

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147484.9 - phase: 0
(125 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
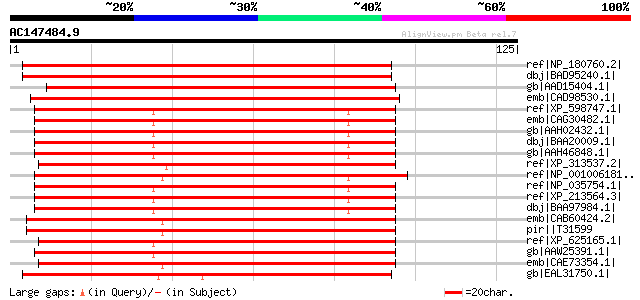
Score E
Sequences producing significant alignments: (bits) Value
ref|NP_180760.2| DNA topoisomerase family protein [Arabidopsis t... 142 2e-33
dbj|BAD95240.1| putative DNA topoisomerase III beta [Arabidopsis... 139 2e-32
gb|AAD15404.1| putative DNA topoisomerase III beta [Arabidopsis ... 136 1e-31
emb|CAD98530.1| DNA topoisomerase III beta-1, probable [Cryptosp... 115 2e-25
ref|XP_598747.1| PREDICTED: similar to topoisomerase (DNA) III b... 101 4e-21
emb|CAG30482.1| TOP3B [Homo sapiens] gi|6686033|sp|O95985|TOP3B_... 96 2e-19
gb|AAH02432.1| Topoisomerase (DNA) III beta [Homo sapiens] 96 2e-19
dbj|BAA20009.1| topoisomerase-III [Homo sapiens] 96 2e-19
gb|AAH46848.1| MGC53016 protein [Xenopus laevis] 94 6e-19
ref|XP_313537.2| ENSANGP00000018921 [Anopheles gambiae str. PEST... 94 7e-19
ref|NP_001006181.1| similar to topoisomerase III beta [Gallus ga... 94 7e-19
ref|NP_035754.1| topoisomerase (DNA) III beta [Mus musculus] gi|... 94 7e-19
ref|XP_213564.3| PREDICTED: similar to topoisomerase III beta [R... 94 7e-19
dbj|BAA97984.1| unnamed protein product [Mus musculus] 94 7e-19
emb|CAB60424.2| Hypothetical protein Y48C3A.14 [Caenorhabditis e... 93 2e-18
pir||T31599 hypothetical protein Y48C3A.q - Caenorhabditis elegans 93 2e-18
ref|XP_625165.1| PREDICTED: similar to MGC53016 protein [Apis me... 92 2e-18
gb|AAW25391.1| unknown [Schistosoma japonicum] 91 8e-18
emb|CAE73354.1| Hypothetical protein CBG20786 [Caenorhabditis br... 91 8e-18
gb|EAL31750.1| GA17464-PA [Drosophila pseudoobscura] 87 1e-16
>ref|NP_180760.2| DNA topoisomerase family protein [Arabidopsis thaliana]
Length = 865
Score = 142 bits (358), Expect = 2e-33
Identities = 70/91 (76%), Positives = 78/91 (84%)
Query: 4 LPKVLMVAEKPSIALSIATVLSHGKMNTRRGSTEVHEFDGKFRKAPARFKVTSVIGHVFS 63
L +VLMVAEKPSIALSIA+VLSHG+M+TRRGSTEVHEFDG FR A ++VTSVIGHVFS
Sbjct: 4 LLRVLMVAEKPSIALSIASVLSHGQMSTRRGSTEVHEFDGMFRGFKAHYRVTSVIGHVFS 63
Query: 64 VDFPGKYQDWAATDPLDLFQAQVIKNESNPK 94
VDFP KYQ+WA DP DLF A +IK ESNPK
Sbjct: 64 VDFPEKYQNWATIDPQDLFDAPIIKKESNPK 94
>dbj|BAD95240.1| putative DNA topoisomerase III beta [Arabidopsis thaliana]
