

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148219.4 + phase: 0 /pseudo
(1109 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
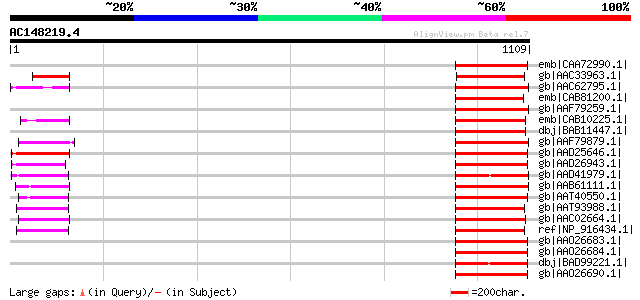
Score E
Sequences producing significant alignments: (bits) Value
emb|CAA72990.1| unnamed protein product [Brassica oleracea] gi|7... 173 3e-41
gb|AAC33963.1| contains similarity to reverse transcriptases (Pf... 173 3e-41
gb|AAC62795.1| contains similarity to retroviral aspartyl protea... 170 2e-40
emb|CAB81200.1| putative retrotransposon polyprotein [Arabidopsi... 167 2e-39
gb|AAF79259.1| F12K21.14 [Arabidopsis thaliana] gi|25301710|pir|... 166 3e-39
emb|CAB10225.1| retrovirus-related like polyprotein [Arabidopsis... 164 2e-38
dbj|BAB11447.1| polyprotein-like [Arabidopsis thaliana] 164 2e-38
gb|AAF79879.1| T7N9.5 [Arabidopsis thaliana] 163 3e-38
gb|AAD25646.1| putative retroelement pol polyprotein [Arabidopsi... 162 6e-38
gb|AAD26943.1| putative retroelement pol polyprotein [Arabidopsi... 159 4e-37
gb|AAD41979.1| putative retroelement pol polyprotein [Arabidopsi... 155 7e-36
gb|AAB61111.1| Strong similarity to Zea mays retrotransposon Hop... 155 7e-36
gb|AAT40550.1| putative receptor kinase [Solanum demissum] 149 7e-34
gb|AAT93988.1| putative polyprotein [Oryza sativa (japonica cult... 145 6e-33
gb|AAC02664.1| polyprotein [Arabidopsis thaliana] 145 7e-33
ref|NP_916434.1| putative gag/pol polyprotein [Oryza sativa (jap... 145 1e-32
gb|AAO26683.1| gag-pol polyprotein [Vitis vinifera] 145 1e-32
gb|AAO26684.1| gag-pol polyprotein [Vitis vinifera] 144 1e-32
dbj|BAD99221.1| polypepetide with reverse transcriptase and RNas... 144 2e-32
gb|AAO26690.1| gag-pol polyprotein [Vitis vinifera] 143 3e-32
>emb|CAA72990.1| unnamed protein product [Brassica oleracea] gi|7488559|pir||T14518
hypothetical protein 2 - wild cabbage transposon Melmoth
Length = 253
Score = 173 bits (438), Expect = 3e-41
Identities = 82/153 (53%), Positives = 111/153 (71%)
Query: 954 FPDVDWAGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWL 1013
F D DWA C+DSRRS G F+G+SLISWR+ KQ T+SRSS+EAEYRAL+ A+ E+ WL
Sbjct: 98 FADADWAACVDSRRSTTGFTMFVGDSLISWRSKKQPTVSRSSAEAEYRALALASCEMMWL 157
Query: 1014 LYLLNDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFLL 1073
LL +LHV + +PVL+ D+ +A++I TNP+FHE TKHIE+ CH VR+K+ G++K L
Sbjct: 158 CILLRELHVASSSVPVLFSDSTAAIYIATNPVFHERTKHIELDCHTVREKIDKGLLKTLH 217
Query: 1074 VSSKDQLADIFTNPLLPQPFSTLLSNLGMLNSY 1106
V ++DQ+ADI T PL P F L S + + N +
Sbjct: 218 VRTEDQVADILTKPLFPYQFEHLKSKMCIRNIF 250
>gb|AAC33963.1| contains similarity to reverse transcriptases (Pfam; rvt.hmm, score:
11.19) [Arabidopsis thaliana] gi|7486705|pir||T01879
hypothetical protein F8M12.17 - Arabidopsis thaliana
Length = 1633
Score = 173 bits (438), Expect = 3e-41
Identities = 83/145 (57%), Positives = 106/145 (72%)
Query: 956 DVDWAGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWLLY 1015
D DW C DSRRS+ G C +LG SLI+W++ KQ +SRSS+E+EYR+L+ AT E+ WL
Sbjct: 1284 DADWGTCKDSRRSVTGFCIYLGTSLITWKSKKQSVVSRSSTESEYRSLAQATCEIIWLQQ 1343
