbjct: 510 QVHAFQAKDRSHILNKEIQTTLAKLRNEMEAAGYKPDLKLSLYDLEEEEKAAEVSHHSEK 569
Query: 488 LAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKIMSRIVGRELIVRDNKRFHHFKDGKCS 547
LA+A+GL+S P P+RI KNLR+CGDCH+ K +S V RE+IVRDN RFH FKDG CS
Sbjct: 570 LALAFGLLSLPLSVPIRITKNLRICGDCHSFFKFVSGSVKREIIVRDNNRFHRFKDGICS 629
Query: 548 CGDYW 552
C DYW
Sbjct: 630 CKDYW 634
Score = 69.7 bits (169), Expect = 2e-10
Identities = 57/251 (22%), Positives = 117/251 (45%), Gaps = 17/251 (6%)
Query: 189 EGKVKEALELMEKGIKADANCFEILF----DLCGKSKSVEDAKKVHDYFLQSTFRSDFKM 244
+G+ +EA+E + + D + I F + C + ++H L+S F +D +
Sbjct: 137 DGRPREAIEAFIEFRRIDGHPNSITFCAFLNACSDWLHLNLGMQLHGLVLRSGFDTDVSV 196
Query: 245 HNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGL 304
N +I+ YG CK + + +F M +N SW ++ Y + ++ L+ + + +
Sbjct: 197 CNGLIDFYGKCKQIRSSEIIFTEMGTKNAVSWCSLVAAYVQNHEDEKASVLYLRSRKDIV 256
Query: 305 EITSETMLAVLSACGSAEAVE-DAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEA 363
E + + +VLSAC +E I+ ++K+ +E + L+D+ G+ G ++++
Sbjct: 257 ETSDFMISSVLSACAGMAGLELGRSIHAHAVKA--CVERTIFVGSALVDMYGKCGCIEDS 314
Query: 364 EEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSKAVANKIPTPPPKKY 423
E+ +++P + VT + YA G VD+ +L + +A + P P
Sbjct: 315 EQAFDEMPEKNLVTRNSLIGGYAH-QGQVDM---------ALALFEEMAPRGCGPTPNYM 364
Query: 424 TAISMLDGKNR 434
T +S+L +R
Sbjct: 365 TFVSLLSACSR 375
Score = 55.5 bits (132), Expect = 4e-06
Identities = 40/198 (20%), Positives = 81/198 (40%), Gaps = 1/198 (0%)
Query: 194 EALELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYG 253
E E+ +G+ + F F + K++H ++ D + +MY
Sbjct: 45 EFFEMRREGVVPNDFTFPCAFKAVASLRLPVTGKQIHALAVKCGRILDVFVGCSAFDMYC 104
Query: 254 NCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLA 313
+ DAR++FD +P RN+++W+ I E ++ F + + S T A
Sbjct: 105 KTRLRDDARKLFDEIPERNLETWNAFISNSVTDGRPREAIEAFIEFRRIDGHPNSITFCA 164
Query: 314 VLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFE 373
L+AC + + + L + + G + V GL+D G+ ++ +E ++ +
Sbjct: 165 FLNACSDWLHL-NLGMQLHGLVLRSGFDTDVSVCNGLIDFYGKCKQIRSSEIIFTEMGTK 223
Query: 374 PTVTVFETLKNYARIHGD 391
V+ + Y + H D
Sbjct: 224 NAVSWCSLVAAYVQNHED 241
>dbj|BAB09416.1| selenium-binding protein-like [Arabidopsis thaliana]
gi|15242550|ref|NP_196557.1| pentatricopeptide (PPR)
repeat-containing protein [Arabidopsis thaliana]
Length = 995
Score = 244 bits (622), Expect = 7e-63
Identities = 140/386 (36%), Positives = 217/386 (55%), Gaps = 31/386 (8%)
Query: 198 LMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKS 257
+++ G + D+ + + ++E +VH +++ SD + + +++MY C
Sbjct: 610 MLQTGQRLDSFMYATVLSAFASVATLERGMEVHACSVRACLESDVVVGSALVDMYSKCGR 669
