

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147496.2 - phase: 0 /pseudo
(550 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
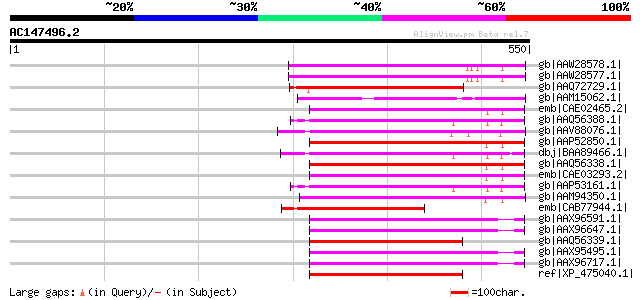
Score E
Sequences producing significant alignments: (bits) Value
gb|AAW28578.1| putative gag-pol polyprotein [Solanum demissum] 191 7e-47
gb|AAW28577.1| putative gag-pol polyprotein [Solanum demissum] 191 7e-47
gb|AAQ72729.1| putative gag-pol polyprotein [Petunia x hybrida] 178 3e-43
gb|AAM15062.1| putative retroelement integrase [Arabidopsis thal... 178 5e-43
emb|CAE02465.2| OSJNBa0042D13.18 [Oryza sativa (japonica cultiva... 177 6e-43
gb|AAQ56388.1| putative gag-pol polyprotein [Oryza sativa (japon... 177 8e-43
gb|AAV88076.1| putative retrotransposon polyprotein [Ipomoea bat... 176 2e-42
gb|AAP52850.1| putative retroelement [Oryza sativa (japonica cul... 175 3e-42
dbj|BAA89466.1| gag-pol polyprotein [Oryza sativa (indica cultiv... 175 3e-42
gb|AAQ56338.1| putative gag-pol polyprotein [Oryza sativa (japon... 175 3e-42
emb|CAE03293.2| OSJNBb0046P18.9 [Oryza sativa (japonica cultivar... 174 5e-42
gb|AAP53161.1| putative retroelement [Oryza sativa (japonica cul... 172 2e-41
gb|AAM94350.1| gag-pol polyprotein [Zea mays] 171 4e-41
emb|CAB77944.1| putative polyprotein [Arabidopsis thaliana] gi|4... 171 7e-41
gb|AAX96591.1| retrotransposon protein, putative, Ty3-gypsy sub-... 171 7e-41
gb|AAX96647.1| retrotransposon protein, putative, Ty3-gypsy sub-... 171 7e-41
gb|AAQ56339.1| putative gag-pol polyprotein [Oryza sativa (japon... 168 4e-40
gb|AAX95495.1| Retrotransposon gag protein, putative [Oryza sati... 168 5e-40
gb|AAX96717.1| retrotransposon protein, putative, Ty3-gypsy sub-... 168 5e-40
ref|XP_475040.1| OSJNBa0042F21.10 [Oryza sativa (japonica cultiv... 167 8e-40
>gb|AAW28578.1| putative gag-pol polyprotein [Solanum demissum]
Length = 1588
Score = 191 bits (484), Expect = 7e-47
Identities = 114/303 (37%), Positives = 161/303 (52%), Gaps = 52/303 (17%)
Query: 296 EVENHEGYVEEEAEEIPS-GDLFMIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLII 354
E E EG EE+ E+P+ G + ++RR + + QRE LFHTRC ++GK +II
Sbjct: 379 EEEEGEGVSEEDDLELPNDGIIGVVRRIMTINLGSVDEGQRENLFHTRCGIKGKTYSMII 438
Query: 355 DGGSRTNVASTRLVSKMELETKPHPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLC 414
DGGS NV S+ LV K+ + PY+LQWLN+ E+ V+KQ + F +G+YED++LC
Sbjct: 439 DGGSCANVVSSYLVDKLGIACMKRSTPYRLQWLNDCGEVQVNKQCMISFNVGRYEDEILC 498
Query: 415 DVVPMEASHLLLGRP*QFDRSVLHDGRTNKYSFMHSGQKISLAPLSPSEVRDDQKKRKEK 474
DVVPM+A H+LLGRP Q+DR H GR N+YS +H+G+K +LAPLSPS+V +DQK+ +E
Sbjct: 499 DVVPMQACHVLLGRPWQYDRDTTHHGRKNRYSLLHNGKKYTLAPLSPSQVFEDQKRLRET 558
Query: 475 YEKEKRKIK-------------------RKERE------------------FA*KKK--- 494
K+K ++K KERE F +K+
Sbjct: 559 MGKQKGEVKSELEGKEKGQEIEKKKDGREKEREGSDLSKEVKEGVSKKIKCFGERKEAKE 618
Query: 495 --------KRRVSRNSHEQTAPIFTTLQKVALLT---NTQNFPSCTKFLLQECEDVFPKK 543
K + N+ + PI K L+ T + P+ LLQ EDVFP
Sbjct: 619 KKNESLYIKAKECLNARREGLPIILLTYKEILINFEQLTPSLPNSVSSLLQNFEDVFPDD 678
Query: 544 VPQ 546
P+
Sbjct: 679 TPK 681
>gb|AAW28577.1| putative gag-pol polyprotein [Solanum demissum]