Length = 865
Score = 139 bits (349), Expect = 2e-32
Identities = 69/91 (75%), Positives = 77/91 (83%)
Query: 4 LPKVLMVAEKPSIALSIATVLSHGKMNTRRGSTEVHEFDGKFRKAPARFKVTSVIGHVFS 63
L +VLMVAEKPSIALSIA+VLSHG+M+TR GSTEVHEFDG FR A ++VTSVIGHVFS
Sbjct: 4 LLRVLMVAEKPSIALSIASVLSHGQMSTRGGSTEVHEFDGMFRGFKAHYRVTSVIGHVFS 63
Query: 64 VDFPGKYQDWAATDPLDLFQAQVIKNESNPK 94
VDFP KYQ+WA DP DLF A +IK ESNPK
Sbjct: 64 VDFPEKYQNWATIDPQDLFGAPIIKKESNPK 94
>gb|AAD15404.1| putative DNA topoisomerase III beta [Arabidopsis thaliana]
gi|25408202|pir||G84727 probable DNA topoisomerase III
beta [imported] - Arabidopsis thaliana
Length = 836
Score = 136 bits (343), Expect = 1e-31
Identities = 66/86 (76%), Positives = 74/86 (85%)
Query: 10 VAEKPSIALSIATVLSHGKMNTRRGSTEVHEFDGKFRKAPARFKVTSVIGHVFSVDFPGK 69
VAEKPSIALSIA+VLSHG+M+TRRGSTEVHEFDG FR A ++VTSVIGHVFSVDFP K
Sbjct: 40 VAEKPSIALSIASVLSHGQMSTRRGSTEVHEFDGMFRGFKAHYRVTSVIGHVFSVDFPEK 99
Query: 70 YQDWAATDPLDLFQAQVIKNESNPKL 95
YQ+WA DP DLF A +IK ESNPK+
Sbjct: 100 YQNWATIDPQDLFDAPIIKKESNPKV 125
>emb|CAD98530.1| DNA topoisomerase III beta-1, probable [Cryptosporidium parvum]
Length = 835
Score = 115 bits (289), Expect = 2e-25
Identities = 54/91 (59%), Positives = 70/91 (76%)
Query: 6 KVLMVAEKPSIALSIATVLSHGKMNTRRGSTEVHEFDGKFRKAPARFKVTSVIGHVFSVD 65
KVLMVAEKPSI+ +I+ +LS+GK++TRRG T VHEF+G F +F+VTSV GHVF +D
Sbjct: 5 KVLMVAEKPSISDTISRILSNGKLDTRRGKTPVHEFNGTFMGMNVQFRVTSVAGHVFEID 64
Query: 66 FPGKYQDWAATDPLDLFQAQVIKNESNPKLV 96
FP Y +W TDP+ LF A +IKNES+ K+V
Sbjct: 65 FPQSYSNWEKTDPVSLFDAPIIKNESSSKMV 95
>ref|XP_598747.1| PREDICTED: similar to topoisomerase (DNA) III beta, partial [Bos
taurus]
Length = 197
Score = 101 bits (252), Expect = 4e-21
Identities = 53/93 (56%), Positives = 65/93 (68%), Gaps = 4/93 (4%)
Query: 7 VLMVAEKPSIALSIATVLSHGKMNTRRG---STEVHEFDGKFRKAPARFKVTSVIGHVFS 63
VLMVAEKPS+A SIA +LS G M++R+G + VHE+ G F PARFK+TSV GHV +
Sbjct: 4 VLMVAEKPSLAQSIARILSRGSMSSRKGLNGTCSVHEYSGAFEGQPARFKMTSVCGHVMT 63
Query: 64 VDFPGKYQDWAATDPLDLF-QAQVIKNESNPKL 95
+DF GKY W DP +LF QA K E+NPKL
Sbjct: 64 LDFLGKYNKWDKVDPAELFSQAPTEKKEANPKL 96
>emb|CAG30482.1| TOP3B [Homo sapiens] gi|6686033|sp|O95985|TOP3B_HUMAN DNA
topoisomerase III beta-1 gi|4768877|gb|AAD29670.1| DNA
topoisomerase III beta [Homo sapiens]
