

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148241.10 - phase: 1 /pseudo
(611 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
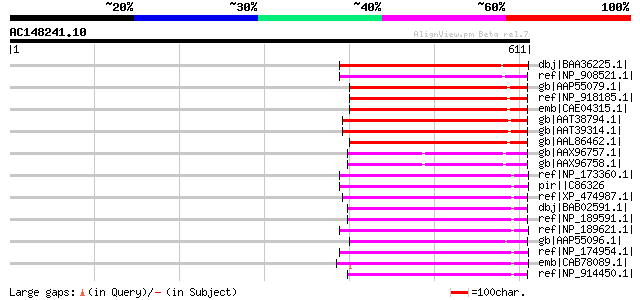
Score E
Sequences producing significant alignments: (bits) Value
dbj|BAA36225.1| transposase [Ipomoea purpurea] 230 1e-58
ref|NP_908521.1| putative transposable element Tip100 protein [O... 169 2e-40
gb|AAP55079.1| putative transposase [Oryza sativa (japonica cult... 156 2e-36
ref|NP_918185.1| putative transposase [Oryza sativa (japonica cu... 156 2e-36
emb|CAE04315.1| OSJNBb0016D16.6 [Oryza sativa (japonica cultivar... 156 2e-36
gb|AAT38794.1| putative hAT family dimerisation domain containin... 156 2e-36
gb|AAT39314.1| putative transposase [Solanum demissum] 156 2e-36
gb|AAL86462.1| putative transposase protein, 5'-partial [Oryza s... 156 2e-36
gb|AAX96757.1| hAT family dimerisation domain, putative [Oryza s... 155 4e-36
gb|AAX96758.1| hAT family dimerisation domain, putative [Oryza s... 155 5e-36
ref|NP_173360.1| hAT dimerisation domain-containing protein [Ara... 151 5e-35
pir||C86326 hypothetical protein T29M8.13 [imported] - Arabidops... 151 5e-35
ref|XP_474987.1| OSJNBa0065B15.7 [Oryza sativa (japonica cultiva... 150 9e-35
dbj|BAB02591.1| unnamed protein product [Arabidopsis thaliana] 150 9e-35
ref|NP_189591.1| hypothetical protein [Arabidopsis thaliana] 150 9e-35
ref|NP_189621.1| hAT dimerisation domain-containing protein [Ara... 150 1e-34
gb|AAP55096.1| putative transposase [Oryza sativa (japonica cult... 150 2e-34
ref|NP_174954.1| hypothetical protein [Arabidopsis thaliana] 149 2e-34
emb|CAB78089.1| putative protein [Arabidopsis thaliana] gi|45388... 146 2e-33
ref|NP_914450.1| putative transposable element [Oryza sativa (ja... 145 5e-33
>dbj|BAA36225.1| transposase [Ipomoea purpurea]
Length = 808
Score = 230 bits (586), Expect = 1e-58
Identities = 111/222 (50%), Positives = 163/222 (73%), Gaps = 1/222 (0%)
Query: 389 MTASREVASIHKFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQV 448
+ +S+EV +H+FF KL ++NVVG+S KR+D+L+AA + I +LL E+ +G+G NQ+
Sbjct: 410 VASSKEVIPVHQFFTKLNSIINVVGASCKRNDQLKAAHASNISHLLSIDELESGRGLNQI 469
Query: 449 GTVKRAGDTRWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDF 508
G+++R GDTRW SH SI SL+ M+ ATC VL I +D A R DAD++Y L SF+F
Sbjct: 470 GSLQRPGDTRWSSHLKSISSLMRMFSATCEVLLNIIEDGTTHAHRGDADAAYEVLTSFEF 529
Query: 509 IFILHLMKEIMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLFANVVSF 568
+FI+HLMK+++ +++LCQALQ QSQD++NA+ LV STK LIQ LR++GWD+L A+V SF
Sbjct: 530 VFIMHLMKKVLEISNMLCQALQLQSQDILNAMHLVSSTKLLIQTLRDSGWDELVASVKSF 589
Query: 569 CEKHDIEVPDLNDSHSTTRFGRSHLEENQVTIEHYFRVDFFY 610
CE +I VPD D+ R GR+ +++++TI H+++VD FY
Sbjct: 590 CETVNITVPDF-DAQYIARRGRARHQQDELTIGHHYKVDIFY 630
>ref|NP_908521.1| putative transposable element Tip100 protein [Oryza sativa
(japonica cultivar-group)]
Length = 802