Length = 1588
Score = 191 bits (484), Expect = 7e-47
Identities = 114/303 (37%), Positives = 161/303 (52%), Gaps = 52/303 (17%)
Query: 296 EVENHEGYVEEEAEEIPS-GDLFMIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLII 354
E E EG EE+ E+P+ G + ++RR + + QRE LFHTRC ++GK +II
Sbjct: 379 EEEEGEGVSEEDDLELPNDGIIGVVRRIMTINLGSVDEGQRENLFHTRCGIKGKTYSMII 438
Query: 355 DGGSRTNVASTRLVSKMELETKPHPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLC 414
DGGS NV S+ LV K+ + PY+LQWLN+ E+ V+KQ + F +G+YED++LC
Sbjct: 439 DGGSCANVVSSYLVDKLGIACMKRSTPYRLQWLNDCGEVQVNKQCMISFNVGRYEDEILC 498
Query: 415 DVVPMEASHLLLGRP*QFDRSVLHDGRTNKYSFMHSGQKISLAPLSPSEVRDDQKKRKEK 474
DVVPM+A H+LLGRP Q+DR H GR N+YS +H+G+K +LAPLSPS+V +DQK+ +E
Sbjct: 499 DVVPMQACHVLLGRPWQYDRDTTHHGRKNRYSLLHNGKKYTLAPLSPSQVFEDQKRLRET 558
Query: 475 YEKEKRKIK-------------------RKERE------------------FA*KKK--- 494
K+K ++K KERE F +K+
Sbjct: 559 MGKQKGEVKSELEGKEKGQEIEKKKDGREKEREGSDLSKEVKEGVSKKIKCFGERKEAKE 618
Query: 495 --------KRRVSRNSHEQTAPIFTTLQKVALLT---NTQNFPSCTKFLLQECEDVFPKK 543
K + N+ + PI K L+ T + P+ LLQ EDVFP
Sbjct: 619 KKNESLYIKAKECLNARREGLPIILLTYKEILINFEQLTPSLPNSVSSLLQNFEDVFPDD 678
Query: 544 VPQ 546
P+
Sbjct: 679 TPK 681
>gb|AAQ72729.1| putative gag-pol polyprotein [Petunia x hybrida]
Length = 803
Score = 178 bits (452), Expect = 3e-43
Identities = 96/191 (50%), Positives = 123/191 (64%), Gaps = 7/191 (3%)
Query: 297 VENHEGYVEEEAEEIPSGD-----LFMIRRFLGNQAKEE-ESNQRETLFHTRCLVQGKVC 350
VE EG VEE+ E+ D ++RR LG E+ E+ QRE LFH RC V+G VC
Sbjct: 421 VEGDEG-VEEDGEDDTVEDEEPHTFLVVRRSLGAMIAEDGETLQRENLFHARCRVKGVVC 479
Query: 351 FLIIDGGSRTNVASTRLVSKMELETKPHPKPYKLQWLNENVEILVDKQVEVCFKIGKYED 410
+IID GS TNV S L+++++L T HPKPY+LQWL+E E+ V KQ + FK+G Y D
Sbjct: 480 SMIIDSGSCTNVVSQSLITQLKLPTMKHPKPYRLQWLSECGELKVVKQAMIKFKMGSYVD 539
Query: 411 DVLCDVVPMEASHLLLGRP*QFDRSVLHDGRTNKYSFMHSGQKISLAPLSPSEVRDDQKK 470
+ L DVVPM+A HLL GRP QFDR V+H+GR N Y+ H G+K L PL+P +V DD +
Sbjct: 540 EHLFDVVPMQACHLLFGRPWQFDRQVVHNGRVNNYTLQHEGRKFVLKPLTPKQVSDDYNR 599
Query: 471 RKEKYEKEKRK 481
KE E K K
Sbjct: 600 MKELREAAKAK 610
>gb|AAM15062.1| putative retroelement integrase [Arabidopsis thaliana]
gi|25411256|pir||E84480 probable retroelement integrase
[imported] - Arabidopsis thaliana
Length = 1215
Score = 178 bits (451), Expect = 5e-43
Identities = 109/243 (44%), Positives = 145/243 (58%), Gaps = 19/243 (7%)
Query: 306 EEAEEIPS-GDLFMIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVAS 364
+E EE+P+ G+L + RR L Q K +E QR+ LFHTRC V GKVC LIIDGGS TNVAS
Sbjct: 200 KENEELPAQGELLVARRTLSVQTKTDEQEQRKNLFHTRCHVHGKVCSLIIDGGSCTNVAS 259
Query: 365 TRLVSKMELETKPHPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVVPMEASHL 424
+V K+ L+ WLN++ ++ V QV V IGKYED++LCDV+PMEA H+
Sbjct: 260 ETMVKKLGLK-----------WLNDSGKMRVKNQVVVPIVIGKYEDEILCDVLPMEAGHI 308
Query: 425 LLGRP*QFDRSVLHDGRTNKYSFMHSGQKISLAPLSPSEVRDDQKKRKEKYEKEKRKIKR 484
LLGRP Q DR V+HDG TN++SF G K L ++P EV DQ K+K K ++ +
Sbjct: 309 LLGRPWQSDRKVMHDGFTNRHSFEFKGGKTILVSMTPHEVYQDQIHLKQK----KEQVVK 364
Query: 485 KEREFA*KKKKRRVSRNSHEQTAPIFTTLQKVALLTN-TQNFPSCTKFLLQECEDVFPKK 543
