

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147538.3 - phase: 0
(123 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
Score E
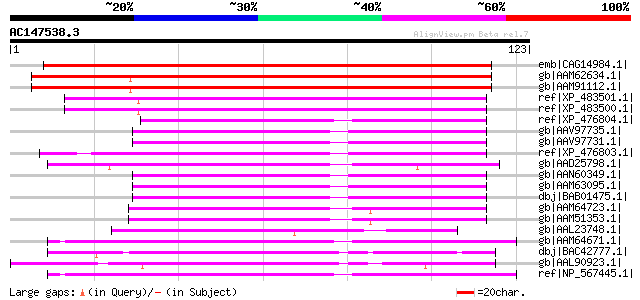
Sequences producing significant alignments: (bits) Value
emb|CAG14984.1| putative lipid transfer protein GPI-anchored [Ci... 188 2e-47
gb|AAM62634.1| lipid transfer protein, putative [Arabidopsis tha... 114 5e-25
gb|AAM91112.1| lipid transfer protein, putative [Arabidopsis tha... 112 3e-24
ref|XP_483501.1| lipid transfer protein-like [Oryza sativa (japo... 87 9e-17
ref|XP_483500.1| lipid transfer protein-like [Oryza sativa (japo... 87 9e-17
ref|XP_476804.1| protease inhibitor-like protein [Oryza sativa (... 68 4e-11
gb|AAV97735.1| lipid transfer protein [Capsicum annuum] gi|56549... 58 5e-08
gb|AAV97731.1| lipid transfer protein [Capsicum annuum] 58 5e-08
ref|XP_476803.1| protease inhibitor-like protein [Oryza sativa (... 58 6e-08
gb|AAD25798.1| F10O3.7 [Arabidopsis thaliana] gi|25406727|pir||A... 58 6e-08
gb|AAN60349.1| unknown [Arabidopsis thaliana] 56 2e-07
gb|AAM63095.1| unknown [Arabidopsis thaliana] 56 2e-07
dbj|BAB01475.1| unnamed protein product [Arabidopsis thaliana] g... 56 2e-07
gb|AAM64723.1| unknown [Arabidopsis thaliana] 56 2e-07
gb|AAM51353.1| unknown protein [Arabidopsis thaliana] gi|1752932... 56 2e-07
gb|AAL23748.1| nonspecific lipid transfer protein [Bromus inermis] 56 2e-07
gb|AAM64671.1| unknown [Arabidopsis thaliana] 55 3e-07
dbj|BAC42777.1| putative non-specific lipid transfer protein nLT... 55 3e-07
gb|AAL90923.1| At1g55260/F7A10_16 [Arabidopsis thaliana] gi|1840... 55 3e-07
ref|NP_567445.1| protease inhibitor/seed storage/lipid transfer ... 55 3e-07
>emb|CAG14984.1| putative lipid transfer protein GPI-anchored [Cicer arietinum]
Length = 185
Score = 188 bits (478), Expect = 2e-47
Identities = 88/106 (83%), Positives = 94/106 (88%)
Query: 9 FMCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATD 68
F+C+ L +IIGG GAEDLA KCG VVQKVIPCLDFATGKA TPKKECCDAANSIK TD
Sbjct: 5 FVCVLGLIMIIGGSEGAEDLAQKCGQVVQKVIPCLDFATGKALTPKKECCDAANSIKETD 64
Query: 69 PECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDCP 114
PECLCYIIQQTHKGSPESKS+GIQEDKLLQLPTVC V AN++DCP
Sbjct: 65 PECLCYIIQQTHKGSPESKSLGIQEDKLLQLPTVCKVKNANLTDCP 110
>gb|AAM62634.1| lipid transfer protein, putative [Arabidopsis thaliana]
Length = 193
Score = 114 bits (285), Expect = 5e-25
Identities = 53/111 (47%), Positives = 71/111 (63%), Gaps = 2/111 (1%)
Query: 6 HQMFMCLCVLALIIGGCNGAED--LASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANS 63
H + + + ++A I A LA +C QKV CLDFATGKA TP K+CCDA
Sbjct: 7 HLVLVTMTIVASIAAAAPAAPGGALADECNQDFQKVTLCLDFATGKATTPSKKCCDAVED 66