Query: 1016 LLNDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFLLVS 1075
LL DLHVT L+CDN+SALH+ TNP+FHE TKHIEI CH VRD+++AG +K L V
Sbjct: 1344 LLKDLHVTMTCPAKLFCDNKSALHLATNPVFHERTKHIEIDCHTVRDQIKAGKLKTLHVP 1403
Query: 1076 SKDQLADIFTNPLLPQPFSTLLSNL 1100
+ +QLADI T PL P PF +LL +
Sbjct: 1404 TGNQLADILTKPLHPGPFHSLLKRI 1428
Score = 67.8 bits (164), Expect = 2e-09
Identities = 30/79 (37%), Positives = 51/79 (63%)
Query: 50 CDICHFARQKQLPFHLSSSVASNKFELLHFDI*GPLIVPFIHNHKYFLTILDDSSRFVWI 109
C I A+QK+L + +++AS+ F+L+H DI GP + + +YFLT++DD +R W+
Sbjct: 532 CRISPLAKQKRLAYVSHNNLASSPFDLIHLDIWGPFSIESVDGFRYFLTLVDDCTRTTWV 591
Query: 110 VLLKSKAEVSQHVKNFITL 128
++K+K+EVS F+ L
Sbjct: 592 YMMKNKSEVSNIFPVFVKL 610
>gb|AAC62795.1| contains similarity to retroviral aspartyl proteases (Pfam: rvp.hmm,
score: 11.80) [Arabidopsis thaliana]
gi|7487550|pir||T01956 hypothetical protein T2L5.9 -
Arabidopsis thaliana
Length = 1244
Score = 170 bits (431), Expect = 2e-40
Identities = 82/155 (52%), Positives = 112/155 (71%)
Query: 954 FPDVDWAGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWL 1013
F D DW C D+RRS G FLG+SLI+WR+ KQ T+SRSS+EAEYRAL+ A+ E+ WL
Sbjct: 1089 FADSDWGTCPDTRRSTTGLTMFLGSSLITWRSKKQPTVSRSSAEAEYRALALASCEMVWL 1148
Query: 1014 LYLLNDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFLL 1073
LL DL + T +P+++ D+ +A++I TNP+FHE TKHIEI CHLVR++L G+I+ L
Sbjct: 1149 ASLLLDLKIITGSVPIVFSDSTAAIYIATNPVFHERTKHIEIDCHLVRERLDKGLIRMLH 1208
Query: 1074 VSSKDQLADIFTNPLLPQPFSTLLSNLGMLNSYHS 1108
V ++DQ+ADI T PL P FS L+S + + N + S
Sbjct: 1209 VRTEDQVADILTKPLFPHQFSYLMSKMSLHNIFAS 1243
Score = 70.5 bits (171), Expect = 3e-10
Identities = 45/127 (35%), Positives = 67/127 (52%), Gaps = 21/127 (16%)
Query: 2 VNNVSVNKLIAILSFAL*HFRLGHVSYERLAHMSQLYPF*LFFDS-NATCDICHFARQKQ 60
+++VSVN A++ +L H R+GH +Y RL +S + + +A C ICH A+QK+
Sbjct: 533 LDSVSVN---AVVDVSLWHMRMGHPAYSRLDAISDILRTTKHKNKGSAYCHICHLAKQKK 589
Query: 61 LPFHLSSSVASNKFELLHFDI*GPLIVPFIHNHKYFLTILDDSSRFVWIVLLKSKAEVSQ 120
L F S+++ I KYFLTI+DD SR W+ LLK+K+EV
Sbjct: 590 LSFQSSNNILET-----------------IDGFKYFLTIVDDHSRATWVYLLKTKSEVLS 632
Query: 121 HVKNFIT 127
F+T
Sbjct: 633 VFPAFVT 639
>emb|CAB81200.1| putative retrotransposon polyprotein [Arabidopsis thaliana]
gi|4539373|emb|CAB40067.1| putative retrotransposon
polyprotein [Arabidopsis thaliana] gi|7486142|pir||T04294
hypothetical protein F25I24.200 - Arabidopsis thaliana
Length = 1203
Score = 167 bits (423), Expect = 2e-39
Identities = 83/146 (56%), Positives = 105/146 (71%), Gaps = 2/146 (1%)
Query: 954 FPDVDWAGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWL 1013
F D DW C DSRRS+ G C +LG SLI+W++ KQ +SRSS+E+EYR+L+ AT E+ WL
Sbjct: 884 FSDADWGTCKDSRRSVTGFCIYLGTSLITWKSKKQSVVSRSSTESEYRSLAQATCEIIWL 943
Query: 1014 LYLLNDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFLL 1073
LL DLHVT L+CDN+SALH+ TNP+FHE TKHIEI CH VRD+++AG +K L
Sbjct: 944 QQLLKDLHVTMTCPAKLFCDNKSALHLATNPVFHERTKHIEIDCHTVRDQIKAGKLKTLH 1003
Query: 1074 VSSKDQLADIFTNPLLP--QPFSTLL 1097
V + +QLADI T PL P P +LL
Sbjct: 1004 VPTGNQLADILTKPLHPVQSPIFSLL 1029
Score = 38.1 bits (87), Expect = 1.7
