

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148347.2 + phase: 0 /pseudo
(813 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
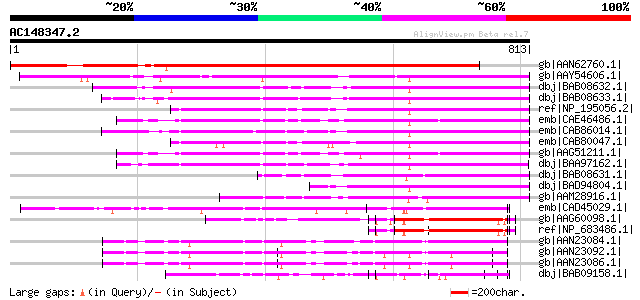
Score E
Sequences producing significant alignments: (bits) Value
gb|AAN62760.1| disease resistance protein-like protein MsR1 [Med... 1121 0.0
gb|AAY54606.1| NRG1 [Nicotiana benthamiana] 432 e-119
dbj|BAB08632.1| disease resistance protein-like [Arabidopsis tha... 403 e-110
dbj|BAB08633.1| disease resistance protein-like [Arabidopsis tha... 380 e-103
ref|NP_195056.2| disease resistance protein (CC-NBS-LRR class), ... 325 5e-87
emb|CAE46486.1| CC-NBS-LRR disease resistance protein [Arabidops... 309 2e-82
emb|CAB86014.1| disease resistance-like protein [Arabidopsis tha... 305 3e-81
emb|CAB80047.1| putative protein [Arabidopsis thaliana] gi|44902... 305 3e-81
gb|AAG51211.1| disease resistance protein, putative; 92850-95636... 303 2e-80
dbj|BAA97162.1| disease resistance protein-like [Arabidopsis tha... 300 1e-79
dbj|BAB08631.1| unnamed protein product [Arabidopsis thaliana] g... 253 1e-65
dbj|BAD94804.1| putative protein [Arabidopsis thaliana] 236 2e-60
gb|AAM28916.1| NBS/LRR [Pinus taeda] 225 5e-57
emb|CAD45029.1| NBS-LRR disease resistance protein homologue [Ho... 152 3e-35
gb|AAG60098.1| disease resistance protein, putative [Arabidopsis... 128 9e-28
ref|NP_683486.1| leucine-rich repeat family protein [Arabidopsis... 128 9e-28
gb|AAN23084.1| putative rp3 protein [Zea mays] 119 5e-25
gb|AAN23092.1| putative rp3 protein [Zea mays] 118 7e-25
gb|AAN23086.1| putative rp3 protein [Zea mays] 118 9e-25
dbj|BAB09158.1| disease resistance protein-like [Arabidopsis tha... 114 2e-23
>gb|AAN62760.1| disease resistance protein-like protein MsR1 [Medicago sativa]
Length = 704
Score = 1121 bits (2899), Expect = 0.0
Identities = 596/739 (80%), Positives = 631/739 (84%), Gaps = 41/739 (5%)
Query: 1 MSSATHIITLTTRFQFNDVLNKISQILQKAQKNQSSRRSLRLILKDLSPVVQDIKQYNDH 60
MSSATHIITL TRFQF+DVLNKISQI +KAQ+NQSSRRSLRLIL++LSPVVQDIKQYNDH
Sbjct: 1 MSSATHIITLATRFQFHDVLNKISQIAEKAQENQSSRRSLRLILRNLSPVVQDIKQYNDH 60
Query: 61 LDHPREEINSLIEENDAGESACTCSSENDSYGYVVEENQSLTVNDVEETLYKTREILELL 120
LDHPREEINSLIEENDA ESAC CSSENDS ++E L
Sbjct: 61 LDHPREEINSLIEENDAEESACKCSSENDS------------------------NVVEKL 96
Query: 121 NHEFNEDGQLFKRPFNVPENQKFTVGLDIPFSKLKMELLRGGSSTLVLTGLGGLGKTTLA 180
+E E G FK PF+VPEN KFTVGLDIPFSKLKMELLR GSSTLVLTGLGGLGKTTLA
Sbjct: 97 KNETFEAGPPFKCPFDVPENPKFTVGLDIPFSKLKMELLRDGSSTLVLTGLGGLGKTTLA 156
Query: 181 TKLCWDEEVNA*KSKIHCWVGYTI*QVENGASEGW*LHTCVDWFRRIGENHSGYKALLG* 240
TKLCWD+EVN + +V + S+ L T V+ RI E H GY
Sbjct: 157 TKLCWDQEVNGKFMENIIFVTF---------SKTPMLKTIVE---RIHE-HCGYPVPEFQ 203
Query: 241 RS*----WLGLLLKKVEGSPLLLVLDDVWPISEPLVEKIQFQISDFKILVTSRVAFPRFS 296
LGLLLKKVEGSPLLLVLDDVWP SE LVEK+QFQISDFKILVTSRVAFPRFS
Sbjct: 204 NDEDAVNRLGLLLKKVEGSPLLLVLDDVWPSSESLVEKLQFQISDFKILVTSRVAFPRFS 263
Query: 297 TTCILKPLAHEEAVTLFHHYAQMEKNSSDIINKNLVEKVVRSCQGLPLTIKVIATSLKNR 356
TTCILKPLAHE+AVTLFHHYA MEKNSSDII+KNLVEKVVRSCQGLPLTIKVIATSLKNR
Sbjct: 264 TTCILKPLAHEDAVTLFHHYALMEKNSSDIIDKNLVEKVVRSCQGLPLTIKVIATSLKNR 323
Query: 357 PRDMWRKIGKELSQGHSILDSNTDLLTRLQKIFDVLEDNPTIMECFMDIALFPEDHRIPV 416
P D+WRKI KELSQGHSILDSNT+LLTRLQKIFDVLEDNPTI+ECFMD+ALFPEDHRIPV
Sbjct: 324 PHDLWRKIVKELSQGHSILDSNTELLTRLQKIFDVLEDNPTIIECFMDLALFPEDHRIPV 383
Query: 417 AALVDMWAELYRLDDNGIQAMEIINKLGIMNLANVIIPRKNASDTDNNNYNNHFIILHDI 476
AALVDMWAELYRLDD GIQAMEIINKLGIMNLANVIIPRK+ASDTDNNNYNNHFI+LHDI
Sbjct: 384 AALVDMWAELYRLDDTGIQAMEIINKLGIMNLANVIIPRKDASDTDNNNYNNHFIMLHDI 443
Query: 477 LRELGIYQSAKEPFEQRKRLIIDINKNKSGLAEKQQGLMTCILSKFMRLCVKRNPQQLTA 536
LRELGIY+S KEPFEQRKRLIID++KNKSGL EKQQGLM ILSKFMRLCVK+NPQQL A
Sbjct: 444 LRELGIYRSTKEPFEQRKRLIIDMHKNKSGLTEKQQGLMIRILSKFMRLCVKQNPQQLAA 503
Query: 537 RILSVSADETCAFDWSQMEPAQVEVLILNLHTKQYSLPEWIAKMSKLRVLIITNYDFHPS 596
RILSVS DETCA DWSQMEPAQVEVLILNLHTKQYSLPEWI KMSKL+VLIITNY HPS
Sbjct: 504 RILSVSTDETCALDWSQMEPAQVEVLILNLHTKQYSLPEWIGKMSKLKVLIITNYTVHPS 563
Query: 597 KLNNIELLGSLQNLERIRLERIYVPSFGTLKNLKKLSLYMCNTILAFEKGSILISDAFPN 656
+L N ELL SLQNLE+IRLERI VPSF T+KNLKKLSLYMCNT LAFEKGSILISDAFPN
Sbjct: 564 ELTNFELLSSLQNLEKIRLERISVPSFATVKNLKKLSLYMCNTRLAFEKGSILISDAFPN 623
Query: 657 LEELNIDYCKDLVVLPIGICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDL 716
LEELNIDYCKDL+VLP GICDI LKKL VTNCHK+SSLP+DIGKLENLELLSLSSCTDL
Sbjct: 624 LEELNIDYCKDLLVLPTGICDIISLKKLSVTNCHKISSLPEDIGKLENLELLSLSSCTDL 683
Query: 717 EAIPTSIGKLLNLKHLDIS 735
EAIPTSI KLLNLKHLDIS
Sbjct: 684 EAIPTSIEKLLNLKHLDIS 702
>gb|AAY54606.1| NRG1 [Nicotiana benthamiana]
Length = 850
Score = 432 bits (1112), Expect = e-119
Identities = 305/868 (35%), Positives = 439/868 (50%), Gaps = 108/868 (12%)
Query: 16 FNDVLNKISQILQKAQKNQSSRRSLRLILKDLSPVVQDIKQYNDHLDHPREEINSLIEEN 75
F+ +L + + K +S +SL L D+ PV DI++ N LD EI ++
Sbjct: 15 FDILLKAVLDVGIKIATFRSKFQSLIKTLNDIKPVFDDIERLNKALDGRDYEIEMFKKQL 74
Query: 76 DAGESACTCSSENDSYGYVVEENQSLTVNDVEETLY------------------------ 111
AGE S+ Y + + N S + +E +L
Sbjct: 75 FAGEELVRKCSKTKCYDALKKWNYSRKLTKLENSLVRFCQVHGFIQVCRDSKIILVNVIE 134
Query: 112 ---KTREILELL-------------------------NHEFNEDGQLFKRPFNVPENQKF 143
K +I +L N + +G F +VP+
Sbjct: 135 HGKKLDQITSMLRGISLRNGSSIGFTNSNGSSGWMNGNSFGSTNGSGFSGWSDVPQFSDS 194
Query: 144 TVGLDIPFSKLKMELLRGGSSTLVLTGLGGLGKTTLATKLCWDEEVNA*KSKIHCWVGYT 203
VG D+P +LK++LL +VL+ G GKTTLA LC ++++ K K T
Sbjct: 195 VVGFDLPLQELKVKLLEEKEKVVVLSAPAGCGKTTLAAMLCQEDDI---KDKYRDIFFVT 251
Query: 204 I*QVENGASEGW*LHTCVDWFRRIGE--NHSGYK----ALLG*RS*WLGLLLKKVEGSPL 257
+ + N R +GE GYK A L LL++ P+
Sbjct: 252 VSKKANIK-------------RIVGEIFEMKGYKGPDFASEHAAVCQLNNLLRRSTSQPV 298
Query: 258 LLVLDDVWPISEPLVEKIQFQISDFKILVTSRVAFPRFSTTCILKPLAHEEAVTLFHHYA 317
LLVLDDVW S+ ++E FQI FKILVTSR FP+F T L L+ ++A LF Y+
Sbjct: 299 LLVLDDVWSESDFVIESFIFQIPGFKILVTSRSVFPKFDTYK-LNLLSEKDAKALF--YS 355
Query: 318 QMEKNSSDIINKNLVEKVVRSCQGLPLTIKVIATSLKNRPRDMWRKIGKELSQGHSILDS 377
K+S + +LV K VRSC G PL +KV+ SL +P +W S+ + +
Sbjct: 356 SAFKDSIPYVQLDLVHKAVRSCCGFPLALKVVGRSLCGQPELIWFNRVMLQSKRQILFPT 415
Query: 378 NTDLLTRLQKIFDVLED-------NPTIMECFMDIALFPEDHRIPVAALVDMWAELYRLD 430
DLL L+ D L++ T+ +C++D+ FPEDHRI AA++DMW E Y LD
Sbjct: 416 ENDLLRTLRASIDALDEIDLYSSEATTLRDCYLDLGSFPEDHRIHAAAILDMWVERYNLD 475
Query: 431 DNGIQAMEIINKLGIMNLANVIIPRKNASDTDNNNYNNHFIILHDILRELGIYQSAKEPF 490
++G++AM I +L NL N+ + RK+A +N H+I HD+LREL I+Q ++
Sbjct: 476 EDGMKAMAIFFQLSSQNLVNLALARKDAPAV-LGLHNLHYIQQHDLLRELVIHQCDEKTV 534
Query: 491 EQRKRLIIDINKNKSGLAEKQQGLMTCILSKFMRLCVKRNPQQLTARILSVSADETCAFD 550
E+RKRL I+I N QQ L Q L A +LS+ DE
Sbjct: 535 EERKRLYINIKGNDFPKWWSQQRL-----------------QPLQAEVLSIFTDEHFESV 577
Query: 551 WSQMEPAQVEVLILNLHTKQYSLPEWIAKMSKLRVLIITNYDFHPSKLNNIELLGSLQNL 610
W + +VEVL+LN TK Y+ P ++ +MS+L+ LI+ N F P+KLNN +L SL NL
Sbjct: 578 WYDVRFPKVEVLVLNFETKTYNFPPFVEQMSQLKTLIVANNYFFPTKLNNFQLC-SLLNL 636
Query: 611 ERIRLERIYVPSFGT----LKNLKKLSLYMCNTILAFEKGSILISDAFPNLEELNIDYCK 666
+RI LERI V S T L NL+K+S MC AFE + +S +P L E+NI+YC
Sbjct: 637 KRISLERISVTSIFTANLQLPNLRKISFIMCEIGEAFENYAANMSYMWPKLVEMNIEYCS 696
Query: 667 DLVVLPIGICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKL 726
DLV +P CD+ LKKL + CH+L +LP+++GKL NLE+L L SCT++ +P S+ KL
Sbjct: 697 DLVEVPAETCDLVGLKKLSICYCHELVALPEELGKLSNLEVLRLHSCTNVSKLPESVVKL 756
Query: 727 LNLKHLDISNCISLSSLPEEFGNLCNLKNLDMAS-CASIELPFSVVNLQNLKTITCDEET 785
L LD+ +C+ L LP E LC+L+ + M S ELP SV+ L L+ + CDEET
Sbjct: 757 NRLGFLDVYDCVELDFLPREMDQLCSLRTICMGSRLGFTELPDSVLRLVKLEDVVCDEET 816
Query: 786 AATWEDFQHMLPNMKIEVLHVDVNLNWL 813
A+ WE ++ L N++I V+ D+NLN L
Sbjct: 817 ASLWEYYKEHLRNLRITVIKEDINLNLL 844
>dbj|BAB08632.1| disease resistance protein-like [Arabidopsis thaliana]
gi|15240126|ref|NP_201491.1| disease resistance protein
(CC-NBS-LRR class), putative [Arabidopsis thaliana]
gi|46395984|sp|Q9FKZ1|DRL42_ARATH Probable disease
resistance protein At5g66900
Length = 809
Score = 403 bits (1036), Expect = e-110
Identities = 258/698 (36%), Positives = 384/698 (54%), Gaps = 59/698 (8%)
Query: 130 LFKRPFNVPENQKFTVGLDIPFSKLKMELLRGGSSTLVLTGLGGLGKTTLATKLCWDEEV 189
+F+ +VP+ K VGLD P +LK LL TLV++ G GKTTL ++LC D ++
Sbjct: 154 VFRDLCSVPKLDKVIVGLDWPLGELKKRLLDDSVVTLVVSAPPGCGKTTLVSRLCDDPDI 213
Query: 190 NA*KSKIHCWVGYTI*QVENGASEGW*LHTCVDWFRRIGEN---HSGYKALLG*RS*WLG 246
I V V N + FR I +N H+GY AL
Sbjct: 214 KGKFKHIFFNV------VSNTPN-----------FRVIVQNLLQHNGYNALTFENDSQAE 256
Query: 247 LLLKKV-----EGSPLLLVLDDVWPISEPLVEKIQFQISDFKILVTSRVAFPRFSTTCIL 301
+ L+K+ E P+LLVLDDVW ++ ++K Q ++ ++KILVTSR FP F + L
Sbjct: 257 VGLRKLLEELKENGPILLVLDDVWRGADSFLQKFQIKLPNYKILVTSRFDFPSFDSNYRL 316
Query: 302 KPLAHEEAVTLFHHYAQMEKNSSDIINKNLVEKVVRSCQGLPLTIKVIATSLKNRPRDMW 361
KPL ++A L H+A N+S ++L++K+++ C G P+ I+V+ SLK R + W
