

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147484.3 - phase: 0 /pseudo
(526 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
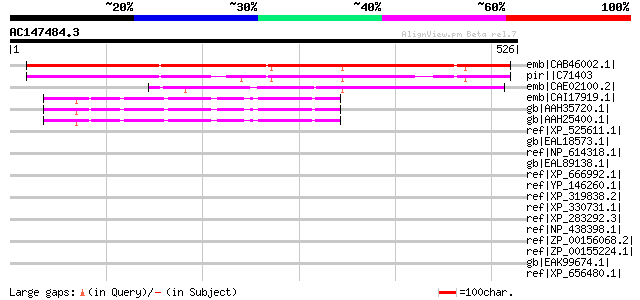
Score E
Sequences producing significant alignments: (bits) Value
emb|CAB46002.1| hypothetical protein [Arabidopsis thaliana] gi|7... 489 e-137
pir||C71403 hypothetical 15.1K protein - Arabidopsis thaliana 428 e-118
emb|CAE02100.2| OSJNBa0020I02.9 [Oryza sativa (japonica cultivar... 227 8e-58
emb|CAI17919.1| novel protein similar to mouse meiosis defective... 48 7e-04
gb|AAH35720.1| MGC40042 protein [Homo sapiens] 48 7e-04
gb|AAH25400.1| MGC40042 protein [Homo sapiens] 48 7e-04
ref|XP_525611.1| PREDICTED: similar to RIKEN cDNA 2310042L06 [Pa... 43 0.021
gb|EAL18573.1| hypothetical protein CNBJ2140 [Cryptococcus neofo... 41 0.11
ref|NP_614318.1| Lhr-like Superfamily II helicase [Methanopyrus ... 38 0.90
gb|EAL89138.1| spindle pole body component, putative [Aspergillu... 37 2.0
ref|XP_666992.1| hypothetical protein Chro.20292 [Cryptosporidiu... 36 2.6
ref|YP_146260.1| hypothetical protein GK0407 [Geobacillus kausto... 35 4.5
ref|XP_319838.2| ENSANGP00000016531 [Anopheles gambiae str. PEST... 35 5.8
ref|XP_330731.1| hypothetical protein [Neurospora crassa] gi|289... 35 5.8
ref|XP_283292.3| PREDICTED: similar to MGC40042 protein [Mus mus... 35 5.8
ref|NP_438398.1| branched-chain amino acid transport system II c... 35 7.6
ref|ZP_00156068.2| COG1114: Branched-chain amino acid permeases ... 35 7.6
ref|ZP_00155224.1| COG1114: Branched-chain amino acid permeases ... 35 7.6
gb|EAK99674.1| hypothetical protein CaO19.10025 [Candida albican... 34 10.0
ref|XP_656480.1| Rab GTPase activating protein, putative [Entamo... 34 10.0
>emb|CAB46002.1| hypothetical protein [Arabidopsis thaliana]
gi|7268123|emb|CAB78460.1| hypothetical protein
[Arabidopsis thaliana] gi|15236434|ref|NP_193154.1|
expressed protein [Arabidopsis thaliana]
gi|25333913|pir||G85154 hypothetical protein dl3130w
[imported] - Arabidopsis thaliana
Length = 1268
Score = 489 bits (1259), Expect = e-137
Identities = 268/533 (50%), Positives = 356/533 (66%), Gaps = 35/533 (6%)
Query: 18 CSQGHRSSLNLQTNQGGTICLVCFSNLISNPSSPTIHVSYALSQLSRSLSIPQFLHSIVT 77
C+ GHRS+++L+ +QGGT CL+CFSNL+S+P PT+HVSYAL QLS ++S P FL ++++
Sbjct: 31 CANGHRSTISLRDDQGGTFCLICFSNLVSDPRIPTVHVSYALHQLSIAISEPIFLRTLLS 90
Query: 78 FHPHFLVSPLVAALSSFDDEPIAEQLVHLILNLSASSDPSVCREFVARVSDRVSSGALGW 137
H HFLVSPLV ALSS DD PIA Q++ +I L + + S+ +FV R+SD++SSGALGW
