

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147496.7 + phase: 0
(110 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
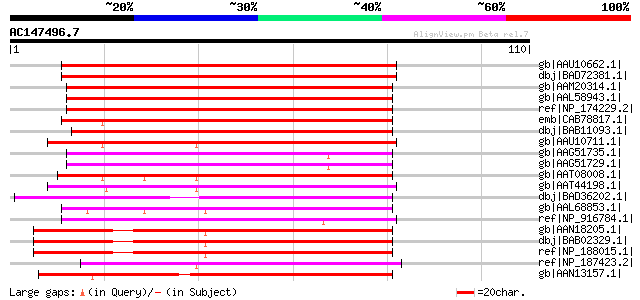
Score E
Sequences producing significant alignments: (bits) Value
gb|AAU10662.1| unknown protein [Oryza sativa (japonica cultivar-... 80 1e-14
dbj|BAD72381.1| TA9 protein-like [Oryza sativa (japonica cultiva... 79 2e-14
gb|AAM20314.1| unknown protein [Arabidopsis thaliana] gi|1738076... 79 3e-14
gb|AAL58943.1| At1g29350/F15D2_27 [Arabidopsis thaliana] 79 3e-14
ref|NP_174229.2| expressed protein [Arabidopsis thaliana] 79 3e-14
emb|CAB78817.1| hypothetical protein [Arabidopsis thaliana] gi|5... 77 8e-14
dbj|BAB11093.1| unnamed protein product [Arabidopsis thaliana] g... 71 7e-12
gb|AAU10711.1| unknown protein [Oryza sativa (japonica cultivar-... 65 5e-10
gb|AAG51735.1| unknown protein; 25451-20507 [Arabidopsis thalian... 64 1e-09
gb|AAG51729.1| unknown protein; 16040-11188 [Arabidopsis thalian... 64 1e-09
gb|AAT08008.1| unknown [Zea mays] 61 7e-09
gb|AAT44198.1| unknown protein [Oryza sativa (japonica cultivar-... 60 1e-08
dbj|BAD36202.1| hydroxyproline-rich glycoprotein-like [Oryza sat... 60 2e-08
gb|AAL68853.1| gb protein [Sorghum bicolor] 59 3e-08
ref|NP_916784.1| P0003E08.3 [Oryza sativa (japonica cultivar-gro... 57 1e-07
gb|AAN18205.1| At3g13990/MDC16_11 [Arabidopsis thaliana] gi|2202... 57 1e-07
dbj|BAB02329.1| unnamed protein product [Arabidopsis thaliana] 57 1e-07
ref|NP_188015.1| hydroxyproline-rich glycoprotein family protein... 57 1e-07
ref|NP_187423.2| expressed protein [Arabidopsis thaliana] 55 4e-07
gb|AAN13157.1| unknown protein [Arabidopsis thaliana] gi|1738085... 53 2e-06
>gb|AAU10662.1| unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 852
Score = 80.1 bits (196), Expect = 1e-14
Identities = 40/71 (56%), Positives = 52/71 (72%)
Query: 12 GGGGGVKAVKEVVTTAKNVVAAVKEIVNCTEQEIYDVLRECDMDPNLAVEKLLSQDTFRE 71
GGG G A V ++ +V ++KEIVN E EIY LR+C MDP+ AV +LLSQDTF+E
Sbjct: 11 GGGKGGAAAGPVPAASRKLVQSLKEIVNRPEAEIYAALRDCGMDPDEAVSRLLSQDTFQE 70
Query: 72 VRSKREKRKEV 82
V+SKR+K+KEV
Sbjct: 71 VKSKRDKKKEV 81
>dbj|BAD72381.1| TA9 protein-like [Oryza sativa (japonica cultivar-group)]
gi|55773800|dbj|BAD72338.1| TA9 protein-like [Oryza
sativa (japonica cultivar-group)]
Length = 851
Score = 79.3 bits (194), Expect = 2e-14
Identities = 40/71 (56%), Positives = 52/71 (72%)
Query: 12 GGGGGVKAVKEVVTTAKNVVAAVKEIVNCTEQEIYDVLRECDMDPNLAVEKLLSQDTFRE 71
GGGG A V +A+ +V +KEIVN + EIY LR+C MDP+ AV +LLSQDTF+E
