

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147714.16 - phase: 0
(322 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
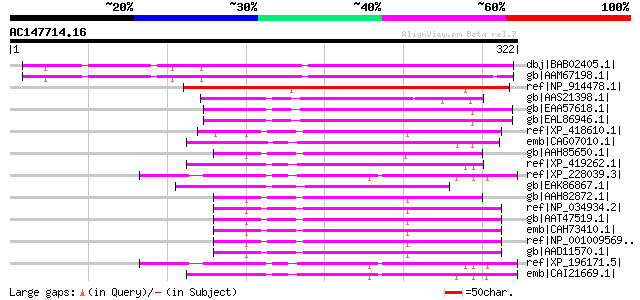
Score E
Sequences producing significant alignments: (bits) Value
dbj|BAB02405.1| unnamed protein product [Arabidopsis thaliana] g... 310 5e-83
gb|AAM67198.1| similar to zinc-finger protein [Arabidopsis thali... 305 1e-81
ref|NP_914478.1| P0489A01.16 [Oryza sativa (japonica cultivar-gr... 261 2e-68
gb|AAS21398.1| Brpf1 protein-like protein [Oikopleura dioica] 154 3e-36
gb|EAA57618.1| hypothetical protein AN6675.2 [Aspergillus nidula... 146 7e-34
gb|EAL86946.1| PHD finger domain protein, putative [Aspergillus ... 143 6e-33
ref|XP_418610.1| PREDICTED: similar to AF-10 protein [Gallus gal... 139 9e-32
emb|CAG07010.1| unnamed protein product [Tetraodon nigroviridis] 139 1e-31
gb|AAH85650.1| Zgc:92208 [Danio rerio] gi|55925532|ref|NP_001007... 138 2e-31
ref|XP_419262.1| PREDICTED: similar to Bromodomain and PHD finge... 138 2e-31
ref|XP_228039.3| PREDICTED: similar to BRPF3 [Rattus norvegicus] 137 3e-31
gb|EAK86867.1| hypothetical protein UM06029.1 [Ustilago maydis 5... 137 3e-31
gb|AAH82872.1| LOC494767 protein [Xenopus laevis] 137 3e-31
ref|NP_034934.2| myeloid/lymphoid or mixed lineage-leukemia tran... 137 4e-31
gb|AAT47519.1| AF10 [Homo sapiens] gi|55664632|emb|CAH73411.1| m... 137 4e-31
emb|CAH73410.1| myeloid\/lymphoid or mixed-lineage leukemia (tri... 137 4e-31
ref|NP_001009569.1| myeloid/lymphoid or mixed-lineage leukemia t... 137 4e-31
gb|AAD11570.1| maf10 [Mus musculus] gi|12229633|sp|O54826|AF10_M... 137 4e-31
ref|XP_196171.5| PREDICTED: bromodomain and PHD finger containin... 137 4e-31
emb|CAI21669.1| BRPF3 [Homo sapiens] 137 6e-31
>dbj|BAB02405.1| unnamed protein product [Arabidopsis thaliana]
gi|42572435|ref|NP_974313.1| PHD finger family protein
[Arabidopsis thaliana]
Length = 343
Score = 310 bits (793), Expect = 5e-83
Identities = 160/343 (46%), Positives = 208/343 (59%), Gaps = 40/343 (11%)
Query: 9 DLPPNKRLRFIYQ-----QQQQQEEQDLSHCSLLPTKKRKESRNSSLFHTPPSSPTPPPS 63
DLPP KRLR + + QQ Q Q LP KKRK++R + + + P+
Sbjct: 7 DLPPLKRLRLLQRDLEAAQQHQLPNQPEMKSLQLPAKKRKQTR---VDYDDDDAENSNPT 63
Query: 64 TYSLPTKKRITALQPHLHHHNNIPNDAVPLIDLNVEYSP-------------SLPSATPI 110
+ LP KKRI A+ P L P DLNVEY P ++ S+ +
Sbjct: 64 YHCLPAKKRIWAIDPDLLSGGGNPFSP---FDLNVEYKPYVEEKSIEKKSTLNVESSLEV 120
Query: 111 EKQSQKQDIE------------FEVDDDE-ILCCVCHSTDANAEDPIVFCDGCNLMVHAS 157
E+ K++I+ EV+D++ I+C VC STD + +PIVFCDGC+LMVHAS
Sbjct: 121 EEDDDKENIDPLGKGKALDLSDREVEDEDGIMCAVCQSTDGDPLNPIVFCDGCDLMVHAS 180
Query: 158 CYGNPLVKQIPDGDWFCDQCRFKNDIDTDTGPIRCSLCPTKEGAMKQTTDGKWVHLVCAL 217
CYGNPLVK IP+GDWFC QC + + C LC TK GAMK T DG+W H+ CAL
Sbjct: 181 CYGNPLVKAIPEGDWFCRQCLSSKNREKI---FSCCLCTTKGGAMKPTNDGRWAHITCAL 237
Query: 218 LVPEVFFVDPEGREGIDCSKIPKKRWLEKCYVCGCFDGCALVCSEQKCGLGFHITCGIKE 277
VPEV+F DPEGREGI CS++ KRW ++CY+C GC + CSE +C L FH+TCG+KE