Sbjct: 317 KPLEDDDARALLIHWASRPCNTSPDEYEDLLQKILKRCNGFPIVIEVVGVSLKGRSLNTW 376
Query: 362 RKIGKELSQGHSILDSN-TDLLTRLQKIFDVLEDNPTIMECFMDIALFPEDHRIPVAALV 420
+ + S+G IL +L LQ FD L+ P + ECF+D+ F ED +I + ++
Sbjct: 377 KGQVESWSEGEKILGKPYPTVLECLQPSFDALD--PNLKECFLDMGSFLEDQKIRASVII 434
Query: 421 DMWAELYRLDDNGIQAMEIINKLGIMNLANVIIPRKNASDTDNNNYNNHFIILHDILREL 480
DMW ELY + + + L NL ++ N + ++ YN+ + HDILREL
Sbjct: 435 DMWVELYGKGSSILYMY--LEDLASQNLLKLVPLGTN--EHEDGFYNDFLVTQHDILREL 490
Query: 481 GIYQSAKEPFEQRKRLIIDINKNKSGLAEKQQGLMTCILSKFMRLCVKRNPQQLTARILS 540
I QS + +RKRL ++I +N F C+ + A +LS
Sbjct: 491 AICQSEFKENLERKRLNLEILENT-----------------FPDWCLNT----INASLLS 529
Query: 541 VSADETCAFDWSQMEPAQVEVLILNLHTKQYSLPEWIAKMSKLRVLIITNYDFHPSKLNN 600
+S D+ + W +M+ VE L+LNL + Y+LP +I+ M KL+VL ITN+ F+P++L+N
Sbjct: 530 ISTDDLFSSKWLEMDCPNVEALVLNLSSSDYALPSFISGMKKLKVLTITNHGFYPARLSN 589
Query: 601 IELLGSLQNLERIRLERIYVPSFGT----LKNLKKLSLYMCNTILAF-EKGSILISDAFP 655
L SL NL+RIRLE++ + L +LKKLSL MC+ F + I++S+A
Sbjct: 590 FSCLSSLPNLKRIRLEKVSITLLDIPQLQLSSLKKLSLVMCSFGEVFYDTEDIVVSNALS 649
Query: 656 NLEELNIDYCKDLVVLPIGICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTD 715
L+E++IDYC DL LP I +I LK L +TNC+KLS LP+ IG L LE+L L S +
Sbjct: 650 KLQEIDIDYCYDLDELPYWISEIVSLKTLSITNCNKLSQLPEAIGNLSRLEVLRLCSSMN 709
Query: 716 LEAIPTSIGKLLNLKHLDISNCISLSSLPEEFGNLCNLKNLDMASCASIELPFSVVNLQN 775
L +P + L NL+ LDIS+C+ L LP+E G L NLK + M C+ ELP SV NL+N
Sbjct: 710 LSELPEATEGLSNLRFLDISHCLGLRKLPQEIGKLQNLKKISMRKCSGCELPESVTNLEN 769
Query: 776 LKTITCDEETAATWEDFQHMLPNMKIEVLHVDVNLNWL 813
L+ + CDEET WE + + N++++ ++ NLN L
Sbjct: 770 LE-VKCDEETGLLWERLKPKMRNLRVQEEEIEHNLNLL 806
>dbj|BAB08633.1| disease resistance protein-like [Arabidopsis thaliana]
gi|18087526|gb|AAL58897.1| AT5g66910/MUD21_17
[Arabidopsis thaliana] gi|15240127|ref|NP_201492.1|
disease resistance protein (CC-NBS-LRR class), putative
[Arabidopsis thaliana] gi|46395983|sp|Q9FKZ0|DRL43_ARATH
Probable disease resistance protein At5g66910
Length = 815
Score = 380 bits (975), Expect = e-103
Identities = 251/688 (36%), Positives = 377/688 (54%), Gaps = 67/688 (9%)
Query: 145 VGLDIPFSKLKMELLRGGSSTLVLTGLGGLGKTTLATKLCWDEEVNA*KSKIHCWVGYTI 204
VGLD P +LK +LL +S +V++G G GKTTL TKLC D E+ KI Y++
Sbjct: 173 VGLDWPLVELKKKLL--DNSVVVVSGPPGCGKTTLVTKLCDDPEIEGEFKKIF----YSV 226
Query: 205 *QVENGASEGW*LHTCVDWFRRIGEN------------HSGYKALLG*RS*WLGLLLKKV 252
V N + FR I +N +A G R LL +
Sbjct: 227 --VSNTPN-----------FRAIVQNLLQDNGCGAITFDDDSQAETGLRD----LLEELT 269
Query: 253 EGSPLLLVLDDVWPISEPLVEKIQFQISDFKILVTSRVAFPRFSTTCILKPLAHEEAVTL 312
+ +LLVLDDVW SE L+ K Q + D+KILVTS+ F T L PL +E A +L
Sbjct: 270 KDGRILLVLDDVWQGSEFLLRKFQIDLPDYKILVTSQFDFTSLWPTYHLVPLKYEYARSL 329
Query: 313 FHHYAQMEKNSSDIINKNLVEKVVRSCQGLPLTIKVIATSLKNRPRDMWRKIGKELSQGH 372
+A ++S ++L++K+++ C G PL I+V+ SLK + +W+ + S+G
Sbjct: 330 LIQWASPPLHTSPDEYEDLLQKILKRCNGFPLVIEVVGISLKGQALYLWKGQVESWSEGE 389
Query: 373 SIL-DSNTDLLTRLQKIFDVLEDNPTIMECFMDIALFPEDHRIPVAALVDMWAELY-RLD 430
+IL ++N + RLQ F+VL+ P + ECFMD+ F +D +I + ++D+W ELY R
Sbjct: 390 TILGNANPTVRQRLQPSFNVLK--PHLKECFMDMGSFLQDQKIRASLIIDIWMELYGRGS 447
Query: 431 DNGIQAMEIINKLGIMNLANVIIPRKNASDTDNNNYNNHFIILHDILRELGIYQSAKEPF 490
+ + M +N+L NL ++ + ++ YN + H+ILREL I+QS EP
Sbjct: 448 SSTNKFMLYLNELASQNLLKLV--HLGTNKREDGFYNELLVTQHNILRELAIFQSELEPI 505
Query: 491 EQRKRLIIDINKNKSGLAEKQQGLMTCILSKFMRLCVKRNPQQLTARILSVSADETCAFD 550
QRK+L ++I ++ F C+ Q + AR+LS+ D+ +
Sbjct: 506 MQRKKLNLEIREDN-----------------FPDECLN---QPINARLLSIYTDDLFSSK 545
Query: 551 WSQMEPAQVEVLILNLHTKQYSLPEWIAKMSKLRVLIITNYDFHPSKLNNIELLGSLQNL 610
W +M+ VE L+LN+ + Y+LP +IA+M KL+VL I N+ F+P++L+N L SL NL
Sbjct: 546 WLEMDCPNVEALVLNISSLDYALPSFIAEMKKLKVLTIANHGFYPARLSNFSCLSSLPNL 605
Query: 611 ERIRLERIYVPSFGT----LKNLKKLSLYMCNTILAF-EKGSILISDAFPNLEELNIDYC 665
+RIR E++ V L +LKKLS +MC+ F + I +S A NL+E++IDYC
Sbjct: 606 KRIRFEKVSVTLLDIPQLQLGSLKKLSFFMCSFGEVFYDTEDIDVSKALSNLQEIDIDYC 665
Query: 666 KDLVVLPIGICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGK 725
DL LP I ++ LK L +TNC+KLS LP+ IG L LE+L + SC +L +P + +
Sbjct: 666 YDLDELPYWIPEVVSLKTLSITNCNKLSQLPEAIGNLSRLEVLRMCSCMNLSELPEATER 725
Query: 726 LLNLKHLDISNCISLSSLPEEFGNLCNLKNLDMASCASIELPFSVVNLQNLKTITCDEET 785
L NL+ LDIS+C+ L LP+E G L L+N+ M C+ ELP SV L+NL+ + CDE T
Sbjct: 726 LSNLRSLDISHCLGLRKLPQEIGKLQKLENISMRKCSGCELPDSVRYLENLE-VKCDEVT 784
Query: 786 AATWEDFQHMLPNMKIEVLHVDVNLNWL 813
WE + N+++ + NL L
Sbjct: 785 GLLWERLMPEMRNLRVHTEETEHNLKLL 812
>ref|NP_195056.2| disease resistance protein (CC-NBS-LRR class), putative
[Arabidopsis thaliana]
Length = 816
Score = 325 bits (832), Expect = 5e-87
Identities = 194/569 (34%), Positives = 310/569 (54%), Gaps = 35/569 (6%)
Query: 253 EGSPLLLVLDDVWPISEPLVEKIQFQISDFKILVTSRVAFPRFSTTCILKPLAHEEAVTL 312
+G+ L++LDDVW ++ L F+ LV SR T ++ L+ +EA++L
Sbjct: 273 DGARKLVILDDVWT-TQALDRLTSFKFPGCTTLVVSRSKLTEPKFTYDVEVLSEDEAISL 331
Query: 313 FHHYAQMEKNSSDIINKNLVEKVVRSCQGLPLTIKVIATSLKNRPRDMWRKIGKELSQGH 372
F A +K+ K+LV++V C+GLPL +KV SL +P W+ + + LS+G
Sbjct: 332 FCLCAFGQKSIPLGFCKDLVKQVANECKGLPLALKVTGASLNGKPEMYWKGVLQRLSKGE 391
Query: 373 SILDSNTD-LLTRLQKIFDVLEDNPTIMECFMDIALFPEDHRIPVAALVDMWAELYRLDD 431
DS+ LL +++ D L+ T +CF+D+ FPED +IP+ L+++W EL+ +D+
Sbjct: 392 PADDSHESRLLRQMEASLDNLDQ--TTKDCFLDLGAFPEDRKIPLDVLINIWIELHDIDE 449
Query: 432 NGIQAMEIINKLGIMNLANVIIPRKNASDTDNNNYNNH---FIILHDILRELGIYQSAKE 488
A I+ L NL + + Y +H F+ HD+LR+L ++ S
Sbjct: 450 GN--AFAILVDLSHKNLLTL-----GKDPRLGSLYASHYDIFVTQHDVLRDLALHLSNAG 502
Query: 489 PFEQRKRLIIDINKNKSGLAEKQQGLMTCILSKFMRLCVKRNPQQLTARILSVSADETCA 548
+RKRL++ K + L + + N + A+I+S+ E
Sbjct: 503 KVNRRKRLLMP--KRELDLPGDWE---------------RNNDEHYIAQIVSIHTGEMNE 545
Query: 549 FDWSQMEPAQVEVLILNLHTKQYSLPEWIAKMSKLRVLIITNYDFHPSKLNNIELLGSLQ 608
W ME + E+LILN + +Y LP +I+KMS+L+VL+I N P+ L++ + L
Sbjct: 546 MQWFDMEFPKAEILILNFSSDKYVLPPFISKMSRLKVLVIINNGMSPAVLHDFSIFAHLS 605
Query: 609 NLERIRLERIYVPSFGT----LKNLKKLSLYMCNTILAFEKGSILISDAFPNLEELNIDY 664
L + LER++VP LKNL K+SL +C +F++ + ++D FP L +L ID+
Sbjct: 606 KLRSLWLERVHVPQLSNSTTPLKNLHKMSLILCKINKSFDQTGLDVADIFPKLGDLTIDH 665
Query: 665 CKDLVVLPIGICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIG 724
C DLV LP IC + L L +TNC +L LP+++ KL+ LE+L L +C +L+ +P I
Sbjct: 666 CDDLVALPSSICGLTSLSCLSITNCPRLGELPKNLSKLQALEILRLYACPELKTLPGEIC 725
Query: 725 KLLNLKHLDISNCISLSSLPEEFGNLCNLKNLDMASCASIELPFSVVNLQNLKTITCDEE 784
+L LK+LDIS C+SLS LPEE G L L+ +DM C + P S V+L++L+ + CD +
Sbjct: 726 ELPGLKYLDISQCVSLSCLPEEIGKLKKLEKIDMRECCFSDRPSSAVSLKSLRHVICDTD 785
Query: 785 TAATWEDFQHMLPNMKIEVLHVDVNLNWL 813
A WE+ + +P +KIE +L+WL
Sbjct: 786 VAFMWEEVEKAVPGLKIEAAEKCFSLDWL 814
Score = 41.6 bits (96), Expect = 0.11
Identities = 20/51 (39%), Positives = 32/51 (62%)
Query: 139 ENQKFTVGLDIPFSKLKMELLRGGSSTLVLTGLGGLGKTTLATKLCWDEEV 189
+++KF VGL++ K+K + ++G+GG+GKTTLA +L D EV
Sbjct: 176 DSEKFGVGLELGKVKVKKMMFESQGGVFGISGMGGVGKTTLAKELQRDHEV 226
>emb|CAE46486.1| CC-NBS-LRR disease resistance protein [Arabidopsis thaliana]
gi|30692890|ref|NP_174620.2| disease resistance protein
(CC-NBS-LRR class), putative [Arabidopsis thaliana]
gi|46395988|sp|Q9FW44|ADR1_ARATH Disease resistance
protein ADR1 (Activated disease resistance protein 1)
Length = 787
Score = 309 bits (792), Expect = 2e-82
Identities = 205/653 (31%), Positives = 341/653 (51%), Gaps = 65/653 (9%)
Query: 168 LTGLGGLGKTTLATKLCWDEEVNA*-KSKIHCWVGYTI*QVENGASEGW*LHTCVDWFRR 226
++G+ G GKTTLA +L D++V K+K+ EN L +C+ F
Sbjct: 191 ISGMSGSGKTTLAIELSKDDDVRGLFKNKVLFLTVSRSPNFEN-------LESCIREFLY 243
Query: 227 IGENHSGYKALLG*RS*WLGLLLKKVEGSPLLLVLDDVWPISEPLVEKIQFQISDFKILV 286
G + L++LDDVW + ++++ +I LV
Sbjct: 244 DGVHQRK------------------------LVILDDVW--TRESLDRLMSKIRGSTTLV 277
Query: 287 TSRVAFPRFSTTCILKPLAHEEAVTLFHHYAQMEKNSSDIINKNLVEKVVRSCQGLPLTI 346
SR TT ++ L +EA++L A +K+ NK LV++VV C+GLPL++
Sbjct: 278 VSRSKLADPRTTYNVELLKKDEAMSLLCLCAFEQKSPPSPFNKYLVKQVVDECKGLPLSL 337
Query: 347 KVIATSLKNRPRDMWRKIGKELSQGHSILDSNTD-LLTRLQKIFDVLEDNPTIMECFMDI 405
KV+ SLKN+P W + K L +G + +++ + +++ + L+ P I +CF+D+
Sbjct: 338 KVLGASLKNKPERYWEGVVKRLLRGEAADETHESRVFAHMEESLENLD--PKIRDCFLDM 395
Query: 406 ALFPEDHRIPVAALVDMWAELYRLDDNGIQAMEIINKLGIMNLANVIIPRKNASDTDNN- 464
FPED +IP+ L +W E + +D+ A + +L NL ++ N D +
Sbjct: 396 GAFPEDKKIPLDLLTSVWVERHDIDEE--TAFSFVLRLADKNLLTIV---NNPRFGDVHI 450
Query: 465 NYNNHFIILHDILRELGIYQSAKEPFEQRKRLIIDINKNKSGLAEKQQGLMTCILSKFMR 524
Y + F+ HD+LR+L ++ S + +D+N+ + L K + ++ R
Sbjct: 451 GYYDVFVTQHDVLRDLALHMSNR----------VDVNRRERLLMPKTEPVLP-------R 493
Query: 525 LCVKRNPQQLTARILSVSADETCAFDWSQMEPAQVEVLILNLHTKQYSLPEWIAKMSKLR 584
K + A+I+S+ E +W M+ + EVLILN + Y LP +I KMS+LR
Sbjct: 494 EWEKNKDEPFDAKIVSLHTGEMDEMNWFDMDLPKAEVLILNFSSDNYVLPPFIGKMSRLR 553
Query: 585 VLIITNYDFHPSKLNNIELLGSLQNLERIRLERIYVPSFGT----LKNLKKLSLYMCNTI 640
VL+I N P++L+ + +L L + L+R++VP + LKNL K+ L C