Query: 258 MTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSE-TMLAVLS 316
+ A R F+ MP RN SW+ MI GYA G+E L+LFE M G T + VLS
Sbjct: 670 LDYALRFFNTMPVRNSYSWNSMISGYARHGQGEEALKLFETMKLDGQTPPDHVTFVGVLS 729
Query: 317 ACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTV 376
AC A +E+ + + ESM YG+ P +EH+ + DVLG++G L + E+FIE++P +P V
Sbjct: 730 ACSHAGLLEEGFKHFESMSDSYGLAPRIEHFSCMADVLGRAGELDKLEDFIEKMPMKPNV 789
Query: 377 TVFET-LKNYARIHG-DVDLEDHVEELIVSLDPSKAV---------------------AN 413
++ T L R +G +L E++ L+P AV
Sbjct: 790 LIWRTVLGACCRANGRKAELGKKAAEMLFQLEPENAVNYVLLGNMYAAGGRWEDLVKARK 849
Query: 414 KIPTPPPKK---YTAISMLDGKNRII--EYKNPTLYKDDEKLIAMN-SMKDAGYVPDTRY 467
K+ KK Y+ ++M DG + + + +P +KL +N M+DAGYVP T +
Sbjct: 850 KMKDADVKKEAGYSWVTMKDGVHMFVAGDKSHPDADVIYKKLKELNRKMRDAGYVPQTGF 909
Query: 468 VLHDIDQEAKEQALLYHSERLAIAYGLISTPPRT-PLRIIKNLRVCGDCHNAIKIMSRIV 526
L+D++QE KE+ L YHSE+LA+A+ L + T P+RI+KNLRVCGDCH+A K +S+I
Sbjct: 910 ALYDLEQENKEEILSYHSEKLAVAFVLAAQRSSTLPIRIMKNLRVCGDCHSAFKYISKIE 969
Query: 527 GRELIVRDNKRFHHFKDGKCSCGDYW 552
GR++I+RD+ RFHHF+DG CSC D+W
Sbjct: 970 GRQIILRDSNRFHHFQDGACSCSDFW 995
Score = 63.2 bits (152), Expect = 2e-08
Identities = 44/152 (28%), Positives = 79/152 (51%), Gaps = 6/152 (3%)
Query: 223 VEDAKKVHDYFLQSTFRSDFKMH--NKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMI 280
++ ++VH + + +T DF + N ++ MY C S+ DARRVF M +++ SW+ MI
Sbjct: 329 LKKGREVHGHVI-TTGLVDFMVGIGNGLVNMYAKCGSIADARRVFYFMTDKDSVSWNSMI 387
Query: 281 RGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAE-AVEDAYIYLESMKSKYG 339
G + E ++ ++ M + S T+++ LS+C S + A I+ ES+ K G
Sbjct: 388 TGLDQNGCFIEAVERYKSMRRHDILPGSFTLISSLSSCASLKWAKLGQQIHGESL--KLG 445
Query: 340 IEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLP 371
I+ V L+ + ++GYL E + +P
Sbjct: 446 IDLNVSVSNALMTLYAETGYLNECRKIFSSMP 477
Score = 57.8 bits (138), Expect = 9e-07
Identities = 47/196 (23%), Positives = 79/196 (39%), Gaps = 3/196 (1%)
Query: 196 LELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNC 255
L G K + F + E K++H L++ + N +I YG C
Sbjct: 506 LNAQRAGQKLNRITFSSVLSAVSSLSFGELGKQIHGLALKNNIADEATTENALIACYGKC 565
Query: 256 KSMTDARRVFDHMPNRNMD-SWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAV 314
M ++F M R + +W+ MI GY ++ + + L L M + G + S V
Sbjct: 566 GEMDGCEKIFSRMAERRDNVTWNSMISGYIHNELLAKALDLVWFMLQTGQRLDSFMYATV 625
Query: 315 LSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEP 374
LSA S +E + + + + +E V L+D+ + G L A F +P
Sbjct: 626 LSAFASVATLERG-MEVHACSVRACLESDVVVGSALVDMYSKCGRLDYALRFFNTMPVRN 684
Query: 375 TVTVFETLKNYARIHG 390
+ + + YAR HG
Sbjct: 685 SYSWNSMISGYAR-HG 699