Sbjct: 12 GGGGRGAAAGPVPGSARKLVQGLKEIVNRPDAEIYAALRDCGMDPDEAVSRLLSQDTFQE 71
Query: 72 VRSKREKRKEV 82
V+SKR+K+KEV
Sbjct: 72 VKSKRDKKKEV 82
>gb|AAM20314.1| unknown protein [Arabidopsis thaliana] gi|17380760|gb|AAL36210.1|
unknown protein [Arabidopsis thaliana]
gi|22329848|ref|NP_174230.2| kinase-related
[Arabidopsis thaliana]
Length = 831
Score = 78.6 bits (192), Expect = 3e-14
Identities = 35/69 (50%), Positives = 52/69 (74%)
Query: 13 GGGGVKAVKEVVTTAKNVVAAVKEIVNCTEQEIYDVLRECDMDPNLAVEKLLSQDTFREV 72
GGG K ++++ + ++ +V ++ EIVN E EIY +L+EC+MDPN V +LLSQD F EV
Sbjct: 7 GGGARKGIQDIPSGSRKIVQSLTEIVNSPEAEIYAMLKECNMDPNETVSRLLSQDPFHEV 66
Query: 73 RSKREKRKE 81
+SK+EK+KE
Sbjct: 67 KSKKEKKKE 75
>gb|AAL58943.1| At1g29350/F15D2_27 [Arabidopsis thaliana]
Length = 832
Score = 78.6 bits (192), Expect = 3e-14
Identities = 35/69 (50%), Positives = 52/69 (74%)
Query: 13 GGGGVKAVKEVVTTAKNVVAAVKEIVNCTEQEIYDVLRECDMDPNLAVEKLLSQDTFREV 72
GGG K ++++ + ++ +V ++ EIVN E EIY +L+EC+MDPN V +LLSQD F EV
Sbjct: 7 GGGARKGIQDIPSGSRKIVQSLTEIVNSPEAEIYAMLKECNMDPNETVSRLLSQDPFHEV 66
Query: 73 RSKREKRKE 81
+SK+EK+KE
Sbjct: 67 KSKKEKKKE 75
>ref|NP_174229.2| expressed protein [Arabidopsis thaliana]
Length = 831
Score = 78.6 bits (192), Expect = 3e-14
Identities = 35/69 (50%), Positives = 52/69 (74%)
Query: 13 GGGGVKAVKEVVTTAKNVVAAVKEIVNCTEQEIYDVLRECDMDPNLAVEKLLSQDTFREV 72
GGG K ++++ + ++ +V ++ EIVN E EIY +L+EC+MDPN V +LLSQD F EV
Sbjct: 7 GGGARKGIQDIPSGSRKIVQSLTEIVNSPEAEIYAMLKECNMDPNETVSRLLSQDPFHEV 66
Query: 73 RSKREKRKE 81
+SK+EK+KE
Sbjct: 67 KSKKEKKKE 75
>emb|CAB78817.1| hypothetical protein [Arabidopsis thaliana]
gi|5817001|emb|CAB53656.1| hypothetical protein
[Arabidopsis thaliana] gi|15236766|ref|NP_193549.1|
hypothetical protein [Arabidopsis thaliana]
gi|7485519|pir||T14815 hypothetical protein F15J5.120 -
Arabidopsis thaliana
Length = 762
Score = 77.4 bits (189), Expect = 8e-14
Identities = 40/71 (56%), Positives = 52/71 (72%), Gaps = 1/71 (1%)
Query: 12 GGGGGVK-AVKEVVTTAKNVVAAVKEIVNCTEQEIYDVLRECDMDPNLAVEKLLSQDTFR 70
GGGG V A V +++ V+ +KEIV C+E EIY +L ECDM+P+ AV +LLSQDTF
Sbjct: 3 GGGGSVSNANGGVPASSRKVIQDLKEIVECSELEIYAMLVECDMNPDEAVNRLLSQDTFH 62
Query: 71 EVRSKREKRKE 81
E +SKREK+KE
Sbjct: 63 EAKSKREKKKE 73
>dbj|BAB11093.1| unnamed protein product [Arabidopsis thaliana]
gi|15237431|ref|NP_199450.1| hypothetical protein
[Arabidopsis thaliana]
Length = 607
Score = 70.9 bits (172), Expect = 7e-12
Identities = 33/68 (48%), Positives = 50/68 (73%)
Query: 14 GGGVKAVKEVVTTAKNVVAAVKEIVNCTEQEIYDVLRECDMDPNLAVEKLLSQDTFREVR 73
G V + + +V ++KEIVNC+EQEIY +L EC+M+ + A+ +LLSQD+F+EV+