Sbjct: 554 VLVIINNGMSPARLHGFSIFANLAKLRSLWLKRVHVPELTSCTIPLKNLHKIHLIFCKVK 613
Query: 641 LAFEKGSILISDAFPNLEELNIDYCKDLVVLPIGICDIFLLKKLRVTNCHKLSSLPQDIG 700
+F + S IS FP+L +L ID+C DL+ L I I L L +TNC ++ LP+++
Sbjct: 614 NSFVQTSFDISKIFPSLSDLTIDHCDDLLELK-SIFGITSLNSLSITNCPRILELPKNLS 672
Query: 701 KLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISLSSLPEEFGNLCNLKNLDMAS 760
+++LE L L +C +L ++P + +L LK++DIS C+SL SLPE+FG L +L+ +DM
Sbjct: 673 NVQSLERLRLYACPELISLPVEVCELPCLKYVDISQCVSLVSLPEKFGKLGSLEKIDMRE 732
Query: 761 CASIELPFSVVNLQNLKTITCDEETAATWEDFQHMLPNMKIEVLHVDVNLNWL 813
C+ + LP SV L +L+ + CDEET++ WE + ++P + IEV ++WL
Sbjct: 733 CSLLGLPSSVAALVSLRHVICDEETSSMWEMVKKVVPELCIEVAKKCFTVDWL 785
>emb|CAB86014.1| disease resistance-like protein [Arabidopsis thaliana]
gi|21281177|gb|AAM45000.1| putative disease resistance
[Arabidopsis thaliana] gi|15292721|gb|AAK92729.1|
putative disease resistance [Arabidopsis thaliana]
gi|9758447|dbj|BAB08976.1| unnamed protein product
[Arabidopsis thaliana] gi|15238281|ref|NP_196092.1|
disease resistance protein (CC-NBS-LRR class), putative
[Arabidopsis thaliana] gi|46396009|sp|Q9LZ25|DRL30_ARATH
Probable disease resistance protein At5g04720
Length = 811
Score = 305 bits (782), Expect = 3e-81
Identities = 213/679 (31%), Positives = 346/679 (50%), Gaps = 59/679 (8%)
Query: 145 VGLDIPFSKLKMELLRG--GSSTLVLTGLGGLGKTTLATKLCWDEEVNA*-KSKIHCWVG 201
VGLD+ K+K L + G + ++G+ G GKTTLA +L DEEV +K+
Sbjct: 180 VGLDLGKRKVKEMLFKSIDGERLIGISGMSGSGKTTLAKELARDEEVRGHFGNKVLFLTV 239
Query: 202 YTI*QVENGASEGW*LHTCVDWFRRIGENHSGYKALLG*RS*WLGLLLKKVEGSPLLLVL 261
+E + W T Y+A +G + S L++L
Sbjct: 240 SQSPNLEELRAHIWGFLT-------------SYEAGVG----------ATLPESRKLVIL 276
Query: 262 DDVWPISEPLVEKIQFQ-ISDFKILVTSRVAFPRFSTTCILKPLAHEEAVTLFHHYAQME 320
DDVW + ++++ F+ I LV SR T ++ L EA LF +
Sbjct: 277 DDVW--TRESLDQLMFENIPGTTTLVVSRSKLADSRVTYDVELLNEHEATALFCLSVFNQ 334
Query: 321 KNSSDIINKNLVEKVVRSCQGLPLTIKVIATSLKNRPRDMWRKIGKELSQGHSILDSNTD 380
K +++LV++VV C+GLPL++KVI SLK RP W + LS+G +++
Sbjct: 335 KLVPSGFSQSLVKQVVGECKGLPLSLKVIGASLKERPEKYWEGAVERLSRGEPADETHES 394
Query: 381 LLTRLQKIFDVLED-NPTIMECFMDIALFPEDHRIPVAALVDMWAELYRLDDNGIQAMEI 439
+ +I LE+ +P +CF+ + FPED +IP+ L+++ EL+ L+D A +
Sbjct: 395 RV--FAQIEATLENLDPKTRDCFLVLGAFPEDKKIPLDVLINVLVELHDLED--ATAFAV 450
Query: 440 INKLGIMNLANVII-PRKNASDTDNNNYNNHFIILHDILRELGIYQSAKEPFEQRKRLII 498
I L NL ++ PR T +Y + F+ HD+LR++ + S R+RL++
Sbjct: 451 IVDLANRNLLTLVKDPRFGHMYT---SYYDIFVTQHDVLRDVALRLSNHGKVNNRERLLM 507
Query: 499 DINKNKSGLAEKQQGLMTCILSKFMRLCVKRNPQQLTARILSVSADETCAFDWSQMEPAQ 558
K +S L + + + N + AR++S+ E DW ME +
Sbjct: 508 P--KRESMLPREWE---------------RNNDEPYKARVVSIHTGEMTQMDWFDMELPK 550
Query: 559 VEVLILNLHTKQYSLPEWIAKMSKLRVLIITNYDFHPSKLNNIELLGSLQNLERIRLERI 618
EVLIL+ + +Y LP +IAKM KL L+I N P++L++ + +L L+ + L+R+
Sbjct: 551 AEVLILHFSSDKYVLPPFIAKMGKLTALVIINNGMSPARLHDFSIFTNLAKLKSLWLQRV 610
Query: 619 YVPSFGT----LKNLKKLSLYMCNTILAFEKGSILISDAFPNLEELNIDYCKDLVVLPIG 674
+VP + L+NL KLSL C + ++ + I+ FP L +L ID+C DL+ LP
Sbjct: 611 HVPELSSSTVPLQNLHKLSLIFCKINTSLDQTELDIAQIFPKLSDLTIDHCDDLLELPST 670
Query: 675 ICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDI 734
IC I L + +TNC ++ LP+++ KL+ L+LL L +C +L ++P I +L LK++DI
Sbjct: 671 ICGITSLNSISITNCPRIKELPKNLSKLKALQLLRLYACHELNSLPVEICELPRLKYVDI 730
Query: 735 SNCISLSSLPEEFGNLCNLKNLDMASCASIELPFSVVNLQNLKTITCDEETAATWEDFQH 794
S C+SLSSLPE+ G + L+ +D C+ +P SVV L +L+ + CD E WE Q
Sbjct: 731 SQCVSLSSLPEKIGKVKTLEKIDTRECSLSSIPNSVVLLTSLRHVICDREALWMWEKVQK 790
Query: 795 MLPNMKIEVLHVDVNLNWL 813
+ +++E + +WL
Sbjct: 791 AVAGLRVEAAEKSFSRDWL 809
>emb|CAB80047.1| putative protein [Arabidopsis thaliana] gi|4490297|emb|CAB38788.1|
putative protein [Arabidopsis thaliana]
gi|46396028|sp|Q9SZA7|DRL29_ARATH Probable disease
resistance protein At4g33300
Length = 855
Score = 305 bits (782), Expect = 3e-81
Identities = 192/599 (32%), Positives = 311/599 (51%), Gaps = 56/599 (9%)
Query: 253 EGSPLLLVLDDVWPISEPLVEKIQFQISDFKILVTSRVAFPRFSTTCILKPLAHEEAVTL 312
+G+ L++LDDVW ++ L F+ LV SR T ++ L+ +EA++L
Sbjct: 273 DGARKLVILDDVWT-TQALDRLTSFKFPGCTTLVVSRSKLTEPKFTYDVEVLSEDEAISL 331
Query: 313 FHHYAQMEKN-----SSDIINKNLVE-----------KVVRSCQGLPLTIKVIATSLKNR 356
F A +K+ D++ ++ ++ +V C+GLPL +KV SL +
Sbjct: 332 FCLCAFGQKSIPLGFCKDLVKQHNIQSFSILRVLCLAQVANECKGLPLALKVTGASLNGK 391
Query: 357 PRDMWRKIGKELSQGHSILDSNTD-LLTRLQKIFDVLEDNPTIMECFMDIALFPEDHRIP 415
P W+ + + LS+G DS+ LL +++ D L+ T +CF+D+ FPED +IP
Sbjct: 392 PEMYWKGVLQRLSKGEPADDSHESRLLRQMEASLDNLDQ--TTKDCFLDLGAFPEDRKIP 449
Query: 416 VAALVDMWAELYRLDDNGIQAMEIINKLGIMNLANVIIPRKNASDTDNNNYNNH---FII 472
+ L+++W EL+ +D+ A I+ L NL + + Y +H F+
Sbjct: 450 LDVLINIWIELHDIDEGN--AFAILVDLSHKNLLTL-----GKDPRLGSLYASHYDIFVT 502
Query: 473 LHDILRELGIYQSAKEPFEQRKRLII---------DINKNK-----SGLAEKQQGLMTCI 518
HD+LR+L ++ S +RKRL++ D +N + + G
Sbjct: 503 QHDVLRDLALHLSNAGKVNRRKRLLMPKRELDLPGDWERNNDEHYIAQIVSIHTGKCFST 562
Query: 519 LSKFMRLCVKRNPQQLTARILSVSADETCAFDWSQMEPAQVEVLILNLHTKQYSLPEWIA 578
L F+ L + + E W ME + E+LILN + +Y LP +I+
Sbjct: 563 LQIFLVLSCNNHQDH--------NVGEMNEMQWFDMEFPKAEILILNFSSDKYVLPPFIS 614
Query: 579 KMSKLRVLIITNYDFHPSKLNNIELLGSLQNLERIRLERIYVPSFGT----LKNLKKLSL 634
KMS+L+VL+I N P+ L++ + L L + LER++VP LKNL K+SL
Sbjct: 615 KMSRLKVLVIINNGMSPAVLHDFSIFAHLSKLRSLWLERVHVPQLSNSTTPLKNLHKMSL 674
Query: 635 YMCNTILAFEKGSILISDAFPNLEELNIDYCKDLVVLPIGICDIFLLKKLRVTNCHKLSS 694
+C +F++ + ++D FP L +L ID+C DLV LP IC + L L +TNC +L
Sbjct: 675 ILCKINKSFDQTGLDVADIFPKLGDLTIDHCDDLVALPSSICGLTSLSCLSITNCPRLGE 734
Query: 695 LPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISLSSLPEEFGNLCNLK 754
LP+++ KL+ LE+L L +C +L+ +P I +L LK+LDIS C+SLS LPEE G L L+
Sbjct: 735 LPKNLSKLQALEILRLYACPELKTLPGEICELPGLKYLDISQCVSLSCLPEEIGKLKKLE 794
Query: 755 NLDMASCASIELPFSVVNLQNLKTITCDEETAATWEDFQHMLPNMKIEVLHVDVNLNWL 813
+DM C + P S V+L++L+ + CD + A WE+ + +P +KIE +L+WL
Sbjct: 795 KIDMRECCFSDRPSSAVSLKSLRHVICDTDVAFMWEEVEKAVPGLKIEAAEKCFSLDWL 853
Score = 41.6 bits (96), Expect = 0.11
Identities = 20/51 (39%), Positives = 32/51 (62%)
Query: 139 ENQKFTVGLDIPFSKLKMELLRGGSSTLVLTGLGGLGKTTLATKLCWDEEV 189
+++KF VGL++ K+K + ++G+GG+GKTTLA +L D EV
Sbjct: 176 DSEKFGVGLELGKVKVKKMMFESQGGVFGISGMGGVGKTTLAKELQRDHEV 226
>gb|AAG51211.1| disease resistance protein, putative; 92850-95636 [Arabidopsis
thaliana] gi|10998939|gb|AAG26078.1| disease resistance
protein, putative [Arabidopsis thaliana]
Length = 797
Score = 303 bits (775), Expect = 2e-80
Identities = 205/663 (30%), Positives = 342/663 (50%), Gaps = 75/663 (11%)
Query: 168 LTGLGGLGKTTLATKLCWDEEVNA*-KSKIHCWVGYTI*QVENGASEGW*LHTCVDWFRR 226
++G+ G GKTTLA +L D++V K+K+ EN L +C+ F
Sbjct: 191 ISGMSGSGKTTLAIELSKDDDVRGLFKNKVLFLTVSRSPNFEN-------LESCIREFLY 243
Query: 227 IGENHSGYKALLG*RS*WLGLLLKKVEGSPLLLVLDDVWPISEPLVEKIQFQISDFKILV 286
G + L++LDDVW + ++++ +I LV
Sbjct: 244 DGVHQRK------------------------LVILDDVW--TRESLDRLMSKIRGSTTLV 277
Query: 287 TSRVAFPRFSTTCILKPLAHEEAVTLFHHYAQMEKNSSDIINKNLVEKVVRSCQGLPLTI 346
SR TT ++ L +EA++L A +K+ NK LV++VV C+GLPL++
Sbjct: 278 VSRSKLADPRTTYNVELLKKDEAMSLLCLCAFEQKSPPSPFNKYLVKQVVDECKGLPLSL 337
Query: 347 KVIATSLKNRPRDMWRKIGKELSQGHSILDSNTD-LLTRLQKIFDVLEDNPTIMECFMDI 405
KV+ SLKN+P W + K L +G + +++ + +++ + L+ P I +CF+D+
Sbjct: 338 KVLGASLKNKPERYWEGVVKRLLRGEAADETHESRVFAHMEESLENLD--PKIRDCFLDM 395
Query: 406 ALFPEDHRIPVAALVDMWAELYRLDDNGIQAMEIINKLGIMNLANVIIPRKNASDTDNN- 464
FPED +IP+ L +W E + +D+ A + +L NL ++ N D +
Sbjct: 396 GAFPEDKKIPLDLLTSVWVERHDIDEE--TAFSFVLRLADKNLLTIV---NNPRFGDVHI 450
Query: 465 NYNNHFIILHDILRELGIYQSAKEPFEQRKRLIIDINKNKSGLAEKQQGLMTCILSKFMR 524
Y + F+ HD+LR+L ++ S + +D+N+ + L K + ++ R
Sbjct: 451 GYYDVFVTQHDVLRDLALHMSNR----------VDVNRRERLLMPKTEPVLP-------R 493
Query: 525 LCVKRNPQQLTARILSVSADETCA----------FDWSQMEPAQVEVLILNLHTKQYSLP 574
K + A+I+S+ +T +W M+ + EVLILN + Y LP
Sbjct: 494 EWEKNKDEPFDAKIVSLHTGKTSLTLNEFGEMDEMNWFDMDLPKAEVLILNFSSDNYVLP 553
Query: 575 EWIAKMSKLRVLIITNYDFHPSKLNNIELLGSLQNLERIRLERIYVPSFGT----LKNLK 630
+I KMS+LRVL+I N P++L+ + +L L + L+R++VP + LKNL
Sbjct: 554 PFIGKMSRLRVLVIINNGMSPARLHGFSIFANLAKLRSLWLKRVHVPELTSCTIPLKNLH 613
Query: 631 KLSLYMCNTILAFEKGSILISDAFPNLEELNIDYCKDLVVLPIGICDIFLLKKLRVTNCH 690
K+ L C +F + S IS FP+L +L ID+C DL+ L I I L L +TNC
Sbjct: 614 KIHLIFCKVKNSFVQTSFDISKIFPSLSDLTIDHCDDLLELK-SIFGITSLNSLSITNCP 672
Query: 691 KLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISLSSLPEEFGNL 750
++ LP+++ +++LE L L +C +L ++P + +L LK++DIS C+SL SLPE+FG L
Sbjct: 673 RILELPKNLSNVQSLERLRLYACPELISLPVEVCELPCLKYVDISQCVSLVSLPEKFGKL 732
Query: 751 CNLKNLDMASCASIELPFSVVNLQNLKTITCDEETAATWEDFQHMLPNMKIEVLHVDVNL 810
+L+ +DM C+ + LP SV L +L+ + CDEET++ WE + ++P + IEV +
Sbjct: 733 GSLEKIDMRECSLLGLPSSVAALVSLRHVICDEETSSMWEMVKKVVPELCIEVAKKCFTV 792
Query: 811 NWL 813
+WL
Sbjct: 793 DWL 795