+ FA K S S +Q +F + + LTN PS LLQ+ +DVFP+
Sbjct: 365 QPNFFA--KSGEVKSAYSSKQPMLLFVFKEALTSLTNFAPVLPSEMTSLLQDYKDVFPED 422
Query: 544 VPQ 546
P+
Sbjct: 423 NPK 425
>emb|CAE02465.2| OSJNBa0042D13.18 [Oryza sativa (japonica cultivar-group)]
gi|50922051|ref|XP_471386.1| OSJNBa0042D13.18 [Oryza
sativa (japonica cultivar-group)]
Length = 2241
Score = 177 bits (450), Expect = 6e-43
Identities = 99/239 (41%), Positives = 143/239 (59%), Gaps = 11/239 (4%)
Query: 318 MIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVASTRLVSKMELETKP 377
+++R L Q ++ E NQR TLF T+C+V+ + C +IIDGGS N+AS+ +V K+ L TKP
Sbjct: 553 IVQRVLSAQMEKAEQNQRHTLFQTKCVVKERCCRMIIDGGSCNNLASSEMVEKLALSTKP 612
Query: 378 HPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVVPMEASHLLLGRP*QFDRSVL 437
HP PY +QWLN + + V K V + F IG Y D V CDVVPM+A ++LLGRP QFDR +
Sbjct: 613 HPHPYCIQWLNNSGKAKVTKLVHINFAIGNYHDVVECDVVPMQACNILLGRPWQFDRDSM 672
Query: 438 HDGRTNKYSFMHSGQKISLAPLSPSEV-RDD-QKKRKEKYEKEKRKIKRKEREFA*KKKK 495
H GR+N+YSF++ +KI L P+SP ++ RDD K K K E +K+ ++ K
Sbjct: 673 HHGRSNQYSFLYHDKKIVLHPMSPEDILRDDVAKAAKSKCESDKKAQSDGKKPETINLKP 732
Query: 496 RRVSRNSHE------QTAPIFTTLQKVALLT---NTQNFPSCTKFLLQECEDVFPKKVP 545
R + + + + + K AL++ + P +LQE DVFPK+VP
Sbjct: 733 RYLLATKSDINELIASPSVAYALVCKDALISLHDMQHSLPPAVANILQEYSDVFPKEVP 791
>gb|AAQ56388.1| putative gag-pol polyprotein [Oryza sativa (japonica
cultivar-group)]
Length = 1616
Score = 177 bits (449), Expect = 8e-43
Identities = 103/276 (37%), Positives = 150/276 (54%), Gaps = 33/276 (11%)
Query: 298 ENHEGYVEEEAEEIPSGDLFMIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGG 357
E H G E AE S +++R L Q + E NQR TLF T+C+++ + C +IIDGG
Sbjct: 455 EEHIG--AEAAEHYES---LVVQRVLSAQMERAEQNQRHTLFQTKCVIKERSCRVIIDGG 509
Query: 358 SRTNVASTRLVSKMELETKPHPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVV 417
S N+AS +V K+ L T+PHP+PY +QWLN + ++ V + V V F IG Y D + CDVV
Sbjct: 510 SCNNLASAEMVEKLALSTQPHPQPYYIQWLNSSGKVKVTRLVRVHFAIGSYHDSINCDVV 569
Query: 418 PMEASHLLLGRP*QFDRSVLHDGRTNKYSFMHSGQKISLAPLSPSEVRDDQ-----KKRK 472
PM+A +LLGRP QFD+ LH G++N+YSF+H+G+K+ L P+SP + D+ K++
Sbjct: 570 PMQACSMLLGRPWQFDKDSLHFGKSNQYSFVHNGKKLVLHPMSPEVILKDELARASKQKN 629
Query: 473 EKYEKEKRKIKRKEREFA*KKKKRRVSRNSHE--------------------QTAPIFTT 512
+++ + + I E E KK V N +E T +
Sbjct: 630 QEHTRSEHLIAANELEKHKKKPTNSVQNNKNEIKLKGSCFIATKSDLDEVDTDTVVCYAL 689
Query: 513 LQKVALL---TNTQNFPSCTKFLLQECEDVFPKKVP 545
+ K L + P LLQE D+FPK+VP
Sbjct: 690 VCKETLFPIEDTPISLPPPVTNLLQEYADIFPKEVP 725
>gb|AAV88076.1| putative retrotransposon polyprotein [Ipomoea batatas]
Length = 1358
Score = 176 bits (445), Expect = 2e-42
Identities = 102/291 (35%), Positives = 160/291 (54%), Gaps = 33/291 (11%)
Query: 285 ENYDDLGE*NVEVENHEGYVEEEAEEIPSGDLFMIRRFLGNQAKEEESNQRETLFHTRCL 344
E + +GE + + E E++A ++ + L ++ QRE +F+ +C
Sbjct: 355 EELEPIGERQKDDHSEEEVQEDDALHFNC----VVHKALSTLVVLDQEEQRENIFYGKCK 410
Query: 345 VQGKVCFLIIDGGSRTNVASTRLVSKMELETKPHPKPYKLQWLNENVEILVDKQVEVCFK 404
+ G C IIDGGS TNV S +V+ M++ T HP+PYKLQWLN++ E+ V KQ +
Sbjct: 411 IPGATCSFIIDGGSCTNVISEDVVNAMKIPTIQHPQPYKLQWLNDDGELKVHKQALISIS 470