Sbjct: 238 FVPEVYFEDPEGREGICCSEVLSKRWKDRCYLCKVRRGCVIECSEMRCKLAFHVTCGLKE 297
Query: 278 DLCIEYKEGKKGATVVAGFCKTHSQIWEKNKGSGKYKIVAVED 320
DLCIEY+EGKK +V GFC H+++WE+ + SGKYKIVA E+
Sbjct: 298 DLCIEYREGKKSGGIVVGFCNEHTKLWERQQESGKYKIVAREE 340
>gb|AAM67198.1| similar to zinc-finger protein [Arabidopsis thaliana]
gi|18400507|ref|NP_566491.1| PHD finger family protein
[Arabidopsis thaliana]
Length = 341
Score = 305 bits (781), Expect = 1e-81
Identities = 160/343 (46%), Positives = 207/343 (59%), Gaps = 42/343 (12%)
Query: 9 DLPPNKRLRFIYQ-----QQQQQEEQDLSHCSLLPTKKRKESRNSSLFHTPPSSPTPPPS 63
DLPP KRLR + + QQ Q Q LP KKRK++R + + + P+
Sbjct: 7 DLPPLKRLRLLQRDLEAAQQHQLPNQPEMKSLQLPAKKRKQTR---VDYDDDDAENSNPT 63
Query: 64 TYSLPTKKRITALQPHLHHHNNIPNDAVPLIDLNVEYSP-------------SLPSATPI 110
+ LP KKRI A+ P L P DLNVEY P ++ S+ +
Sbjct: 64 YHCLPAKKRIWAIDPDLLSGGGNPFSP---FDLNVEYKPYVEEKSIEKKSTLNVESSLEV 120
Query: 111 EKQSQKQDIE------------FEVDDDE-ILCCVCHSTDANAEDPIVFCDGCNLMVHAS 157
E+ K++I+ EV+D++ I+C VC STD + +PIVFCDGC+LMVHAS
Sbjct: 121 EEDDDKENIDPLGKGKALDLSDREVEDEDGIMCAVCQSTDGDPLNPIVFCDGCDLMVHAS 180
Query: 158 CYGNPLVKQIPDGDWFCDQCRFKNDIDTDTGPIRCSLCPTKEGAMKQTTDGKWVHLVCAL 217
CYGNPLVK IP+GDWFC QC + + C LC TK GAMK T DG+W H+ CAL
Sbjct: 181 CYGNPLVKAIPEGDWFCRQCLSSKNREKI---FSCCLCTTKGGAMKPTNDGRWAHITCAL 237
Query: 218 LVPEVFFVDPEGREGIDCSKIPKKRWLEKCYVCGCFDGCALVCSEQKCGLGFHITCGIKE 277
VPEV+F DPEGREGI CS++ KRW ++CY+C GC + CSE +C L FH+TCG+KE
Sbjct: 238 FVPEVYFEDPEGREGICCSEVLSKRWKDRCYLCKVRRGCVIECSEMRCKLAFHVTCGLKE 297
Query: 278 DLCIEYKEGKKGATVVAGFCKTHSQIWEKNKGSGKYKIVAVED 320
DLCIEY+EGKK +V GFC H+++WE+ SGKYKIVA E+
Sbjct: 298 DLCIEYREGKKSGGIVVGFCNEHTKLWERE--SGKYKIVAREE 338
>ref|NP_914478.1| P0489A01.16 [Oryza sativa (japonica cultivar-group)]
Length = 607
Score = 261 bits (667), Expect = 2e-68
Identities = 118/215 (54%), Positives = 147/215 (67%), Gaps = 8/215 (3%)
Query: 111 EKQSQKQDIEFEVDDDEILCCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDG 170
E + + E E +DD + C VC STD + DPIVFCDGC+LMVHASCYGNPL IPDG
Sbjct: 389 EAMAAAAEEEVEEEDDGVHCAVCGSTDGDPSDPIVFCDGCDLMVHASCYGNPLASFIPDG 448
Query: 171 DWFCDQC-----RFKNDIDTDTGPIRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFV 225
DWFC C + P RC LCP + GAMK+TTD +W H+ CALLVPEVFF
Sbjct: 449 DWFCSVCTAAAAKKSKGNKPPPPPPRCCLCPARGGAMKRTTDARWAHIACALLVPEVFFR 508
Query: 226 DPEGREGIDCSKIPKKRWLEKCYVCGCFDGCALVCSEQKCGLGFHITCGIKEDLCIEYKE 285
DP+GR+G+DCS++P R+ CYVC GCAL CS+ +CGLGFH++CG+ LCIEY+E
Sbjct: 509 DPDGRDGVDCSRVPAHRFATACYVCESGGGCALECSQPRCGLGFHVSCGLDAGLCIEYQE 568
Query: 286 GKK---GATVVAGFCKTHSQIWEKNKGSGKYKIVA 317
K G VVAGFC H+++WEK + +GKYKIV+
Sbjct: 569 AKAGGGGGGVVAGFCLEHTKLWEKQQLTGKYKIVS 603
>gb|AAS21398.1| Brpf1 protein-like protein [Oikopleura dioica]
Length = 1062
Score = 154 bits (389), Expect = 3e-36
Identities = 75/189 (39%), Positives = 109/189 (56%), Gaps = 18/189 (9%)
Query: 122 EVDDDEILCCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFKN 181
+++D++++CCVC+ + + I+FCD CNL VH CYG P IP+G W C +C+F