>dbj|BAA97162.1| disease resistance protein-like [Arabidopsis thaliana]
gi|15238054|ref|NP_199539.1| disease resistance protein
(NBS-LRR class), putative [Arabidopsis thaliana]
gi|46396005|sp|Q9LVT1|DRL39_ARATH Putative disease
resistance protein At5g47280
Length = 623
Score = 300 bits (768), Expect = 1e-79
Identities = 205/652 (31%), Positives = 341/652 (51%), Gaps = 52/652 (7%)
Query: 168 LTGLGGLGKTTLATKLCWDEEVNA*KSKIHCWVGYTI*QVENGASEGW*LHTCVDWFRRI 227
++G+ G GKT LA +L DEEV G+ +V L V +
Sbjct: 14 ISGMIGSGKTILAKELARDEEVR----------GHFANRV---------LFLTVSQSPNL 54
Query: 228 GENHSGYKALLG*RS*WLGLLLKKVEG-SPLLLVLDDVWPISEPLVEKIQFQISDFKILV 286
E S + L G L + G + L++LDDV + ++++ F I LV
Sbjct: 55 EELRSLIRDFLTGHEAGFGTALPESVGHTRKLVILDDVR--TRESLDQLMFNIPGTTTLV 112
Query: 287 TSRVAFPRFSTTCILKPLAHEEAVTLFHHYAQMEKNSSDIINKNLVEKVVRSCQGLPLTI 346
S+ TT ++ L +A +LF A +K+ +K+LV++VV +GLPL++
Sbjct: 113 VSQSKLVDPRTTYDVELLNEHDATSLFCLSAFNQKSVPSGFSKSLVKQVVGESKGLPLSL 172
Query: 347 KVIATSLKNRPRDMWRKIGKELSQGHSILDSN-TDLLTRLQKIFDVLEDNPTIMECFMDI 405
KV+ SL +RP W + LS+G + +++ + + +++ + L+ P ECF+D+
Sbjct: 173 KVLGASLNDRPETYWAIAVERLSRGEPVDETHESKVFAQIEATLENLD--PKTKECFLDM 230
Query: 406 ALFPEDHRIPVAALVDMWAELYRLDDNGIQAMEIINKLGIMNLANVII-PRKNASDTDNN 464
FPE +IPV L++M +++ L+D A +++ L NL ++ P A T
Sbjct: 231 GAFPEGKKIPVDVLINMLVKIHDLEDAA--AFDVLVDLANRNLLTLVKDPTFVAMGT--- 285
Query: 465 NYNNHFIILHDILRELGIYQSAKEPFEQRKRLIIDINKNKSGLAEKQQGLMTCILSKFMR 524
+Y + F+ HD+LR++ ++ + + +R RL++ + T + S++ R
Sbjct: 286 SYYDIFVTQHDVLRDVALHLTNRGKVSRRDRLLMPKRE-------------TMLPSEWER 332
Query: 525 LCVKRNPQQLTARILSVSADETCAFDWSQMEPAQVEVLILNLHTKQYSLPEWIAKMSKLR 584
N + AR++S+ E DW M+ + EVLI+N + Y LP +IAKM LR
Sbjct: 333 ----SNDEPYNARVVSIHTGEMTEMDWFDMDFPKAEVLIVNFSSDNYVLPPFIAKMGMLR 388
Query: 585 VLIITNYDFHPSKLNNIELLGSLQNLERIRLERIYVPSFGT----LKNLKKLSLYMCNTI 640
V +I N P+ L++ + SL NL + LER++VP + LKNL KL L +C
Sbjct: 389 VFVIINNGTSPAHLHDFPIPTSLTNLRSLWLERVHVPELSSSMIPLKNLHKLYLIICKIN 448
Query: 641 LAFEKGSILISDAFPNLEELNIDYCKDLVVLPIGICDIFLLKKLRVTNCHKLSSLPQDIG 700
+F++ +I I+ FP L ++ IDYC DL LP IC I L + +TNC + LP++I
Sbjct: 449 NSFDQTAIDIAQIFPKLTDITIDYCDDLAELPSTICGITSLNSISITNCPNIKELPKNIS 508
Query: 701 KLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISLSSLPEEFGNLCNLKNLDMAS 760
KL+ L+LL L +C +L+++P I +L L ++DIS+C+SLSSLPE+ GN+ L+ +DM
Sbjct: 509 KLQALQLLRLYACPELKSLPVEICELPRLVYVDISHCLSLSSLPEKIGNVRTLEKIDMRE 568
Query: 761 CASIELPFSVVNLQNLKTITCDEETAATWEDFQHMLPNMKIEVLHVDVNLNW 812
C+ +P S V+L +L +TC E W++ + +P ++IE N+ W
Sbjct: 569 CSLSSIPSSAVSLTSLCYVTCYREALWMWKEVEKAVPGLRIEATEKWFNMTW 620
>dbj|BAB08631.1| unnamed protein product [Arabidopsis thaliana]
gi|15240125|ref|NP_201490.1| disease resistance protein
(CC-NBS-LRR class), putative [Arabidopsis thaliana]
gi|46395985|sp|Q9FKZ2|DRL41_ARATH Probable disease
resistance protein At5g66890
Length = 415
Score = 253 bits (647), Expect = 1e-65
Identities = 159/433 (36%), Positives = 239/433 (54%), Gaps = 37/433 (8%)
Query: 389 FDVLEDNPTIMECFMDIALFPEDHRIPVAALVDMWAELYRLDDNGIQAMEIINKLGIMNL 448
FD L N + ECF+D+A F ED RI + ++D+W+ Y G + M + L NL
Sbjct: 9 FDALPHN--LRECFLDMASFLEDQRIIASTIIDLWSASY-----GKEGMNNLQDLASRNL 61
Query: 449 ANVIIPRKNASDTDNNNYNNHFIILHDILRELGIYQSAKEPFE--QRKRLIIDINKNKSG 506
++ +N + ++ YN + ++LRE I Q KE +RKRL ++I NK
Sbjct: 62 LKLLPIGRN--EYEDGFYNELLVKQDNVLREFAINQCLKESSSIFERKRLNLEIQDNK-- 117
Query: 507 LAEKQQGLMTCILSKFMRLCVK-RNPQQLTARILSVSADETCAFDWSQMEPAQVEVLILN 565
F C+ + P + A + S+S D++ A W +M+ VE L+LN
Sbjct: 118 ---------------FPNWCLNPKQPIVINASLFSISTDDSFASSWFEMDCPNVEALVLN 162
Query: 566 LHTKQYSLPEWIAKMSKLRVLIITNYDFHPSKLNNIELLGSLQNLERIRLERIYV----- 620
+ + Y+LP +IA M +L+V+II N+ P+KL N+ L SL NL+RIR E++ +
Sbjct: 163 ISSSNYALPNFIATMKELKVVIIINHGLEPAKLTNLSCLSSLPNLKRIRFEKVSISLLDI 222
Query: 621 PSFGTLKNLKKLSLYMCNTILAFEKGSILISDAFPNLEELNIDYCKDLVVLPIGICDIFL 680
P G LK+L+KLSL+ C+ + A + +S+ +L+E+ IDYC +L LP I +
Sbjct: 223 PKLG-LKSLEKLSLWFCHVVDALNELED-VSETLQSLQEIEIDYCYNLDELPYWISQVVS 280
Query: 681 LKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISL 740
LKKL VTNC+KL + + IG L +LE L LSSC L +P +I +L NL+ LD+S L
Sbjct: 281 LKKLSVTNCNKLCRVIEAIGDLRDLETLRLSSCASLLELPETIDRLDNLRFLDVSGGFQL 340
Query: 741 SSLPEEFGNLCNLKNLDMASCASIELPFSVVNLQNLKTITCDEETAATWEDFQHMLPNMK 800
+LP E G L L+ + M C ELP SV NL+NL+ + CDE+TA W+ + + N+
Sbjct: 341 KNLPLEIGKLKKLEKISMKDCYRCELPDSVKNLENLE-VKCDEDTAFLWKILKPEMKNLT 399
Query: 801 IEVLHVDVNLNWL 813
I + NLN L
Sbjct: 400 ITEEKTEHNLNLL 412
>dbj|BAD94804.1| putative protein [Arabidopsis thaliana]
Length = 366
Score = 236 bits (602), Expect = 2e-60
Identities = 127/348 (36%), Positives = 198/348 (56%), Gaps = 21/348 (6%)
Query: 470 FIILHDILRELGIYQSAKEPFEQRKRLIIDINKNKSGLAEKQQGLMTCILSKFMRLCVKR 529
F+ HD+LR+L ++ S +RKRL++ K + L + +
Sbjct: 34 FVTQHDVLRDLALHLSNAGKVNRRKRLLMP--KRELDLPGDWE---------------RN 76
Query: 530 NPQQLTARILSVSADETCAFDWSQMEPAQVEVLILNLHTKQYSLPEWIAKMSKLRVLIIT 589
N + A+I+S+ E W ME + E+LILN + +Y LP +I+KMS+L+VL+I
Sbjct: 77 NDEHYIAQIVSIHTGEMNEMQWFDMEFPKAEILILNFSSDKYVLPPFISKMSRLKVLVII 136
Query: 590 NYDFHPSKLNNIELLGSLQNLERIRLERIYVPSFGT----LKNLKKLSLYMCNTILAFEK 645
N P+ L++ + L L + LER++VP LKNL K+SL +C +F++
Sbjct: 137 NNGMSPAVLHDFSIFAHLSKLRSLWLERVHVPQLSNSTTPLKNLHKMSLILCKINKSFDQ 196
Query: 646 GSILISDAFPNLEELNIDYCKDLVVLPIGICDIFLLKKLRVTNCHKLSSLPQDIGKLENL 705
+ ++D FP L +L ID+C DLV LP IC + L L +TNC +L LP+++ KL+ L
Sbjct: 197 TGLDVADIFPKLGDLTIDHCDDLVALPSSICGLTSLSCLSITNCPRLGELPKNLSKLQAL 256
Query: 706 ELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISLSSLPEEFGNLCNLKNLDMASCASIE 765
E+L L +C +L+ +P I +L LK+LDIS C+SLS LPEE G L L+ +DM C +
Sbjct: 257 EILRLYACPELKTLPGEICELPGLKYLDISQCVSLSCLPEEIGKLKKLEKIDMRECCFSD 316
Query: 766 LPFSVVNLQNLKTITCDEETAATWEDFQHMLPNMKIEVLHVDVNLNWL 813
P S V+L++L+ + CD + A WE+ + +P +KIE +L+WL
Sbjct: 317 RPSSAVSLKSLRHVICDTDVAFMWEEVEKAVPGLKIEAAEKCFSLDWL 364
>gb|AAM28916.1| NBS/LRR [Pinus taeda]
Length = 479
Score = 225 bits (573), Expect = 5e-57
Identities = 154/500 (30%), Positives = 260/500 (51%), Gaps = 42/500 (8%)
Query: 329 KNLVEKVVRSCQGLPLTIKVIATSLKNRPRDMWRKIGKELSQGHSILDSNTD-LLTRLQK 387
++LV++V C+GLPL +KVI +SL+ + R +W ++LS+ SI + + LL RL+
Sbjct: 5 EDLVKQVAAECKGLPLALKVIGSSLRGKRRPIWINAERKLSKSESISEYYKESLLKRLET 64
Query: 388 IFDVLEDNPTIMECFMDIALFPEDHRIPVAALVDMWAELYRLDDNGIQAMEIINKLGIMN 447
DVL+D +CF+D+ FP+ + V L+D+W + +++ A E++ +L N
Sbjct: 65 SIDVLDDKHK--QCFLDLGAFPKGRKFSVETLLDIWVYVRQME--WTDAFEVLLELASRN 120
Query: 448 LANVI-IPRKNASDTDNNNYNNHFIILHDILRELGIYQSAKEPFEQRKRLIIDINKNKSG 506
L N+ P A D + + HD++R+L +Y ++++ KRL ++K
Sbjct: 121 LLNLTGYPGSGA--IDYSCASELTFSQHDVMRDLALYLASQDNIISPKRLFTPSKEDK-- 176
Query: 507 LAEKQQGLMTCILSKFMRLCVKRNPQQLTARILSVSADETCAFDWSQMEPAQVEVLILNL 566
+ T LS Q A+ +S+ DW Q++ +VE L L
Sbjct: 177 -------IPTEWLSTL-------KDQASRAQFVSIYTGAMQEQDWCQIDFPEVEALALFF 222
Query: 567 HTKQYSLPEWIAKMSKLRVLIITNYDFHPSKLNNIELLGSLQNLERIRLERIYVPSF--- 623
QY LP ++ + KL+VLI+ NY + + + S + + L ++ VP+
Sbjct: 223 SANQYCLPTFLHRTPKLKVLIVYNYSSMRANIIGLPRFSSPIQIRSVFLHKLIVPASLYE 282
Query: 624 --GTLKNLKKLSLYMCNTILAFEKGSILISDA------FPNLEELNIDYCKDLVVLPIGI 675
+ + L+KLS+ +C + G+ + D FPN+ E+NID+C DL LP+ +
Sbjct: 283 NCRSWERLEKLSVCLCEGL-----GNSSLVDMELEPLNFPNITEINIDHCSDLGELPLKL 337
Query: 676 CDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDIS 735
C++ L++L VTNCH + +LP D+G+L++L +L LS+C L +P SI KL L++LDIS
Sbjct: 338 CNLTSLQRLSVTNCHLIQNLPDDMGRLKSLRVLRLSACPSLSRLPPSICKLGQLEYLDIS 397
Query: 736 NCISLSSLPEEFGNLCNLKNLDMASCASIELPFSVV--NLQNLKTITCDEETAATWEDFQ 793
C L LP EF L NL+ LDM C+ ++ +V+ +L+ + D+E A
Sbjct: 398 LCRCLQDLPSEFDQLSNLETLDMRECSGLKKVPTVIQSSLKRVVISDSDKEYEAWXSIKA 457
Query: 794 HMLPNMKIEVLHVDVNLNWL 813
L + I+V+ +L WL
Sbjct: 458 STLHTLTIDVVPEIFSLAWL 477
>emb|CAD45029.1| NBS-LRR disease resistance protein homologue [Hordeum vulgare]
Length = 1262
Score = 152 bits (385), Expect = 3e-35
Identities = 192/808 (23%), Positives = 347/808 (42%), Gaps = 81/808 (10%)
Query: 18 DVLNKISQILQKAQKNQSSRRSLRLILKDLSPVVQDIKQYNDHLDHPREEINSLIEENDA 77
D L + +L+ A++ +RL L L DI D E + +
Sbjct: 43 DTLESMEAVLKDAERRSVKEELVRLWLNRLKHAAYDISYMLDEFQANSEPASRKM----I 98
Query: 78 GESACTCSSENDSYGYVVEE--NQSLTVNDVEETLYKTREILELLN-HEFNEDGQLFKRP 134
G+ C + + Y +++ Q + + E+ T L+N H+ + +
Sbjct: 99 GKLDCFAIAPKITLAYKMKKMRGQLRKIKEDHESFKFTHANSSLINVHQLPDPRETSS-- 156
Query: 135 FNVPENQKFTVGLDIPFSKLKMELL---------RGGSSTLVLTGLGGLGKTTLATKLCW 185
NV E+ L I K +M +L + + L + GLGG+GKTTLA +
Sbjct: 157 -NVVES------LIIGREKDRMNVLSLLSTSNNIKEDFTVLPICGLGGIGKTTLAQLVFN 209
Query: 186 DEEVNA*KSKIHCWVGYTI*QVENGASEGW*LHTCVDWFRRIGENHSGYKALLG*RS*WL 245
D + N WV + QV + G ++ + G HS + +
Sbjct: 210 DAQFN---DYHRVWVYVS--QVFDLNKIG---NSIISQVSGKGSEHSHTLQHISKQ---- 257
Query: 246 GLLLKKVEGSPLLLVLDDVWPISEPLVEKIQFQIS---DFKILVTSR-VAFPRFSTTC-- 299