Query: 64 IKATDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDCP 114
IK DP+CLC++IQQ G K +G+QEDKL+QLPT C ++ A+I++CP
Sbjct: 67 IKERDPKCLCFVIQQAKTGGQALKDLGVQEDKLIQLPTSCQLHNASITNCP 117
>gb|AAM91112.1| lipid transfer protein, putative [Arabidopsis thaliana]
gi|20260114|gb|AAM12955.1| lipid transfer protein,
putative [Arabidopsis thaliana]
gi|15217777|ref|NP_174116.1| lipid transfer
protein-related [Arabidopsis thaliana]
gi|12322995|gb|AAG51485.1| lipid transfer protein,
putative [Arabidopsis thaliana] gi|25403045|pir||H86404
probable lipid transfer protein [imported] - Arabidopsis
thaliana gi|38258836|sp|Q9C7F7|UGP5_ARATH
Uncharacterized GPI-anchored protein At1g27950 precursor
Length = 193
Score = 112 bits (279), Expect = 3e-24
Identities = 52/111 (46%), Positives = 70/111 (62%), Gaps = 2/111 (1%)
Query: 6 HQMFMCLCVLALIIGGCNGAED--LASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANS 63
H + + + ++A I A LA +C QKV CLDFATGKA P K+CCDA
Sbjct: 7 HLVLVTMTIVASIAAAAPAAPGGALADECNQDFQKVTLCLDFATGKATIPSKKCCDAVED 66
Query: 64 IKATDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDCP 114
IK DP+CLC++IQQ G K +G+QEDKL+QLPT C ++ A+I++CP
Sbjct: 67 IKERDPKCLCFVIQQAKTGGQALKDLGVQEDKLIQLPTSCQLHNASITNCP 117
>ref|XP_483501.1| lipid transfer protein-like [Oryza sativa (japonica
cultivar-group)] gi|51979701|ref|XP_507597.1| PREDICTED
P0702E04.24-2 gene product [Oryza sativa (japonica
cultivar-group)] gi|42761387|dbj|BAD11655.1| lipid
transfer protein-like [Oryza sativa (japonica
cultivar-group)]
Length = 179
Score = 87.0 bits (214), Expect = 9e-17
Identities = 42/102 (41%), Positives = 58/102 (56%), Gaps = 2/102 (1%)
Query: 14 VLALIIGGCNGAEDLA--SKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPEC 71
V+A + A D A SKC K+ C+D+ATG P CC ++ + PEC
Sbjct: 12 VVAWCVAAAAAAPDAALQSKCQQDFTKLTDCMDYATGHEEAPSSTCCGDMSATQQARPEC 71
Query: 72 LCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDC 113
LCYIIQQ H G E +S+G++ D+LL +PT C + AN+S C
Sbjct: 72 LCYIIQQVHGGRNEVQSLGLRFDRLLAMPTACKLPNANVSLC 113
>ref|XP_483500.1| lipid transfer protein-like [Oryza sativa (japonica
cultivar-group)] gi|51978926|ref|XP_507304.2| PREDICTED
P0702E04.24-2 gene product [Oryza sativa (japonica
cultivar-group)] gi|42761388|dbj|BAD11656.1| lipid
transfer protein-like [Oryza sativa (japonica
cultivar-group)]
Length = 178
Score = 87.0 bits (214), Expect = 9e-17
Identities = 42/102 (41%), Positives = 58/102 (56%), Gaps = 2/102 (1%)
Query: 14 VLALIIGGCNGAEDLA--SKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPEC 71
V+A + A D A SKC K+ C+D+ATG P CC ++ + PEC
Sbjct: 12 VVAWCVAAAAAAPDAALQSKCQQDFTKLTDCMDYATGHEEAPSSTCCGDMSATQQARPEC 71