Score = 169 bits (428), Expect = 2e-40
Identities = 86/221 (38%), Positives = 131/221 (58%), Gaps = 2/221 (0%)
Query: 389 MTASREVASIHKFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQV 448
+ S +I FF + +VN VG+S R D L A + +E GEI TG+G NQ
Sbjct: 390 VAVSTSTPAIADFFNYVPLIVNTVGASCMRKDALLAKHHDVLLEKVENGEITTGRGLNQE 449
Query: 449 GTVKRAGDTRWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDF 508
++ R GDTRWGSH ++ ++ M+ A VL+I+ KD+ K A ++SFDF
Sbjct: 450 SSLARPGDTRWGSHLKTLLRILVMWEAIIDVLEIVKKDSTKPTFNGGAFGLMGKMQSFDF 509
Query: 509 IFILHLMKEIMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLFANVVSF 568
+FI+HLM +++ TD L +ALQ++ QD+V A+ L+ K L+QD+RENGW+ L V+SF
Sbjct: 510 VFIMHLMIDMLSITDDLSRALQRKDQDIVEAMSLLIDVKELLQDMRENGWEPLLNRVISF 569
Query: 569 CEKHDIEVPDLNDSHSTTRFGRSHLEENQVTIEHYFRVDFF 609
C KH+I+VP ++ G S ++VT +HY+ V+ +
Sbjct: 570 CNKHEIKVPKMD--KEVNERGTSTHRRHKVTNKHYYHVEIY 608
>gb|AAP55079.1| putative transposase [Oryza sativa (japonica cultivar-group)]
gi|37536980|ref|NP_922792.1| putative transposase [Oryza
sativa (japonica cultivar-group)]
gi|18854992|gb|AAL79684.1| putative transposase [Oryza
sativa]
Length = 1283
Score = 156 bits (395), Expect = 2e-36
Identities = 85/210 (40%), Positives = 130/210 (61%), Gaps = 3/210 (1%)
Query: 401 FFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQVGTVKRAGDTRWG 460
FFE+L F++N++G+S K+ + L+ AQ++ I L+ G+I TGKG NQ + R DTRWG
Sbjct: 639 FFEQLRFLLNLLGNSCKKTEMLRVAQAQRIVEELDLGDIETGKGLNQEMGLGRPADTRWG 698
Query: 461 SHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFIFILHLMKEIMG 520
SH+ ++ ++S+Y + VL ++ KD A+ A+A + +SF+F+F+ HLM+ ++G
Sbjct: 699 SHYKTVMHVLSLYPSIQKVLIMVGKDRSFGAECANAQTVLTIFQSFEFVFMAHLMQTVLG 758
Query: 521 TTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLREN-GWDKLFANVVSFCEKHDIEVPDL 579
T L ALQK+ QD+VNAV L+ TK +Q LRE+ GWD V SFC KH I++PD+
Sbjct: 759 FTSDLNHALQKRDQDIVNAVGLILLTKFQLQQLREDPGWDDFLQEVQSFCVKHKIKIPDM 818
Query: 580 NDSHSTTRFGRSHLEENQVTIEHYFRVDFF 609
+ + GR ++ H F VD F
Sbjct: 819 DSFYRPV--GRDRRFFIKIKNMHRFHVDMF 846
>ref|NP_918185.1| putative transposase [Oryza sativa (japonica cultivar-group)]
Length = 897
Score = 156 bits (395), Expect = 2e-36
Identities = 85/210 (40%), Positives = 130/210 (61%), Gaps = 3/210 (1%)
Query: 401 FFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQVGTVKRAGDTRWG 460
FFE+L F++N++G+S K+ + L+ AQ++ I L+ G+I TGKG NQ + R DTRWG
Sbjct: 499 FFEQLRFLLNLLGNSCKKTEMLRVAQAQRIVEELDLGDIETGKGLNQEMGLGRPADTRWG 558
Query: 461 SHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFIFILHLMKEIMG 520
SH+ ++ ++S+Y + VL ++ KD A+ A+A + +SF+F+F+ HLM+ ++G
Sbjct: 559 SHYKTVMHVLSLYPSIQKVLIMVGKDRSFGAECANAQTVLTIFQSFEFVFMAHLMQTVLG 618
Query: 521 TTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLREN-GWDKLFANVVSFCEKHDIEVPDL 579
T L ALQK+ QD+VNAV L+ TK +Q LRE+ GWD V SFC KH I++PD+
Sbjct: 619 FTSDLNHALQKRDQDIVNAVGLILLTKFQLQQLREDPGWDDFLQEVQSFCVKHKIKIPDM 678