L ++ L+VLDD+W +++++ ++ K+LVT+R + R
Sbjct: 258 --LKDLLQDKKTLIVLDDLWETGYFQLDQLKLMLNVSTKMKVLVTTRSIDIARKMGNVGV 315
Query: 300 ---ILKPLAHEEAVTLFHHYAQMEKNSSDIINKNLVEKVVRSCQGLPLTIKVIATSLKNR 356
+L PL ++ + ++ + + +K+ R C GLPL + + L
Sbjct: 316 EPYMLDPLDNDMCWRIIKQSSRFQSRPDKEQLEPNGQKIARKCGGLPLAAQALGFLLSGM 375
Query: 357 PRDMWRKIGKELSQGHSILDSNTDLLTRLQKIFDVLEDNPTIMECFMDIALFPEDHRIPV 416
W I +S S++ +L L+ ++ L P + CF +FP+ H I
Sbjct: 376 DLSEWEAIC--ISDIWDEPFSDSTVLPSLKLSYNTL--TPYMRLCFAYCGIFPKGHNISK 431
Query: 417 AALVDMWAELYRLD-DNGIQAMEIINKLGIMNLANVIIPRKNASDTDNNNYNNHFIILHD 475
L+ W L ++ N A+++ K L + +T N ++HD
Sbjct: 432 DYLIHQWIALGFIEPSNKFSAIQLGGKYVRQFLGMSFLHHSKLPETFGNAMFTMHDLVHD 491
Query: 476 ILR-----ELGIYQSAKEPFEQRKRLIIDINKNKSGLAEKQQ-GLMTCILSKFMRL---- 525
+ R EL ++ + + K I + +++ + MT I +R+
Sbjct: 492 LARSVITEELVVFDAEIVSDNRIKEYCIYASLTNCNISDHNKVRKMTTIFPPKLRVMHFS 551
Query: 526 -CVKRNPQ---QLTARILSVSADETCAFDWSQMEPAQVEVLILN-LHTKQYSLPEWIAKM 580
C Q R+L +S F + + Q+EVLI L +Q+ PE I ++
Sbjct: 552 DCKLHGSAFSFQKCLRVLDLSGCSIKDFASALGQLKQLEVLIAQKLQDRQF--PESITRL 609
Query: 581 SKLRVLIITNYDFHPSKLNNIELLGSLQNLERIRLERIYV--PSFGTLKNLKKLSLYMCN 638
SKL L ++ +++ L SL +L+ + V + G L+NL+ L L C
Sbjct: 610 SKLHYLNLSGSRGISEIPSSVGKLVSLVHLDLSYCTNVKVIPKALGILRNLQTLDLSWCE 669
Query: 639 TILAFEK--GSILISDAFPNLEELNIDYCKDLVVLPIGICDIFLLKKLRVTNCHKLSSLP 696
+ + + GS+ NL+ LN+ C +L LP + + ++ L +++C+KL SLP
Sbjct: 670 KLESLPESLGSV------QNLQRLNLSNCFELEALPESLGSLKDVQTLDLSSCYKLESLP 723
Query: 697 QDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISLSSLPEEFGNLCNLKNL 756
+ +G L+N++ L LS C L ++P ++G+L NL+ +D+S C L + PE FG+L NL+ L
Sbjct: 724 ESLGSLKNVQTLDLSRCYKLVSLPKNLGRLKNLRTIDLSGCKKLETFPESFGSLENLQIL 783
Query: 757 DMASCASIE-LPFSVVNLQNLKTITCDE 783
++++C +E LP S +L+NL+T+ E
Sbjct: 784 NLSNCFELESLPESFGSLKNLQTLNLVE 811
Score = 127 bits (319), Expect = 1e-27
Identities = 86/234 (36%), Positives = 133/234 (56%), Gaps = 16/234 (6%)
Query: 559 VEVLILNLHTKQYSLPEWIAKMSKLRVLIITNYDFHPSKLNNIELLGSLQNLERIRLE-- 616
++ L ++ K S+PE + ++ L+ L ++ D S L + LGSL+NL+ + L
Sbjct: 828 LQTLDFSVCHKLESVPESLGGLNNLQTLKLSVCDNLVSLLKS---LGSLKNLQTLDLSGC 884
Query: 617 ---RIYVPSFGTLKNLKKLSLYMCNTILAFEKGSILISDAFPNLEELNIDYCKDLVVLPI 673
S G+L+NL+ L+L C + + + NL+ LNI +C +LV LP
Sbjct: 885 KKLESLPESLGSLENLQILNLSNCFKLESLPESL----GRLKNLQTLNISWCTELVFLPK 940
Query: 674 GICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLD 733
+ ++ L +L ++ C KL SLP +G LENLE L+LS C LE++P S+G L NL+ LD
Sbjct: 941 NLGNLKNLPRLDLSGCMKLESLPDSLGSLENLETLNLSKCFKLESLPESLGGLQNLQTLD 1000
Query: 734 ISNCISLSSLPEEFGNLCNLKNLDMASCASIE-LPFSVVNLQNLKTIT---CDE 783
+ C L SLPE G L NL+ L ++ C +E LP S+ L+NL+T+T CD+
Sbjct: 1001 LLVCHKLESLPESLGGLKNLQTLQLSFCHKLESLPESLGGLKNLQTLTLSVCDK 1054
Score = 126 bits (316), Expect = 3e-27
Identities = 84/228 (36%), Positives = 130/228 (56%), Gaps = 15/228 (6%)
Query: 559 VEVLILNLHTKQYSLPEWIAKMSKLRVLIITNYDFHPSKLNNI-ELLGSLQNLERIRLER 617
+E L L+ K SLPE + + L+ L + KL ++ E LG L+NL+ ++L
Sbjct: 972 LETLNLSKCFKLESLPESLGGLQNLQTLDLLVCH----KLESLPESLGGLKNLQTLQLSF 1027
Query: 618 IYV-----PSFGTLKNLKKLSLYMCNTILAFEKGSILISDAFPNLEELNIDYCKDLVVLP 672
+ S G LKNL+ L+L +C+ + + + + NL L + C L LP
Sbjct: 1028 CHKLESLPESLGGLKNLQTLTLSVCDKLESLPESL----GSLKNLHTLKLQVCYKLKSLP 1083
Query: 673 IGICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHL 732
+ I L L ++ CH L S+P+ +G LENL++L+LS+C LE+IP S+G L NL+ L
Sbjct: 1084 ESLGSIKNLHTLNLSVCHNLESIPESVGSLENLQILNLSNCFKLESIPKSLGSLKNLQTL 1143
Query: 733 DISNCISLSSLPEEFGNLCNLKNLDMASCASIE-LPFSVVNLQNLKTI 779
+S C L SLP+ GNL NL+ LD++ C +E LP S+ +L+NL+T+
Sbjct: 1144 ILSWCTRLVSLPKNLGNLKNLQTLDLSGCKKLESLPDSLGSLENLQTL 1191
Score = 123 bits (308), Expect = 3e-26
Identities = 80/228 (35%), Positives = 134/228 (58%), Gaps = 15/228 (6%)
Query: 559 VEVLILNLHTKQYSLPEWIAKMSKLRVLIITNYDFHPSKLNNI-ELLGSLQNLERIRLER 617
++ L L+ K SLPE + + L+ L ++ D KL ++ E LGSL+NL ++L+
Sbjct: 1020 LQTLQLSFCHKLESLPESLGGLKNLQTLTLSVCD----KLESLPESLGSLKNLHTLKLQV 1075
Query: 618 IYV-----PSFGTLKNLKKLSLYMCNTILAFEKGSILISDAFPNLEELNIDYCKDLVVLP 672
Y S G++KNL L+L +C+ + + + + NL+ LN+ C L +P
Sbjct: 1076 CYKLKSLPESLGSIKNLHTLNLSVCHNLESIPESV----GSLENLQILNLSNCFKLESIP 1131
Query: 673 IGICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHL 732
+ + L+ L ++ C +L SLP+++G L+NL+ L LS C LE++P S+G L NL+ L
Sbjct: 1132 KSLGSLKNLQTLILSWCTRLVSLPKNLGNLKNLQTLDLSGCKKLESLPDSLGSLENLQTL 1191
Query: 733 DISNCISLSSLPEEFGNLCNLKNLDMASCASIE-LPFSVVNLQNLKTI 779
++SNC L SLPE G+L L+ L++ C +E LP S+ +L++L+T+
Sbjct: 1192 NLSNCFKLESLPEILGSLKKLQTLNLFRCGKLESLPESLGSLKHLQTL 1239
Score = 119 bits (298), Expect = 4e-25
Identities = 85/230 (36%), Positives = 130/230 (55%), Gaps = 19/230 (8%)
Query: 559 VEVLILNLHTKQYSLPEWIAKMSKLRVLIITNYDFHPSKLNNI-ELLGSLQNLERIRL-- 615
++ L L+ K SLPE + + L++L ++N KL ++ E LG L+NL+ + +
Sbjct: 876 LQTLDLSGCKKLESLPESLGSLENLQILNLSNC----FKLESLPESLGRLKNLQTLNISW 931
Query: 616 --ERIYVP-SFGTLKNLKKLSLYMCNTI--LAFEKGSILISDAFPNLEELNIDYCKDLVV 670
E +++P + G LKNL +L L C + L GS+ NLE LN+ C L
Sbjct: 932 CTELVFLPKNLGNLKNLPRLDLSGCMKLESLPDSLGSL------ENLETLNLSKCFKLES 985
Query: 671 LPIGICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLK 730
LP + + L+ L + CHKL SLP+ +G L+NL+ L LS C LE++P S+G L NL+
Sbjct: 986 LPESLGGLQNLQTLDLLVCHKLESLPESLGGLKNLQTLQLSFCHKLESLPESLGGLKNLQ 1045
Query: 731 HLDISNCISLSSLPEEFGNLCNLKNLDMASCASIE-LPFSVVNLQNLKTI 779
L +S C L SLPE G+L NL L + C ++ LP S+ +++NL T+
Sbjct: 1046 TLTLSVCDKLESLPESLGSLKNLHTLKLQVCYKLKSLPESLGSIKNLHTL 1095
Score = 117 bits (294), Expect = 1e-24
Identities = 85/228 (37%), Positives = 128/228 (55%), Gaps = 15/228 (6%)
Query: 559 VEVLILNLHTKQYSLPEWIAKMSKLRVLIITNYDFHPSKLNNIELLGSLQNLERIRLE-- 616
V+ L L+ K SLPE + + ++ L ++ S N LG L+NL I L
Sbjct: 708 VQTLDLSSCYKLESLPESLGSLKNVQTLDLSRCYKLVSLPKN---LGRLKNLRTIDLSGC 764
Query: 617 ---RIYVPSFGTLKNLKKLSLYMCNTILAFEKGSILIS-DAFPNLEELNIDYCKDLVVLP 672
+ SFG+L+NL+ L+L C FE S+ S + NL+ LN+ CK L LP
Sbjct: 765 KKLETFPESFGSLENLQILNLSNC-----FELESLPESFGSLKNLQTLNLVECKKLESLP 819
Query: 673 IGICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHL 732
+ + L+ L + CHKL S+P+ +G L NL+ L LS C +L ++ S+G L NL+ L
Sbjct: 820 ESLGGLKNLQTLDFSVCHKLESVPESLGGLNNLQTLKLSVCDNLVSLLKSLGSLKNLQTL 879
Query: 733 DISNCISLSSLPEEFGNLCNLKNLDMASCASIE-LPFSVVNLQNLKTI 779
D+S C L SLPE G+L NL+ L++++C +E LP S+ L+NL+T+
Sbjct: 880 DLSGCKKLESLPESLGSLENLQILNLSNCFKLESLPESLGRLKNLQTL 927
Score = 117 bits (292), Expect = 2e-24
Identities = 80/230 (34%), Positives = 130/230 (55%), Gaps = 19/230 (8%)
Query: 559 VEVLILNLHTKQYSLPEWIAKMSKLRVLIITNYDFHPSKLNNI-ELLGSLQNLERIRLE- 616
++ L L + K SLPE + + L+ L ++ KL ++ E LG L+NL+ + L
Sbjct: 996 LQTLDLLVCHKLESLPESLGGLKNLQTLQLS----FCHKLESLPESLGGLKNLQTLTLSV 1051
Query: 617 ----RIYVPSFGTLKNLKKLSLYMCNTILAFEK--GSILISDAFPNLEELNIDYCKDLVV 670
S G+LKNL L L +C + + + GSI NL LN+ C +L
Sbjct: 1052 CDKLESLPESLGSLKNLHTLKLQVCYKLKSLPESLGSI------KNLHTLNLSVCHNLES 1105
Query: 671 LPIGICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLK 730
+P + + L+ L ++NC KL S+P+ +G L+NL+ L LS CT L ++P ++G L NL+
Sbjct: 1106 IPESVGSLENLQILNLSNCFKLESIPKSLGSLKNLQTLILSWCTRLVSLPKNLGNLKNLQ 1165
Query: 731 HLDISNCISLSSLPEEFGNLCNLKNLDMASCASIE-LPFSVVNLQNLKTI 779
LD+S C L SLP+ G+L NL+ L++++C +E LP + +L+ L+T+
Sbjct: 1166 TLDLSGCKKLESLPDSLGSLENLQTLNLSNCFKLESLPEILGSLKKLQTL 1215
Score = 117 bits (292), Expect = 2e-24
Identities = 79/227 (34%), Positives = 122/227 (52%), Gaps = 16/227 (7%)
Query: 559 VEVLILNLHTKQYSLPEWIAKMSKLRVLIITNYDFHPSKLNNI-ELLGSLQNLERIRLER 617
V+ L L+ K SLP+ + ++ LR + ++ KL E GSL+NL+ + L
Sbjct: 732 VQTLDLSRCYKLVSLPKNLGRLKNLRTIDLSGC----KKLETFPESFGSLENLQILNLSN 787
Query: 618 IYV-----PSFGTLKNLKKLSLYMCNTILAFEKGSILISDAFPNLEELNIDYCKDLVVLP 672
+ SFG+LKNL+ L+L C + + + NL+ L+ C L +P
Sbjct: 788 CFELESLPESFGSLKNLQTLNLVECKKLESLPESL----GGLKNLQTLDFSVCHKLESVP 843
Query: 673 IGICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHL 732
+ + L+ L+++ C L SL + +G L+NL+ L LS C LE++P S+G L NL+ L
Sbjct: 844 ESLGGLNNLQTLKLSVCDNLVSLLKSLGSLKNLQTLDLSGCKKLESLPESLGSLENLQIL 903
Query: 733 DISNCISLSSLPEEFGNLCNLKNLDMASCASIELPFSVVNLQNLKTI 779
++SNC L SLPE G L NL+ L+++ C EL F NL NLK +
Sbjct: 904 NLSNCFKLESLPESLGRLKNLQTLNISWCT--ELVFLPKNLGNLKNL 948
>gb|AAG60098.1| disease resistance protein, putative [Arabidopsis thaliana]
Length = 1398
Score = 128 bits (321), Expect = 9e-28
Identities = 85/219 (38%), Positives = 122/219 (54%), Gaps = 17/219 (7%)
Query: 573 LPEWIAKMSKLRVLIITNYDFHPSKLNNIELLGSLQNLERIRLER----IYVPS-FGTLK 627
LP I + L++L + S + +G+L NL+ + L + +PS G L
Sbjct: 826 LPSSIGNLISLKILYLKRIS---SLVEIPSSIGNLINLKLLNLSGCSSLVELPSSIGNLI 882
Query: 628 NLKKLSLYMCNTI--LAFEKGSILISDAFPNLEELNIDYCKDLVVLPIGICDIFLLKKLR 685
NLKKL L C+++ L G+++ NL+EL + C LV LP I ++ LK L
Sbjct: 883 NLKKLDLSGCSSLVELPLSIGNLI------NLQELYLSECSSLVELPSSIGNLINLKTLN 936