Query: 405 IGKYEDDVLCDVVPMEASHLLLGRP*QFDRSVLHDGRTNKYSFMHSGQKISLAPLSPSEV 464
IGKY+DDVLCDV+PM A H+LLGRP Q+DR LH G+TNKY+ G+K +L PL+P EV
Sbjct: 471 IGKYQDDVLCDVIPMHACHILLGRPWQYDRDTLHHGKTNKYTIHKGGKKYTLTPLAPKEV 530
Query: 465 RD----DQKKRKEKYEKEKRKIKR----KEREFA*KKKKRR--VSRNSHEQTAPIFTTLQ 514
+ +K R+E +K K +K K+ A +KK+R+ + +++ + + + T +
Sbjct: 531 YNLQVQSKKLREELAQKAKEAMKETTSGKQNTIAHEKKQRKEGMKKDTTQSSHNLLMTKR 590
Query: 515 KVAL-------------------LTNTQNFPSCTKFLLQECEDVFPKKVPQ 546
+V + ++ PS LL E DVFP+++P+
Sbjct: 591 EVEQALRRGEGVFLLYPIDFCLNVIKSEIIPSDVSALLSEFADVFPEELPK 641
>gb|AAP52850.1| putative retroelement [Oryza sativa (japonica cultivar-group)]
gi|37532522|ref|NP_920563.1| putative retroelement
[Oryza sativa (japonica cultivar-group)]
gi|13992688|gb|AAK51582.1| Putative retroelement [Oryza
sativa]
Length = 2447
Score = 175 bits (444), Expect = 3e-42
Identities = 101/239 (42%), Positives = 145/239 (60%), Gaps = 11/239 (4%)
Query: 318 MIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVASTRLVSKMELETKP 377
+++R L Q ++ E NQR TLF T+C+V+ + C +IIDGGS N+AS+ +V K+ L TKP
Sbjct: 459 IVQRVLSAQMEKAEQNQRHTLFQTKCVVKERCCRMIIDGGSCNNLASSEMVEKLALSTKP 518
Query: 378 HPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVVPMEASHLLLGRP*QFDRSVL 437
HP PY +QWLN + + V K V + F IG Y D V CDVVPM+A ++LLGRP QFDR +
Sbjct: 519 HPHPYYIQWLNNSGKAKVTKLVHINFAIGNYHDVVECDVVPMQACNILLGRPWQFDRDSM 578
Query: 438 HDGRTNKYSFMHSGQKISLAPLSPSEV-RDD-QKKRKEKYEKEKR-KIKRKEREFA*KKK 494
H GR+N+YSF++ +KI L +SP ++ RDD K K K E +K+ + K+ E K
Sbjct: 579 HHGRSNQYSFLYHDKKIVLHSMSPEDILRDDVAKAAKSKCESDKKAQSDGKKPETINLKP 638
Query: 495 KRRVSRNSH-----EQTAPIFTTLQKVALLT---NTQNFPSCTKFLLQECEDVFPKKVP 545
K ++ S + + + K AL++ + P +LQE DVFPK+VP
Sbjct: 639 KCLLATKSDINELIASPSVAYALVCKDALISLHDMQHSLPPAVANILQEYSDVFPKEVP 697
>dbj|BAA89466.1| gag-pol polyprotein [Oryza sativa (indica cultivar-group)]
Length = 1587
Score = 175 bits (444), Expect = 3e-42
Identities = 104/287 (36%), Positives = 155/287 (53%), Gaps = 33/287 (11%)
Query: 288 DDLGE*NVEVENHEGYVEEEAEEIPSGDLFMIRRFLGNQAKEEESNQRETLFHTRCLVQG 347
++ GE + ++ E E AE S +++R L Q + E NQ TLF T+C+++
Sbjct: 443 NNAGEGDAPHQDEEHIRAEAAEHYES---LVVQRVLSAQMERAEQNQLHTLFQTKCVIKE 499
Query: 348 KVCFLIIDGGSRTNVASTRLVSKMELETKPHPKPYKLQWLNENVEILVDKQVEVCFKIGK 407
+ C +IIDGGS N+AS +V K+ L T+PHP+PY +QWLN + ++ V + V V F IG
Sbjct: 500 RSCHVIIDGGSCNNLASAEMVEKLALSTQPHPQPYYIQWLNSSGKVKVTRLVRVHFAIGS 559
Query: 408 YEDDVLCDVVPMEASHLLLGRP*QFDRSVLHDGRTNKYSFMHSGQKISLAPLSPSEVRDD 467
Y D + CDVVPM+A +LLGRP QFD+ LH G++N+YSF+H+G+K+ L P+SP + D
Sbjct: 560 YHDSINCDVVPMQACSMLLGRPWQFDKDSLHFGKSNQYSFVHNGKKLVLHPMSPEVILKD 619
Query: 468 Q-----KKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHE------------------ 504
+ K++ +++ + + I E E KK V N +E
Sbjct: 620 ELARASKQKNQEHTRSEHLIAANELEKHKKKPTNSVQNNKNEIKLKGSCFIATKSDLDEV 679
Query: 505 --QTAPIFTTLQKVALL----TNTQNFPSCTKFLLQECEDVFPKKVP 545
T + + K L T P T LLQE D+FPK+VP
Sbjct: 680 DTDTVFCYALVCKETLFPIEDTPISLRPPVTN-LLQEYADIFPKEVP 725
>gb|AAQ56338.1| putative gag-pol polyprotein [Oryza sativa (japonica
cultivar-group)]
Length = 1619
Score = 175 bits (444), Expect = 3e-42