gi|4321652|gb|AAD15791.1| DNA topoisomerase III beta
[Homo sapiens] gi|4102879|gb|AAD01614.1| DNA
topoisomerase III beta [Homo sapiens]
gi|4507635|ref|NP_003926.1| topoisomerase (DNA) III
beta [Homo sapiens]
Length = 862
Score = 96.3 bits (238), Expect = 2e-19
Identities = 50/93 (53%), Positives = 63/93 (66%), Gaps = 4/93 (4%)
Query: 7 VLMVAEKPSIALSIATVLSHGKMNTRRG---STEVHEFDGKFRKAPARFKVTSVIGHVFS 63
VLMVAEKPS+A SIA +LS G +++ +G + VHE+ G F P RFK+TSV GHV +
Sbjct: 4 VLMVAEKPSLAQSIAKILSRGSLSSHKGLNGACSVHEYTGTFAGQPVRFKMTSVCGHVMT 63
Query: 64 VDFPGKYQDWAATDPLDLF-QAQVIKNESNPKL 95
+DF GKY W DP +LF QA K E+NPKL
Sbjct: 64 LDFLGKYNKWDKVDPAELFSQAPTEKKEANPKL 96
>gb|AAH02432.1| Topoisomerase (DNA) III beta [Homo sapiens]
Length = 862
Score = 96.3 bits (238), Expect = 2e-19
Identities = 50/93 (53%), Positives = 63/93 (66%), Gaps = 4/93 (4%)
Query: 7 VLMVAEKPSIALSIATVLSHGKMNTRRG---STEVHEFDGKFRKAPARFKVTSVIGHVFS 63
VLMVAEKPS+A SIA +LS G +++ +G + VHE+ G F P RFK+TSV GHV +
Sbjct: 4 VLMVAEKPSLAQSIAKILSRGSLSSHKGLNGACSVHEYTGTFAGQPVRFKMTSVCGHVMT 63
Query: 64 VDFPGKYQDWAATDPLDLF-QAQVIKNESNPKL 95
+DF GKY W DP +LF QA K E+NPKL
Sbjct: 64 LDFLGKYNKWDKVDPAELFSQAPTEKKEANPKL 96
>dbj|BAA20009.1| topoisomerase-III [Homo sapiens]
Length = 753
Score = 96.3 bits (238), Expect = 2e-19
Identities = 50/93 (53%), Positives = 63/93 (66%), Gaps = 4/93 (4%)
Query: 7 VLMVAEKPSIALSIATVLSHGKMNTRRG---STEVHEFDGKFRKAPARFKVTSVIGHVFS 63
VLMVAEKPS+A SIA +LS G +++ +G + VHE+ G F P RFK+TSV GHV +
Sbjct: 4 VLMVAEKPSLAQSIAKILSRGSLSSHKGLNGACSVHEYTGTFAGQPVRFKMTSVCGHVMT 63
Query: 64 VDFPGKYQDWAATDPLDLF-QAQVIKNESNPKL 95
+DF GKY W DP +LF QA K E+NPKL
Sbjct: 64 LDFLGKYNKWDKVDPAELFSQAPTEKKEANPKL 96
>gb|AAH46848.1| MGC53016 protein [Xenopus laevis]
Length = 858
Score = 94.4 bits (233), Expect = 6e-19
Identities = 50/93 (53%), Positives = 63/93 (66%), Gaps = 4/93 (4%)
Query: 7 VLMVAEKPSIALSIATVLSHGKMNTRRG---STEVHEFDGKFRKAPARFKVTSVIGHVFS 63
VLMVAEKPS+A SIA +LS G ++R+G + VHE+ G F RFK+TSV GHV S
Sbjct: 4 VLMVAEKPSLAQSIAKILSRGNASSRKGLNGACSVHEYTGSFMGQNVRFKMTSVCGHVMS 63
Query: 64 VDFPGKYQDWAATDPLDLF-QAQVIKNESNPKL 95
+DF GKY +W DP +LF +A K E+NPKL
Sbjct: 64 LDFIGKYNNWDKVDPAELFSKAPTEKKEANPKL 96
>ref|XP_313537.2| ENSANGP00000018921 [Anopheles gambiae str. PEST]
gi|55240964|gb|EAA08994.2| ENSANGP00000018921
[Anopheles gambiae str. PEST]