Identities = 15/38 (39%), Positives = 26/38 (67%)
Query: 91 HNHKYFLTILDDSSRFVWIVLLKSKAEVSQHVKNFITL 128
+ + YFLT++DD +R W+ ++K+K+EVS F+ L
Sbjct: 248 YRNLYFLTLVDDCTRTTWVYMMKNKSEVSNIFPVFVKL 285
>gb|AAF79259.1| F12K21.14 [Arabidopsis thaliana] gi|25301710|pir||C86469 protein
F12K21.14 [imported] - Arabidopsis thaliana
Length = 427
Score = 166 bits (421), Expect = 3e-39
Identities = 85/155 (54%), Positives = 108/155 (68%), Gaps = 1/155 (0%)
Query: 954 FPDVDWAGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWL 1013
+ D DW C DSRRS G F+G+SLISWR+ KQ T+SRSS+EAEYRAL+ A+ E+ WL
Sbjct: 273 YTDADWGTCPDSRRSTTGFTMFVGSSLISWRSKKQPTVSRSSAEAEYRALALASCEMAWL 332
Query: 1014 LYLLNDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFLL 1073
LL L V T +P+LY D+ +A++I NP+FHE TKHIEI CH VRDKL G +K L
Sbjct: 333 FTLLIALRVAT-SVPILYSDSTAAVYIAINPVFHERTKHIEIDCHTVRDKLDNGQLKLLH 391
Query: 1074 VSSKDQLADIFTNPLLPQPFSTLLSNLGMLNSYHS 1108
V ++DQ+ADI T PL P F+ LLS + + N Y S
Sbjct: 392 VKTEDQVADILTKPLFPHQFTHLLSKMSLHNIYAS 426
>emb|CAB10225.1| retrovirus-related like polyprotein [Arabidopsis thaliana]
gi|7268152|emb|CAB78488.1| retrovirus-related like
polyprotein [Arabidopsis thaliana] gi|7488175|pir||G71406
probable retrovirus-related polyprotein - Arabidopsis
thaliana
Length = 1489
Score = 164 bits (414), Expect = 2e-38
Identities = 80/149 (53%), Positives = 103/149 (68%)
Query: 954 FPDVDWAGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWL 1013
F D DWA C D+RRSI G C +LG SLISW++ KQ SRSS+E+EYR+++ AT E+ WL
Sbjct: 1329 FSDADWAACKDTRRSISGFCIYLGTSLISWKSKKQAVASRSSTESEYRSMAQATCEIIWL 1388
Query: 1014 LYLLNDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFLL 1073
LL DLH+ L+CDN+SALH NP+FHE TKHIEI CH VRD+++AG +K L
Sbjct: 1389 QQLLKDLHIPLTCPAKLFCDNKSALHSSLNPVFHERTKHIEIDCHTVRDQIKAGNLKALH 1448
Query: 1074 VSSKDQLADIFTNPLLPQPFSTLLSNLGM 1102
V +++Q ADI T L P PF LL + +
Sbjct: 1449 VPTENQHADILTKALHPGPFHHLLRQMSL 1477
Score = 65.1 bits (157), Expect = 1e-08
Identities = 34/105 (32%), Positives = 56/105 (52%), Gaps = 16/105 (15%)
Query: 24 GHVSYERLAHMSQLYPF*LFFDSNATCDICHFARQKQLPFHLSSSVASNKFELLHFDI*G 83
GH+ ++RL H S + + K+L + +++ASN F+L+H DI G
Sbjct: 599 GHLWHQRLGHPSSVV----------------LQKLKRLAYISHNNLASNPFDLVHLDIWG 642
Query: 84 PLIVPFIHNHKYFLTILDDSSRFVWIVLLKSKAEVSQHVKNFITL 128
P + I +YFLT++DD +R W+ +L++K +VS FI L
Sbjct: 643 PFSIESIEGFRYFLTVVDDCTRTTWVYMLRNKKDVSSVFPEFIKL 687
>dbj|BAB11447.1| polyprotein-like [Arabidopsis thaliana]
Length = 509
Score = 164 bits (414), Expect = 2e-38
Identities = 78/149 (52%), Positives = 102/149 (68%)
Query: 954 FPDVDWAGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWL 1013
F D DW CLDSRRS+ G C FLG SLI+W++ KQ S SS+EAEYR+++ AT EL WL
Sbjct: 345 FADADWGTCLDSRRSVSGVCVFLGTSLITWKSKKQEVASGSSTEAEYRSMAVATKELLWL 404
Query: 1014 LYLLNDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFLL 1073
+L DLHV L+CDN+SA+HI N +FHE TKH+EI CH RD+++ G +K L
Sbjct: 405 AQMLKDLHVEMEFQVKLFCDNKSAMHIANNSVFHERTKHVEIDCHTTRDRVKNGFLKVLH 464
Query: 1074 VSSKDQLADIFTNPLLPQPFSTLLSNLGM 1102
V +++QLADI T L P PF ++L L +
Sbjct: 465 VDTENQLADILTKALQPGPFRSILGRLSV 493