Sbjct: 91 SHIHFLVSPLVHALSSIDDAPIAIQIMDMISLLCSVEESSIGEDFVERISDQLSSGALGW 150
Query: 138 SSRQLHSLHCLGVLLNCEKEDDLHVHIKDVYSLISVLVTGLQLPSEEIRGEVLFVLYKLF 197
S RQLH LHC GVL++CE +++ HI+D +L+ LV GLQLPSEEIRGE+LF LYK
Sbjct: 151 SRRQLHMLHCFGVLMSCE-NININSHIRDKEALVCQLVEGLQLPSEEIRGEILFALYKFS 209
Query: 198 ALHSTSDEGDGSDMLIPFCPKLLYLLGDVLLKTQNDDVRLNCIALLTMLAQRQLLREEPA 257
AL T DG ++L CPKLL L + L KTQ DDVRLNC+ALLT+LAQ+ LL +
Sbjct: 210 ALQFTEQNVDGIEVLSLLCPKLLCLSLEALAKTQRDDVRLNCVALLTILAQQGLLANSHS 269
Query: 258 YDTGNMSLSGEVN---FKETEDVPEGASLVNLFAEAIKGPLLSSVSQVQIGALDLLFHYL 314
+MSL EV+ + E+V L LFAEAIKGPLLS+ S+VQI LDL+FHY+
Sbjct: 270 NSASSMSLD-EVDDDPMQTAENVAARPCLNVLFAEAIKGPLLSTDSEVQIKTLDLIFHYI 328
Query: 315 SSVGTSGNQIQVLVEENIADYLFEILRLS-------------------------VPFHPV 349
S T QIQV+VEEN+ADY+FEILRLS VP HP
Sbjct: 329 SQESTPSKQIQVMVEENVADYIFEILRLSAEHSFRKRLVIGFPSVIRVLHYVGEVPCHPF 388
Query: 350 QYETLKLIYECISECPGAVSTSQMEELVLVLIKMLTKHSDGEMGMIPETFIMACSVFVAL 409
Q +TLKLI CIS+ PG S+SQ++E+ LVL KML ++ EMG+ P+ F + CSVFV+L
Sbjct: 389 QIQTLKLISSCISDFPGIASSSQVQEIALVLKKMLERYYSQEMGLFPDAFAIICSVFVSL 448
Query: 410 MRSPSCNGALDLSKSIEEAVKQAILACLYVSERNINQILQCFYLLKEAYAYSHDENSTHI 469
M++PS D+ S++E+++ +ILA L + E++ QIL YLL E Y Y ST I
Sbjct: 449 MKTPSFGETADVLTSLQESLRHSILASLSLPEKDSTQILHAVYLLNEVYVYC--TASTSI 506
Query: 470 SK---LELRSGILDICRTHLLPWLATGIMNGEEELFLGLLEIFHSILLLHCSI 519
+K +ELR ++D+C +HLLPW + + EE LG++E FHSILL + I
Sbjct: 507 NKTICIELRHCVIDVCTSHLLPWFLSDVNEVNEEATLGIMETFHSILLQNSDI 559
>pir||C71403 hypothetical 15.1K protein - Arabidopsis thaliana
Length = 1361
Score = 428 bits (1101), Expect = e-118
Identities = 255/571 (44%), Positives = 335/571 (58%), Gaps = 105/571 (18%)
Query: 18 CSQGHRSSLNLQTNQGGTICLVCFSNLISNPSSPTIHVSYALSQLSRSLSIPQFLHSIVT 77
C+ GHRS+++L+ +QGGT CL+CFSNL+S+P PT+HVSYAL QLS ++S P FL ++++
Sbjct: 13 CANGHRSTISLRDDQGGTFCLICFSNLVSDPRIPTVHVSYALHQLSIAISEPIFLRTLLS 72
Query: 78 FHPHFLVSPLVAALSSFDDEPIAEQLVHLILNLSASSDPSVCREFVARVSDRVSSGALGW 137
H HFLVSPLV ALSS DD PIA Q++ +I L + + S+ +FV R+SD++SSGALGW
Sbjct: 73 SHIHFLVSPLVHALSSIDDAPIAIQIMDMISLLCSVEESSIGEDFVERISDQLSSGALGW 132
Query: 138 SSRQLHSLHCLGVLLNCEKEDDLHVHIKDVYSLISVLVTGLQLPSEEIRGEVLFVLYKLF 197
S RQLH LHC GVL++CE +++ HI+D +L+ LV GLQLPSEEIRGE+LF LYK
Sbjct: 133 SRRQLHMLHCFGVLMSCE-NININSHIRDKEALVCQLVEGLQLPSEEIRGEILFALYKFS 191
Query: 198 ALHSTSDEGDGSDMLIPFCPKLLYLLGDVLLKTQNDDVRLNC------------------ 239
AL T DG + L KTQ DDVRLNC
Sbjct: 192 ALQFTEQNVDGI---------------EALAKTQRDDVRLNCVGSYELIYLLRSYARNLN 236
Query: 240 -IALLTMLAQRQLLREEPAYDTGNMSLSGEVN---FKETEDVPEGASLVNLFAEAIKGPL 295