Query: 72 LCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDC 113
LCYIIQQ H G E +S+G++ D+LL +PT C + AN+S C
Sbjct: 72 LCYIIQQVHGGRNEVQSLGLRFDRLLAMPTACKLPNANVSLC 113
>ref|XP_476804.1| protease inhibitor-like protein [Oryza sativa (japonica
cultivar-group)] gi|25553596|dbj|BAC24861.1| protease
inhibitor-like protein [Oryza sativa (japonica
cultivar-group)]
Length = 181
Score = 68.2 bits (165), Expect = 4e-11
Identities = 31/82 (37%), Positives = 44/82 (52%), Gaps = 4/82 (4%)
Query: 32 CGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPECLCYIIQQTHKGSPESKSMGI 91
C SV+ + PCLD+ +GK+P P+ CC + +DP CLC ++ GS S + I
Sbjct: 37 CSSVMMTLSPCLDYISGKSPIPEFTCCTTLAGVVQSDPRCLCMVLD----GSAASFGISI 92
Query: 92 QEDKLLQLPTVCHVNGANISDC 113
+ L+LP VC V IS C
Sbjct: 93 NHTRALELPGVCKVQAPPISQC 114
>gb|AAV97735.1| lipid transfer protein [Capsicum annuum] gi|56549233|gb|AAV97734.1|
lipid transfer protein [Capsicum annuum]
gi|56549231|gb|AAV97733.1| lipid transfer protein
[Capsicum chinense] gi|56549229|gb|AAV97732.1| lipid
transfer protein [Capsicum chinense]
Length = 172
Score = 58.2 bits (139), Expect = 5e-08
Identities = 27/84 (32%), Positives = 42/84 (49%), Gaps = 4/84 (4%)
Query: 30 SKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPECLCYIIQQTHKGSPESKSM 89
S C S + + CL F TG A TP CC + + + ++P CLC I+ G S +
Sbjct: 29 SDCTSTLITMASCLSFVTGSAKTPPASCCSSLSGVLQSNPRCLCVIV----NGGGSSLGV 84
Query: 90 GIQEDKLLQLPTVCHVNGANISDC 113
I + + L LP+ C++ +S C
Sbjct: 85 QINQTQALALPSACNLQTPPVSRC 108
>gb|AAV97731.1| lipid transfer protein [Capsicum annuum]
Length = 172
Score = 58.2 bits (139), Expect = 5e-08
Identities = 27/84 (32%), Positives = 42/84 (49%), Gaps = 4/84 (4%)
Query: 30 SKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPECLCYIIQQTHKGSPESKSM 89
S C S + + CL F TG A TP CC + + + ++P CLC I+ G S +
Sbjct: 29 SDCTSTLITMASCLSFVTGSAKTPPASCCSSLSGVLQSNPRCLCVIV----NGGGSSLGV 84
Query: 90 GIQEDKLLQLPTVCHVNGANISDC 113
I + + L LP+ C++ +S C
Sbjct: 85 QINQTQALALPSACNLQTPPVSRC 108
>ref|XP_476803.1| protease inhibitor-like protein [Oryza sativa (japonica
cultivar-group)] gi|25553595|dbj|BAC24860.1| protease
inhibitor-like protein [Oryza sativa (japonica
cultivar-group)]
Length = 171
Score = 57.8 bits (138), Expect = 6e-08
Identities = 33/106 (31%), Positives = 48/106 (45%), Gaps = 7/106 (6%)
Query: 8 MFMCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKAT 67
MF + V+A G S C S + + PCLD+ G A P CC A +S+ +
Sbjct: 10 MFAAVVVVA---GALVAGAAAQSGCTSEMVSLAPCLDYMQGNASRPTASCCAALSSVVKS 66
Query: 68 DPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDC 113
PECLC ++ G S + + + L+LP C V S+C
Sbjct: 67 RPECLCAVL----GGGASSLGVTVNTTRALELPAACGVKTPPPSEC 108