>gb|AAF79879.1| T7N9.5 [Arabidopsis thaliana]
Length = 1436
Score = 163 bits (412), Expect = 3e-38
Identities = 80/155 (51%), Positives = 103/155 (65%)
Query: 954 FPDVDWAGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWL 1013
F D DW C D+RR + G F+GNSL+SWR+ KQ +S SS+EAEYRA+S AT EL WL
Sbjct: 1280 FSDSDWQTCPDTRRCVTGFAIFVGNSLVSWRSKKQDVVSMSSAEAEYRAMSVATKELIWL 1339
Query: 1014 LYLLNDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFLL 1073
Y+L + LYCDN++ALHI N +FHE TKHIE CH VR+ ++AGI+K +
Sbjct: 1340 GYILTAFKIPFTHPAYLYCDNEAALHIANNSVFHERTKHIENDCHKVRECIEAGILKTIF 1399
Query: 1074 VSSKDQLADIFTNPLLPQPFSTLLSNLGMLNSYHS 1108
V + +QLAD T PL P+PF S LG+LN Y +
Sbjct: 1400 VRTDNQLADTLTKPLYPKPFRENNSKLGLLNIYEA 1434
Score = 86.7 bits (213), Expect = 4e-15
Identities = 44/121 (36%), Positives = 69/121 (56%), Gaps = 4/121 (3%)
Query: 20 HFRLGHVSYERLAHMSQ--LYPF*LFFDSNATCDICHFARQKQLPFHLSSSVASNKFELL 77
H RLGH S+ ++ +S + P ++ C +CH ++QK LPF + + FEL+
Sbjct: 559 HKRLGHPSFAKIDTLSDVLMLPKQKINKDSSHCHVCHLSKQKHLPFKSVNHIREKAFELV 618
Query: 78 HFDI*GPLIVPFIHNHKYFLTILDDSSRFVWIVLLKSKAEVSQHVKNFITLC*KSISYHS 137
H D GP VP + +++YFLTI+DD SR WI LLK K++V +F+ + YH+
Sbjct: 619 HIDTWGPFSVPTVDSYRYFLTIVDDFSRATWIYLLKQKSDVLTVFPSFLKMV--ETQYHT 676
Query: 138 Q 138
+
Sbjct: 677 K 677
>gb|AAD25646.1| putative retroelement pol polyprotein [Arabidopsis thaliana]
gi|25301701|pir||E84589 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana
Length = 1461
Score = 162 bits (410), Expect = 6e-38
Identities = 83/153 (54%), Positives = 108/153 (70%), Gaps = 1/153 (0%)
Query: 954 FPDVDWAGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWL 1013
+ D DW C DSRRS G F+G+SLISWR+ KQ T+SRSS+EAEYRAL+ A+ E+ WL
Sbjct: 1306 YTDADWGTCPDSRRSTTGFTMFVGSSLISWRSKKQPTVSRSSAEAEYRALALASCEMAWL 1365
Query: 1014 LYLLNDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFLL 1073
LL L V + +P+LY D+ +A++I TNP+FHE TKHIEI CH VR+KL G +K L
Sbjct: 1366 STLLLALRVHS-GVPILYSDSTAAVYIATNPVFHERTKHIEIDCHTVREKLDNGQLKLLH 1424
Query: 1074 VSSKDQLADIFTNPLLPQPFSTLLSNLGMLNSY 1106
V +KDQ+ADI T PL P F+ LLS + + N +
Sbjct: 1425 VKTKDQVADILTKPLFPYQFAHLLSKMSIQNIF 1457
Score = 95.1 bits (235), Expect = 1e-17
Identities = 52/125 (41%), Positives = 78/125 (61%), Gaps = 4/125 (3%)
Query: 5 VSVNKLIAILSFAL*HFRLGHVSYERLAHMSQLYPF*LFFDS-NATCDICHFARQKQLPF 63
+SVN A++ ++ H RLGH S+ RL +S++ + +A C +CH A+QK+L F
Sbjct: 558 ISVN---AVVDVSVWHKRLGHPSFSRLDSLSEVLGTTRHKNKKSAYCHVCHLAKQKKLSF 614
Query: 64 HLSSSVASNKFELLHFDI*GPLIVPFIHNHKYFLTILDDSSRFVWIVLLKSKAEVSQHVK 123
++++ ++ FELLH D+ GP V + +KYFLTI+DD SR WI LLKSK++V
Sbjct: 615 PSANNICNSTFELLHIDVWGPFSVETVEGYKYFLTIVDDHSRATWIYLLKSKSDVLTVFP 674
Query: 124 NFITL 128
FI L
Sbjct: 675 AFIDL 679
>gb|AAD26943.1| putative retroelement pol polyprotein [Arabidopsis thaliana]
gi|25301694|pir||E84535 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana
Length = 1454
Score = 159 bits (403), Expect = 4e-37
Identities = 80/153 (52%), Positives = 106/153 (68%), Gaps = 1/153 (0%)
Query: 954 FPDVDWAGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWL 1013