Query: 580 NDSHSTTRFGRSHLEENQVTIEHYFRVDFF 609
+ + GR ++ H F VD F
Sbjct: 679 DSFYRPV--GRDRRFFIKIKNMHRFHVDMF 706
>emb|CAE04315.1| OSJNBb0016D16.6 [Oryza sativa (japonica cultivar-group)]
gi|50928319|ref|XP_473687.1| OSJNBb0016D16.6 [Oryza
sativa (japonica cultivar-group)]
Length = 897
Score = 156 bits (395), Expect = 2e-36
Identities = 85/210 (40%), Positives = 130/210 (61%), Gaps = 3/210 (1%)
Query: 401 FFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQVGTVKRAGDTRWG 460
FFE+L F++N++G+S K+ + L+ AQ++ I L+ G+I TGKG NQ + R DTRWG
Sbjct: 499 FFEQLRFLLNLLGNSCKKTEMLRVAQAQRIVEELDLGDIETGKGLNQEMGLGRPADTRWG 558
Query: 461 SHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFIFILHLMKEIMG 520
SH+ ++ ++S+Y + VL ++ KD A+ A+A + +SF+F+F+ HLM+ ++G
Sbjct: 559 SHYKTVMHVLSLYPSIQKVLIMVGKDRSFGAECANAQTVLTIFQSFEFVFMAHLMQTVLG 618
Query: 521 TTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLREN-GWDKLFANVVSFCEKHDIEVPDL 579
T L ALQK+ QD+VNAV L+ TK +Q LRE+ GWD V SFC KH I++PD+
Sbjct: 619 FTSDLNHALQKRDQDIVNAVGLILLTKFQLQQLREDPGWDDFLQEVQSFCVKHKIKIPDM 678
Query: 580 NDSHSTTRFGRSHLEENQVTIEHYFRVDFF 609
+ + GR ++ H F VD F
Sbjct: 679 DSFYRPV--GRDRRFFIKIKNMHRFHVDMF 706
>gb|AAT38794.1| putative hAT family dimerisation domain containing protein [Solanum
demissum]
Length = 805
Score = 156 bits (395), Expect = 2e-36
Identities = 83/218 (38%), Positives = 132/218 (60%), Gaps = 2/218 (0%)
Query: 392 SREVASIHKFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQVGTV 451
S++ + K + + NV+GSS KR D L+ +Q I+ L+ GE+ TG G NQ +
Sbjct: 410 SKKCVEVGKLVVLISNIFNVLGSSFKRMDNLRDSQKSTIQVALDMGELTTGSGLNQQLGL 469
Query: 452 KRAGDTRWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFIFI 511
RA DTRWGSH+ S + I M+G+ VL+ +A DA+ + +RA A ++F+ F+
Sbjct: 470 SRACDTRWGSHYKSFNNFIIMFGSILEVLESLALDARSMDERAKAMGHLEACQTFEIAFM 529
Query: 512 LHLMKEIMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLFANVVSFCEK 571
LHLM++++ T+ L + LQK+ QD+ NA+LLV K +Q LR++ WD L A V +FC K
Sbjct: 530 LHLMRDVLAITNELNKCLQKKEQDIANAMLLVEVAKRRLQVLRDDEWDSLIAKVSTFCIK 589
Query: 572 HDIEVPDLNDSHSTTRFGRSHLEENQVTIEHYFRVDFF 609
H+I +P+ + + ++ RS + TI H++RV+ F
Sbjct: 590 HNILIPNFEEPYVSSL--RSRRKLANYTILHHYRVEVF 625
>gb|AAT39314.1| putative transposase [Solanum demissum]
Length = 705
Score = 156 bits (395), Expect = 2e-36
Identities = 83/218 (38%), Positives = 132/218 (60%), Gaps = 2/218 (0%)
Query: 392 SREVASIHKFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQVGTV 451
S++ + K + + NV+GSS KR D L+ +Q I+ L+ GE+ TG G NQ +
Sbjct: 310 SKKCVEVGKLVVLISNIFNVLGSSFKRMDNLRDSQKSTIQVALDMGELTTGSGLNQQLGL 369
Query: 452 KRAGDTRWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFIFI 511
RA DTRWGSH+ S + I M+G+ VL+ +A DA+ + +RA A ++F+ F+
Sbjct: 370 SRACDTRWGSHYKSFNNFIIMFGSILEVLESLALDARSMDERAKAMGHLEACQTFEIAFM 429
Query: 512 LHLMKEIMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLFANVVSFCEK 571
LHLM++++ T+ L + LQK+ QD+ NA+LLV K +Q LR++ WD L A V +FC K