+ALLT+LAQ+ LL + +MSL EV+ + E+V L LFAEAIKGPL
Sbjct: 237 FVALLTILAQQGLLANSHSNSASSMSLD-EVDDDPMQTAENVAARPCLNVLFAEAIKGPL 295
Query: 296 LSSVSQVQIGALDLLFHYLSSVGTSGNQIQVLVEENIADYLFEILRLS------------ 343
LS+ S+VQI LDL+FHY+S T QIQV+VEEN+ADY+FEILRLS
Sbjct: 296 LSTDSEVQIKTLDLIFHYISQESTPSKQIQVMVEENVADYIFEILRLSECKDQVVNSCLR 355
Query: 344 --------------------------------VPFHPVQYETLKLIYECISECPGAVSTS 371
VP HP Q +TLKLI CIS+ PG S+S
Sbjct: 356 VLDLFSLAEHSFRKRLVIGFPSVIRVLHYVGEVPCHPFQIQTLKLISSCISDFPGIASSS 415
Query: 372 QMEELVLVLIKMLTKHSDGEMGMIPETFIMACSVFVALMRSPSCNGALDLSKSIEEAVKQ 431
Q++E+ LVL KML ++ EMG+ P+ F + CSVFV+LM++PS D+
Sbjct: 416 QVQEIALVLKKMLERYYSQEMGLFPDAFAIICSVFVSLMKTPSFGETADV---------- 465
Query: 432 AILACLYVSERNINQILQCFYLLKEAYAYSHDENSTHISK---LELRSGILDICRTHLLP 488
+ E++ QIL YLL E Y Y ST I+K +ELR ++D+C +HLLP
Sbjct: 466 -------LPEKDSTQILHAVYLLNEVYVYC--TASTSINKTICIELRHCVIDVCTSHLLP 516
Query: 489 WLATGIMNGEEELFLGLLEIFHSILLLHCSI 519
W + + EE LG++E FHSILL + I
Sbjct: 517 WFLSDVNEVNEEATLGIMETFHSILLQNSDI 547
>emb|CAE02100.2| OSJNBa0020I02.9 [Oryza sativa (japonica cultivar-group)]
gi|50923297|ref|XP_472009.1| OSJNBa0020I02.9 [Oryza
sativa (japonica cultivar-group)]
Length = 1176
Score = 227 bits (578), Expect = 8e-58
Identities = 167/446 (37%), Positives = 226/446 (50%), Gaps = 86/446 (19%)
Query: 145 LHCLGVLLNCEKEDDLHVHIKDVYSLISVLVTGLQLP----------------------- 181
LHCLG+LLN K D +I D SL LV L+LP
Sbjct: 5 LHCLGILLNSTK--DAATYIGDKQSLYLNLVNNLRLPRLIPLHIDTFLALRITLSDSILN 62
Query: 182 ----SEEIRGEVLFVLYKLFALHST--SDEGDGSDM-LIPFCPKLLYLLGDVLLKTQNDD 234
S+EIRGE+LFVLYKL L++T D D ++ L LL +VLLKTQNDD
Sbjct: 63 LFWYSDEIRGEILFVLYKLSLLNATPWDDICDNDNVDLSAIGRSLLQFSLEVLLKTQNDD 122
Query: 235 VRLNCIALLTMLAQRQLLREEPAYDTGNMSLSGEVNFKETED-VPEGASLVNLFAEAIKG 293
VRLNCIALL LA++ A+D +S +N E ED VP SLV LFAEA+KG
Sbjct: 123 VRLNCIALLLTLAKKG------AFDILLLSDPSLINSAEAEDNVPLNDSLVILFAEAVKG 176
Query: 294 PLLSSVSQVQIGALDLLFHYLSSVGTSGNQIQVLVEENIADYLFEILRLS---------- 343
LLS+ +VQ G L+L+FH+LSS + ++ L+++N+ADY+FE+LRLS
Sbjct: 177 SLLSTNIEVQTGTLELIFHFLSS-DANIFVLKTLIDQNVADYVFEVLRLSGNNDPLVISS 235
Query: 344 ----------------------------------VPFHPVQYETLKLIYECISECPGAVS 369
+PFHPVQ + L+L+ I C G +S
Sbjct: 236 IKVLSILANSEERFKEKLAIAVSTLLPVLHYVSEIPFHPVQSQVLRLVCISIINCSGILS 295
Query: 370 TSQMEELVLVLIKMLTKHSDGEMGMIPETFIMACSVFVALMRSPSCNGALDLSKSIEEAV 429
SQ E++ L +L +H +GE+GM ETF + CS+ V +++ PS + L I EA
Sbjct: 296 LSQEEQIACTLSAILRRHGNGELGMSSETFALVCSMLVEILKLPSADDIQKLPSFIVEAS 355