Length = 839
Score = 94.0 bits (232), Expect = 7e-19
Identities = 46/91 (50%), Positives = 60/91 (65%), Gaps = 3/91 (3%)
Query: 8 LMVAEKPSIALSIATVLSHGKMNTRRGSTE---VHEFDGKFRKAPARFKVTSVIGHVFSV 64
LMVAEKPS+A S+A++LS+GK + R+GS VHE+ G+FR RFK+TSV GH+ +
Sbjct: 5 LMVAEKPSLAASLASILSNGKCSVRKGSNSACSVHEWVGQFRGETTRFKMTSVCGHIMGL 64
Query: 65 DFPGKYQDWAATDPLDLFQAQVIKNESNPKL 95
+F GKY W DP +LF K ES P L
Sbjct: 65 EFVGKYNSWDRVDPAELFACPTEKKESTPNL 95
>ref|NP_001006181.1| similar to topoisomerase III beta [Gallus gallus]
gi|66796187|dbj|BAD99111.1| topoisomerase III beta
[Gallus gallus] gi|53132094|emb|CAG31872.1|
hypothetical protein [Gallus gallus]
Length = 862
Score = 94.0 bits (232), Expect = 7e-19
Identities = 51/96 (53%), Positives = 63/96 (65%), Gaps = 4/96 (4%)
Query: 7 VLMVAEKPSIALSIATVLSHGKMNTRRGST---EVHEFDGKFRKAPARFKVTSVIGHVFS 63
VLMVAEKPS+A SIA +LS G M++R+G VHE+ G F A FK+TSV GHV +
Sbjct: 4 VLMVAEKPSLAQSIAKILSRGNMSSRKGLNGVCSVHEYTGSFIGQSAHFKMTSVCGHVMT 63
Query: 64 VDFPGKYQDWAATDPLDLF-QAQVIKNESNPKLVDV 98
+DF GKY W DP +LF +A K E+NPKL V
Sbjct: 64 LDFIGKYNSWDKVDPAELFSKAPTEKKEANPKLTMV 99
>ref|NP_035754.1| topoisomerase (DNA) III beta [Mus musculus]
gi|21594767|gb|AAH31723.1| Topoisomerase (DNA) III beta
[Mus musculus] gi|6686042|sp|Q9Z321|TOP3B_MOUSE DNA
topoisomerase III beta-1 gi|3868800|dbj|BAA34227.1|
topoisomerase III beta [Mus musculus]
Length = 862
Score = 94.0 bits (232), Expect = 7e-19
Identities = 49/93 (52%), Positives = 62/93 (65%), Gaps = 4/93 (4%)
Query: 7 VLMVAEKPSIALSIATVLSHGKMNTRRG---STEVHEFDGKFRKAPARFKVTSVIGHVFS 63
VLMVAEKPS+A SIA +LS G M++ +G + VH++ G F P FK+TSV GHV +
Sbjct: 4 VLMVAEKPSLAQSIAKILSRGNMSSHKGLNGACSVHKYTGTFAGQPVHFKMTSVCGHVMT 63
Query: 64 VDFPGKYQDWAATDPLDLF-QAQVIKNESNPKL 95
+DF GKY W DP +LF QA K E+NPKL
Sbjct: 64 LDFLGKYNKWDKVDPAELFSQAPTEKKEANPKL 96
>ref|XP_213564.3| PREDICTED: similar to topoisomerase III beta [Rattus norvegicus]
Length = 862
Score = 94.0 bits (232), Expect = 7e-19
Identities = 49/93 (52%), Positives = 62/93 (65%), Gaps = 4/93 (4%)
Query: 7 VLMVAEKPSIALSIATVLSHGKMNTRRG---STEVHEFDGKFRKAPARFKVTSVIGHVFS 63
VLMVAEKPS+A SIA +LS G M++ +G + VH++ G F P FK+TSV GHV +
Sbjct: 4 VLMVAEKPSLAQSIAKILSRGNMSSHKGLNGACSVHKYTGTFAGQPVHFKMTSVCGHVMT 63
Query: 64 VDFPGKYQDWAATDPLDLF-QAQVIKNESNPKL 95