>gb|AAD25798.1| F10O3.7 [Arabidopsis thaliana] gi|25406727|pir||A86162 protein
F10O3.7 [imported] - Arabidopsis thaliana
Length = 129
Score = 57.8 bits (138), Expect = 6e-08
Identities = 35/111 (31%), Positives = 55/111 (49%), Gaps = 9/111 (8%)
Query: 10 MCLCVLALIIGGC--NGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKAT 67
M L +LAL+I GA + + C + + PCL + G + +P CC +++ +
Sbjct: 1 MILAILALVIATFLYGGATTVQAGCRDTLTSLSPCLYYLNGGSSSPSWSCCRQFSTVVQS 60
Query: 68 DPECLCYIIQQTHKGSPESKSMGIQEDK--LLQLPTVCHVNGANISDCPSK 116
PECLC ++ S ES G + ++ L LPT C+V + S C SK
Sbjct: 61 SPECLCSVV-----NSNESSFYGFKFNRTLALNLPTACNVQTPSPSLCNSK 106
>gb|AAN60349.1| unknown [Arabidopsis thaliana]
Length = 168
Score = 56.2 bits (134), Expect = 2e-07
Identities = 22/84 (26%), Positives = 44/84 (52%), Gaps = 4/84 (4%)
Query: 30 SKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPECLCYIIQQTHKGSPESKSM 89
S C + + + PCL++ TG + +P ++CC+ + + + P+CLC ++ G +
Sbjct: 24 SSCTNALISMSPCLNYITGNSTSPNQQCCNQLSRVVQSSPDCLCQVL----NGGGSQLGI 79
Query: 90 GIQEDKLLQLPTVCHVNGANISDC 113
+ + + L LP C+V +S C
Sbjct: 80 NVNQTQALGLPRACNVQTPPVSRC 103
>gb|AAM63095.1| unknown [Arabidopsis thaliana]
Length = 166
Score = 56.2 bits (134), Expect = 2e-07
Identities = 22/84 (26%), Positives = 44/84 (52%), Gaps = 4/84 (4%)
Query: 30 SKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPECLCYIIQQTHKGSPESKSM 89
S C + + + PCL++ TG + +P ++CC+ + + + P+CLC ++ G +
Sbjct: 22 SSCTNALISMSPCLNYITGNSTSPNQQCCNQLSRVVQSSPDCLCQVL----NGGGSQLGI 77
Query: 90 GIQEDKLLQLPTVCHVNGANISDC 113
+ + + L LP C+V +S C
Sbjct: 78 NVNQTQALGLPRACNVQTPPVSRC 101
>dbj|BAB01475.1| unnamed protein product [Arabidopsis thaliana]
gi|18958062|gb|AAL79604.1| AT3g22600/F16J14_17
[Arabidopsis thaliana] gi|15010698|gb|AAK74008.1|
AT3g22600/F16J14_17 [Arabidopsis thaliana]
gi|18403453|ref|NP_566712.1| protease inhibitor/seed
storage/lipid transfer protein (LTP) family protein
[Arabidopsis thaliana]
Length = 170
Score = 56.2 bits (134), Expect = 2e-07
Identities = 22/84 (26%), Positives = 44/84 (52%), Gaps = 4/84 (4%)
Query: 30 SKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPECLCYIIQQTHKGSPESKSM 89
S C + + + PCL++ TG + +P ++CC+ + + + P+CLC ++ G +
Sbjct: 26 SSCTNALISMSPCLNYITGNSTSPNQQCCNQLSRVVQSSPDCLCQVL----NGGGSQLGI 81
Query: 90 GIQEDKLLQLPTVCHVNGANISDC 113
+ + + L LP C+V +S C
Sbjct: 82 NVNQTQALGLPRACNVQTPPVSRC 105
>gb|AAM64723.1| unknown [Arabidopsis thaliana]
Length = 183
Score = 55.8 bits (133), Expect = 2e-07
Identities = 25/86 (29%), Positives = 44/86 (51%), Gaps = 6/86 (6%)
Query: 29 ASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPECLCYIIQQTHKGSP-ESK 87
+S C S + + PCL + TG + TP + CC +S+ + P+C+C + SP +