Identities = 101/239 (42%), Positives = 145/239 (60%), Gaps = 11/239 (4%)
Query: 318 MIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVASTRLVSKMELETKP 377
+++R L Q ++ E NQR TLF T+C+V+ + C +IIDGGS N+AS+ +V K+ L TKP
Sbjct: 438 IVQRVLSAQMEKAEQNQRHTLFQTKCVVKERCCRMIIDGGSCNNLASSEMVEKLALSTKP 497
Query: 378 HPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVVPMEASHLLLGRP*QFDRSVL 437
HP Y +QWLN + + V K V + F IG Y D V CDVVPM+A ++LLGRP QFDR +
Sbjct: 498 HPHSYYIQWLNNSGKAKVTKLVHINFAIGNYHDVVECDVVPMQACNILLGRPWQFDRDSM 557
Query: 438 HDGRTNKYSFMHSGQKISLAPLSPSEV-RDD-QKKRKEKYEKEKR-KIKRKEREFA*KKK 494
H GR+N+YSF++ +KI L P+SP ++ RDD K K K E +K+ + K+ E K
Sbjct: 558 HHGRSNQYSFLYHDKKIVLHPMSPEDILRDDVAKAAKSKCESDKKAQSDGKKPETINLKP 617
Query: 495 KRRVSRNSH-----EQTAPIFTTLQKVALLT---NTQNFPSCTKFLLQECEDVFPKKVP 545
K ++ S + + + K AL++ + P +LQE DVFPK+VP
Sbjct: 618 KCLLATKSDINELIASPSVAYALVCKDALISLHDMQHSLPPAVANILQEYSDVFPKEVP 676
>emb|CAE03293.2| OSJNBb0046P18.9 [Oryza sativa (japonica cultivar-group)]
gi|38344456|emb|CAE04927.2| OSJNBa0017P10.4 [Oryza
sativa (japonica cultivar-group)]
gi|50921961|ref|XP_471341.1| OSJNBb0046P18.9 [Oryza
sativa (japonica cultivar-group)]
Length = 1134
Score = 174 bits (442), Expect = 5e-42
Identities = 99/239 (41%), Positives = 143/239 (59%), Gaps = 11/239 (4%)
Query: 318 MIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVASTRLVSKMELETKP 377
+++R L Q ++ E NQR TLF T+C+V+ + C +IIDGGS N+AS+ +V K+ L TKP
Sbjct: 642 IVQRVLSTQMEKAEQNQRHTLFQTKCVVKERCCRMIIDGGSCNNLASSEMVEKLALSTKP 701
Query: 378 HPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVVPMEASHLLLGRP*QFDRSVL 437
HP PY +QWLN + + V K V + F IG Y D V CDVVPM+A ++LLGRP QFD+ L
Sbjct: 702 HPHPYYIQWLNNSGKAKVTKLVHINFAIGNYHDVVECDVVPMQACNILLGRPWQFDKDSL 761
Query: 438 HDGRTNKYSFMHSGQKISLAPLSPSEVRDDQ--KKRKEKYEKEKR-KIKRKEREFA*KKK 494
H GR+N+YSF++ +KI L P+S ++ D K K K E +K+ + K+ E K
Sbjct: 762 HHGRSNQYSFLYHDKKIVLHPMSSEDILHDDVAKAAKSKCESDKKAQSDGKKPETINLKP 821
Query: 495 KRRVSRNSH-----EQTAPIFTTLQKVALLT---NTQNFPSCTKFLLQECEDVFPKKVP 545
K ++ S + + + K AL++ + P +LQE DVFPK+VP
Sbjct: 822 KCLLATKSDINELIASPSVAYALVCKDALISLHDMQHSLPPAIANILQEYSDVFPKEVP 880
>gb|AAP53161.1| putative retroelement [Oryza sativa (japonica cultivar-group)]
gi|37533144|ref|NP_920874.1| putative retroelement
[Oryza sativa (japonica cultivar-group)]
gi|15187182|gb|AAK91332.1| Putative gag-pol polyprotein
[Oryza sativa] gi|15217296|gb|AAK92640.1| Putative
retroelement [Oryza sativa]
Length = 1708
Score = 172 bits (437), Expect = 2e-41
Identities = 101/276 (36%), Positives = 148/276 (53%), Gaps = 33/276 (11%)
Query: 298 ENHEGYVEEEAEEIPSGDLFMIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGG 357
E H G E AE S +++R L Q + E NQR TLF T+C+++ + C +IID G
Sbjct: 455 EEHIG--AEAAEHYES---LVVQRVLSAQMERAEQNQRHTLFQTKCVIKERSCRVIIDRG 509
Query: 358 SRTNVASTRLVSKMELETKPHPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVV 417
S N+AS +V K+ L T+PHP+PY +QWLN + ++ V + V V F IG Y D + CDVV
Sbjct: 510 SCNNLASAEMVEKLALSTQPHPQPYYIQWLNSSGKVKVTRLVRVHFAIGSYHDSINCDVV 569
Query: 418 PMEASHLLLGRP*QFDRSVLHDGRTNKYSFMHSGQKISLAPLSPSEVRDDQ-----KKRK 472