+DF GKY W DP +LF QA K E+NPKL
Sbjct: 64 LDFLGKYNKWDKVDPAELFSQAPTEKKEANPKL 96
>dbj|BAA97984.1| unnamed protein product [Mus musculus]
Length = 464
Score = 94.0 bits (232), Expect = 7e-19
Identities = 49/93 (52%), Positives = 62/93 (65%), Gaps = 4/93 (4%)
Query: 7 VLMVAEKPSIALSIATVLSHGKMNTRRG---STEVHEFDGKFRKAPARFKVTSVIGHVFS 63
VLMVAEKPS+A SIA +LS G M++ +G + VH++ G F P FK+TSV GHV +
Sbjct: 4 VLMVAEKPSLAQSIAKILSRGNMSSHKGLNGACSVHKYTGTFAGQPVHFKMTSVCGHVMT 63
Query: 64 VDFPGKYQDWAATDPLDLF-QAQVIKNESNPKL 95
+DF GKY W DP +LF QA K E+NPKL
Sbjct: 64 LDFLGKYNKWDKVDPAELFSQAPTEKKEANPKL 96
>emb|CAB60424.2| Hypothetical protein Y48C3A.14 [Caenorhabditis elegans]
gi|32563869|ref|NP_496822.2| TOPRIM and DNA
topoisomerase I (2N691) [Caenorhabditis elegans]
Length = 855
Score = 92.8 bits (229), Expect = 2e-18
Identities = 47/94 (50%), Positives = 62/94 (65%), Gaps = 3/94 (3%)
Query: 5 PKVLMVAEKPSIALSIATVLSHGKMNTRRGST---EVHEFDGKFRKAPARFKVTSVIGHV 61
P VLMVAEKP +A SIA +LS+G+ R+G V E+DG+F ARFKVTS GHV
Sbjct: 3 PTVLMVAEKPLLADSIANLLSNGQAKKRKGWNGVCSVSEYDGQFNGRAARFKVTSTCGHV 62
Query: 62 FSVDFPGKYQDWAATDPLDLFQAQVIKNESNPKL 95
S+DFP K+ +W DP +L+ A + E+NPK+
Sbjct: 63 MSLDFPPKFNNWERVDPAELYSAPTQRIEANPKM 96
>pir||T31599 hypothetical protein Y48C3A.q - Caenorhabditis elegans
Length = 646
Score = 92.8 bits (229), Expect = 2e-18
Identities = 47/94 (50%), Positives = 62/94 (65%), Gaps = 3/94 (3%)
Query: 5 PKVLMVAEKPSIALSIATVLSHGKMNTRRGST---EVHEFDGKFRKAPARFKVTSVIGHV 61
P VLMVAEKP +A SIA +LS+G+ R+G V E+DG+F ARFKVTS GHV
Sbjct: 3 PTVLMVAEKPLLADSIANLLSNGQAKKRKGWNGVCSVSEYDGQFNGRAARFKVTSTCGHV 62
Query: 62 FSVDFPGKYQDWAATDPLDLFQAQVIKNESNPKL 95
S+DFP K+ +W DP +L+ A + E+NPK+
Sbjct: 63 MSLDFPPKFNNWERVDPAELYSAPTQRIEANPKM 96
>ref|XP_625165.1| PREDICTED: similar to MGC53016 protein [Apis mellifera]
Length = 851
Score = 92.4 bits (228), Expect = 2e-18
Identities = 45/91 (49%), Positives = 61/91 (66%), Gaps = 3/91 (3%)
Query: 8 LMVAEKPSIALSIATVLSHGKMNTRRG---STEVHEFDGKFRKAPARFKVTSVIGHVFSV 64
LMVAEKPS+A S+A +LS+G+ +R+G S VHE+ G+FR FK+TSV GHV ++
Sbjct: 5 LMVAEKPSLAASLANILSNGRCTSRKGLNGSCSVHEWIGQFRSETVNFKMTSVFGHVMTL 64
Query: 65 DFPGKYQDWAATDPLDLFQAQVIKNESNPKL 95
DF GKY +W DP +LF K E+ P+L
Sbjct: 65 DFIGKYNNWDKVDPAELFSCPTEKKEAVPRL 95
>gb|AAW25391.1| unknown [Schistosoma japonicum]