Sbjct: 430 LHLMRDVLAITNELNKCLQKKEQDIANAMLLVEVAKRRLQVLRDDEWDSLIAKVSTFCIK 489
Query: 572 HDIEVPDLNDSHSTTRFGRSHLEENQVTIEHYFRVDFF 609
H+I +P+ + + ++ RS + TI H++RV+ F
Sbjct: 490 HNILIPNFEEPYVSSL--RSRRKLANYTILHHYRVEVF 525
>gb|AAL86462.1| putative transposase protein, 5'-partial [Oryza sativa (japonica
cultivar-group)]
Length = 881
Score = 156 bits (395), Expect = 2e-36
Identities = 85/210 (40%), Positives = 130/210 (61%), Gaps = 3/210 (1%)
Query: 401 FFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQVGTVKRAGDTRWG 460
FFE+L F++N++G+S K+ + L+ AQ++ I L+ G+I TGKG NQ + R DTRWG
Sbjct: 133 FFEQLRFLLNLLGNSCKKTEMLRVAQAQRIVEELDLGDIETGKGLNQEMGLGRPADTRWG 192
Query: 461 SHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFIFILHLMKEIMG 520
SH+ ++ ++S+Y + VL ++ KD A+ A+A + +SF+F+F+ HLM+ ++G
Sbjct: 193 SHYKTVMHVLSLYPSIQKVLIMVGKDRSFGAECANAQTVLTIFQSFEFVFMAHLMQTVLG 252
Query: 521 TTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLREN-GWDKLFANVVSFCEKHDIEVPDL 579
T L ALQK+ QD+VNAV L+ TK +Q LRE+ GWD V SFC KH I++PD+
Sbjct: 253 FTSDLNHALQKRDQDIVNAVGLILLTKFQLQQLREDPGWDDFLQEVQSFCVKHKIKIPDM 312
Query: 580 NDSHSTTRFGRSHLEENQVTIEHYFRVDFF 609
+ + GR ++ H F VD F
Sbjct: 313 DSFYRPV--GRDRRFFIKIKNMHRFHVDMF 340
>gb|AAX96757.1| hAT family dimerisation domain, putative [Oryza sativa (japonica
cultivar-group)]
Length = 1071
Score = 155 bits (392), Expect = 4e-36
Identities = 82/213 (38%), Positives = 125/213 (58%), Gaps = 5/213 (2%)
Query: 398 IHKFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQVGTVKRAGDT 457
+ FF +VN V S KRHD+L + + LETGEI +G+GKNQ + R GDT
Sbjct: 460 VSDFFNYTPMIVNTVNGSCKRHDQLAQEHHDNLVSSLETGEIFSGRGKNQATNLARPGDT 519
Query: 458 RWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFIFILHLMKE 517
RWGSH ++C L M+ A VL+ I +D + A ++SF+F+ I+HLM
Sbjct: 520 RWGSHHKTLCRLQHMWKAVLEVLENICED--NPTTKTTATGLLKQMESFEFVLIMHLMIR 577
Query: 518 IMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLFANVVSFCEKHDIEVP 577
++G T+ L Q LQK+ Q+++ A+ L+ +T I +R++GW +LF +FC K++I VP
Sbjct: 578 LLGKTNDLSQCLQKKDQNIIRAIGLIGTTLQKINHIRQHGWQELFEETKTFCLKYNIIVP 637
Query: 578 DLNDSHSTTRFGRSHLEENQ-VTIEHYFRVDFF 609
D++D +TT GRS Q +T +H+F + F
Sbjct: 638 DMSD--TTTTRGRSRGRGGQLITYQHHFNNEIF 668
>gb|AAX96758.1| hAT family dimerisation domain, putative [Oryza sativa (japonica
cultivar-group)]
Length = 1050
Score = 155 bits (391), Expect = 5e-36
Identities = 82/213 (38%), Positives = 125/213 (58%), Gaps = 5/213 (2%)
Query: 398 IHKFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQVGTVKRAGDT 457
+ FF +VN V S KRHD+L + + LETGEI +G+GKNQ + R GDT
Sbjct: 636 VSDFFNYTPMIVNTVNGSCKRHDQLAQEHHDNLVSSLETGEIFSGRGKNQATNLARPGDT 695
Query: 458 RWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFIFILHLMKE 517
RWGSH ++C L M+ A VL+ I +D + A ++SF+F+ I+HLM
Sbjct: 696 RWGSHHKTLCRLQHMWKAVLEVLENICED--NPTTKITATGLLKQMESFEFVLIMHLMIR 753