Query: 686 VTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISLSSLPE 745
++ C L LP IG L NL+ L LS C+ L +P+SIG L+NLK LD+S C SL LP
Sbjct: 937 LSECSSLVELPSSIGNLINLQELYLSECSSLVELPSSIGNLINLKKLDLSGCSSLVELPL 996
Query: 746 EFGNLCNLKNLDMASCAS-IELPFSVVNLQNLKTITCDE 783
GNL NLK L+++ C+S +ELP S+ NL NL+ + E
Sbjct: 997 SIGNLINLKTLNLSECSSLVELPSSIGNLINLQELYLSE 1035
Score = 121 bits (303), Expect = 1e-25
Identities = 75/184 (40%), Positives = 112/184 (60%), Gaps = 15/184 (8%)
Query: 604 LGSLQNLERIRLER----IYVPS-FGTLKNLKKLSLYMCNTILAFEK--GSILISDAFPN 656
+G+L NL+ + L + +PS G L NL++L L C++++ G+++ N
Sbjct: 926 IGNLINLKTLNLSECSSLVELPSSIGNLINLQELYLSECSSLVELPSSIGNLI------N 979
Query: 657 LEELNIDYCKDLVVLPIGICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDL 716
L++L++ C LV LP+ I ++ LK L ++ C L LP IG L NL+ L LS C+ L
Sbjct: 980 LKKLDLSGCSSLVELPLSIGNLINLKTLNLSECSSLVELPSSIGNLINLQELYLSECSSL 1039
Query: 717 EAIPTSIGKLLNLKHLDISNCISLSSLPEEFGNLCNLKNLDMASCAS-IELPFSVVNLQN 775
+P+SIG L+NLK LD+S C SL LP GNL NLK L+++ C+S +ELP S+ NL N
Sbjct: 1040 VELPSSIGNLINLKKLDLSGCSSLVELPLSIGNLINLKTLNLSGCSSLVELPSSIGNL-N 1098
Query: 776 LKTI 779
LK +
Sbjct: 1099 LKKL 1102
Score = 115 bits (287), Expect = 8e-24
Identities = 82/230 (35%), Positives = 121/230 (51%), Gaps = 17/230 (7%)
Query: 562 LILNLHTKQYSLPEWIAKMSKLRVLIITNYDFHPSKLNNIELLGSLQNLERIRL----ER 617
++L+ + LP I + ++ L I S L +G+L L R+ L
Sbjct: 719 MVLSDCSSLIELPSSIGNATNIKSLDIQGCS---SLLKLPSSIGNLITLPRLDLMGCSSL 775
Query: 618 IYVPS-FGTLKNLKKLSLYMCNTILAFEK--GSILISDAFPNLEELNIDYCKDLVVLPIG 674
+ +PS G L NL +L L C++++ G+++ NLE C L+ LP
Sbjct: 776 VELPSSIGNLINLPRLDLMGCSSLVELPSSIGNLI------NLEAFYFHGCSSLLELPSS 829
Query: 675 ICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDI 734
I ++ LK L + L +P IG L NL+LL+LS C+ L +P+SIG L+NLK LD+
Sbjct: 830 IGNLISLKILYLKRISSLVEIPSSIGNLINLKLLNLSGCSSLVELPSSIGNLINLKKLDL 889
Query: 735 SNCISLSSLPEEFGNLCNLKNLDMASCAS-IELPFSVVNLQNLKTITCDE 783
S C SL LP GNL NL+ L ++ C+S +ELP S+ NL NLKT+ E
Sbjct: 890 SGCSSLVELPLSIGNLINLQELYLSECSSLVELPSSIGNLINLKTLNLSE 939
Score = 114 bits (284), Expect = 2e-23
Identities = 85/251 (33%), Positives = 127/251 (49%), Gaps = 29/251 (11%)
Query: 563 ILNLHTKQYS-------LPEWIAKMSKLRVLIITNYDFHPSKLNNIELLGSLQNLERIRL 615
++NL T S LP I + L+ L ++ S + +G+L NL+++ L
Sbjct: 1001 LINLKTLNLSECSSLVELPSSIGNLINLQELYLSECS---SLVELPSSIGNLINLKKLDL 1057
Query: 616 ER----IYVP-SFGTLKNLKKLSLYMCNTILAFEKGSILISDAFPNLEELNIDYCKDLVV 670
+ +P S G L NLK L+L C++++ S NL++L++ C LV
Sbjct: 1058 SGCSSLVELPLSIGNLINLKTLNLSGCSSLVELPS-----SIGNLNLKKLDLSGCSSLVE 1112
Query: 671 LPIGICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLK 730
LP I ++ LKKL ++ C L LP IG L NL+ L LS C+ L +P+SIG L+NL+
Sbjct: 1113 LPSSIGNLINLKKLDLSGCSSLVELPLSIGNLINLQELYLSECSSLVELPSSIGNLINLQ 1172
Query: 731 HLDISNCISLSSLPEEFGNLCNLKNLDMASCASI----ELPFSVVNL-----QNLKTITC 781
L +S C SL LP GNL NLK LD+ C + +LP S+ L ++L+T+ C
Sbjct: 1173 ELYLSECSSLVELPSSIGNLINLKKLDLNKCTKLVSLPQLPDSLSVLVAESCESLETLAC 1232
Query: 782 DEETAATWEDF 792
W F
Sbjct: 1233 SFPNPQVWLKF 1243
Score = 101 bits (251), Expect = 1e-19
Identities = 124/480 (25%), Positives = 200/480 (40%), Gaps = 64/480 (13%)
Query: 308 EAVTLFHHYAQMEKNSSDIINKNLVEKVVRSCQGLPLTIKVIATSLKNRPRDMWRKIGKE 367
EA+ +F +A + D K L KV R LPL ++V+ + + ++ W+ E
Sbjct: 416 EALQIFCIHAFGHDSPDDGFEK-LATKVTRLAGNLPLGLRVMGSHFRGMSKEDWKG---E 471
Query: 368 LSQGHSILDSNTDLLTRLQKIFDVLEDNPTIMECFMDIALFPEDHRIPVAALVDMWAELY 427
L + LD + + +DVL+D + F+ IA F D I D +
Sbjct: 472 LPRLRIRLDGEIGSILKFS--YDVLDDEDK--DLFLHIACFFNDEGID-HTFEDTLRHKF 526
Query: 428 RLDDNGIQAMEIINKLGIMNLANVIIPRKNASDTDNNNYNNHFIIL-HDILRELGIYQSA 486
G+Q V++ R S+ +N + L +I+R +Y+
Sbjct: 527 SNVQRGLQ---------------VLVQRSLISEDLTQPMHNLLVQLGREIVRNQSVYEPG 571
Query: 487 KEPFEQRKRLIIDINKNKSGLAEKQQGLMTCILSKFMRLCVKRNPQQLTARILSVSADET 546
K F + I ++ + +G +E G+ + L + + + + DE
Sbjct: 572 KRQFLVDGKEICEVLTSHTG-SESVIGINFEVYWSMDELNISDRVFEGMSNLQFFRFDEN 630
Query: 547 CAFDWSQMEPAQVEVLILNLHTKQYSLPEWIAKMS-KLRVLIITNYDFHP-----SKLNN 600
LH LP+ + + KLR+L ++D++P SK N
Sbjct: 631 SYG---------------RLH-----LPQGLNYLPPKLRIL---HWDYYPMTSLPSKFNL 667
Query: 601 IELLGSLQNLERIRLERIYVPSFGTLKNLKKLSLYMCNTILAFEKGSILISDAFPNLEEL 660
L+ + L+ LE+++ L NLK + L + + S I NL E+
Sbjct: 668 KFLVKII--LKHSELEKLW-EGIQPLVNLKVMDLRYSSHLKELPNLSTAI-----NLLEM 719
Query: 661 NIDYCKDLVVLPIGICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIP 720
+ C L+ LP I + +K L + C L LP IG L L L L C+ L +P
Sbjct: 720 VLSDCSSLIELPSSIGNATNIKSLDIQGCSSLLKLPSSIGNLITLPRLDLMGCSSLVELP 779
Query: 721 TSIGKLLNLKHLDISNCISLSSLPEEFGNLCNLKNLDMASCAS-IELPFSVVNLQNLKTI 779
+SIG L+NL LD+ C SL LP GNL NL+ C+S +ELP S+ NL +LK +
Sbjct: 780 SSIGNLINLPRLDLMGCSSLVELPSSIGNLINLEAFYFHGCSSLLELPSSIGNLISLKIL 839
>ref|NP_683486.1| leucine-rich repeat family protein [Arabidopsis thaliana]
Length = 703
Score = 128 bits (321), Expect = 9e-28
Identities = 85/219 (38%), Positives = 122/219 (54%), Gaps = 17/219 (7%)
Query: 573 LPEWIAKMSKLRVLIITNYDFHPSKLNNIELLGSLQNLERIRLER----IYVPS-FGTLK 627
LP I + L++L + S + +G+L NL+ + L + +PS G L
Sbjct: 131 LPSSIGNLISLKILYLKRIS---SLVEIPSSIGNLINLKLLNLSGCSSLVELPSSIGNLI 187
Query: 628 NLKKLSLYMCNTI--LAFEKGSILISDAFPNLEELNIDYCKDLVVLPIGICDIFLLKKLR 685
NLKKL L C+++ L G+++ NL+EL + C LV LP I ++ LK L
Sbjct: 188 NLKKLDLSGCSSLVELPLSIGNLI------NLQELYLSECSSLVELPSSIGNLINLKTLN 241
Query: 686 VTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISLSSLPE 745
++ C L LP IG L NL+ L LS C+ L +P+SIG L+NLK LD+S C SL LP
Sbjct: 242 LSECSSLVELPSSIGNLINLQELYLSECSSLVELPSSIGNLINLKKLDLSGCSSLVELPL 301
Query: 746 EFGNLCNLKNLDMASCAS-IELPFSVVNLQNLKTITCDE 783
GNL NLK L+++ C+S +ELP S+ NL NL+ + E
Sbjct: 302 SIGNLINLKTLNLSECSSLVELPSSIGNLINLQELYLSE 340
Score = 121 bits (303), Expect = 1e-25
Identities = 75/184 (40%), Positives = 112/184 (60%), Gaps = 15/184 (8%)
Query: 604 LGSLQNLERIRLER----IYVPS-FGTLKNLKKLSLYMCNTILAFEK--GSILISDAFPN 656
+G+L NL+ + L + +PS G L NL++L L C++++ G+++ N
Sbjct: 231 IGNLINLKTLNLSECSSLVELPSSIGNLINLQELYLSECSSLVELPSSIGNLI------N 284
Query: 657 LEELNIDYCKDLVVLPIGICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDL 716
L++L++ C LV LP+ I ++ LK L ++ C L LP IG L NL+ L LS C+ L
Sbjct: 285 LKKLDLSGCSSLVELPLSIGNLINLKTLNLSECSSLVELPSSIGNLINLQELYLSECSSL 344
Query: 717 EAIPTSIGKLLNLKHLDISNCISLSSLPEEFGNLCNLKNLDMASCAS-IELPFSVVNLQN 775
+P+SIG L+NLK LD+S C SL LP GNL NLK L+++ C+S +ELP S+ NL N
Sbjct: 345 VELPSSIGNLINLKKLDLSGCSSLVELPLSIGNLINLKTLNLSGCSSLVELPSSIGNL-N 403
Query: 776 LKTI 779
LK +
Sbjct: 404 LKKL 407
Score = 115 bits (287), Expect = 8e-24
Identities = 82/230 (35%), Positives = 121/230 (51%), Gaps = 17/230 (7%)
Query: 562 LILNLHTKQYSLPEWIAKMSKLRVLIITNYDFHPSKLNNIELLGSLQNLERIRL----ER 617
++L+ + LP I + ++ L I S L +G+L L R+ L
Sbjct: 24 MVLSDCSSLIELPSSIGNATNIKSLDIQGCS---SLLKLPSSIGNLITLPRLDLMGCSSL 80
Query: 618 IYVPS-FGTLKNLKKLSLYMCNTILAFEK--GSILISDAFPNLEELNIDYCKDLVVLPIG 674
+ +PS G L NL +L L C++++ G+++ NLE C L+ LP
Sbjct: 81 VELPSSIGNLINLPRLDLMGCSSLVELPSSIGNLI------NLEAFYFHGCSSLLELPSS 134
Query: 675 ICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDI 734
I ++ LK L + L +P IG L NL+LL+LS C+ L +P+SIG L+NLK LD+
Sbjct: 135 IGNLISLKILYLKRISSLVEIPSSIGNLINLKLLNLSGCSSLVELPSSIGNLINLKKLDL 194
Query: 735 SNCISLSSLPEEFGNLCNLKNLDMASCAS-IELPFSVVNLQNLKTITCDE 783
S C SL LP GNL NL+ L ++ C+S +ELP S+ NL NLKT+ E
Sbjct: 195 SGCSSLVELPLSIGNLINLQELYLSECSSLVELPSSIGNLINLKTLNLSE 244
Score = 114 bits (284), Expect = 2e-23
Identities = 85/251 (33%), Positives = 127/251 (49%), Gaps = 29/251 (11%)
Query: 563 ILNLHTKQYS-------LPEWIAKMSKLRVLIITNYDFHPSKLNNIELLGSLQNLERIRL 615
++NL T S LP I + L+ L ++ S + +G+L NL+++ L
Sbjct: 306 LINLKTLNLSECSSLVELPSSIGNLINLQELYLSECS---SLVELPSSIGNLINLKKLDL 362
Query: 616 ER----IYVP-SFGTLKNLKKLSLYMCNTILAFEKGSILISDAFPNLEELNIDYCKDLVV 670
+ +P S G L NLK L+L C++++ S NL++L++ C LV
Sbjct: 363 SGCSSLVELPLSIGNLINLKTLNLSGCSSLVELPS-----SIGNLNLKKLDLSGCSSLVE 417
Query: 671 LPIGICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLK 730
LP I ++ LKKL ++ C L LP IG L NL+ L LS C+ L +P+SIG L+NL+
Sbjct: 418 LPSSIGNLINLKKLDLSGCSSLVELPLSIGNLINLQELYLSECSSLVELPSSIGNLINLQ 477
Query: 731 HLDISNCISLSSLPEEFGNLCNLKNLDMASCASI----ELPFSVVNL-----QNLKTITC 781
L +S C SL LP GNL NLK LD+ C + +LP S+ L ++L+T+ C
Sbjct: 478 ELYLSECSSLVELPSSIGNLINLKKLDLNKCTKLVSLPQLPDSLSVLVAESCESLETLAC 537
Query: 782 DEETAATWEDF 792
W F
Sbjct: 538 SFPNPQVWLKF 548
Score = 90.9 bits (224), Expect = 2e-16
Identities = 51/125 (40%), Positives = 68/125 (53%), Gaps = 1/125 (0%)
Query: 656 NLEELNIDYCKDLVVLPIGICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTD 715
NL E+ + C L+ LP I + +K L + C L LP IG L L L L C+
Sbjct: 20 NLLEMVLSDCSSLIELPSSIGNATNIKSLDIQGCSSLLKLPSSIGNLITLPRLDLMGCSS 79
Query: 716 LEAIPTSIGKLLNLKHLDISNCISLSSLPEEFGNLCNLKNLDMASCAS-IELPFSVVNLQ 774