F D DWA C DSRRS F+G+SLISWR+ KQ T+SRSS+EAEYRAL+ AT E+ WL
Sbjct: 1299 FADSDWASCQDSRRSTTSFTMFVGDSLISWRSKKQHTVSRSSAEAEYRALALATCEMVWL 1358
Query: 1014 LYLLNDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFLL 1073
LL L + +P+LY D+ +A++I TNP+FHE TKHI++ CH VR++L G +K L
Sbjct: 1359 FTLLVSLQASP-PVPILYSDSTAAIYIATNPVFHERTKHIKLDCHTVRERLDNGELKLLH 1417
Query: 1074 VSSKDQLADIFTNPLLPQPFSTLLSNLGMLNSY 1106
V ++DQ+ADI T PL P F L S + +LN +
Sbjct: 1418 VRTEDQVADILTKPLFPYQFEHLKSKMSILNIF 1450
Score = 87.8 bits (216), Expect = 2e-15
Identities = 47/116 (40%), Positives = 70/116 (59%), Gaps = 4/116 (3%)
Query: 4 NVSVNKLIAILSFAL*HFRLGHVSYERLAHMSQLYPF*LFFDSNAT-CDICHFARQKQLP 62
++SVN A++ ++ H RLGH S +RL +S + + C +CH A+Q++L
Sbjct: 546 SISVN---AVVDISMWHRRLGHASLQRLDAISDSLGTTRHKNKGSDFCHVCHLAKQRKLS 602
Query: 63 FHLSSSVASNKFELLHFDI*GPLIVPFIHNHKYFLTILDDSSRFVWIVLLKSKAEV 118
F S+ V F+LLH D+ GP V + +KYFLTI+DD SR W+ LLK+K+EV
Sbjct: 603 FPTSNKVCKEIFDLLHIDVWGPFSVETVEGYKYFLTIVDDHSRATWMYLLKTKSEV 658
>gb|AAD41979.1| putative retroelement pol polyprotein [Arabidopsis thaliana]
gi|25301676|pir||B84534 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana
Length = 1264
Score = 155 bits (392), Expect = 7e-36
Identities = 81/155 (52%), Positives = 104/155 (66%), Gaps = 1/155 (0%)
Query: 954 FPDVDWAGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWL 1013
F D DWA C DSR S G F+G+SLIS R+ KQ +SRSS+EAEYRAL+ AT EL WL
Sbjct: 1109 FADSDWASCPDSRHSTTGFTMFVGDSLISLRSKKQHVVSRSSAEAEYRALALATCELVWL 1168
Query: 1014 LYLLNDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFLL 1073
LL L T +P+L+ D+ +A++I NP+FHE TKHIEI CH VR+K+ G +K L
Sbjct: 1169 HTLLASLTAATT-IPILFSDSTAAIYIAINPVFHERTKHIEIDCHTVREKIDDGELKLLH 1227
Query: 1074 VSSKDQLADIFTNPLLPQPFSTLLSNLGMLNSYHS 1108
V ++DQ+ADI T PL P F L S + +LN + S
Sbjct: 1228 VRTEDQVADIMTKPLFPNQFEHLKSKMSILNIFES 1262
Score = 84.7 bits (208), Expect = 2e-14
Identities = 49/124 (39%), Positives = 73/124 (58%), Gaps = 4/124 (3%)
Query: 4 NVSVNKLIAILSFAL*HFRLGHVSYERLAHMSQLYPF*LFFDSNAT-CDICHFARQKQLP 62
NVSVN ++ I ++ H RL H S +RL +S+ + + C +CH A+ ++L
Sbjct: 405 NVSVNAVVDISTW---HNRLRHASLQRLDVISESLGTTKHKNKGSDYCHVCHLAKHRKLS 461
Query: 63 FHLSSSVASNKFELLHFDI*GPLIVPFIHNHKYFLTILDDSSRFVWIVLLKSKAEVSQHV 122
F ++V + FE+LH DI GP V + ++YFLTI+DD SR WI LLK+K+EV
Sbjct: 462 FPSQNNVCNEIFEMLHIDIWGPFSVETVDGYQYFLTIVDDHSRATWIYLLKTKSEVLTIF 521
Query: 123 KNFI 126
+FI
Sbjct: 522 HDFI 525
>gb|AAB61111.1| Strong similarity to Zea mays retrotransposon Hopscotch polyprotein
(gb|U12626). [Arabidopsis thaliana]
gi|25301690|pir||G96722 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana
Length = 1315
Score = 155 bits (392), Expect = 7e-36
Identities = 75/155 (48%), Positives = 105/155 (67%)
Query: 954 FPDVDWAGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWL 1013
+ + D+ C DSRRS G C FLG+SLI W++ KQ +S+SS+EAEYR+LS AT EL WL
Sbjct: 1160 YANADYNSCRDSRRSTSGYCMFLGDSLICWKSRKQDVVSKSSAEAEYRSLSVATDELVWL 1219
Query: 1014 LYLLNDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFLL 1073
L +L V K +L+CDN++A+HI N +FHE TKHIE CH VR++L G+ +
Sbjct: 1220 TNFLKELQVPLSKPTLLFCDNEAAIHIANNHVFHERTKHIESDCHSVRERLLKGLFELYH 1279