Query: 518 IMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLFANVVSFCEKHDIEVP 577
++G T+ L Q LQK+ Q+++ A+ L+ +T I +R++GW +LF +FC K++I VP
Sbjct: 754 LLGKTNDLSQCLQKKDQNIIRAIGLIGTTLQKINHIRQHGWQELFEETKTFCLKYNIIVP 813
Query: 578 DLNDSHSTTRFGRSHLEENQ-VTIEHYFRVDFF 609
D++D +TT GRS Q +T +H+F + F
Sbjct: 814 DMSD--TTTTRGRSRGRGGQLITYQHHFNNEIF 844
>ref|NP_173360.1| hAT dimerisation domain-containing protein [Arabidopsis thaliana]
Length = 769
Score = 151 bits (382), Expect = 5e-35
Identities = 76/222 (34%), Positives = 133/222 (59%), Gaps = 3/222 (1%)
Query: 389 MTASREVASIHKFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQV 448
M +++ + +FF + ++NVVG+S KR D+++ +++E + GEI TG G NQ
Sbjct: 373 MAVAKKHVEVGEFFYMISVLLNVVGASCKRKDKIREIHRQKVEEKISNGEIKTGTGLNQE 432
Query: 449 GTVKRAGDTRWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDF 508
+++R G+TRWGSH+ ++ L ++ + VL+ I + +R A + +FDF
Sbjct: 433 LSLQRPGNTRWGSHYKTLLRLEELFSSIVIVLEYIQDEGTDTTKRQQAYGILKYFHTFDF 492
Query: 509 IFILHLMKEIMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLFANVVSF 568
+F L LM +MG TD L +ALQ++ QD++NA+ LV++TK +Q +R++GWD A V SF
Sbjct: 493 VFYLELMLLVMGLTDSLSKALQRKDQDILNAISLVKTTKCQLQKVRDDGWDAFMAKVSSF 552
Query: 569 CEKHDIEVPDLNDSHSTTRFGRSHLEENQVTIEHYFRVDFFY 610
EK++ + + + +R R ++ +T H+++VD FY
Sbjct: 553 SEKNNTGMLKMEEEFVDSRRPR---KKTGITNLHHYKVDCFY 591
>pir||C86326 hypothetical protein T29M8.13 [imported] - Arabidopsis thaliana
gi|8954063|gb|AAF82236.1| Contains similarity to a
transposable element Tip100 protein for transposase from
Ipomoea purpurea gb|4063769 and is a member of the
transmembrane 4 family PF|00335. [Arabidopsis thaliana]
Length = 811
Score = 151 bits (382), Expect = 5e-35
Identities = 76/222 (34%), Positives = 133/222 (59%), Gaps = 3/222 (1%)
Query: 389 MTASREVASIHKFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQV 448
M +++ + +FF + ++NVVG+S KR D+++ +++E + GEI TG G NQ
Sbjct: 415 MAVAKKHVEVGEFFYMISVLLNVVGASCKRKDKIREIHRQKVEEKISNGEIKTGTGLNQE 474
Query: 449 GTVKRAGDTRWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDF 508
+++R G+TRWGSH+ ++ L ++ + VL+ I + +R A + +FDF
Sbjct: 475 LSLQRPGNTRWGSHYKTLLRLEELFSSIVIVLEYIQDEGTDTTKRQQAYGILKYFHTFDF 534
Query: 509 IFILHLMKEIMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLFANVVSF 568
+F L LM +MG TD L +ALQ++ QD++NA+ LV++TK +Q +R++GWD A V SF
Sbjct: 535 VFYLELMLLVMGLTDSLSKALQRKDQDILNAISLVKTTKCQLQKVRDDGWDAFMAKVSSF 594
Query: 569 CEKHDIEVPDLNDSHSTTRFGRSHLEENQVTIEHYFRVDFFY 610
EK++ + + + +R R ++ +T H+++VD FY
Sbjct: 595 SEKNNTGMLKMEEEFVDSRRPR---KKTGITNLHHYKVDCFY 633
>ref|XP_474987.1| OSJNBa0065B15.7 [Oryza sativa (japonica cultivar-group)]
gi|38344561|emb|CAD39903.2| OSJNBa0065B15.7 [Oryza
sativa (japonica cultivar-group)]
Length = 639
Score = 150 bits (380), Expect = 9e-35
Identities = 77/217 (35%), Positives = 127/217 (58%), Gaps = 3/217 (1%)
Query: 393 REVASIHKFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQVGTVK 452