Sbjct: 27 SSSCVSTLTTLSPCLSYITGNSTTPSQPCCSRLDSVIKSSPQCICSAV-----NSPIPNI 81
Query: 88 SMGIQEDKLLQLPTVCHVNGANISDC 113
+ I + LQLP C++ ++ C
Sbjct: 82 GLNINRTQALQLPNACNIQTPPLTQC 107
>gb|AAM51353.1| unknown protein [Arabidopsis thaliana] gi|17529320|gb|AAL38887.1|
unknown protein [Arabidopsis thaliana]
gi|25409096|pir||G84923 hypothetical protein At2g48130
[imported] - Arabidopsis thaliana
gi|18407534|ref|NP_566126.1| protease inhibitor/seed
storage/lipid transfer protein (LTP) family protein
[Arabidopsis thaliana]
Length = 183
Score = 55.8 bits (133), Expect = 2e-07
Identities = 25/86 (29%), Positives = 44/86 (51%), Gaps = 6/86 (6%)
Query: 29 ASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPECLCYIIQQTHKGSP-ESK 87
+S C S + + PCL + TG + TP + CC +S+ + P+C+C + SP +
Sbjct: 27 SSSCVSTLTTLSPCLSYITGNSTTPSQPCCSRLDSVIKSSPQCICSAV-----NSPIPNI 81
Query: 88 SMGIQEDKLLQLPTVCHVNGANISDC 113
+ I + LQLP C++ ++ C
Sbjct: 82 GLNINRTQALQLPNACNIQTPPLTQC 107
>gb|AAL23748.1| nonspecific lipid transfer protein [Bromus inermis]
Length = 124
Score = 55.8 bits (133), Expect = 2e-07
Identities = 30/87 (34%), Positives = 42/87 (47%), Gaps = 10/87 (11%)
Query: 25 AEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKA-----TDPECLCYIIQQT 79
AE S CG VV + PC+ +ATG+AP P CC S+ A D + C ++Q
Sbjct: 28 AEGAVSNCGQVVSYLAPCITYATGRAPGPNGGCCSGVRSLNAAASTTADRQATCSCLKQQ 87
Query: 80 HKGSPESKSMGIQEDKLLQLPTVCHVN 106
G GI+ D + +P+ C VN
Sbjct: 88 TSGMG-----GIKPDLVAGIPSKCGVN 109
>gb|AAM64671.1| unknown [Arabidopsis thaliana]
Length = 156
Score = 55.5 bits (132), Expect = 3e-07
Identities = 31/111 (27%), Positives = 47/111 (41%), Gaps = 5/111 (4%)
Query: 10 MCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDP 69
MCL +L + + S C +V+ + PCL F T P ++CC+ +
Sbjct: 5 MCL-ILFIALMRVMSIVSAQSSCTNVLISMAPCLSFITQNTSLPSQQCCNQLAHVVRYSS 63
Query: 70 ECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDCPSKSKTD 120
ECLC ++ G + + E + L LP CHV S C S S +
Sbjct: 64 ECLCQVLD----GGGSQLGINVNETQALALPKACHVQTPPASRCHSGSSVN 110
>dbj|BAC42777.1| putative non-specific lipid transfer protein nLTP [Arabidopsis
thaliana] gi|28973215|gb|AAO63932.1| unknown protein
[Arabidopsis thaliana] gi|3128175|gb|AAC16079.1| unknown
protein [Arabidopsis thaliana]
gi|15224862|ref|NP_181958.1| protease inhibitor/seed
storage/lipid transfer protein (LTP) family protein
(YLS3) [Arabidopsis thaliana] gi|7486420|pir||T02385
hypothetical protein At2g44290 [imported] - Arabidopsis
thaliana
Length = 205
Score = 55.5 bits (132), Expect = 3e-07
Identities = 31/108 (28%), Positives = 58/108 (53%), Gaps = 7/108 (6%)
Query: 10 MCLCVLALII--GGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKAT 67