Sbjct: 310 DIEDEDVVCCVCNDGECTNTNAILFCDLCNLAVHQECYGVPY---IPEGQWLCRRCQF-- 364
Query: 182 DIDTDTGPIRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFVDPEGREGID-CSKIPK 240
+ + P+ C LCP+ GA KQT D +W H+VCAL VPEV F +P E ID SK+PK
Sbjct: 365 ---SPSRPVDCVLCPSLNGAFKQTHDNRWCHVVCALWVPEVSFQNPVFLEPIDGFSKVPK 421
Query: 241 KRWLEKCYVCGCFDGCALVCSEQKCGLGFHITC----GIKEDLCIEYKEGKKGAT----V 292
RW KCY+C C + C++ C FH+TC G+ ++ ++ +G G T
Sbjct: 422 ARWNLKCYICKKAGAC-IQCNKNACYTAFHVTCAQQAGLYMEMKVQKSDGINGITSNKVT 480
Query: 293 VAGFCKTHS 301
FC H+
Sbjct: 481 QTAFCHNHT 489
>gb|EAA57618.1| hypothetical protein AN6675.2 [Aspergillus nidulans FGSC A4]
gi|67541010|ref|XP_664279.1| hypothetical protein
AN6675_2 [Aspergillus nidulans FGSC A4]
gi|49099886|ref|XP_410812.1| hypothetical protein
AN6675.2 [Aspergillus nidulans FGSC A4]
Length = 1173
Score = 146 bits (369), Expect = 7e-34
Identities = 74/203 (36%), Positives = 102/203 (49%), Gaps = 15/203 (7%)
Query: 124 DDDEILCCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDI 183
+D + C +C D + IVFCDGC+L VH CYG P IP+G W C +C+
Sbjct: 426 EDQDTKCAICDDGDCENANAIVFCDGCDLAVHQECYGVPF---IPEGQWLCRKCQL---- 478
Query: 184 DTDTGPIRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFVDPEGREGI-DCSKIPKKR 242
G C CP EGA KQTT KW HL+CA+ +PEV +P E I D K+P+ R
Sbjct: 479 -IGRGSPNCIFCPNIEGAFKQTTTSKWSHLLCAIWIPEVSIGNPSLMEPITDVEKVPRSR 537
Query: 243 WLEKCYVCGCFDGCALVCSEQKCGLGFHITCGIKEDLCIEYKEGKKGATV-----VAGFC 297
W +CY+C G ++ CS + C + FH+TC + L ++ K G V + FC
Sbjct: 538 WKLQCYICRQKMGASIQCSNKNCFVAFHVTCARRAQLYLKMKSGHGNLAVMDSHLLKAFC 597
Query: 298 KTH-SQIWEKNKGSGKYKIVAVE 319
H W + G+ A+E
Sbjct: 598 DKHVPPEWRREHGTDAATADAIE 620
>gb|EAL86946.1| PHD finger domain protein, putative [Aspergillus fumigatus Af293]
Length = 1205
Score = 143 bits (361), Expect = 6e-33
Identities = 71/203 (34%), Positives = 100/203 (48%), Gaps = 15/203 (7%)
Query: 124 DDDEILCCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDI 183
++ + C +C D + IVFCDGC+L VH CYG P IP+G W C +C+
Sbjct: 436 EEQDSKCAICDDGDCENSNAIVFCDGCDLAVHQECYGVPF---IPEGQWLCRKCQL---- 488
Query: 184 DTDTGPIRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFVDPEGREGI-DCSKIPKKR 242
G + C CP EGA KQTT KW HL+CA+ +PEV +P E + D K+P+ R
Sbjct: 489 -IGRGSVNCIFCPNTEGAFKQTTSSKWSHLLCAIWIPEVSLGNPSLMEPVTDVEKVPRSR 547
Query: 243 WLEKCYVCGCFDGCALVCSEQKCGLGFHITCGIKEDLCIEYKEGKKGATV-----VAGFC 297
W CY+C G ++ CS + C + FH TC + L ++ K G + + FC
Sbjct: 548 WKLTCYICKQRMGASIQCSNKNCFVAFHPTCARRAQLYLKMKSGHGAPAIMDSHLLKAFC 607
Query: 298 KTH-SQIWEKNKGSGKYKIVAVE 319
H W G+ A+E
Sbjct: 608 DKHVPDDWRLEHGTDAATADAIE 630
>ref|XP_418610.1| PREDICTED: similar to AF-10 protein [Gallus gallus]
Length = 1249
Score = 139 bits (351), Expect = 9e-32
Identities = 72/205 (35%), Positives = 106/205 (51%), Gaps = 19/205 (9%)
Query: 120 EFEVDDDEIL--CCVCHSTDANAEDPIVFCDG--CNLMVHASCYGNPLVKQIPDGDWFCD 175
EF E++ CCVC AE+P+V+CDG CN+ VH +CYG + Q+P G WFC
Sbjct: 215 EFSTSMKEMIGGCCVCSDERGWAENPLVYCDGHGCNVAVHQACYG---IVQVPTGPWFCR 271