Query: 1074 VSSKDQLADIFTNPLLPQPFSTLLSNLGMLNSYHS 1108
++++ Q+AD FT PL P F L+S +G+LN + S
Sbjct: 1280 INTELQIADPFTKPLYPSHFHRLISKMGLLNIFVS 1314
Score = 86.7 bits (213), Expect = 4e-15
Identities = 47/120 (39%), Positives = 70/120 (58%), Gaps = 5/120 (4%)
Query: 12 AILSFAL*HFRLGHVSYERLAHMSQLYPF*LFFDSNAT---CDICHFARQKQLPFHLSSS 68
++ S L H RLGH S ++L MS L F N T C +CH ++QK LPF ++
Sbjct: 400 SVTSHDLWHKRLGHPSVQKLQPMSSLLSFPK--QKNNTDFHCRVCHISKQKHLPFVSHNN 457
Query: 69 VASNKFELLHFDI*GPLIVPFIHNHKYFLTILDDSSRFVWIVLLKSKAEVSQHVKNFITL 128
+S F+L+H D GP V ++YFLTI+DD SR W+ LL++K++V + F+T+
Sbjct: 458 KSSRPFDLIHIDTWGPFSVQTHDGYRYFLTIVDDYSRATWVYLLRNKSDVLTVIPTFVTM 517
>gb|AAT40550.1| putative receptor kinase [Solanum demissum]
Length = 1358
Score = 149 bits (375), Expect = 7e-34
Identities = 74/153 (48%), Positives = 98/153 (63%)
Query: 954 FPDVDWAGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWL 1013
+ D DWAG RRS G C +G +L+SW++ KQ ++RSS+E+EYRA++ AT EL W+
Sbjct: 1203 YTDADWAGSPSDRRSTSGYCVLVGGNLVSWKSKKQNVVARSSAESEYRAMATATCELVWI 1262
Query: 1014 LYLLNDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFLL 1073
LL +L V L CDNQ+ALHI +NP+FHE TKHIEI CH VR+K+ +G I
Sbjct: 1263 KQLLGELKFGKVDKMELVCDNQAALHIASNPVFHERTKHIEIDCHFVREKILSGDIVTKF 1322
Query: 1074 VSSKDQLADIFTNPLLPQPFSTLLSNLGMLNSY 1106
V S DQLADIFT L + + + LG + Y
Sbjct: 1323 VKSNDQLADIFTKSLTCPRINYICNKLGTYDLY 1355
Score = 54.3 bits (129), Expect = 2e-05
Identities = 34/107 (31%), Positives = 51/107 (46%), Gaps = 5/107 (4%)
Query: 20 HFRLGHVSYERLAHMSQLYPF*LFFDSNATCDICHFARQKQLPFHLSSSVASNK-FELLH 78
H RLGH S +L M L S C+ C + + F S+ S F L+H
Sbjct: 489 HKRLGHSSLSKLQKMVPS----LSSLSTLDCESCQLGKHTRATFSRSTEGRSESIFSLVH 544
Query: 79 FDI*GPLIVPFIHNHKYFLTILDDSSRFVWIVLLKSKAEVSQHVKNF 125
DI GP V +YF++ +DD S+ W+ L+K ++E+ K+F
Sbjct: 545 SDIWGPSRVSSTLGFRYFVSFIDDYSKCTWVFLMKDRSELFSIFKSF 591
>gb|AAT93988.1| putative polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1480
Score = 145 bits (367), Expect = 6e-33
Identities = 71/147 (48%), Positives = 97/147 (65%)
Query: 954 FPDVDWAGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWL 1013
+ D DWAGC DS RS G C FLG++L+SW + +Q T+SRSS+EAEYRA++ A E WL
Sbjct: 1321 YSDADWAGCPDSHRSTSGYCVFLGDNLVSWSSKRQTTVSRSSAEAEYRAVAHAIAECCWL 1380
Query: 1014 LYLLNDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFLL 1073
LL++LH++ V+YCDN SA+++ NP+ H TKHIEI H VR+K+ G ++ L
Sbjct: 1381 RQLLSELHISLTSATVVYCDNVSAVYMAANPVHHRRTKHIEIDIHFVREKVALGQVRVLH 1440
Query: 1074 VSSKDQLADIFTNPLLPQPFSTLLSNL 1100
+ S Q ADI T L Q F+ S+L
Sbjct: 1441 LPSSHQFADIMTKGLPVQLFTEFRSSL 1467
Score = 68.6 bits (166), Expect = 1e-09
Identities = 38/112 (33%), Positives = 58/112 (50%), Gaps = 1/112 (0%)
Query: 15 SFAL*HFRLGHVSYERLAHMSQLYPF*LFFDSNATCDICHFARQKQLPFHLSSSVASNKF 74
S L H RLGH S + + +L +N C CH + +L F SSS S+ F
Sbjct: 486 SSTLWHHRLGHPSPAAVQTLHKLAILSCTRSNNKLCHACHLGKHTRLSFSKSSSSTSSPF 545
Query: 75 ELLHFDI*GPLIVPFIHNHKYFLTILDDSSRFVWIVLLKSKAEVSQHVKNFI 126
EL+H D+ ++ + KY+L +LDD + F W L+ K++V QH+ F+
Sbjct: 546 ELVHCDVWTSPVLS-LSGFKYYLVVLDDFTHFCWTFPLRHKSDVHQHLLEFV 596