R+ + FF K+ ++NVVG S+KR D ++ KE+ L +G++ TG NQ ++
Sbjct: 254 RKHKDVSDFFTKISILLNVVGGSSKRRDLIRDINVKEMSKALGSGQLQTGTRLNQEQCLQ 313
Query: 453 RAGDTRWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFIFIL 512
R GDTRW SH+ ++ SL+ M+ VL+I+ KD R + + +SFDF+F L
Sbjct: 314 RPGDTRWSSHYKTLKSLVGMFATIVKVLEIVEKDKNDWKIRDQGSNLLEYFQSFDFVFYL 373
Query: 513 HLMKEIMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLFANVVSFCEKH 572
HLM I+ T+ L ALQ++ QD+VNA+ V+ST+ + +LR W+K+ V FC+ +
Sbjct: 374 HLMLTILTITNSLSLALQRKDQDIVNAMKCVKSTRLNLDELRREKWEKVLDEVSDFCDNY 433
Query: 573 DIEVPDLNDSHSTTRFGRSHLEENQVTIEHYFRVDFF 609
DI ++ D++ + R ++ +T +HY++VD F
Sbjct: 434 DIVKLEIEDTYIDPKKRR---HKSGITNKHYYQVDCF 467
>dbj|BAB02591.1| unnamed protein product [Arabidopsis thaliana]
Length = 571
Score = 150 bits (380), Expect = 9e-35
Identities = 78/212 (36%), Positives = 127/212 (59%), Gaps = 3/212 (1%)
Query: 398 IHKFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQVGTVKRAGDT 457
I FF+ + ++NVVG+S KR D ++ K++E + GEI TGKG NQ +++R G+T
Sbjct: 315 IGDFFDMISVLINVVGASCKRKDRVRDEFRKKLEERINQGEIKTGKGLNQKLSLQRPGNT 374
Query: 458 RWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFIFILHLMKE 517
RWG+H+ ++ L+ ++ VL+ I D +R A+ + +FDF+F L LM
Sbjct: 375 RWGTHYTTLLRLVDLFSVIIKVLEWIEDDGTDSTKRRQANGLLKYFNTFDFVFYLQLMLL 434
Query: 518 IMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLFANVVSFCEKHDIEVP 577
I+G T+ L ALQ++ QD++NA+ LV+STK + LR++GWD V SFC+ HDIE
Sbjct: 435 ILGLTNSLSVALQRKDQDILNAMSLVKSTKQQLFKLRDDGWDSFLNEVFSFCKDHDIEFV 494
Query: 578 DLNDSHSTTRFGRSHLEENQVTIEHYFRVDFF 609
++ R R +++ +T H+++V+ F
Sbjct: 495 IMDGEFVDPRKPR---KKSNMTNLHHYQVECF 523
>ref|NP_189591.1| hypothetical protein [Arabidopsis thaliana]
Length = 522
Score = 150 bits (380), Expect = 9e-35
Identities = 78/212 (36%), Positives = 127/212 (59%), Gaps = 3/212 (1%)
Query: 398 IHKFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQVGTVKRAGDT 457
I FF+ + ++NVVG+S KR D ++ K++E + GEI TGKG NQ +++R G+T
Sbjct: 266 IGDFFDMISVLINVVGASCKRKDRVRDEFRKKLEERINQGEIKTGKGLNQKLSLQRPGNT 325
Query: 458 RWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFIFILHLMKE 517
RWG+H+ ++ L+ ++ VL+ I D +R A+ + +FDF+F L LM
Sbjct: 326 RWGTHYTTLLRLVDLFSVIIKVLEWIEDDGTDSTKRRQANGLLKYFNTFDFVFYLQLMLL 385
Query: 518 IMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLFANVVSFCEKHDIEVP 577
I+G T+ L ALQ++ QD++NA+ LV+STK + LR++GWD V SFC+ HDIE
Sbjct: 386 ILGLTNSLSVALQRKDQDILNAMSLVKSTKQQLFKLRDDGWDSFLNEVFSFCKDHDIEFV 445
Query: 578 DLNDSHSTTRFGRSHLEENQVTIEHYFRVDFF 609
++ R R +++ +T H+++V+ F
Sbjct: 446 IMDGEFVDPRKPR---KKSNMTNLHHYQVECF 474
>ref|NP_189621.1| hAT dimerisation domain-containing protein [Arabidopsis thaliana]
Length = 536
Score = 150 bits (379), Expect = 1e-34
Identities = 76/222 (34%), Positives = 132/222 (59%), Gaps = 3/222 (1%)
Query: 389 MTASREVASIHKFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQV 448