Query: 176 QCRFKNDIDTDTGPIRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFVDPEGREGIDC 235
+C + +RC LCP K+GA+K+T +G W H+VCAL +PEV F + E I
Sbjct: 272 KCESQER----AARVRCELCPHKDGALKRTDNGGWAHVVCALYIPEVQFANVATMEPIVL 327
Query: 236 SKIPKKRWLEKCYVCG-------CFDGCALVCSEQKCGLGFHITCGIKEDLCIEYKEGKK 288
+P R+ + CY+C G + C++ C FH+TC L E +
Sbjct: 328 QSVPHDRYNKTCYICDEQGRESKAATGACMTCNKHGCRQAFHVTCAQFAGLLCEEEGNGA 387
Query: 289 GATVVAGFCKTH-SQIWEKNKGSGK 312
G+CK H S++ + +GS +
Sbjct: 388 DNVQYCGYCKYHFSKLKKSKRGSNR 412
>emb|CAG07010.1| unnamed protein product [Tetraodon nigroviridis]
Length = 1523
Score = 139 bits (350), Expect = 1e-31
Identities = 75/208 (36%), Positives = 106/208 (50%), Gaps = 17/208 (8%)
Query: 113 QSQKQDIEFEVDDDEILCCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDW 172
+S+ Q + D++ CCVC + + I+FCD CNL VH CYG P V P+G W
Sbjct: 213 ESRSQALSHSAVDEDAFCCVCLDDECLNSNVILFCDICNLAVHQECYGVPYV---PEGQW 269
Query: 173 FCDQCRFKNDIDTDTGPIRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFVDPEGREG 232
C C + + + P+ C LCP + GA KQT+DG+W H+VCA+ +PEV F + E
Sbjct: 270 LCRCC-----LQSPSRPVDCVLCPNRGGAFKQTSDGRWAHVVCAIWIPEVCFANTVFLEP 324
Query: 233 IDCSK-IPKKRWLEKCYVCGCFD-GCALVCSEQKCGLGFHITCGIKEDLCIEY----KEG 286
++ K IP RW CY+C G ++ C + C FH+TC K L ++ + G
Sbjct: 325 VEGIKNIPPARWKLTCYLCKQKGRGASIQCHKANCYRAFHVTCAQKAGLFMKIDPVRETG 384
Query: 287 KKGATV---VAGFCKTHSQIWEKNKGSG 311
G T FC+ HS GSG
Sbjct: 385 VNGTTFSVKKTAFCEHHSPAGSHRDGSG 412
>gb|AAH85650.1| Zgc:92208 [Danio rerio] gi|55925532|ref|NP_001007309.1| zgc:92208
[Danio rerio]
Length = 352
Score = 138 bits (347), Expect = 2e-31
Identities = 66/180 (36%), Positives = 94/180 (51%), Gaps = 16/180 (8%)
Query: 130 CCVCHSTDANAEDPIVFCDG--CNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTDT 187
CCVC AE+P+V+CDG CN+ VH +CYG + Q+P G WFC +C +
Sbjct: 26 CCVCSDERGWAENPLVYCDGHGCNVAVHQACYG---IVQVPTGPWFCRKCESQER----A 78
Query: 188 GPIRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFVDPEGREGIDCSKIPKKRWLEKC 247
+RC LCP K+GA+K+T +G W H+VCAL +PEV F + E I +P +R+ + C
Sbjct: 79 ARVRCELCPQKDGALKRTDNGGWAHVVCALYIPEVEFANVSTMEPIVLQSVPHERYNKTC 138
Query: 248 YVC-------GCFDGCALVCSEQKCGLGFHITCGIKEDLCIEYKEGKKGATVVAGFCKTH 300
Y+C G + C++ C FH+TC L E + G+CK H
Sbjct: 139 YICEEQGRESKAATGACMTCNKHGCRQAFHVTCAQLAGLLCEEQGSDADNVKYCGYCKYH 198
>ref|XP_419262.1| PREDICTED: similar to Bromodomain and PHD finger-containing protein
3 [Gallus gallus]
Length = 1071
Score = 138 bits (347), Expect = 2e-31
Identities = 70/198 (35%), Positives = 106/198 (53%), Gaps = 17/198 (8%)
Query: 113 QSQKQDIEFEVDDDEILCCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDW 172
+S+ + + D++ CCVC + + + I+FCD CNL VH CYG P IP+G W
Sbjct: 198 ESRNNGTQHSIIDEDAFCCVCMDDECHNSNVILFCDICNLAVHQECYGVPY---IPEGQW 254
Query: 173 FCDQCRFKNDIDTDTGPIRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFVDPEGREG 232
C C + + + P+ C LCP K GA KQT+DG+W H+VCA+ +PEV F + E
Sbjct: 255 LCRCC-----LQSPSRPVDCVLCPNKGGAFKQTSDGRWAHVVCAIWIPEVCFANTVFLEP 309