Sbjct: 2 GSSAIEVNAIPEYTRAIVRSMKEIVNCSEQEIYAMLVECNMNADEAITRLLSQDSFQEVK 61
Query: 74 SKREKRKE 81
SKR+K+KE
Sbjct: 62 SKRDKKKE 69
>gb|AAU10711.1| unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 852
Score = 64.7 bits (156), Expect = 5e-10
Identities = 35/80 (43%), Positives = 51/80 (63%), Gaps = 6/80 (7%)
Query: 9 RNSGGGGGVK-----AVKEVVTTAKNVVAAVKEIV-NCTEQEIYDVLRECDMDPNLAVEK 62
R GGGGG + A ++ +V +K I+ + +E EIY L +C MDP++AVE+
Sbjct: 7 RGGGGGGGGRVPFYAAAAAAEPASRKLVQGLKGILTDRSEAEIYATLLDCGMDPDVAVER 66
Query: 63 LLSQDTFREVRSKREKRKEV 82
L+SQD F EVR KR+K+KE+
Sbjct: 67 LISQDPFHEVRRKRDKKKEI 86
>gb|AAG51735.1| unknown protein; 25451-20507 [Arabidopsis thaliana]
gi|25403055|pir||C86416 unknown protein, 25451-20507
[imported] - Arabidopsis thaliana
Length = 858
Score = 63.5 bits (153), Expect = 1e-09
Identities = 35/97 (36%), Positives = 52/97 (53%), Gaps = 28/97 (28%)
Query: 13 GGGGVKAVKEVVTTAKNVVAAVKEIVNCTEQEIYDVLRECDMDPNLAVEKLLSQ------ 66
GGG K ++++ + ++ +V ++ EIVN E EIY +L+EC+MDPN V +LLSQ
Sbjct: 7 GGGARKGIQDIPSGSRKIVQSLTEIVNSPEAEIYAMLKECNMDPNETVSRLLSQAVAFNM 66
Query: 67 ----------------------DTFREVRSKREKRKE 81
D F EV+SK+EK+KE
Sbjct: 67 VPLVIVQLSSWFGFIEKKDVVNDPFHEVKSKKEKKKE 103
>gb|AAG51729.1| unknown protein; 16040-11188 [Arabidopsis thaliana]
gi|25403053|pir||B86416 unknown protein, 16040-11188
[imported] - Arabidopsis thaliana
Length = 858
Score = 63.5 bits (153), Expect = 1e-09
Identities = 35/97 (36%), Positives = 52/97 (53%), Gaps = 28/97 (28%)
Query: 13 GGGGVKAVKEVVTTAKNVVAAVKEIVNCTEQEIYDVLRECDMDPNLAVEKLLSQ------ 66
GGG K ++++ + ++ +V ++ EIVN E EIY +L+EC+MDPN V +LLSQ
Sbjct: 7 GGGARKGIQDIPSGSRKIVQSLTEIVNSPEAEIYAMLKECNMDPNETVSRLLSQAVAFNM 66
Query: 67 ----------------------DTFREVRSKREKRKE 81
D F EV+SK+EK+KE
Sbjct: 67 VPLVIVQLSSWFGFIEKKDVVNDPFHEVKSKKEKKKE 103
>gb|AAT08008.1| unknown [Zea mays]
Length = 942
Score = 60.8 bits (146), Expect = 7e-09
Identities = 34/80 (42%), Positives = 51/80 (63%), Gaps = 9/80 (11%)
Query: 11 SGGGGGVK-------AVKEVVTTA-KNVVAAVKEIV-NCTEQEIYDVLRECDMDPNLAVE 61
+GGGGG + A TTA ++ + ++KE+V ++ +I D LRE +MDPN +
Sbjct: 2 AGGGGGARGSDRAGAARSPTTTTAIQSTIQSIKEVVVGHSDADILDTLRESNMDPNETAQ 61
Query: 62 KLLSQDTFREVRSKREKRKE 81
KLL+QD F EV+ KR+K+KE
Sbjct: 62 KLLNQDPFHEVKRKRDKKKE 81
>gb|AAT44198.1| unknown protein [Oryza sativa (japonica cultivar-group)]
Length = 860
Score = 60.5 bits (145), Expect = 1e-08
Identities = 34/90 (37%), Positives = 51/90 (55%), Gaps = 16/90 (17%)
Query: 9 RNSGGGGGVKA---------------VKEVVTTAKNVVAAVKEIV-NCTEQEIYDVLREC 52
R GGGGG + + ++ +V +K I+ + +E EIY L +C