M +++ + +FF + ++NVVG+S KR D+++ +++E + GEI TG G NQ
Sbjct: 140 MAVAKKHVEVGEFFYMISVLLNVVGASCKRKDKIREIDRQKVEEKISNGEIKTGTGLNQE 199
Query: 449 GTVKRAGDTRWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDF 508
+++R G+TRWGSH+ ++ L ++ + VL+ I + +R A + +FDF
Sbjct: 200 LSLQRPGNTRWGSHYKTLLRLEELFSSIVIVLEYIQDEGTDTTKRQQAYGILKYFHTFDF 259
Query: 509 IFILHLMKEIMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLFANVVSF 568
+F L LM +MG TD L +ALQ++ QD++NA+ LV++TK +Q +R++GWD A V SF
Sbjct: 260 VFYLELMLLVMGLTDSLSKALQRKDQDILNAISLVKTTKCQLQKVRDDGWDAFMAEVSSF 319
Query: 569 CEKHDIEVPDLNDSHSTTRFGRSHLEENQVTIEHYFRVDFFY 610
EK + + + + +R R ++ +T H+++VD FY
Sbjct: 320 SEKTNTGMLKMEEEFVDSRRPR---KKTGITNLHHYKVDCFY 358
>gb|AAP55096.1| putative transposase [Oryza sativa (japonica cultivar-group)]
gi|37537014|ref|NP_922809.1| putative transposase [Oryza
sativa (japonica cultivar-group)]
gi|19225003|gb|AAL86479.1| putative transposase [Oryza
sativa (japonica cultivar-group)]
Length = 811
Score = 150 bits (378), Expect = 2e-34
Identities = 85/210 (40%), Positives = 121/210 (57%), Gaps = 3/210 (1%)
Query: 401 FFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQVGTVKRAGDTRWG 460
FF ++ ++N+VG+S KRHD L+ ++++I LE GEI +G G NQ + R GDTRWG
Sbjct: 424 FFTQVSHLLNIVGTSCKRHDMLRDVRAQKIMEALELGEIESGVGLNQEMGLARPGDTRWG 483
Query: 461 SHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFIFILHLMKEIMG 520
SH+ +I +I MY VL + KD + + +SFDF+F HLM I+G
Sbjct: 484 SHYKTILHIIGMYPTIHEVLITLGKDPTQRDDWPRIHAVVGAFESFDFVFSAHLMLVILG 543
Query: 521 TTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLF-ANVVSFCEKHDIEVPDL 579
T+ LC LQK+ QD+VNA+ LV K +Q LR GW++ F VVSFC KH I+VP L
Sbjct: 544 YTNELCLCLQKRDQDIVNAMSLVTLAKERMQKLRSEGWEEFFQGTVVSFCNKHSIQVPTL 603
Query: 580 NDSHSTTRFGRSHLEENQVTIEHYFRVDFF 609
+ + GRS T + +FR + +
Sbjct: 604 DGKY--VPHGRSPRFYPDQTNDDHFRREVY 631
>ref|NP_174954.1| hypothetical protein [Arabidopsis thaliana]
Length = 496
Score = 149 bits (377), Expect = 2e-34
Identities = 77/222 (34%), Positives = 131/222 (58%), Gaps = 3/222 (1%)
Query: 389 MTASREVASIHKFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQV 448
+ +++ + FF+ + ++NVVG+S K D ++ KEIE + GEI TGKG NQ
Sbjct: 198 VAVAKKQFEVGDFFDMISALLNVVGASCKGKDRIREEYLKEIEEGINQGEIKTGKGLNQE 257
Query: 449 GTVKRAGDTRWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDF 508
+++R G+TRWG+H+ ++ L ++ +L+ + ++ +R A+ + +FDF
Sbjct: 258 LSLQRPGNTRWGTHYTTLHRLAHLFSVIIKLLEFVEEEGTDSTKRRQANGLLKYFNTFDF 317
Query: 509 IFILHLMKEIMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLFANVVSF 568
F L LM ++G T+ L ALQK+ QD++N + LV+STK + LR++GWD L V SF
Sbjct: 318 AFYLQLMLLLLGLTNSLSVALQKKDQDILNPMSLVKSTKQQLCKLRDDGWDSLVNEVFSF 377
Query: 569 CEKHDIEVPDLNDSHSTTRFGRSHLEENQVTIEHYFRVDFFY 610
C+KHDIE+ ++ R R + + +T H+++V+ FY
Sbjct: 378 CKKHDIELVIMDGEFVNPRNPR---KRSNMTNFHHYQVECFY 416
>emb|CAB78089.1| putative protein [Arabidopsis thaliana] gi|4538896|emb|CAB39633.1|