Query: 233 ID-CSKIPKKRWLEKCYVCGCFD-GCALVCSEQKCGLGFHITCGIKEDLCIEYKEGKK-- 288
I+ + IP RW CY+C G A+ C + C FH+TC + L ++ + ++
Sbjct: 310 IEGINNIPPARWKLTCYICKQKGMGAAIQCHKVNCYTAFHVTCAQRAGLFMKIEPMRETS 369
Query: 289 --GATVV---AGFCKTHS 301
G T +C++HS
Sbjct: 370 INGTTFTVRKTAYCESHS 387
>ref|XP_228039.3| PREDICTED: similar to BRPF3 [Rattus norvegicus]
Length = 1266
Score = 137 bits (346), Expect = 3e-31
Identities = 84/253 (33%), Positives = 123/253 (48%), Gaps = 31/253 (12%)
Query: 83 HNNIPNDAVPLIDLNVEYSPSLPSATPIEKQSQKQDIEFEVDDDEILCCVCHSTDANAED 142
H+++ D L+ +E L S + +QS + D++ CCVC + + +
Sbjct: 176 HSSVSADTFELLVDRLEKESYLESRSSGAQQS--------LIDEDAFCCVCLDDECHNSN 227
Query: 143 PIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTDTGPIRCSLCPTKEGAM 202
I+FCD CNL VH CYG P IP+G W C C + + + P+ C LCP K GA
Sbjct: 228 VILFCDICNLAVHQECYGVPY---IPEGQWLCRCC-----LQSPSRPVDCVLCPNKGGAF 279
Query: 203 KQTTDGKWVHLVCALLVPEVFFVDP---EGREGIDCSKIPKKRWLEKCYVCGCFD-GCAL 258
KQT+DG W H+VCA+ +PEV F + E EGID IP RW CY+C G A+
Sbjct: 280 KQTSDGHWAHVVCAIWIPEVCFANTVFLEPIEGID--SIPPARWKLTCYICKQKGLGAAI 337
Query: 259 VCSEQKCGLGFHITCGIKEDLCIE---YKEGKKGATVV----AGFCKTHSQ--IWEKNKG 309
C + C FH+TC + L ++ +E T+ +C+ HS + KG
Sbjct: 338 QCHKVNCYTAFHVTCAQRAGLFMKIEPMRETSLNGTIFTVRKTAYCEAHSPSVATARRKG 397
Query: 310 SGKYKIVAVEDKE 322
+ V D++
Sbjct: 398 DSPRSLSEVGDED 410
>gb|EAK86867.1| hypothetical protein UM06029.1 [Ustilago maydis 521]
gi|49080186|ref|XP_403644.1| hypothetical protein
UM06029.1 [Ustilago maydis 521]
Length = 1227
Score = 137 bits (346), Expect = 3e-31
Identities = 68/175 (38%), Positives = 89/175 (50%), Gaps = 9/175 (5%)
Query: 106 SATPIEKQSQKQDIEFEVDDDEILCCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVK 165
+AT + + D D+ C +C + + IVFCDGCNL VH CYG P
Sbjct: 134 AATEVAAEGADADDADSEGGDDSKCAICDDGECENSNAIVFCDGCNLAVHQDCYGIPY-- 191
Query: 166 QIPDGDWFCDQCRFKNDIDTDTGPIRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFV 225
IP+G W C +C D + C LCP + GA KQTT GKW HL+CA+ +PE
Sbjct: 192 -IPEGQWLCRKCTVSPD-----RAVSCILCPHEGGAFKQTTTGKWAHLLCAMWIPETGVS 245
Query: 226 DPEGREGID-CSKIPKKRWLEKCYVCGCFDGCALVCSEQKCGLGFHITCGIKEDL 279
+P E ID +IPK RW +CY+C G + C + C FH+TC K L
Sbjct: 246 NPVYMEPIDSVERIPKARWKLQCYLCRYRMGACIQCDNRSCFTAFHVTCARKAGL 300
>gb|AAH82872.1| LOC494767 protein [Xenopus laevis]
Length = 963
Score = 137 bits (346), Expect = 3e-31
Identities = 66/180 (36%), Positives = 94/180 (51%), Gaps = 16/180 (8%)
Query: 130 CCVCHSTDANAEDPIVFCDG--CNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTDT 187
CCVC AE+P+V+CDG CN+ VH +CYG + Q+P G WFC +C +
Sbjct: 8 CCVCSDERGWAENPLVYCDGHGCNVAVHQACYG---IVQVPTGPWFCRKCESQER----A 60
Query: 188 GPIRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFVDPEGREGIDCSKIPKKRWLEKC 247
+RC LCP K+GA+K+T +G W H+VCAL +PEV F + E I +P +R+ + C
Sbjct: 61 ARVRCELCPHKDGALKRTDNGGWAHVVCALYIPEVQFANVSTMEPIVLQSVPHERYNKTC 120
Query: 248 YVCG-------CFDGCALVCSEQKCGLGFHITCGIKEDLCIEYKEGKKGATVVAGFCKTH 300
Y+C G + C++ C FH+TC L E + G+CK H