Length = 114
Score = 90.5 bits (223), Expect = 8e-18
Identities = 46/92 (50%), Positives = 61/92 (66%), Gaps = 3/92 (3%)
Query: 7 VLMVAEKPSIALSIATVLSHGKMNTRRG---STEVHEFDGKFRKAPARFKVTSVIGHVFS 63
V MVAEKPS+A SIA LS+ + ++RRG + +HE+ G+F RFK+TSV GHV +
Sbjct: 4 VFMVAEKPSLAESIAKSLSNKRNSSRRGFNLACSIHEWTGQFMSEQVRFKMTSVCGHVMN 63
Query: 64 VDFPGKYQDWAATDPLDLFQAQVIKNESNPKL 95
VDF +Y W DP +LF A + K E+NPKL
Sbjct: 64 VDFLSRYNSWDKVDPAELFVAPIEKKEANPKL 95
>emb|CAE73354.1| Hypothetical protein CBG20786 [Caenorhabditis briggsae]
Length = 591
Score = 90.5 bits (223), Expect = 8e-18
Identities = 45/91 (49%), Positives = 60/91 (65%), Gaps = 3/91 (3%)
Query: 8 LMVAEKPSIALSIATVLSHGKMNTRRGST---EVHEFDGKFRKAPARFKVTSVIGHVFSV 64
LMVAEKP +A SIA +LS+G+ R+G V E+DG+F A+FKVTS GHV S+
Sbjct: 1 LMVAEKPMLADSIANLLSNGQARKRKGWNGVCSVSEYDGQFNGRNAKFKVTSTCGHVMSL 60
Query: 65 DFPGKYQDWAATDPLDLFQAQVIKNESNPKL 95
DFP KY +W DP +L+ A +K E+N K+
Sbjct: 61 DFPSKYNNWERVDPAELYSAPAVKIEANAKM 91
>gb|EAL31750.1| GA17464-PA [Drosophila pseudoobscura]
Length = 816
Score = 86.7 bits (213), Expect = 1e-16
Identities = 44/95 (46%), Positives = 60/95 (62%), Gaps = 4/95 (4%)
Query: 4 LPKVLMVAEKPSIALSIATVLSHGKMNTRRGS---TEVHEFDGKFR-KAPARFKVTSVIG 59
+ VLMVAEKPS+A S+AT+LS+G+ +RG+ HE+ G FR + F++TSV G
Sbjct: 1 MKNVLMVAEKPSLAASLATILSNGRCTVKRGTGNGCSTHEWTGTFRNEGTVHFRMTSVCG 60
Query: 60 HVFSVDFPGKYQDWAATDPLDLFQAQVIKNESNPK 94
HV S+DF KY W DP+ LF K E++PK
Sbjct: 61 HVMSLDFISKYNSWDKVDPVQLFGCPTEKKETSPK 95
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.323 0.137 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 197,557,221
Number of Sequences: 2540612
Number of extensions: 7181284
Number of successful extensions: 12998
Number of sequences better than 10.0: 105
Number of HSP's better than 10.0 without gapping: 80
Number of HSP's successfully gapped in prelim test: 25
Number of HSP's that attempted gapping in prelim test: 12846
Number of HSP's gapped (non-prelim): 108
length of query: 125
length of database: 863,360,394
effective HSP length: 101
effective length of query: 24
effective length of database: 606,758,582
effective search space: 14562205968
effective search space used: 14562205968
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 68 (30.8 bits)
Medicago: description of AC147484.9