PM+A + LGRP QFD+ LH G++N+YSF+H+G+K+ L P+SP + D+ K++
Sbjct: 570 PMQACSIFLGRPWQFDKDSLHFGKSNQYSFVHNGKKLVLHPMSPEVILKDELARASKQKN 629
Query: 473 EKYEKEKRKIKRKEREFA*KKKKRRVSRNSHE--------------------QTAPIFTT 512
+++ + + I E E KK V N +E T +
Sbjct: 630 QEHTRSEHLIAANELEKHKKKPTNSVQNNKNEIKLKGSCFIATKSDLDEVDTDTVVCYAL 689
Query: 513 LQKVALL---TNTQNFPSCTKFLLQECEDVFPKKVP 545
+ K L + P LLQE D+FPK+VP
Sbjct: 690 VCKETLFPIEDTPISLPPPVTNLLQEYADIFPKEVP 725
>gb|AAM94350.1| gag-pol polyprotein [Zea mays]
Length = 1618
Score = 171 bits (434), Expect = 4e-41
Identities = 99/250 (39%), Positives = 145/250 (57%), Gaps = 11/250 (4%)
Query: 308 AEEIPSGDLFMIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVASTRL 367
A++ + +++R L Q ++ E NQR TLF T+C+++ + C LIIDGGS N+AS+ +
Sbjct: 452 ADDAEHYESLIVQRVLSAQMEKAEQNQRHTLFQTKCVIKERSCRLIIDGGSCNNLASSDM 511
Query: 368 VSKMELETKPHPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVVPMEASHLLLG 427
V K+ L TKPHP PY +QWLN + ++ V K V + F IG Y D V CDVVPM+A ++LLG
Sbjct: 512 VEKLALTTKPHPHPYHIQWLNNSGKVKVTKLVRINFAIGSYRDVVDCDVVPMDACNILLG 571
Query: 428 RP*QFDRSVLHDGRTNKYSFMHSGQKISLAPLSPSE-VRDD-QKKRKEKYEKEKR-KIKR 484
RP QFD +H GR+N+YS +H +KI L P+SP VRDD K K K E K K+
Sbjct: 572 RPWQFDSDCMHHGRSNQYSLIHHDKKIILLPMSPEAIVRDDVAKATKAKTENNKNIKVVG 631
Query: 485 KEREFA*KKKKRRVSRNS-----HEQTAPIFTTLQKVALLT---NTQNFPSCTKFLLQEC 536
++ K ++ + T + + K AL++ + P +LQE
Sbjct: 632 NNKDGIKLKGHCLLATKTDVNELFASTTVAYALVCKDALISIQDMQHSLPPVITNILQEY 691
Query: 537 EDVFPKKVPQ 546
DVFP ++P+
Sbjct: 692 SDVFPSEIPE 701
>emb|CAB77944.1| putative polyprotein [Arabidopsis thaliana]
gi|4325354|gb|AAD17351.1| contains similarity to
retrovirus-related polyproteins and to CCHC zinc finger
protein (Pfam: PF00098, Score=16.3, E=0.051, E= 1)
[Arabidopsis thaliana] gi|25407415|pir||G85077 probable
polyprotein [imported] - Arabidopsis thaliana
Length = 1138
Score = 171 bits (432), Expect = 7e-41
Identities = 88/151 (58%), Positives = 105/151 (69%), Gaps = 3/151 (1%)
Query: 289 DLGE*NVEVENHEGYVEEEAEEIPSGDLFMIRRFLGNQAKEEESNQRETLFHTRCLVQGK 348
D GE E E E E + EE P G+L + R L K EE QRE LFHTRCL++GK
Sbjct: 303 DSGEVESEDEKPE---ESDVEEAPKGELLVTMRVLSVLNKAEEQAQRENLFHTRCLIKGK 359
Query: 349 VCFLIIDGGSRTNVASTRLVSKMELETKPHPKPYKLQWLNENVEILVDKQVEVCFKIGKY 408
VC LIIDGGS TNVAS +V K+ LE PHPKPYKLQWLNE+ E+ V +QV+V IGKY
Sbjct: 360 VCSLIIDGGSCTNVASETMVQKLGLEEFPHPKPYKLQWLNESGEMAVTRQVQVPLAIGKY 419
Query: 409 EDDVLCDVVPMEASHLLLGRP*QFDRSVLHD 439
ED++LCD++P+EASH+LLGRP Q DR D
Sbjct: 420 EDEILCDILPLEASHVLLGRPWQSDRKDYQD 450
>gb|AAX96591.1| retrotransposon protein, putative, Ty3-gypsy sub-class [Oryza
sativa (japonica cultivar-group)]
Length = 1297
Score = 171 bits (432), Expect = 7e-41
Identities = 96/230 (41%), Positives = 135/230 (57%), Gaps = 18/230 (7%)
Query: 318 MIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVASTRLVSKMELETKP 377
+++R L Q ++ E NQR TLF T+C+V+ + C +IIDGGS N+AS+ +V K+ L TKP
Sbjct: 400 IVQRVLSAQMEKAEQNQRHTLFQTKCVVKERCCRMIIDGGSCNNLASSEMVEKLALSTKP 459
Query: 378 HPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVVPMEASHLLLGRP*QFDRSVL 437