Sbjct: 121 YICDEQGRESKAATGACMTCNKHGCRQAFHVTCAQLAGLLCEEEGNGADNVQYCGYCKYH 180
>ref|NP_034934.2| myeloid/lymphoid or mixed lineage-leukemia translocation to 10
homolog [Mus musculus] gi|47125547|gb|AAH70475.1|
Myeloid/lymphoid or mixed lineage-leukemia translocation
to 10 homolog [Mus musculus]
Length = 1068
Score = 137 bits (345), Expect = 4e-31
Identities = 68/193 (35%), Positives = 101/193 (52%), Gaps = 17/193 (8%)
Query: 130 CCVCHSTDANAEDPIVFCDG--CNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTDT 187
CCVC AE+P+V+CDG C++ VH +CYG + Q+P G WFC +C +
Sbjct: 25 CCVCSDERGWAENPLVYCDGHGCSVAVHQACYG---IVQVPTGPWFCRKCESQER----A 77
Query: 188 GPIRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFVDPEGREGIDCSKIPKKRWLEKC 247
+RC LCP K+GA+K+T +G W H+VCAL +PEV F + E I +P R+ + C
Sbjct: 78 ARVRCELCPHKDGALKRTDNGGWAHVVCALYIPEVQFANVSTMEPIVLQSVPHDRYNKTC 137
Query: 248 YVCG-------CFDGCALVCSEQKCGLGFHITCGIKEDLCIEYKEGKKGATVVAGFCKTH 300
Y+C G + C++ C FH+TC L E + G+CK H
Sbjct: 138 YICDEQGRESKAATGACMTCNKHGCRQAFHVTCAQFAGLLCEEEGNGADNVQYCGYCKYH 197
Query: 301 -SQIWEKNKGSGK 312
S++ + +GS +
Sbjct: 198 FSKLKKSKRGSNR 210
>gb|AAT47519.1| AF10 [Homo sapiens] gi|55664632|emb|CAH73411.1| myeloid\/lymphoid
or mixed-lineage leukemia (trithorax homolog,
Drosophila)\; translocated to, 10 (MLLT10) [Homo
sapiens] gi|55959122|emb|CAI13980.1| myeloid\/lymphoid
or mixed-lineage leukemia (trithorax homolog,
Drosophila)\; translocated to, 10 (MLLT10) [Homo
sapiens] gi|57209054|emb|CAI41349.1| myeloid\/lymphoid
or mixed-lineage leukemia (trithorax homolog,
Drosophila)\; translocated to, 10 (MLLT10) [Homo
sapiens] gi|57162213|emb|CAI39664.1| myeloid\/lymphoid
or mixed-lineage leukemia (trithorax homolog,
Drosophila)\; translocated to, 10 (MLLT10) [Homo
sapiens]
Length = 1068
Score = 137 bits (345), Expect = 4e-31
Identities = 68/193 (35%), Positives = 101/193 (52%), Gaps = 17/193 (8%)
Query: 130 CCVCHSTDANAEDPIVFCDG--CNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTDT 187
CCVC AE+P+V+CDG C++ VH +CYG + Q+P G WFC +C +
Sbjct: 25 CCVCSDERGWAENPLVYCDGHGCSVAVHQACYG---IVQVPTGPWFCRKCESQER----A 77
Query: 188 GPIRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFVDPEGREGIDCSKIPKKRWLEKC 247
+RC LCP K+GA+K+T +G W H+VCAL +PEV F + E I +P R+ + C
Sbjct: 78 ARVRCELCPHKDGALKRTDNGGWAHVVCALYIPEVQFANVSTMEPIVLQSVPHDRYNKTC 137
Query: 248 YVCG-------CFDGCALVCSEQKCGLGFHITCGIKEDLCIEYKEGKKGATVVAGFCKTH 300
Y+C G + C++ C FH+TC L E + G+CK H
Sbjct: 138 YICDEQGRESKAATGACMTCNKHGCRQAFHVTCAQFAGLLCEEEGNGADNVQYCGYCKYH 197
Query: 301 -SQIWEKNKGSGK 312
S++ + +GS +
Sbjct: 198 FSKLKKSKRGSNR 210
>emb|CAH73410.1| myeloid\/lymphoid or mixed-lineage leukemia (trithorax homolog,
Drosophila)\; translocated to, 10 (MLLT10) [Homo
sapiens] gi|55959121|emb|CAI13979.1| myeloid\/lymphoid
or mixed-lineage leukemia (trithorax homolog,
Drosophila)\; translocated to, 10 (MLLT10) [Homo
sapiens] gi|57209053|emb|CAI41348.1| myeloid\/lymphoid
or mixed-lineage leukemia (trithorax homolog,