Sbjct: 7 RGGGGGGGGRVPFYAAAAAAEPRAGDAAAIPPASRKLVQGLKGILTDRSEAEIYATLLDC 66
Query: 53 DMDPNLAVEKLLSQDTFREVRSKREKRKEV 82
MDP++AVE+L+SQD F EVR KR+K+KE+
Sbjct: 67 GMDPDVAVERLISQDPFHEVRRKRDKKKEI 96
>dbj|BAD36202.1| hydroxyproline-rich glycoprotein-like [Oryza sativa (japonica
cultivar-group)] gi|51091151|dbj|BAD35846.1|
hydroxyproline-rich glycoprotein-like [Oryza sativa
(japonica cultivar-group)]
Length = 854
Score = 59.7 bits (143), Expect = 2e-08
Identities = 29/80 (36%), Positives = 46/80 (57%), Gaps = 6/80 (7%)
Query: 2 GSESVVNRNSGGGGGVKAVKEVVTTAKNVVAAVKEIVNCTEQEIYDVLRECDMDPNLAVE 61
G + + + G G A++ + + K VV ++ +IY LREC+MDPN +
Sbjct: 4 GGRAGTEKGAAAGRGQTAIQSTIQSIKEVVGGH------SDADIYAALRECNMDPNETTQ 57
Query: 62 KLLSQDTFREVRSKREKRKE 81
KLL+QD F EV+ KR+K++E
Sbjct: 58 KLLNQDPFHEVKRKRDKKRE 77
>gb|AAL68853.1| gb protein [Sorghum bicolor]
Length = 1009
Score = 58.9 bits (141), Expect = 3e-08
Identities = 34/82 (41%), Positives = 49/82 (59%), Gaps = 12/82 (14%)
Query: 12 GGGG---------GVKAVKEVVTTA--KNVVAAVKEIVNC-TEQEIYDVLRECDMDPNLA 59
GGGG G A + TTA + + ++KE+V ++ +I D LRE +MDPN
Sbjct: 5 GGGGAGARGSDRAGAGAARSPTTTAAIQTTIQSIKEVVGGHSDADILDTLRESNMDPNET 64
Query: 60 VEKLLSQDTFREVRSKREKRKE 81
+KLL+QD F EV+ KR+K+KE
Sbjct: 65 AQKLLNQDPFHEVKRKRDKKKE 86
>ref|NP_916784.1| P0003E08.3 [Oryza sativa (japonica cultivar-group)]
Length = 793
Score = 57.0 bits (136), Expect = 1e-07
Identities = 39/113 (34%), Positives = 52/113 (45%), Gaps = 42/113 (37%)
Query: 12 GGGGGVKAVKEVVTTAKNVVAAVKEIVNCTEQEIYDVLRECDMDPNLAVEKLLS------ 65
GGGG A V +A+ +V +KEIVN + EIY LR+C MDP+ AV +LLS
Sbjct: 399 GGGGRGAAAGPVPGSARKLVQGLKEIVNRPDAEIYAALRDCGMDPDEAVSRLLSQVFVAA 458
Query: 66 ------------------------------------QDTFREVRSKREKRKEV 82
Q+TF+EV+SKR+K+KEV
Sbjct: 459 LVALGLFRGDWIGLDWIEVLGSSVPVLDVRRRALFIQNTFQEVKSKRDKKKEV 511
>gb|AAN18205.1| At3g13990/MDC16_11 [Arabidopsis thaliana]
gi|22022592|gb|AAM83252.1| AT3g13990/MDC16_11
[Arabidopsis thaliana]
Length = 848
Score = 57.0 bits (136), Expect = 1e-07
Identities = 31/77 (40%), Positives = 48/77 (62%), Gaps = 5/77 (6%)
Query: 6 VVNRNSGGGGGVKAVKEVVTTAKNVVAAVKEIVNC-TEQEIYDVLRECDMDPNLAVEKLL 64
V + G GV E AK ++ ++KE+V+ ++ +IY L+E +MD N AVEKL+
Sbjct: 2 VTGSRTSGNRGVGLDDE----AKKMIQSIKEVVDSHSDADIYTALKEANMDANEAVEKLI 57
Query: 65 SQDTFREVRSKREKRKE 81
QD F EV+ KR+++KE
Sbjct: 58 HQDPFHEVKRKRDRKKE 74
>dbj|BAB02329.1| unnamed protein product [Arabidopsis thaliana]
Length = 870
Score = 57.0 bits (136), Expect = 1e-07
Identities = 31/77 (40%), Positives = 48/77 (62%), Gaps = 5/77 (6%)