>gb|AAC02664.1| polyprotein [Arabidopsis thaliana]
Length = 1451
Score = 145 bits (366), Expect = 7e-33
Identities = 69/149 (46%), Positives = 100/149 (66%)
Query: 954 FPDVDWAGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWL 1013
+ D DWAG +D+ S +LG++ ISW + KQ ++RSS+EAEYRA++ AT E++W+
Sbjct: 1297 YSDADWAGDIDNYNSTNAYILYLGSNPISWSSKKQKGVARSSTEAEYRAVANATSEIRWV 1356
Query: 1014 LYLLNDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFLL 1073
LL +L +T PV+YCDN A ++ NP+FH KHI + H VR+ +QAG ++
Sbjct: 1357 CSLLTELGITLSSPPVVYCDNVGATYLSANPVFHSRMKHIALDFHFVRESVQAGALRVTH 1416
Query: 1074 VSSKDQLADIFTNPLLPQPFSTLLSNLGM 1102
VS+KDQLAD T PL QPF+TL+S +G+
Sbjct: 1417 VSTKDQLADALTKPLPRQPFTTLISKIGV 1445
Score = 52.4 bits (124), Expect = 8e-05
Identities = 32/111 (28%), Positives = 54/111 (47%), Gaps = 3/111 (2%)
Query: 20 HFRLGHVSYERLAHMSQLY--PF*LFFDSNATCDICHFARQKQLPFHLSSSVASNKFELL 77
H RLGH S L + + P + C C + +LPF+ ++ +S+ E L
Sbjct: 470 HARLGHPSLPILKALISKFSLPISHSLQNQLLCSDCSINKSHKLPFYSNTIASSHPLEYL 529
Query: 78 HFDI*GPLIVPFIHNHKYFLTILDDSSRFVWIVLLKSKAEVSQHVKNFITL 128
+ D+ I I N+KY+L I+D +R+ W+ L+ K++V + F L
Sbjct: 530 YTDVWTSPITS-IDNYKYYLVIVDHYTRYTWLYPLRKKSQVREMFITFTAL 579
>ref|NP_916434.1| putative gag/pol polyprotein [Oryza sativa (japonica cultivar-group)]
Length = 1090
Score = 145 bits (365), Expect = 1e-32
Identities = 71/147 (48%), Positives = 96/147 (65%)
Query: 954 FPDVDWAGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWL 1013
+ D DWAGC +SRRS G C +LG++L+SW + +Q T+SRSS+EAEYRA++ A E WL
Sbjct: 919 YSDADWAGCPNSRRSTSGYCVYLGDNLVSWSSKRQTTVSRSSAEAEYRAVAHAVAECCWL 978
Query: 1014 LYLLNDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFLL 1073
LL +LHV ++YCDN SA+++ NP+ H TKHIEI H VR+K+ G ++ L
Sbjct: 979 RQLLQELHVPIASATIVYCDNVSAVYMTANPVHHRRTKHIEIDIHFVREKVALGQVRVLY 1038
Query: 1074 VSSKDQLADIFTNPLLPQPFSTLLSNL 1100
V S Q ADI T L Q F+ S+L
Sbjct: 1039 VPSSHQFADIMTKGLPVQLFTDFRSSL 1065
Score = 65.5 bits (158), Expect = 1e-08
Identities = 37/112 (33%), Positives = 55/112 (49%), Gaps = 1/112 (0%)
Query: 15 SFAL*HFRLGHVSYERLAHMSQLYPF*LFFDSNATCDICHFARQKQLPFHLSSSVASNKF 74
S L H RLGH + + + + C C + +LPFH SSS S F
Sbjct: 121 SSTLWHCRLGHPGPAAIHGLRNIASISCNKIDTSLCHACQLGKHTRLPFHNSSSRTSVPF 180
Query: 75 ELLHFDI*GPLIVPFIHNHKYFLTILDDSSRFVWIVLLKSKAEVSQHVKNFI 126
EL+H D+ ++ KY+L +LDD S F W LL+ K++V +H+ F+
Sbjct: 181 ELVHCDVWTSPVMS-TSGFKYYLVVLDDFSHFCWTFLLRLKSDVHRHIVEFV 231
>gb|AAO26683.1| gag-pol polyprotein [Vitis vinifera]
Length = 326
Score = 145 bits (365), Expect = 1e-32
Identities = 68/153 (44%), Positives = 97/153 (62%)
Query: 954 FPDVDWAGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWL 1013
F D DWAG RRS G C F G +L++W++ KQ +SRSS+E+EYRA+S AT E+ W+
Sbjct: 171 FSDADWAGSKFDRRSTTGYCVFFGGNLVAWKSKKQSVVSRSSAESEYRAMSRATCEIIWI 230
Query: 1014 LYLLNDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFLL 1073
LL ++ + L+CDNQ+ALHI NP++HE TKHIE+ CH +R+K++ ++
Sbjct: 231 HQLLCEVGMKCTMPAKLWCDNQAALHIAANPVYHERTKHIEVDCHFIREKIEENLVSTGY 290
Query: 1074 VSSKDQLADIFTNPLLPQPFSTLLSNLGMLNSY 1106
V + +QL DIFT L + LGM+N Y
