

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148240.10 - phase: 0 /pseudo
(1012 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
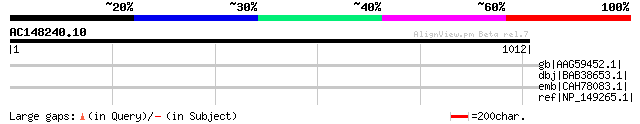
Score E
Sequences producing significant alignments: (bits) Value
gb|AAG59452.1| orf; Unknown function [Escherichia coli O157:H7 E... 42 0.080
dbj|BAB38653.1| hypothetical protein [Escherichia coli O157:H7] ... 39 0.88
emb|CAH78083.1| conserved hypothetical protein [Plasmodium chaba... 37 4.4
ref|NP_149265.1| Membrane protein [Clostridium acetobutylicum AT... 36 7.4
>gb|AAG59452.1| orf; Unknown function [Escherichia coli O157:H7 EDL933]
gi|25501232|pir||H86123 hypothetical protein yjgL
[imported] - Escherichia coli (strain O157:H7,
substrain EDL933) gi|15804845|ref|NP_290886.1|
hypothetical protein Z5865 [Escherichia coli O157:H7
EDL933]
Length = 596
Score = 42.4 bits (98), Expect = 0.080
Identities = 22/46 (47%), Positives = 31/46 (66%)
Query: 436 QVQLILKMHQNLINLLILKSIQKLNLVQKLKSLQKLSLIQKQKLVQ 481
++ +L HQ L NL L ++QKLN +QKL +LQKL+ IQK +Q
Sbjct: 181 EIARLLNNHQKLNNLQKLNNLQKLNNLQKLNNLQKLNNIQKLNNIQ 226
>dbj|BAB38653.1| hypothetical protein [Escherichia coli O157:H7]
gi|25391903|pir||F91282 hypothetical protein ECs5230
[imported] - Escherichia coli (strain O157:H7,
substrain RIMD 0509952) gi|15834484|ref|NP_313257.1|
hypothetical protein ECs5230 [Escherichia coli O157:H7]
Length = 590
Score = 38.9 bits (89), Expect = 0.88
Identities = 19/41 (46%), Positives = 29/41 (70%)
Query: 436 QVQLILKMHQNLINLLILKSIQKLNLVQKLKSLQKLSLIQK 476
++ +L HQ L NL L ++QKLN +QKL ++QKL+ IQ+
Sbjct: 181 EIARLLNNHQKLNNLQKLNNLQKLNNLQKLNNIQKLNNIQE 221
>emb|CAH78083.1| conserved hypothetical protein [Plasmodium chabaudi]
Length = 768
Score = 36.6 bits (83), Expect = 4.4
Identities = 29/99 (29%), Positives = 51/99 (51%), Gaps = 6/99 (6%)
Query: 437 VQLILKMHQNLINLLILKSIQKLNLVQKLKSLQKLSLIQKQK-LVQ*NRMKMLLRMFKIT 495
+ +LK INLL LK IQ + L+ K SL+ +S+ +K + L++ + + L+ K
Sbjct: 381 INKVLKECNKEINLLYLKKIQAIKLI-KYISLKIVSITEKNRILIEIRKKERKLKKQKKI 439
Query: 496 LSRLFSQNSNTNLHILRS*SLEAKIVLEEQDHISDKKSL 534
+ +Q N N+ E KI+LE+ + + KK +
Sbjct: 440 YGKQINQIENDNIACAE----EIKILLEQLESHNQKKQM 474
>ref|NP_149265.1| Membrane protein [Clostridium acetobutylicum ATCC 824]
gi|14994417|gb|AAK76847.1| Membrane protein [Clostridium
acetobutylicum ATCC 824]
Length = 1058
Score = 35.8 bits (81), Expect = 7.4
Identities = 19/54 (35%), Positives = 26/54 (47%)
Query: 959 NISWKTIRLMLTVFPFFVIILLLSVCQRIQFYIQEPSILRSNTILSETMFKKGF 1012
N +W+ + T +PFF +IL+L+VC FY S I S KGF
Sbjct: 359 NKAWEKAATISTKYPFFALILVLAVCGLSYFYTSSLSFNNLKEIASNYPSVKGF 412
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.359 0.160 0.523
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,236,172,943
Number of Sequences: 2540612
Number of extensions: 40707635
Number of successful extensions: 147131
Number of sequences better than 10.0: 4
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 147119
Number of HSP's gapped (non-prelim): 8
length of query: 1012
length of database: 863,360,394
effective HSP length: 138
effective length of query: 874
effective length of database: 512,755,938
effective search space: 448148689812
effective search space used: 448148689812
T: 11
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 80 (35.4 bits)
Medicago: description of AC148240.10