Query: 6 VVNRNSGGGGGVKAVKEVVTTAKNVVAAVKEIVNC-TEQEIYDVLRECDMDPNLAVEKLL 64
V + G GV E AK ++ ++KE+V+ ++ +IY L+E +MD N AVEKL+
Sbjct: 2 VTGSRTSGNRGVGLDDE----AKKMIQSIKEVVDSHSDADIYTALKEANMDANEAVEKLI 57
Query: 65 SQDTFREVRSKREKRKE 81
QD F EV+ KR+++KE
Sbjct: 58 HQDPFHEVKRKRDRKKE 74
>ref|NP_188015.1| hydroxyproline-rich glycoprotein family protein [Arabidopsis
thaliana]
Length = 883
Score = 57.0 bits (136), Expect = 1e-07
Identities = 31/77 (40%), Positives = 48/77 (62%), Gaps = 5/77 (6%)
Query: 6 VVNRNSGGGGGVKAVKEVVTTAKNVVAAVKEIVNC-TEQEIYDVLRECDMDPNLAVEKLL 64
V + G GV E AK ++ ++KE+V+ ++ +IY L+E +MD N AVEKL+
Sbjct: 2 VTGSRTSGNRGVGLDDE----AKKMIQSIKEVVDSHSDADIYTALKEANMDANEAVEKLI 57
Query: 65 SQDTFREVRSKREKRKE 81
QD F EV+ KR+++KE
Sbjct: 58 HQDPFHEVKRKRDRKKE 74
>ref|NP_187423.2| expressed protein [Arabidopsis thaliana]
Length = 841
Score = 55.1 bits (131), Expect = 4e-07
Identities = 29/70 (41%), Positives = 42/70 (59%), Gaps = 2/70 (2%)
Query: 16 GVKAVKEVVTTAKNVVAAVKEIV--NCTEQEIYDVLRECDMDPNLAVEKLLSQDTFREVR 73
G A + T + ++ +KE N +E EI +L EC+MDP+ ++LL QD F EV+
Sbjct: 3 GSGARVSISATTRKMIQNIKETTAGNYSEDEIIAMLHECNMDPDETAQRLLLQDPFHEVK 62
Query: 74 SKREKRKEVI 83
KR+KRKE I
Sbjct: 63 KKRDKRKENI 72
>gb|AAN13157.1| unknown protein [Arabidopsis thaliana] gi|17380854|gb|AAL36239.1|
unknown protein [Arabidopsis thaliana]
gi|42564117|ref|NP_187929.4| expressed protein
[Arabidopsis thaliana]
Length = 567
Score = 52.8 bits (125), Expect = 2e-06
Identities = 29/76 (38%), Positives = 47/76 (61%), Gaps = 3/76 (3%)
Query: 7 VNRNSGGGGG-VKAVKEVVTTAKNVVAAVKEIVNCTEQEIYDVLRECDMDPNLAVEKLLS 65
++R SG GG V +++ T +N+ + ++++I+ V +EC DP+ +KLL
Sbjct: 1 MSRISGDGGSRVSIPADLLQTIQNIREVTGK--QHSDEDIFSVFKECFSDPHETTQKLLY 58
Query: 66 QDTFREVRSKREKRKE 81
DTF EVRSKRE++KE
Sbjct: 59 LDTFHEVRSKRERKKE 74
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.319 0.135 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 159,332,265
Number of Sequences: 2540612
Number of extensions: 5261429
Number of successful extensions: 21682
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 29
Number of HSP's successfully gapped in prelim test: 20
Number of HSP's that attempted gapping in prelim test: 21645
Number of HSP's gapped (non-prelim): 54
length of query: 110
length of database: 863,360,394
effective HSP length: 86
effective length of query: 24
effective length of database: 644,867,762
effective search space: 15476826288
effective search space used: 15476826288
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 68 (30.8 bits)
Medicago: description of AC147496.7