Score = 49.3 bits (116), Expect = 3e-04
Identities = 33/126 (26%), Positives = 53/126 (41%), Gaps = 3/126 (2%)
Query: 226 AKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYAN 285
A+ H ++ D + N +I Y AR+VFD MP RN SW ++ GY+
Sbjct: 20 ARFFHSRLYKNRLDKDVYLCNNLINAYLETGDSVSARKVFDEMPLRNCVSWACIVSGYSR 79
Query: 286 STMGDEGLQLFEQMNELGLEITSETMLAVLSAC---GSAEAVEDAYIYLESMKSKYGIEP 342
+ E L M + G+ ++VL AC GS + I+ K Y ++
Sbjct: 80 NGEHKEALVFLRDMVKEGIFSNQYAFVSVLRACQEIGSVGILFGRQIHGLMFKLSYAVDA 139
Query: 343 GVEHYM 348
V + +
Sbjct: 140 VVSNVL 145
Score = 44.3 bits (103), Expect = 0.010
Identities = 46/201 (22%), Positives = 84/201 (40%), Gaps = 8/201 (3%)
Query: 186 FCQEGKVKEAL----ELMEKGIKADANCFEILFDLCGKSKSVED--AKKVHDYFLQSTFR 239
+ + G+ KEAL +++++GI ++ F + C + SV +++H + ++
Sbjct: 77 YSRNGEHKEALVFLRDMVKEGIFSNQYAFVSVLRACQEIGSVGILFGRQIHGLMFKLSYA 136
Query: 240 SDFKMHNKVIEMYGNC-KSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQ 298
D + N +I MY C S+ A F + +N SW+ +I Y+ + ++F
Sbjct: 137 VDAVVSNVLISMYWKCIGSVGYALCAFGDIEVKNSVSWNSIISVYSQAGDQRSAFRIFSS 196
Query: 299 MNELGLEITSETM-LAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQS 357
M G T T V +AC E + K G+ + GL+ +S
Sbjct: 197 MQYDGSRPTEYTFGSLVTTACSLTEPDVRLLEQIMCTIQKSGLLTDLFVGSGLVSAFAKS 256
Query: 358 GYLKEAEEFIEQLPFEPTVTV 378
G L A + Q+ VT+
Sbjct: 257 GSLSYARKVFNQMETRNAVTL 277
Score = 41.2 bits (95), Expect = 0.087
Identities = 42/194 (21%), Positives = 80/194 (40%), Gaps = 8/194 (4%)
Query: 183 LTRFCQEGKVKEALELMEKGIKADA--NCFEILFDL--CGKSKSVEDAKKVHDYFLQSTF 238
+T Q G EA+E + + D F ++ L C K + +++H L+
Sbjct: 387 ITGLDQNGCFIEAVERYKSMRRHDILPGSFTLISSLSSCASLKWAKLGQQIHGESLKLGI 446
Query: 239 RSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMG-DEGLQLFE 297
+ + N ++ +Y + + R++F MP + SW+ +I A S E + F
Sbjct: 447 DLNVSVSNALMTLYAETGYLNECRKIFSSMPEHDQVSWNSIIGALARSERSLPEAVVCFL 506
Query: 298 QMNELGLEITSETMLAVLSACGSAEAVE-DAYIYLESMKSKYGIEPGVEHYMGLLDVLGQ 356
G ++ T +VLSA S E I+ ++K+ E E+ L+ G+
Sbjct: 507 NAQRAGQKLNRITFSSVLSAVSSLSFGELGKQIHGLALKNNIADEATTEN--ALIACYGK 564
Query: 357 SGYLKEAEEFIEQL 370
G + E+ ++
Sbjct: 565 CGEMDGCEKIFSRM 578
Score = 40.0 bits (92), Expect = 0.19
Identities = 28/134 (20%), Positives = 65/134 (47%), Gaps = 6/134 (4%)
Query: 235 QSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQ 294
+S +D + + ++ + S++ AR+VF+ M RN + + ++ G G+E +
Sbjct: 236 KSGLLTDLFVGSGLVSAFAKSGSLSYARKVFNQMETRNAVTLNGLMVGLVRQKWGEEATK 295
Query: 295 LFEQMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYM-----G 349
LF MN + ++++ E+ + +LS+ E+ + + I G+ +M G
Sbjct: 296 LFMDMNSM-IDVSPESYVILLSSFPEYSLAEEVGLKKGREVHGHVITTGLVDFMVGIGNG 354