L +P+SIG L+NL LD+ C SL LP GNL NL+ C+S +ELP S+ NL
Sbjct: 80 LVELPSSIGNLINLPRLDLMGCSSLVELPSSIGNLINLEAFYFHGCSSLLELPSSIGNLI 139
Query: 775 NLKTI 779
+LK +
Sbjct: 140 SLKIL 144
Score = 45.8 bits (107), Expect = 0.006
Identities = 25/70 (35%), Positives = 42/70 (59%), Gaps = 2/70 (2%)
Query: 708 LSLSSCTDLEAIPTSIGKLLNLKHLDISNCISLSSLPEEFGNLCNLKNLDMASCAS-IEL 766
+ L + L+ +P ++ +NL + +S+C SL LP GN N+K+LD+ C+S ++L
Sbjct: 1 MDLRYSSHLKELP-NLSTAINLLEMVLSDCSSLIELPSSIGNATNIKSLDIQGCSSLLKL 59
Query: 767 PFSVVNLQNL 776
P S+ NL L
Sbjct: 60 PSSIGNLITL 69
>gb|AAN23084.1| putative rp3 protein [Zea mays]
Length = 944
Score = 119 bits (297), Expect = 5e-25
Identities = 168/662 (25%), Positives = 279/662 (41%), Gaps = 78/662 (11%)
Query: 146 GLDIPFSKLKMELLRGGSSTLVLT--GLGGLGKTTLATKLCWDEEVNA*KSKIHCWVGYT 203
G D +++ EL+ S +++ GLGG GKTTLA KL +++ ++ WV
Sbjct: 171 GRDQAKNQIISELIETDSQQKIVSVIGLGGSGKTTLA-KLVFNDGNIIKHFEVVLWVH-- 227
Query: 204 I*QVENGASEGW*LHTCVD-WFRRIGENHSGYKALLG*RS*WLGLLLKKVEGSPLLLVLD 262
S + + V+ F+ I + S + L + K+ G L VLD
Sbjct: 228 -------VSREFAVEKLVEKLFKAIAGDMSDHPPLQHVSR----TISDKLVGKRFLAVLD 276
Query: 263 DVWPISEPLVEKIQFQIS------DFKILVTSR---VAFPRFSTTCILKP-LAHEEAVTL 312
DVW +E VE QF + IL+T+R VA S+ P L+ E++ +
Sbjct: 277 DVW--TEDRVEWEQFMVHLKSGAPGSSILLTTRSRKVAEAVDSSYAYNLPFLSKEDSWKV 334
Query: 313 FHHYAQMEKNSSDIINKNLVEKVVRSCQGLPLTIKVIATSLKN-RPRDMWRKIGKELSQG 371
F + + D +++V C G+PL IKVIA L + + WR I S
Sbjct: 335 FQQCFGIALKALDPEFLQTGKEIVEKCGGVPLAIKVIAGVLHGIKGIEEWRSICD--SNL 392
Query: 372 HSILDSNTDLLTRLQKIFDVLEDNPTIMECFMDIALFPEDHRIPVAALVDMWA------- 424
+ D + L F L D+ + CF+ ++FP + I L+ W
Sbjct: 393 LDVQDDEHRVFACLSLSFVHLPDH--LKPCFLHCSIFPRGYVINRRHLISQWIAHGFVPT 450
Query: 425 -ELYRLDDNGIQAMEIINKLGIMNLANVIIPRKNASDTDNNNYNNHFIILHDILRELGIY 483
+ + +D GI + + K+G + I + ++ ++HD+ R++
Sbjct: 451 NQARQAEDVGIGYFDSLLKVGFLQDHVQIWSTRGEVTCKMHD------LVHDLARQI--- 501
Query: 484 QSAKEPFEQRKRLIIDINKNKSGLAEKQQGLMTCILSKFMRLCVKRNPQQLTARILSVSA 543
R + +I NK + L +C +LC K + R L
Sbjct: 502 --------LRDEFVSEIETNKQIKRCRYLSLTSCTGKLDNKLCGKVRALYVCGRELE--- 550
Query: 544 DETCAFDWSQMEPAQVEVLILNLHTKQYSLPEWIAKMSKLRVLIIT--NYDFHPSKLNNI 601
FD + + V +IL T SLP +++K L L I+ N + P L+
Sbjct: 551 -----FDKTMNKQCCVRTIILKYITDD-SLPLFVSKFEYLGYLEISDVNCEALPEALSRC 604
Query: 602 ELLGSLQNLERIRLERIYVPSFGTLKNLKKLSLYMCNTILAFEKGSILISDAFPNLEELN 661
L +L L RL + S G LK L+ L L ++I + + I D NL L
Sbjct: 605 WNLQALHVLNCSRLA-VVPESIGKLKKLRTLELNGVSSIKSLPQS---IGDC-DNLRRLY 659
Query: 662 IDYCKDLVVLPIGICDIFLLKKLRVTNCHKLSSLP--QDIGKLENLELLSLSSCTDLEAI 719
++ C+ + +P + + L+ L + +C L LP GKL NL+ ++ + C +L +
Sbjct: 660 LEECRGIEDIPNSLGKLENLRILSIVDCVSLQKLPPSDSFGKLLNLQTITFNLCYNLRNL 719
Query: 720 PTSIGKLLNLKHLDISNCISLSSLPEEFGNLCNLKNLDMASCASIE-LPFSVVNLQNLKT 778
P + L++L+ +D+ C L LPE GNL NLK L++ C + LP L L+
Sbjct: 720 PQCMTSLIHLESVDLGYCFQLVELPEGMGNLRNLKVLNLKKCKKLRGLPAGCGKLTRLQQ 779
Query: 779 IT 780
++
Sbjct: 780 LS 781
>gb|AAN23092.1| putative rp3 protein [Zea mays]
Length = 1222
Score = 118 bits (296), Expect = 7e-25
Identities = 170/662 (25%), Positives = 280/662 (41%), Gaps = 78/662 (11%)
Query: 146 GLDIPFSKLKMELLRGGSSTLVLT--GLGGLGKTTLATKLCWDEEVNA*KSKIHCWVGYT 203
G D +++ EL+ S +++ GLGG GKTTLA KL +++ ++ WV
Sbjct: 171 GRDQAKNQIISELIETDSQQKIVSVIGLGGSGKTTLA-KLVFNDGNIIKHFEVVLWVH-- 227
Query: 204 I*QVENGASEGW*LHTCVD-WFRRIGENHSGYKALLG*RS*WLGLLLKKVEGSPLLLVLD 262
S + + V+ F+ I + S + L + K+ G L VLD
Sbjct: 228 -------VSREFAVEKLVEKLFKAIAGDMSDHPPLQHVSR----TISDKLVGKRFLAVLD 276
Query: 263 DVWPISEPLVEKIQFQIS------DFKILVTSR---VAFPRFSTTCILKP-LAHEEAVTL 312
DVW +E VE QF + IL+T+R VA S+ P L+ E++ +
Sbjct: 277 DVW--TEDRVEWEQFMVHLKSGAPGSSILLTTRSRKVAEAVDSSYAYNLPFLSKEDSWKV 334
Query: 313 FHHYAQMEKNSSDIINKNLVEKVVRSCQGLPLTIKVIATSLKN-RPRDMWRKIGKELSQG 371
F + + D +++V C G+PL IKVIA L + + WR I S
Sbjct: 335 FQQCFGIALKALDPEFLQTGKEIVEKCGGVPLAIKVIAGVLHGIKGIEEWRSICD--SNL 392
Query: 372 HSILDSNTDLLTRLQKIFDVLEDNPTIMECFMDIALFPEDHRIPVAALVDMWA------- 424
+ D + L F L D+ + CF+ ++FP + I L+ W
Sbjct: 393 LDVQDDEHRVFACLSLSFVHLPDH--LKPCFLHCSIFPRGYVINRRHLISQWIAHGFVPT 450
Query: 425 -ELYRLDDNGIQAMEIINKLGIMNLANVIIPRKNASDTDNNNYNNHFIILHDILRELGIY 483
+ + +D GI + + K+G + I + ++ ++HD+ R++
Sbjct: 451 NQARQAEDVGIGYFDSLLKVGFLQDHVQIWSTRGEVTCKMHD------LVHDLARQI--- 501
Query: 484 QSAKEPFEQRKRLIIDINKNKSGLAEKQQGLMTCILSKFMRLCVKRNPQQLTARILSVSA 543
R + +I NK + L +C +LC K R L V
Sbjct: 502 --------LRDEFVSEIETNKQIKRCRYLSLTSCTGKLDNKLCGK-------VRALYVCG 546
Query: 544 DETCAFDWSQMEPAQVEVLILNLHTKQYSLPEWIAKMSKLRVLIIT--NYDFHPSKLNNI 601
E FD + + V +IL T SLP +++K L L I+ N + P L+
Sbjct: 547 PEL-EFDKTMNKQCCVRTIILKYITAD-SLPLFVSKFEYLGYLEISSVNCEALPEALSRC 604
Query: 602 ELLGSLQNLERIRLERIYVPSFGTLKNLKKLSLYMCNTILAFEKGSILISDAFPNLEELN 661
L +L L+ RL + S G LK L+ L L ++I + + I D NL L
Sbjct: 605 WNLQALHVLKCSRLA-VVPESIGKLKKLRTLELNGVSSIKSLPQS---IGDC-DNLRRLY 659
Query: 662 IDYCKDLVVLPIGICDIFLLKKLRVTNCHKLSSLP--QDIGKLENLELLSLSSCTDLEAI 719
++ C + +P + + L+ L + +C L LP GKL NL+ ++ + C +L +
Sbjct: 660 LEGCHGIEDIPNSLGKLENLRILNIVHCISLQKLPPSDSFGKLLNLQTITFNLCYNLRNL 719
Query: 720 PTSIGKLLNLKHLDISNCISLSSLPEEFGNLCNLKNLDMASCASIE-LPFSVVNLQNLKT 778
P + L++L+ +D+ C L LPE GNL NLK L++ C + LP L L+
Sbjct: 720 PQCMTSLIHLESVDLGYCFQLVELPEGMGNLRNLKVLNLKKCKKLRGLPAGCGKLTRLQQ 779
Query: 779 IT 780
++
Sbjct: 780 LS 781
Score = 55.8 bits (133), Expect = 5e-06
Identities = 89/361 (24%), Positives = 150/361 (40%), Gaps = 51/361 (14%)
Query: 420 VDMWAELYRLDDNGIQAMEIINKLGIMNLANVIIPRKNASDTDNNNYNNHFIILHDILRE 479
++M EL+ LD ++ I KL I +PR A +D+ + I
Sbjct: 856 LNMEKELHLLDS--LEPPSKIEKLRIRGYRGSQLPRWMAKQSDSCGPADDTHI------- 906
Query: 480 LGIYQSAKEPFEQRKRLIIDINKNKSGLAEKQQGLMTCILSKFMRLCVKRNPQQLTARIL 539
+ Q F L++D N L E L+ L K ++L KR P+ + +L
Sbjct: 907 --VMQRNPSEFSHLTELVLDNLPNLEHLGE----LVELPLVKILKL--KRLPKLV--ELL 956
Query: 540 SVSADETCAFDWSQMEPAQVEVLILNLHTKQYSLPEWIAKMSKLRV------LIITNYDF 593
+ + E + V L++ K P + + LR+ L+ + F
Sbjct: 957 TTTTGEEGVEVLYRFH--HVSTLVIIDCPKLVVKPYFPPSLQSLRLEGNNGQLVSSGCFF 1014
Query: 594 HPSKLN--NIELLGSLQNLERIRLERIYVPSFGT-----LKNLKKLSLYMCNTILAFEKG 646
HP + + +++G+ +LER+ L R+ S G L L L ++ C T L
Sbjct: 1015 HPRHHHAAHADVIGT--HLERLELRRLTGSSSGWEVLQHLTGLHTLEIFKC-TDLTHLPE 1071
Query: 647 SILISDAFPNLEELNIDYCKDLVVLPIGICDIFLLKKLRVTNCH-----------KLSSL 695
SI F L I C +L VLP + ++ L+ L + +C L+ L
Sbjct: 1072 SIHCPTTFCRLL---ITGCHNLRVLPDWLVELKSLQSLNIDSCDALQHLTISSLTSLTCL 1128
Query: 696 PQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISLSSLPEEFGNLCNLKN 755
P+ + L +L L+L C +L +P +G+L L+ L + +C L+SLP+ L L+
Sbjct: 1129 PESMQHLTSLRTLNLCRCNELTHLPEWLGELSVLQKLWLQDCRGLTSLPQSIQRLTALEE 1188
Query: 756 L 756
L
Sbjct: 1189 L 1189
Score = 45.4 bits (106), Expect = 0.007
Identities = 30/98 (30%), Positives = 51/98 (51%), Gaps = 7/98 (7%)
Query: 626 LKNLKKLSLYMCNTILAFEKGSILISDAFP-------NLEELNIDYCKDLVVLPIGICDI 678
LK+L+ L++ C+ + S+ P +L LN+ C +L LP + ++
Sbjct: 1100 LKSLQSLNIDSCDALQHLTISSLTSLTCLPESMQHLTSLRTLNLCRCNELTHLPEWLGEL 1159
Query: 679 FLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDL 716
+L+KL + +C L+SLPQ I +L LE L +S +L
Sbjct: 1160 SVLQKLWLQDCRGLTSLPQSIQRLTALEELYISGNPNL 1197
Score = 40.8 bits (94), Expect = 0.18
Identities = 34/120 (28%), Positives = 58/120 (48%), Gaps = 15/120 (12%)
Query: 681 LKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISL 740
L++L + SS + + L L L + CTDL +P SI L I+ C +L
Sbjct: 1031 LERLELRRLTGSSSGWEVLQHLTGLHTLEIFKCTDLTHLPESIHCPTTFCRLLITGCHNL 1090
Query: 741 SSLPEEFGNLCNLKNLDMASCASIE------------LPFSVVNLQNLKTIT---CDEET 785
LP+ L +L++L++ SC +++ LP S+ +L +L+T+ C+E T
Sbjct: 1091 RVLPDWLVELKSLQSLNIDSCDALQHLTISSLTSLTCLPESMQHLTSLRTLNLCRCNELT 1150
>gb|AAN23086.1| putative rp3 protein [Zea mays]
Length = 1222
Score = 118 bits (295), Expect = 9e-25
Identities = 170/662 (25%), Positives = 279/662 (41%), Gaps = 78/662 (11%)
Query: 146 GLDIPFSKLKMELLRGGSSTLVLT--GLGGLGKTTLATKLCWDEEVNA*KSKIHCWVGYT 203
G D +++ EL+ S +++ GLGG GKTTLA KL +++ ++ WV
Sbjct: 171 GRDQAKNQIISELIETDSQQKIVSVIGLGGSGKTTLA-KLVFNDGNIIKHFEVVLWVH-- 227
Query: 204 I*QVENGASEGW*LHTCVD-WFRRIGENHSGYKALLG*RS*WLGLLLKKVEGSPLLLVLD 262
S + + V+ F+ I + S + L + K+ G L VLD
Sbjct: 228 -------VSREFAVEKLVEKLFKAIAGDMSDHPPLQHVSR----TISDKLVGKRFLAVLD 276
Query: 263 DVWPISEPLVEKIQFQIS------DFKILVTSR---VAFPRFSTTCILKP-LAHEEAVTL 312
DVW E VE QF + IL+T+R VA S+ P L+ E++ +
Sbjct: 277 DVW--IEDRVEWEQFMVHLKSGAPGSSILLTTRSRKVAEAVDSSYAYNLPFLSKEDSWKV 334
Query: 313 FHHYAQMEKNSSDIINKNLVEKVVRSCQGLPLTIKVIATSLKN-RPRDMWRKIGKELSQG 371
F + + D +++V C G+PL IKVIA L + + WR I S
Sbjct: 335 FQQCFGIALKALDPEFLQTGKEIVEKCGGVPLAIKVIAGVLHGIKGIEEWRSICD--SNL 392
Query: 372 HSILDSNTDLLTRLQKIFDVLEDNPTIMECFMDIALFPEDHRIPVAALVDMWA------- 424
+ D + L F L D+ + CF+ ++FP + I L+ W
Sbjct: 393 LDVQDDEHRVFACLSLSFVHLPDH--LKPCFLHCSIFPRGYVINRRHLISQWIAHGFVPT 450