putative protein [Arabidopsis thaliana]
gi|15233987|ref|NP_192704.1| hypothetical protein
[Arabidopsis thaliana] gi|7485592|pir||T04013
hypothetical protein F17A8.10 - Arabidopsis thaliana
Length = 664
Score = 146 bits (368), Expect = 2e-33
Identities = 78/232 (33%), Positives = 137/232 (58%), Gaps = 9/232 (3%)
Query: 385 EENGMTAS--REVASIH----KFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGE 438
E NG+ + +E+A H +FF+ + ++NVVG+S R D+++ +++E + GE
Sbjct: 307 EFNGLRSLILKEIAKKHVEVGEFFDMISVLLNVVGASCTRKDKIREIHRQKVEEKISNGE 366
Query: 439 IVTGKGKNQVGTVKRAGDTRWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADS 498
I TG NQ +++R G+TRWGSH+ ++ L ++ + VL+ I + +R A
Sbjct: 367 IKTGTRLNQELSLQRPGNTRWGSHYKTLLRLEELFSSIVIVLEYIQDEGTDTTKRQQAYG 426
Query: 499 SYNHLKSFDFIFILHLMKEIMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGW 558
+ +FDF+F L LM +MG TD L +ALQ++ QD++N + LV++TK +Q +R++GW
Sbjct: 427 ILKYFHTFDFVFYLELMLLVMGLTDSLSKALQRKDQDILNVISLVKTTKCQLQKVRDDGW 486
Query: 559 DKLFANVVSFCEKHDIEVPDLNDSHSTTRFGRSHLEENQVTIEHYFRVDFFY 610
D A V SF EK++ + + + +R R +++ +T H+++VD FY
Sbjct: 487 DAFMAEVSSFSEKNNTAMLKMEEEFVDSRRPR---KKSGITNLHHYKVDCFY 535
>ref|NP_914450.1| putative transposable element [Oryza sativa (japonica
cultivar-group)]
Length = 796
Score = 145 bits (365), Expect = 5e-33
Identities = 71/212 (33%), Positives = 125/212 (58%), Gaps = 3/212 (1%)
Query: 398 IHKFFEKLIFVVNVVGSSTKRHDELQAAQSKEIENLLETGEIVTGKGKNQVGTVKRAGDT 457
I FF+ + ++NVVG S+KR D ++ ++ + + LE G ++TG G NQ +++R GDT
Sbjct: 406 IGNFFDMISLLLNVVGGSSKRRDMIRDINAEHVHDALECGLLITGSGLNQEISLQRPGDT 465
Query: 458 RWGSHFNSICSLISMYGATCPVLKIIAKDAKKIAQRADADSSYNHLKSFDFIFILHLMKE 517
RW SH+ ++ SL+ ++ T VL+ + ++ + A + + F F+F L LM
Sbjct: 466 RWSSHYKTLKSLVQLFPTTVKVLEYVEENDRDDKNSRQAAGLLVYFQLFQFVFYLQLMLN 525
Query: 518 IMGTTDLLCQALQKQSQDVVNAVLLVRSTKALIQDLRENGWDKLFANVVSFCEKHDIEVP 577
I+ T+ LQ++ QD+VNAV V+ST+ + + R +GW+ + A V +FCEK+DI
Sbjct: 526 ILAITNTFSVTLQRKDQDIVNAVNCVKSTRNHLDEFRRSGWESILAEVHNFCEKYDISEL 585
Query: 578 DLNDSHSTTRFGRSHLEENQVTIEHYFRVDFF 609
++ D++ + R ++ +T +H+F VD F
Sbjct: 586 EMEDAYINPKRPR---QKTGITNKHHFEVDCF 614
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.359 0.157 0.535
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 783,172,948
Number of Sequences: 2540612
Number of extensions: 26386640
Number of successful extensions: 138508
Number of sequences better than 10.0: 37
Number of HSP's better than 10.0 without gapping: 35
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 138442
Number of HSP's gapped (non-prelim): 41
length of query: 611
length of database: 863,360,394
effective HSP length: 134
effective length of query: 477
effective length of database: 522,918,386
effective search space: 249432070122
effective search space used: 249432070122
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 78 (34.7 bits)
Medicago: description of AC148241.10