Query: 350 LLDVLGQSGYLKEA 363
L+++ + G + +A
Sbjct: 355 LVNMYAKCGSIADA 368
>gb|AAT64030.1| putative pentatricopeptide repeat protein [Gossypium hirsutum]
Length = 805
Score = 244 bits (622), Expect = 7e-63
Identities = 126/378 (33%), Positives = 209/378 (54%), Gaps = 28/378 (7%)
Query: 203 IKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDAR 262
+K D+ + C ++E K++H Y L++ + SD + N ++++Y C + AR
Sbjct: 428 LKPDSRTMACILPACASLSALERGKEIHGYILRNGYSSDRHVANALVDLYVKCGVLGLAR 487
Query: 263 RVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAE 322
+FD +P++++ SW +MI GY G+E + F +M + G+E + +++L AC +
Sbjct: 488 LLFDMIPSKDLVSWTVMIAGYGMHGYGNEAIATFNEMRDAGIEPDEVSFISILYACSHSG 547
Query: 323 AVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETL 382
+E + + MK+ + IEP +EHY ++D+L ++G L +A +FIE LP P T++ L
Sbjct: 548 LLEQGWRFFYIMKNDFNIEPKLEHYACMVDLLSRTGNLSKAYKFIETLPIAPDATIWGAL 607
Query: 383 KNYARIHGDVDLEDHVEELIVSLDPS---------------------KAVANKIPTPPPK 421
RI+ D++L + V E + L+P K + KI +
Sbjct: 608 LCGCRIYHDIELAEKVAERVFELEPENTGYYVLLANIYAEAEKREEVKRMREKIGKKGLR 667
Query: 422 KYTAISMLDGKNRIIEY--KNPTLYKDDEKLIAM-----NSMKDAGYVPDTRYVLHDIDQ 474
K S ++ K R+ + N + + +K+ ++ MK+ GY P T+Y L + D+
Sbjct: 668 KNPGCSWIEIKGRVNLFVSGNNSSHPHSKKIESLLKKMRRKMKEEGYFPKTKYALINADE 727
Query: 475 EAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKIMSRIVGRELIVRD 534
KE AL HSE+LA+A+GL++ PPR +R+ KNLRVCGDCH K MS+ RE+++RD
Sbjct: 728 MQKEMALCGHSEKLAMAFGLLTLPPRKTIRVTKNLRVCGDCHEMAKFMSKETRREIVLRD 787
Query: 535 NKRFHHFKDGKCSCGDYW 552
+ RFHHFKDG CSC +W
Sbjct: 788 SNRFHHFKDGYCSCRGFW 805
Score = 72.8 bits (177), Expect = 3e-11
Identities = 40/140 (28%), Positives = 69/140 (48%), Gaps = 3/140 (2%)
Query: 197 ELMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCK 256
++M GI D + C S ++ K VH ++S+F N +++MY C
Sbjct: 241 QMMYLGIDVDLATIISVLVGCANSGTLSLGKAVHSLAIKSSFERRINFSNTLLDMYSKCG 300
Query: 257 SMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLS 316
+ A RVF+ M RN+ SW MI GY D + L +QM + G+++ + ++L
Sbjct: 301 DLDGALRVFEKMGERNVVSWTSMIAGYTRDGWSDGAIILLQQMEKEGVKLDVVAITSILH 360
Query: 317 AC---GSAEAVEDAYIYLES 333
AC GS + +D + Y+++
Sbjct: 361 ACARSGSLDNGKDVHDYIKA 380
Score = 71.2 bits (173), Expect = 8e-11
Identities = 72/337 (21%), Positives = 128/337 (37%), Gaps = 71/337 (21%)
Query: 140 NRGINQNQWNPQNGNLNQFQNPNNQFQTPNVQEQAP-----PPPSIVD---------LTR 185
N+ +N ++ QNG + F +P + P P +D +
Sbjct: 16 NQNRKENFFSSQNGCFIHKPSLKTTFFSPIFRSCIPVRISATPTRTIDHQVTDYNAKILH 75
Query: 186 FCQEGKVKEALEL--MEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFK 243
FCQ G ++ A+EL M + + + + + LC KS+ D KKVH ++ D