Query: 425 -ELYRLDDNGIQAMEIINKLGIMNLANVIIPRKNASDTDNNNYNNHFIILHDILRELGIY 483
+ + +D GI + + K+G + I + ++ ++HD+ R++
Sbjct: 451 NQARQAEDVGIGYFDSLLKVGFLQDHVQIWSTRGEVTCKMHD------LVHDLARQI--- 501
Query: 484 QSAKEPFEQRKRLIIDINKNKSGLAEKQQGLMTCILSKFMRLCVKRNPQQLTARILSVSA 543
R + +I NK + L +C +LC K R L V
Sbjct: 502 --------LRDEFVSEIETNKQIKRCRYLSLTSCTGKLDNKLCGK-------VRALYVCG 546
Query: 544 DETCAFDWSQMEPAQVEVLILNLHTKQYSLPEWIAKMSKLRVLIIT--NYDFHPSKLNNI 601
E FD + + V +IL T SLP +++K L L I+ N + P L+
Sbjct: 547 PEL-EFDKTMNKQCCVRTIILKYITAD-SLPLFVSKFEYLGYLEISDVNCEALPEALSRC 604
Query: 602 ELLGSLQNLERIRLERIYVPSFGTLKNLKKLSLYMCNTILAFEKGSILISDAFPNLEELN 661
L +L L RL + S G LK L+ L L ++I + + I D NL L
Sbjct: 605 WNLQALHVLNCSRLA-VVPESIGKLKKLRTLELNGVSSIKSLPQS---IGDC-DNLRRLY 659
Query: 662 IDYCKDLVVLPIGICDIFLLKKLRVTNCHKLSSLP--QDIGKLENLELLSLSSCTDLEAI 719
++ C+ + +P + + L+ L + +C L LP GKL NL+ ++ + C +L +
Sbjct: 660 LEECRGIEDIPNSLGKLENLRILSIVDCVSLQKLPPSDSFGKLLNLQTITFNLCYNLRNL 719
Query: 720 PTSIGKLLNLKHLDISNCISLSSLPEEFGNLCNLKNLDMASCASIE-LPFSVVNLQNLKT 778
P + L++L+ +D+ C L LPE GNL NLK L++ C + LP L L+
Sbjct: 720 PQCMTSLIHLESVDLGYCFQLVELPEGMGNLRNLKVLNLKKCKKLRGLPAGCGKLTRLQQ 779
Query: 779 IT 780
++
Sbjct: 780 LS 781
Score = 55.8 bits (133), Expect = 5e-06
Identities = 89/361 (24%), Positives = 150/361 (40%), Gaps = 51/361 (14%)
Query: 420 VDMWAELYRLDDNGIQAMEIINKLGIMNLANVIIPRKNASDTDNNNYNNHFIILHDILRE 479
++M EL+ LD ++ I KL I +PR A +D+ + I
Sbjct: 856 LNMEKELHLLDS--LEPPSKIEKLRIRGYRGSQLPRWMAKQSDSCGPADDTHI------- 906
Query: 480 LGIYQSAKEPFEQRKRLIIDINKNKSGLAEKQQGLMTCILSKFMRLCVKRNPQQLTARIL 539
+ Q F L++D N L E L+ L K ++L KR P+ + +L
Sbjct: 907 --VMQRNPSEFSHLTELVLDNLPNLEHLGE----LVELPLVKILKL--KRLPKLV--ELL 956
Query: 540 SVSADETCAFDWSQMEPAQVEVLILNLHTKQYSLPEWIAKMSKLRV------LIITNYDF 593
+ + E + V L++ K P + + LR+ L+ + F
Sbjct: 957 TTTTGEEGVEVLYRFH--HVSTLVIIDCPKLVVKPYFPPSLQSLRLEGNNGQLVSSGCFF 1014
Query: 594 HPSKLN--NIELLGSLQNLERIRLERIYVPSFGT-----LKNLKKLSLYMCNTILAFEKG 646
HP + + +++G+ +LER+ L R+ S G L L L ++ C T L
Sbjct: 1015 HPRHHHAAHADVIGT--HLERLELRRLTGSSSGWEVLQHLTGLHTLEIFKC-TDLTHLPE 1071
Query: 647 SILISDAFPNLEELNIDYCKDLVVLPIGICDIFLLKKLRVTNCH-----------KLSSL 695
SI F L I C +L VLP + ++ L+ L + +C L+ L
Sbjct: 1072 SIHCPTTFCRLL---ITGCHNLRVLPDWLVELKSLQSLNIDSCDALQHLTISSLTSLTCL 1128
Query: 696 PQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISLSSLPEEFGNLCNLKN 755
P+ + L +L L+L C +L +P +G+L L+ L + +C L+SLP+ L L+
Sbjct: 1129 PESMQHLTSLRTLNLCRCNELTHLPEWLGELSVLQKLWLQDCRGLTSLPQSIQRLTALEE 1188
Query: 756 L 756
L
Sbjct: 1189 L 1189
Score = 45.4 bits (106), Expect = 0.007
Identities = 30/98 (30%), Positives = 51/98 (51%), Gaps = 7/98 (7%)
Query: 626 LKNLKKLSLYMCNTILAFEKGSILISDAFP-------NLEELNIDYCKDLVVLPIGICDI 678
LK+L+ L++ C+ + S+ P +L LN+ C +L LP + ++
Sbjct: 1100 LKSLQSLNIDSCDALQHLTISSLTSLTCLPESMQHLTSLRTLNLCRCNELTHLPEWLGEL 1159
Query: 679 FLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDL 716
+L+KL + +C L+SLPQ I +L LE L +S +L
Sbjct: 1160 SVLQKLWLQDCRGLTSLPQSIQRLTALEELYISGNPNL 1197
Score = 40.8 bits (94), Expect = 0.18
Identities = 34/120 (28%), Positives = 58/120 (48%), Gaps = 15/120 (12%)
Query: 681 LKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISL 740
L++L + SS + + L L L + CTDL +P SI L I+ C +L
Sbjct: 1031 LERLELRRLTGSSSGWEVLQHLTGLHTLEIFKCTDLTHLPESIHCPTTFCRLLITGCHNL 1090
Query: 741 SSLPEEFGNLCNLKNLDMASCASIE------------LPFSVVNLQNLKTIT---CDEET 785
LP+ L +L++L++ SC +++ LP S+ +L +L+T+ C+E T
Sbjct: 1091 RVLPDWLVELKSLQSLNIDSCDALQHLTISSLTSLTCLPESMQHLTSLRTLNLCRCNELT 1150
>dbj|BAB09158.1| disease resistance protein-like [Arabidopsis thaliana]
gi|15241520|ref|NP_199264.1| disease resistance protein
(TIR-NBS-LRR class), putative [Arabidopsis thaliana]
Length = 1187
Score = 114 bits (284), Expect = 2e-23
Identities = 144/572 (25%), Positives = 254/572 (44%), Gaps = 87/572 (15%)
Query: 245 LGLLLKKVEGSPLLLVLDDVWPISE--PLVEKIQ-FQISDFKILVTSRVAFPRFSTTCIL 301
LG+ ++++ +LLVLDDV + + + + +Q F + I+VT + + +
Sbjct: 308 LGVAQERLKDKKVLLVLDDVDGLVQLDAMAKDVQWFGLGSRIIVVTQDLKLLKAHGIKYI 367
Query: 302 KPL---AHEEAVTLFHHYAQMEKNSSDIINKNLVEKVVRSCQG-LPLTIKVIATSLKNRP 357
+ +EA+ +F YA EK+ + + + V + G LPL ++V+ + L+
Sbjct: 368 YKVDFPTSDEALEIFCMYAFGEKSPK--VGFEQIARTVTTLAGKLPLGLRVMGSYLRRMS 425
Query: 358 RDMWRKIGKELSQGHSILDSNTDLLTRLQKIFDVLEDNPTIMECFMDIALFPEDHRIPVA 417
+ W K + + + LD D+ + L+ ++ L + + F+ I F RI
Sbjct: 426 KQEW---AKSIPRLRTSLDD--DIESVLKFSYNSLAEQEK--DLFLHITCFFRRERIETL 478
Query: 418 AL------VDMWAELYRLDDNGIQAMEIINKLGIMNLANVIIPRKNASDTDNNNYNNHFI 471
+ VDM L L D + ++ LG + + N+++
Sbjct: 479 EVFLAKKSVDMRQGLQILADKSLLSLN----LGNIEMHNLLVQ----------------- 517
Query: 472 ILHDILRELGIYQSAKEPF-------------EQRKRLIIDINKNKSGLAEKQQGLMTCI 518
+ DI+R+ I++ K F + R +I I+ SG+ E G++
Sbjct: 518 LGLDIVRKQSIHKPGKRQFLVDTEDICEVLTDDTGTRTLIGIDLELSGVIE---GVINIS 574
Query: 519 LSKFMRLCVKRNPQQLTARILSVSADETCAFDWSQMEPAQVEVLILNLHTKQYSL----- 573
F R+C N Q L R D + + + + LH ++Y L
Sbjct: 575 ERAFERMC---NLQFL--RFHHPYGDRCHDILYLPQGLSHISRKLRLLHWERYPLTCLPP 629
Query: 574 ---PEWIAKMSKLRVLIITNYDFHPSKLNNIEL--LGSLQNLERIRLERIYVPSFGTLKN 628
PE++ K++ ++ +D + + N++ L NL+ + P F T N
Sbjct: 630 KFNPEFLVKINMRDSMLEKLWDGN-EPIRNLKWMDLSFCVNLKEL-------PDFSTATN 681
Query: 629 LKKLSLYMCNTILAFEKGSILISDAFPNLEELNIDYCKDLVVLPIGICDIFLLKKLRVTN 688
L++L L C +++ I +A NL EL++ C LV LP I ++ LKKL +
Sbjct: 682 LQELRLINCLSLVELPSS---IGNA-TNLLELDLIDCSSLVKLPSSIGNLTNLKKLFLNR 737
Query: 689 CHKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISLSSLPEEFG 748
C L LP G + +L+ L+LS C+ L IP+SIG ++NLK + C SL LP G
Sbjct: 738 CSSLVKLPSSFGNVTSLKELNLSGCSSLLEIPSSIGNIVNLKKVYADGCSSLVQLPSSIG 797
Query: 749 NLCNLKNLDMASCASI-ELPFSVVNLQNLKTI 779
N NLK L + +C+S+ E P S++NL L+ +
Sbjct: 798 NNTNLKELHLLNCSSLMECPSSMLNLTRLEDL 829
Score = 95.1 bits (235), Expect = 8e-18
Identities = 52/128 (40%), Positives = 77/128 (59%), Gaps = 2/128 (1%)
Query: 656 NLEELNIDYCKDLVVLPIGICDIFLLKKLRVTNCHKLSSLPQDIGKLENLELLSLSSCTD 715
NL+ +++ +C +L LP L++LR+ NC L LP IG NL L L C+
Sbjct: 658 NLKWMDLSFCVNLKELP-DFSTATNLQELRLINCLSLVELPSSIGNATNLLELDLIDCSS 716
Query: 716 LEAIPTSIGKLLNLKHLDISNCISLSSLPEEFGNLCNLKNLDMASCASI-ELPFSVVNLQ 774
L +P+SIG L NLK L ++ C SL LP FGN+ +LK L+++ C+S+ E+P S+ N+
Sbjct: 717 LVKLPSSIGNLTNLKKLFLNRCSSLVKLPSSFGNVTSLKELNLSGCSSLLEIPSSIGNIV 776
Query: 775 NLKTITCD 782
NLK + D
Sbjct: 777 NLKKVYAD 784
Score = 81.3 bits (199), Expect = 1e-13
Identities = 60/174 (34%), Positives = 90/174 (51%), Gaps = 9/174 (5%)
Query: 573 LPEWIAKMSKLRVLIITNYDFH---PSKLNNIELLGSLQNLERIRLERIYVPSFGTLKNL 629
LP I + L+ L + N PS + N+ L L NL L + +PS G + NL
Sbjct: 792 LPSSIGNNTNLKELHLLNCSSLMECPSSMLNLTRLEDL-NLSGC-LSLVKLPSIGNVINL 849
Query: 630 KKLSLYMCNTILAFEKGSILISDAFPNLEELNIDYCKDLVVLPIGICDIFLLKKLRVTNC 689
+ L L C++++ I +A NL+ L +D C +L+ LP I +I L+ L + C
Sbjct: 850 QSLYLSDCSSLMELP---FTIENA-TNLDTLYLDGCSNLLELPSSIWNITNLQSLYLNGC 905
Query: 690 HKLSSLPQDIGKLENLELLSLSSCTDLEAIPTSIGKLLNLKHLDISNCISLSSL 743
L LP + NL+ LSL C+ L +P+SI ++ NL +LD+SNC SL L
Sbjct: 906 SSLKELPSLVENAINLQSLSLMKCSSLVELPSSIWRISNLSYLDVSNCSSLLEL 959
Score = 77.8 bits (190), Expect = 1e-12
Identities = 60/231 (25%), Positives = 106/231 (44%), Gaps = 35/231 (15%)
Query: 562 LILNLHTKQYSLPEWIAKMSKLRVLIITNYDFHPSKLNNIELLGSLQNLERIRLER---- 617
L LN + LP ++ L+ L ++ S L +G++ NL+++ +
Sbjct: 733 LFLNRCSSLVKLPSSFGNVTSLKELNLSGCS---SLLEIPSSIGNIVNLKKVYADGCSSL 789
Query: 618 IYVPS-FGTLKNLKKLSLYMCNTILAFEKGSILISDAFPNLEELNIDYCKDLVVLP---- 672
+ +PS G NLK+L L C++++ + ++ LE+LN+ C LV LP
Sbjct: 790 VQLPSSIGNNTNLKELHLLNCSSLMECPSSMLNLT----RLEDLNLSGCLSLVKLPSIGN 845
Query: 673 -IGICDIFL------------------LKKLRVTNCHKLSSLPQDIGKLENLELLSLSSC 713
I + ++L L L + C L LP I + NL+ L L+ C
Sbjct: 846 VINLQSLYLSDCSSLMELPFTIENATNLDTLYLDGCSNLLELPSSIWNITNLQSLYLNGC 905
Query: 714 TDLEAIPTSIGKLLNLKHLDISNCISLSSLPEEFGNLCNLKNLDMASCASI 764
+ L+ +P+ + +NL+ L + C SL LP + NL LD+++C+S+
Sbjct: 906 SSLKELPSLVENAINLQSLSLMKCSSLVELPSSIWRISNLSYLDVSNCSSL 956
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.323 0.139 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,305,272,937
Number of Sequences: 2540612
Number of extensions: 54375146
Number of successful extensions: 204229
Number of sequences better than 10.0: 3625
Number of HSP's better than 10.0 without gapping: 1517
Number of HSP's successfully gapped in prelim test: 2138
Number of HSP's that attempted gapping in prelim test: 178067
Number of HSP's gapped (non-prelim): 15852
length of query: 813
length of database: 863,360,394
effective HSP length: 136
effective length of query: 677
effective length of database: 517,837,162
effective search space: 350575758674
effective search space used: 350575758674
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 79 (35.0 bits)
Medicago: description of AC148347.2