Sbjct: 291 VKTGEQLGDIFTKALNGTRVECFCNKLGMINIY 323
>gb|AAO26684.1| gag-pol polyprotein [Vitis vinifera]
Length = 326
Score = 144 bits (364), Expect = 1e-32
Identities = 68/153 (44%), Positives = 97/153 (62%)
Query: 954 FPDVDWAGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWL 1013
F D DWAG RRS G C F G +L++W++ KQ +SRSS+E+EYRA+S AT E+ W+
Sbjct: 171 FSDADWAGSKFDRRSTTGYCVFFGGNLVAWKSKKQSVVSRSSAESEYRAMSQATCEIIWI 230
Query: 1014 LYLLNDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFLL 1073
LL ++ + L+CDNQ+ALHI NP++HE TKHIE+ CH +R+K++ ++
Sbjct: 231 HQLLCEVGMKCTMPAKLWCDNQAALHIAANPVYHERTKHIEVDCHFIREKIEENLVSTGY 290
Query: 1074 VSSKDQLADIFTNPLLPQPFSTLLSNLGMLNSY 1106
V + +QL DIFT L + LGM+N Y
Sbjct: 291 VKTGEQLGDIFTKALNGTRVEYFCNKLGMINIY 323
>dbj|BAD99221.1| polypepetide with reverse transcriptase and RNaseH domains [Petunia x
hybrida]
Length = 389
Score = 144 bits (363), Expect = 2e-32
Identities = 76/153 (49%), Positives = 98/153 (63%), Gaps = 2/153 (1%)
Query: 954 FPDVDWAGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWL 1013
F D DWA C DSRRSI G +G +SW++ KQ +S SS+EAEYR+L E+ W+
Sbjct: 231 FCDADWASCPDSRRSISGFFLTMGGCPLSWKSKKQQVVSLSSAEAEYRSLRRLVAEIAWI 290
Query: 1014 LYLLNDLHVTTVKLPV-LYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFL 1072
+ LL+DL V LPV ++CDNQSA+HI NP+FHE TKHIE+ CH VR+KL G+I
Sbjct: 291 VRLLHDLSVD-FSLPVSVHCDNQSAIHIAKNPVFHERTKHIELDCHFVREKLLDGLISLS 349
Query: 1073 LVSSKDQLADIFTNPLLPQPFSTLLSNLGMLNS 1105
V S Q+ADIFT L +L+ LG+ S
Sbjct: 350 FVPSSSQIADIFTKSLTGPLHRSLMGKLGVRKS 382
>gb|AAO26690.1| gag-pol polyprotein [Vitis vinifera]
Length = 326
Score = 143 bits (361), Expect = 3e-32
Identities = 67/153 (43%), Positives = 97/153 (62%)
Query: 954 FPDVDWAGCLDSRRSIFGQCFFLGNSLISWRTMKQLTISRSSSEAEYRALSAATYELQWL 1013
F D DWAG RRS G C F G +L++W++ KQ +SRSS+E+EYRA++ AT E+ W+
Sbjct: 171 FSDADWAGSKFDRRSTTGYCVFFGGNLVAWKSKKQSVVSRSSAESEYRAMAQATCEIIWI 230
Query: 1014 LYLLNDLHVTTVKLPVLYCDNQSALHIGTNPMFHEITKHIEIICHLVRDKLQAGIIKFLL 1073
LL ++ + L+CDNQ+ALHI NP++HE TKHIE+ CH +R+K++ ++
Sbjct: 231 HQLLCEVGMKCTMPAKLWCDNQAALHIAANPVYHERTKHIEVDCHFIREKIEENLVSTDY 290
Query: 1074 VSSKDQLADIFTNPLLPQPFSTLLSNLGMLNSY 1106
V + +QL DIFT L + LGM+N Y
Sbjct: 291 VKTGEQLGDIFTKALNGTRVEYFCNKLGMINIY 323
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.362 0.160 0.603
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,560,096,680
Number of Sequences: 2540612
Number of extensions: 57317876
Number of successful extensions: 247618
Number of sequences better than 10.0: 846
Number of HSP's better than 10.0 without gapping: 714
Number of HSP's successfully gapped in prelim test: 132
Number of HSP's that attempted gapping in prelim test: 245271
Number of HSP's gapped (non-prelim): 1785
length of query: 1109
length of database: 863,360,394
effective HSP length: 139
effective length of query: 970
effective length of database: 510,215,326
effective search space: 494908866220
effective search space used: 494908866220
T: 11
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (22.0 bits)
S2: 81 (35.8 bits)
Medicago: description of AC148219.4