HP PY +QWLN + + V K V + F IG Y D V CDVVPM+A ++LLGRP QFDR +
Sbjct: 460 HPHPYYIQWLNNSGKAKVTKLVHINFAIGNYHDVVECDVVPMQACNILLGRPWQFDRDSM 519
Query: 438 HDGRTNKYSFMHSGQKISLAPLSPSEV-RDD-QKKRKEKYEKEKRKIKRKEREFA*KKKK 495
H GR+N+YSF++ +KI L P+SP ++ RDD K K K E +K+ ++ K
Sbjct: 520 HHGRSNQYSFLYHDKKIVLHPMSPEDILRDDVAKAAKSKCESDKKAQSDGKKPETINLKP 579
Query: 496 RRVSRNSHEQTAPIFTTLQKVALLTNTQNFPSCTKFLLQECEDVFPKKVP 545
+ + + T I + AL E DVFPK+VP
Sbjct: 580 KCLLATKSDITELIASPSVAYAL----------------EYSDVFPKEVP 613
>gb|AAX96647.1| retrotransposon protein, putative, Ty3-gypsy sub-class [Oryza
sativa (japonica cultivar-group)]
Length = 1238
Score = 171 bits (432), Expect = 7e-41
Identities = 96/230 (41%), Positives = 135/230 (57%), Gaps = 18/230 (7%)
Query: 318 MIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVASTRLVSKMELETKP 377
+++R L Q ++ E NQR TLF T+C+V+ + C +IIDGGS N+AS+ +V K+ L TKP
Sbjct: 378 IVQRVLSAQMEKAEQNQRHTLFQTKCVVKERCCRMIIDGGSCNNLASSEMVEKLALSTKP 437
Query: 378 HPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVVPMEASHLLLGRP*QFDRSVL 437
HP PY +QWLN + + V K V + F IG Y D V CDVVPM+A ++LLGRP QFDR +
Sbjct: 438 HPHPYYIQWLNNSGKAKVTKLVHINFAIGNYHDVVECDVVPMQACNILLGRPWQFDRDSM 497
Query: 438 HDGRTNKYSFMHSGQKISLAPLSPSEV-RDD-QKKRKEKYEKEKRKIKRKEREFA*KKKK 495
H GR+N+YSF++ +KI L P+SP ++ RDD K K K E +K+ ++ K
Sbjct: 498 HHGRSNQYSFLYHDKKIVLHPMSPEDILRDDVAKAAKSKCESDKKAQSDGKKPETINLKP 557
Query: 496 RRVSRNSHEQTAPIFTTLQKVALLTNTQNFPSCTKFLLQECEDVFPKKVP 545
+ + + T I + AL E DVFPK+VP
Sbjct: 558 KCLLATKSDITELIASPSVAYAL----------------EYSDVFPKEVP 591
>gb|AAQ56339.1| putative gag-pol polyprotein [Oryza sativa (japonica
cultivar-group)]
Length = 1234
Score = 168 bits (426), Expect = 4e-40
Identities = 83/165 (50%), Positives = 115/165 (69%), Gaps = 2/165 (1%)
Query: 318 MIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVASTRLVSKMELETKP 377
+++R L Q ++ E NQR TLF T+C+V+ + C +IIDGGS N+AS+ +V K+ L TKP
Sbjct: 404 IVQRVLSAQMEKAEQNQRHTLFQTKCVVKERCCRMIIDGGSCKNLASSEMVEKLALSTKP 463
Query: 378 HPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVVPMEASHLLLGRP*QFDRSVL 437
HP PY +QWLN + + V K V + F IG Y D V CDVVPM+A ++LLGRP QFDR +
Sbjct: 464 HPHPYYIQWLNNSGKAKVTKLVHINFAIGNYHDVVECDVVPMQACNILLGRPWQFDRDSM 523
Query: 438 HDGRTNKYSFMHSGQKISLAPLSPSEV-RDD-QKKRKEKYEKEKR 480
H GR+N+YSF++ +KI L P+SP ++ RDD K K K E +K+
Sbjct: 524 HHGRSNQYSFLYHDKKIVLHPMSPEDILRDDVAKAAKSKCESDKK 568
>gb|AAX95495.1| Retrotransposon gag protein, putative [Oryza sativa (japonica
cultivar-group)]
Length = 1739
Score = 168 bits (425), Expect = 5e-40
Identities = 94/230 (40%), Positives = 135/230 (57%), Gaps = 18/230 (7%)
Query: 318 MIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVASTRLVSKMELETKP 377
+++R L Q ++ E NQR TLF T+C+++ + C +IIDGGS N+AS+ +V K+ L TKP
Sbjct: 817 IVQRVLSAQMEKAEQNQRHTLFQTKCVLKERCCRMIIDGGSCNNLASSEMVEKLALSTKP 876
Query: 378 HPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVVPMEASHLLLGRP*QFDRSVL 437
HP PY +QWLN + ++ V K V + F IG Y D V CDVVPM+A ++LLGRP QFDR +
Sbjct: 877 HPHPYYIQWLNNSGKVKVTKLVHINFAIGNYHDVVECDVVPMQACNILLGRPWQFDRDSM 936
Query: 438 HDGRTNKYSFMHSGQKISLAPLSPSEV-RDD-QKKRKEKYEKEKRKIKRKEREFA*KKKK 495