Drosophila)\; translocated to, 10 (MLLT10) [Homo
sapiens] gi|57162212|emb|CAI39663.1| myeloid\/lymphoid
or mixed-lineage leukemia (trithorax homolog,
Drosophila)\; translocated to, 10 (MLLT10) [Homo
sapiens] gi|4757726|ref|NP_004632.1| myeloid/lymphoid or
mixed-lineage leukemia translocated to, 10 isoform a
[Homo sapiens] gi|1703190|sp|P55197|AF10_HUMAN AF-10
protein gi|538277|gb|AAA79972.1| zinc finger/leucine
zipper protein
Length = 1027
Score = 137 bits (345), Expect = 4e-31
Identities = 68/193 (35%), Positives = 101/193 (52%), Gaps = 17/193 (8%)
Query: 130 CCVCHSTDANAEDPIVFCDG--CNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTDT 187
CCVC AE+P+V+CDG C++ VH +CYG + Q+P G WFC +C +
Sbjct: 25 CCVCSDERGWAENPLVYCDGHGCSVAVHQACYG---IVQVPTGPWFCRKCESQER----A 77
Query: 188 GPIRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFVDPEGREGIDCSKIPKKRWLEKC 247
+RC LCP K+GA+K+T +G W H+VCAL +PEV F + E I +P R+ + C
Sbjct: 78 ARVRCELCPHKDGALKRTDNGGWAHVVCALYIPEVQFANVSTMEPIVLQSVPHDRYNKTC 137
Query: 248 YVCG-------CFDGCALVCSEQKCGLGFHITCGIKEDLCIEYKEGKKGATVVAGFCKTH 300
Y+C G + C++ C FH+TC L E + G+CK H
Sbjct: 138 YICDEQGRESKAATGACMTCNKHGCRQAFHVTCAQFAGLLCEEEGNGADNVQYCGYCKYH 197
Query: 301 -SQIWEKNKGSGK 312
S++ + +GS +
Sbjct: 198 FSKLKKSKRGSNR 210
>ref|NP_001009569.1| myeloid/lymphoid or mixed-lineage leukemia translocated to, 10
isoform b [Homo sapiens]
Length = 1011
Score = 137 bits (345), Expect = 4e-31
Identities = 68/193 (35%), Positives = 101/193 (52%), Gaps = 17/193 (8%)
Query: 130 CCVCHSTDANAEDPIVFCDG--CNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTDT 187
CCVC AE+P+V+CDG C++ VH +CYG + Q+P G WFC +C +
Sbjct: 25 CCVCSDERGWAENPLVYCDGHGCSVAVHQACYG---IVQVPTGPWFCRKCESQER----A 77
Query: 188 GPIRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFVDPEGREGIDCSKIPKKRWLEKC 247
+RC LCP K+GA+K+T +G W H+VCAL +PEV F + E I +P R+ + C
Sbjct: 78 ARVRCELCPHKDGALKRTDNGGWAHVVCALYIPEVQFANVSTMEPIVLQSVPHDRYNKTC 137
Query: 248 YVCG-------CFDGCALVCSEQKCGLGFHITCGIKEDLCIEYKEGKKGATVVAGFCKTH 300
Y+C G + C++ C FH+TC L E + G+CK H
Sbjct: 138 YICDEQGRESKAATGACMTCNKHGCRQAFHVTCAQFAGLLCEEEGNGADNVQYCGYCKYH 197
Query: 301 -SQIWEKNKGSGK 312
S++ + +GS +
Sbjct: 198 FSKLKKSKRGSNR 210
>gb|AAD11570.1| maf10 [Mus musculus] gi|12229633|sp|O54826|AF10_MOUSE AF-10 protein
Length = 1068
Score = 137 bits (345), Expect = 4e-31
Identities = 68/193 (35%), Positives = 101/193 (52%), Gaps = 17/193 (8%)
Query: 130 CCVCHSTDANAEDPIVFCDG--CNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTDT 187
CCVC AE+P+V+CDG C++ VH +CYG + Q+P G WFC +C +
Sbjct: 25 CCVCSDERGWAENPLVYCDGHGCSVAVHQACYG---IVQVPTGPWFCRKCESQER----A 77
Query: 188 GPIRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFVDPEGREGIDCSKIPKKRWLEKC 247
+RC LCP K+GA+K+T +G W H+VCAL +PEV F + E I +P R+ + C
Sbjct: 78 ARVRCELCPHKDGALKRTDNGGWAHVVCALYIPEVQFANVSTMEPIVLQSVPHDRYNKTC 137
Query: 248 YVCG-------CFDGCALVCSEQKCGLGFHITCGIKEDLCIEYKEGKKGATVVAGFCKTH 300
Y+C G + C++ C FH+TC L E + G+CK H