Sbjct: 76 FCQLGDLENAMELVCMCQKSELETKTYGSVLQLCAGLKSLTDGKKVHSIIKSNSVGVDEA 135
Query: 244 MHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHM------------------------- 278
+ K++ Y C + + RRVFD M +N+ W+
Sbjct: 136 LGLKLVSFYATCGDLKEGRRVFDTMEKKNVYLWNFMVSEYAKIGDFKESICLFKIMVEKG 195
Query: 279 --------------------------MIRGYANSTMGDEGLQLFEQMNELGLEITSETML 312
MI GY ++ + + GL +++QM LG+++ T++
Sbjct: 196 IEGKRPESASELFDKLCDRDVISWNSMISGYVSNGLTERGLGIYKQMMYLGIDVDLATII 255
Query: 313 AVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPF 372
+VL C ++ + + S+ K E + LLD+ + G L A E++
Sbjct: 256 SVLVGCANSGTLSLGKA-VHSLAIKSSFERRINFSNTLLDMYSKCGDLDGALRVFEKMGE 314
Query: 373 EPTVTVFETLKNYAR---IHGDVDLEDHVEELIVSLD 406
V+ + Y R G + L +E+ V LD
Sbjct: 315 RNVVSWTSMIAGYTRDGWSDGAIILLQQMEKEGVKLD 351
Score = 64.7 bits (156), Expect = 7e-09
Identities = 53/197 (26%), Positives = 90/197 (44%), Gaps = 26/197 (13%)
Query: 196 LELMEK-GIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGN 254
L+ MEK G+K D + C +S S+++ K VHDY + S+ + N +++MY
Sbjct: 340 LQQMEKEGVKLDVVAITSILHACARSGSLDNGKDVHDYIKANNMASNLFVCNALMDMYAK 399
Query: 255 CKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAV 314
C SM A VF M +++ SW+ M+ G+ L+ S TM +
Sbjct: 400 CGSMEGANSVFSTMVVKDIISWNTMV--------GE-------------LKPDSRTMACI 438
Query: 315 LSACGSAEAVE-DAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFE 373
L AC S A+E I+ +++ Y + V + L+D+ + G L A + +P +
Sbjct: 439 LPACASLSALERGKEIHGYILRNGYSSDRHVAN--ALVDLYVKCGVLGLARLLFDMIPSK 496
Query: 374 PTVTVFETLKNYARIHG 390
V+ + Y +HG
Sbjct: 497 DLVSWTVMIAGYG-MHG 512
>ref|NP_173449.1| pentatricopeptide (PPR) repeat-containing protein [Arabidopsis
thaliana]
Length = 760
Score = 244 bits (622), Expect = 7e-63
Identities = 127/396 (32%), Positives = 219/396 (55%), Gaps = 31/396 (7%)
Query: 188 QEGKVKEALELMEK----GIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFK 243
Q GK EALEL + G+K + + CG ++ + H + ++ +
Sbjct: 365 QNGKDIEALELFREMQVAGVKPNHVTIPSMLPACGNIAALGHGRSTHGFAVRVHLLDNVH 424
Query: 244 MHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELG 303
+ + +I+MY C + ++ VF+ MP +N+ W+ ++ G++ E + +FE +
Sbjct: 425 VGSALIDMYAKCGRINLSQIVFNMMPTKNLVCWNSLMNGFSMHGKAKEVMSIFESLMRTR 484
Query: 304 LEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEA 363
L+ + ++LSACG ++ + Y + M +YGI+P +EHY ++++LG++G L+EA
Sbjct: 485 LKPDFISFTSLLSACGQVGLTDEGWKYFKMMSEEYGIKPRLEHYSCMVNLLGRAGKLQEA 544
Query: 364 EEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSK-------------- 409
+ I+++PFEP V+ L N R+ +VDL + E + L+P
Sbjct: 545 YDLIKEMPFEPDSCVWGALLNSCRLQNNVDLAEIAAEKLFHLEPENPGTYVLLSNIYAAK 604