+ L ++A+++ G + +D +C + + + CL + GKA +P +CC + +
Sbjct: 13 IALIMVAMVVDAAGADKGKD-KEECTAQLVGMATCLPYVQGKAKSPTPDCCSGLKQVINS 71
Query: 68 DPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDCPS 115
D +CLC IIQ+ + P+ + + L LP+VCH A+I+ CP+
Sbjct: 72 DMKCLCMIIQE--RNDPD-LGLQVNVSLALALPSVCHAT-ADITKCPA 115
>gb|AAL90923.1| At1g55260/F7A10_16 [Arabidopsis thaliana]
gi|18405294|ref|NP_564682.1| protease inhibitor/seed
storage/lipid transfer protein (LTP) family protein
[Arabidopsis thaliana] gi|15724272|gb|AAL06529.1|
At1g55260/F7A10_16 [Arabidopsis thaliana]
gi|12323173|gb|AAG51569.1| unknown protein; 63629-62263
[Arabidopsis thaliana] gi|25354706|pir||E96594 unknown
protein, 63629-62263 [imported] - Arabidopsis thaliana
Length = 184
Score = 55.5 bits (132), Expect = 3e-07
Identities = 31/120 (25%), Positives = 61/120 (50%), Gaps = 12/120 (10%)
Query: 1 MKNQQHQMFMCLCVLALIIGGCNGAEDLAS---KCGSVVQKVIPCLDFATGKAPTPKKEC 57
M+ +F+ + + ++++G G DLA +C + + ++ C+ + G A P K+C
Sbjct: 1 MEKSTRTLFITIVITSMLLGF--GNSDLAQDREECTNQLIELSTCIPYVGGDAKAPTKDC 58
Query: 58 CDAANSIKATDPECLCYIIQQTHKGSPESKSMGIQEDKLL--QLPTVCHVNGANISDCPS 115
C + +C+C +++ K P+ +GI+ + L LP+ CH+ NI+DC S
Sbjct: 59 CAGFGQVIRKSEKCVCILVRD--KDDPQ---LGIKINATLAAHLPSACHITAPNITDCIS 113
>ref|NP_567445.1| protease inhibitor/seed storage/lipid transfer protein (LTP) family
protein [Arabidopsis thaliana]
Length = 156
Score = 55.5 bits (132), Expect = 3e-07
Identities = 31/111 (27%), Positives = 47/111 (41%), Gaps = 5/111 (4%)
Query: 10 MCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDP 69
MCL +L + + S C +V+ + PCL F T P ++CC+ +
Sbjct: 5 MCL-ILFIALMRVMSIVSAQSSCTNVLISMAPCLSFITQNTSLPSQQCCNQLAHVVRYSS 63
Query: 70 ECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDCPSKSKTD 120
ECLC ++ G + + E + L LP CHV S C S S +
Sbjct: 64 ECLCQVLD----GGGSQLGINVNETQALALPKACHVETPPASRCHSGSSVN 110
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.319 0.134 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 206,358,481
Number of Sequences: 2540612
Number of extensions: 7597203
Number of successful extensions: 14253
Number of sequences better than 10.0: 284
Number of HSP's better than 10.0 without gapping: 115
Number of HSP's successfully gapped in prelim test: 169
Number of HSP's that attempted gapping in prelim test: 14077
Number of HSP's gapped (non-prelim): 295
length of query: 123
length of database: 863,360,394
effective HSP length: 99
effective length of query: 24
effective length of database: 611,839,806
effective search space: 14684155344
effective search space used: 14684155344
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 68 (30.8 bits)
Medicago: description of AC147538.3