Sbjct: 138 YICDEQGRESKAATGACMTCNKHGCRQAFHVTCAQFAGLLCEEEGNGADNVQYCGYCKYH 197
Query: 301 -SQIWEKNKGSGK 312
S++ + +GS +
Sbjct: 198 FSKLKKSKRGSNR 210
>ref|XP_196171.5| PREDICTED: bromodomain and PHD finger containing, 3 [Mus musculus]
Length = 1255
Score = 137 bits (345), Expect = 4e-31
Identities = 84/253 (33%), Positives = 124/253 (48%), Gaps = 31/253 (12%)
Query: 83 HNNIPNDAVPLIDLNVEYSPSLPSATPIEKQSQKQDIEFEVDDDEILCCVCHSTDANAED 142
H+++ D L+ +E L S + +QS + D++ CCVC + + +
Sbjct: 227 HSSVSADTFELLVDRLEKESYLESRSSGAQQS--------LIDEDAFCCVCLDDECHNSN 278
Query: 143 PIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTDTGPIRCSLCPTKEGAM 202
I+FCD CNL VH CYG P IP+G W C C + + + P+ C LCP K GA
Sbjct: 279 VILFCDICNLAVHQECYGVPY---IPEGQWLCRCC-----LQSPSRPVDCVLCPNKGGAF 330
Query: 203 KQTTDGKWVHLVCALLVPEVFFVDP---EGREGIDCSKIPKKRWLEKCYVCGCFD-GCAL 258
KQT+DG W H+VCA+ +PEV F + E EGID IP RW CY+C G A+
Sbjct: 331 KQTSDGHWAHVVCAIWIPEVCFANTVFLEPIEGID--NIPPARWKLTCYICKQKGLGAAI 388
Query: 259 VCSEQKCGLGFHITCGIKEDLCIEYKEGKK----GATVV---AGFCKTHSQ--IWEKNKG 309
C + C FH+TC + L ++ + ++ G T +C+ HS + KG
Sbjct: 389 QCHKVNCYTAFHVTCAQRAGLFMKIEPMRETSLNGTTFTVRKTAYCEAHSPSVAVARRKG 448
Query: 310 SGKYKIVAVEDKE 322
+ V D++
Sbjct: 449 DSPRSLSEVGDED 461
>emb|CAI21669.1| BRPF3 [Homo sapiens]
Length = 1205
Score = 137 bits (344), Expect = 6e-31
Identities = 78/224 (34%), Positives = 111/224 (48%), Gaps = 24/224 (10%)
Query: 113 QSQKQDIEFEVDDDEILCCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDW 172
+S+ + + D++ CCVC + + + I+FCD CNL VH CYG P IP+G W
Sbjct: 198 ESRSSGAQQSLIDEDAFCCVCLDDECHNSNVILFCDICNLAVHQECYGVPY---IPEGQW 254
Query: 173 FCDQCRFKNDIDTDTGPIRCSLCPTKEGAMKQTTDGKWVHLVCALLVPEVFFVDP---EG 229
C C + + + P+ C LCP K GA KQT+DG W H+VCA+ +PEV F + E
Sbjct: 255 LCRCC-----LQSPSRPVDCILCPNKGGAFKQTSDGHWAHVVCAIWIPEVCFANTVFLEP 309
Query: 230 REGIDCSKIPKKRWLEKCYVCGCFD-GCALVCSEQKCGLGFHITCGIKEDLCIE---YKE 285
EGID IP RW CY+C G A+ C + C FH+TC + L ++ +E
Sbjct: 310 IEGID--NIPPARWKLTCYICKQKGLGAAIQCHKVNCYTAFHVTCAQRAGLFMKIEPMRE 367
Query: 286 GKKGATVV----AGFCKTHS---QIWEKNKGSGKYKIVAVEDKE 322
T+ +C+ HS + KG I D+E
Sbjct: 368 TSLNGTIFTVRKTAYCEAHSPPGAATARRKGDSPRSISETGDEE 411
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.319 0.137 0.445
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 652,869,278
Number of Sequences: 2540612
Number of extensions: 31619441
Number of successful extensions: 136465
Number of sequences better than 10.0: 1561
Number of HSP's better than 10.0 without gapping: 364
Number of HSP's successfully gapped in prelim test: 1199
Number of HSP's that attempted gapping in prelim test: 132948
Number of HSP's gapped (non-prelim): 3152
length of query: 322
length of database: 863,360,394
effective HSP length: 128
effective length of query: 194
effective length of database: 538,162,058
effective search space: 104403439252
effective search space used: 104403439252
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 75 (33.5 bits)
Medicago: description of AC147714.16