Query: 410 -------AVANKIPTPPPKKYTAISMLDGKNRII-----EYKNPTLYKDDEKLIAMNS-M 456
++ NK+ + KK S + KNR+ + +P + + EK+ ++ M
Sbjct: 605 GMWTEVDSIRNKMESLGLKKNPGCSWIQVKNRVYTLLAGDKSHPQIDQITEKMDEISKEM 664
Query: 457 KDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCH 516
+ +G+ P+ + LHD++++ +EQ L HSE+LA+ +GL++TP TPL++IKNLR+CGDCH
Sbjct: 665 RKSGHRPNLDFALHDVEEQEQEQMLWGHSEKLAVVFGLLNTPDGTPLQVIKNLRICGDCH 724
Query: 517 NAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
IK +S GRE+ +RD RFHHFKDG CSCGD+W
Sbjct: 725 AVIKFISSYAGREIFIRDTNRFHHFKDGICSCGDFW 760
Score = 52.4 bits (124), Expect = 4e-05
Identities = 29/137 (21%), Positives = 63/137 (45%)
Query: 198 LMEKGIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKS 257
+ G+ D++ LF +C + + + K++H S D + + MY C
Sbjct: 107 MFSHGLIPDSHVLPNLFKVCAELSAFKVGKQIHCVSCVSGLDMDAFVQGSMFHMYMRCGR 166
Query: 258 MTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSA 317
M DAR+VFD M ++++ + ++ YA +E +++ +M G+E + +LS
Sbjct: 167 MGDARKVFDRMSDKDVVTCSALLCAYARKGCLEEVVRILSEMESSGIEANIVSWNGILSG 226
Query: 318 CGSAEAVEDAYIYLESM 334
+ ++A + + +
Sbjct: 227 FNRSGYHKEAVVMFQKI 243
Score = 41.6 bits (96), Expect = 0.067
Identities = 32/165 (19%), Positives = 73/165 (43%), Gaps = 7/165 (4%)
Query: 183 LTRFCQEGKVKEALELMEK----GIKADANCFEILFDLCGKSKSVEDAKKVHDYFLQSTF 238
L+ F + G KEA+ + +K G D + G S+ + + +H Y ++
Sbjct: 224 LSGFNRSGYHKEAVVMFQKIHHLGFCPDQVTVSSVLPSVGDSEMLNMGRLIHGYVIKQGL 283
Query: 239 RSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQ 298
D + + +I+MYG + +F+ + I G + + + D+ L++FE
Sbjct: 284 LKDKCVISAMIDMYGKSGHVYGIISLFNQFEMMEAGVCNAYITGLSRNGLVDKALEMFEL 343
Query: 299 MNELGLEITSETMLAVLSACG-SAEAVEDAYIYLESMKSKYGIEP 342
E +E+ + ++++ C + + +E ++ E + G++P
Sbjct: 344 FKEQTMELNVVSWTSIIAGCAQNGKDIEALELFREMQVA--GVKP 386
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.317 0.136 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,105,543,070
Number of Sequences: 2540612
Number of extensions: 57051759
Number of successful extensions: 675367
Number of sequences better than 10.0: 10355
Number of HSP's better than 10.0 without gapping: 3267
Number of HSP's successfully gapped in prelim test: 7750
Number of HSP's that attempted gapping in prelim test: 411122
Number of HSP's gapped (non-prelim): 72209
length of query: 552
length of database: 863,360,394
effective HSP length: 133
effective length of query: 419
effective length of database: 525,458,998
effective search space: 220167320162
effective search space used: 220167320162
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 78 (34.7 bits)
Medicago: description of AC148348.6