H GR+N+YSF++ +KI L P+S ++ RDD K K K E +K+ ++ K
Sbjct: 937 HHGRSNQYSFLYHDKKIVLHPMSSEDILRDDVAKAAKSKCESDKKAQSDGKKPETINLKP 996
Query: 496 RRVSRNSHEQTAPIFTTLQKVALLTNTQNFPSCTKFLLQECEDVFPKKVP 545
+ + + T I + AL E DVFPK+VP
Sbjct: 997 KCLLATKSDITELIASPSVAYAL----------------EYSDVFPKEVP 1030
>gb|AAX96717.1| retrotransposon protein, putative, Ty3-gypsy sub-class [Oryza sativa
(japonica cultivar-group)]
Length = 1748
Score = 168 bits (425), Expect = 5e-40
Identities = 94/230 (40%), Positives = 135/230 (57%), Gaps = 18/230 (7%)
Query: 318 MIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVASTRLVSKMELETKP 377
+++R L Q ++ E NQR TLF T+C+++ + C +IIDGGS N+AS+ +V K+ L TKP
Sbjct: 826 IVQRVLSAQMEKAEQNQRHTLFQTKCVLKERCCRMIIDGGSCNNLASSEMVEKLALSTKP 885
Query: 378 HPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVVPMEASHLLLGRP*QFDRSVL 437
HP PY +QWLN + ++ V K V + F IG Y D V CDVVPM+A ++LLGRP QFDR +
Sbjct: 886 HPHPYYIQWLNNSGKVKVTKLVHINFAIGNYHDVVECDVVPMQACNILLGRPWQFDRDSM 945
Query: 438 HDGRTNKYSFMHSGQKISLAPLSPSEV-RDD-QKKRKEKYEKEKRKIKRKEREFA*KKKK 495
H GR+N+YSF++ +KI L P+S ++ RDD K K K E +K+ ++ K
Sbjct: 946 HHGRSNQYSFLYHDKKIVLHPMSSEDILRDDVAKAAKSKCESDKKAQSDGKKPETINLKP 1005
Query: 496 RRVSRNSHEQTAPIFTTLQKVALLTNTQNFPSCTKFLLQECEDVFPKKVP 545
+ + + T I + AL E DVFPK+VP
Sbjct: 1006 KCLLATKSDITELIASPSVAYAL----------------EYSDVFPKEVP 1039
>ref|XP_475040.1| OSJNBa0042F21.10 [Oryza sativa (japonica cultivar-group)]
gi|38347308|emb|CAE02303.2| OSJNBa0042F21.10 [Oryza
sativa (japonica cultivar-group)]
Length = 2388
Score = 167 bits (423), Expect = 8e-40
Identities = 82/165 (49%), Positives = 114/165 (68%), Gaps = 2/165 (1%)
Query: 318 MIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTNVASTRLVSKMELETKP 377
+++R L Q ++ E NQR TLF T+C+V+ + C +IIDGGS N+AS+ +V K+ L TKP
Sbjct: 350 IVQRVLSAQMEKAEQNQRHTLFQTKCVVKERCCRMIIDGGSCNNLASSEMVEKLALSTKP 409
Query: 378 HPKPYKLQWLNENVEILVDKQVEVCFKIGKYEDDVLCDVVPMEASHLLLGRP*QFDRSVL 437
HP PY +QWLN + + V K V + F IG Y D V CDVVPM+ ++LLGRP QFDR +
Sbjct: 410 HPHPYYIQWLNNSGKAKVTKLVHINFAIGNYHDVVECDVVPMQVCNILLGRPWQFDRDSM 469
Query: 438 HDGRTNKYSFMHSGQKISLAPLSPSEV-RDD-QKKRKEKYEKEKR 480
H GR+N+YSF++ +KI L P+SP ++ RDD K K K E +K+
Sbjct: 470 HHGRSNQYSFLYHDKKIVLHPMSPEDILRDDVAKAAKSKCESDKK 514
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.358 0.160 0.569
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 768,988,347
Number of Sequences: 2540612
Number of extensions: 29193510
Number of successful extensions: 293744
Number of sequences better than 10.0: 228
Number of HSP's better than 10.0 without gapping: 139
Number of HSP's successfully gapped in prelim test: 101
Number of HSP's that attempted gapping in prelim test: 288933
Number of HSP's gapped (non-prelim): 1983
length of query: 550
length of database: 863,360,394
effective HSP length: 133
effective length of query: 417
effective length of database: 525,458,998
effective search space: 219116402166
effective search space used: 219116402166
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 78 (34.7 bits)
Medicago: description of AC147496.2