Query: 430 KQAILACLYVSERNINQILQCFYLLKEAYAYSHDENSTHI-SKLELRSGILDICRTHLLP 488
K AI + I LLKEA + + N I K L I++ C T+LLP
Sbjct: 356 KHAISLTFSHEYDCLFLIPHSLLLLKEALIFCLEGNKDQILRKKSLEDSIIETCETYLLP 415
Query: 489 WLATGIMNG-EEELFLGLLEIFHSIL 513
WL + I++G +EE G+L+IF IL
Sbjct: 416 WLESAIVDGNDEETLSGILQIFQIIL 441
>emb|CAI17919.1| novel protein similar to mouse meiosis defective 1 gene (Mei1)
[Homo sapiens] gi|55956937|emb|CAI17882.1| novel protein
similar to mouse meiosis defective 1 gene (Mei1) [Homo
sapiens] gi|56417691|emb|CAI22918.1| novel protein
similar to mouse meiosis defective 1 gene (Mei1) [Homo
sapiens]
Length = 1280
Score = 48.1 bits (113), Expect = 7e-04
Identities = 70/317 (22%), Positives = 129/317 (40%), Gaps = 32/317 (10%)
Query: 36 ICLVCFSNLISNPSSPTIHVSYALSQLSRSLS-----IPQFLHSIVTFHPHFLVSPLVAA 90
+CL C L+ +P + + LS +L + Q + HF +S L
Sbjct: 43 LCLACALELLPDPGVSLVRKKHMLSCFQDALVRHTSLVTQLVSQDQRVCIHF-ISVLFGL 101
Query: 91 LSSFDDEPIAEQLVHLILNLSASSDPSVCREFVARVSDRVSSGALGWSSRQ--LHSLHCL 148
L S +D + + + +++ ++ + + + D S + L +L L
Sbjct: 102 LCSMEDGSVTDLCIEVLIQITTQLK---LEQTIRCLLDECHKELCNMPSMRGSLATLTLL 158
Query: 149 GVLLNC--EKEDDLHVHIKDVYSLISVLVTGLQLPSEEIRGEVLFVLYKLFALHSTSDEG 206
G L++ D+L + + +L+ L+ GL PSE I+ V ++ KL++ ++
Sbjct: 159 GKLVDAIPALADEL---VMEHGNLMEHLLRGLVYPSEGIQASVCYLYGKLYSSPVAAEML 215
Query: 207 DGSDMLIPFCPKLLYLLGDVLLKTQNDDVRLNCIALLTMLAQRQLLREEPAYDTGNMSLS 266
G F KL L +L Q ++++NC+ LL RQLL+ YD +
Sbjct: 216 SGH-----FREKLFPLFLSILDGAQTKELQINCLGLL-----RQLLK----YDLFVSMIM 261
Query: 267 GEVNFKETEDVPEGASLVNLFAEAIKGPLLSSVSQVQIGALDLLFHYLSSVGTSGNQIQV 326
+ E+ EG+S +K LLS +Q+ + + L V +
Sbjct: 262 NQDGLGESAKNIEGSSGNTSLPLVLKKLLLSRDETLQVASAHCITAVL--VHSPAKHASA 319
Query: 327 LVEENIADYLFEILRLS 343
+ +I ++LFE L S
Sbjct: 320 FIHADIPEFLFEHLSSS 336
>gb|AAH35720.1| MGC40042 protein [Homo sapiens]
Length = 452
Score = 48.1 bits (113), Expect = 7e-04
Identities = 70/317 (22%), Positives = 129/317 (40%), Gaps = 32/317 (10%)
Query: 36 ICLVCFSNLISNPSSPTIHVSYALSQLSRSLS-----IPQFLHSIVTFHPHFLVSPLVAA 90
+CL C L+ +P + + LS +L + Q + HF +S L
Sbjct: 51 LCLACALELLPDPGVSLVRKKHMLSCFQDALVRHTSLVTQLVSQDQRVCIHF-ISVLFGL 109
Query: 91 LSSFDDEPIAEQLVHLILNLSASSDPSVCREFVARVSDRVSSGALGWSSRQ--LHSLHCL 148
L S +D + + + +++ ++ + + + D S + L +L L
Sbjct: 110 LCSMEDGSVTDLCIEVLIQITTQLK---LEQTIRCLLDECHKELCNMPSMRGSLATLTLL 166
Query: 149 GVLLNC--EKEDDLHVHIKDVYSLISVLVTGLQLPSEEIRGEVLFVLYKLFALHSTSDEG 206
G L++ D+L + + +L+ L+ GL PSE I+ V ++ KL++ ++
Sbjct: 167 GKLVDAIPALADEL---VMEHGNLMEHLLRGLVYPSEGIQASVCYLYGKLYSSPVAAEML 223
Query: 207 DGSDMLIPFCPKLLYLLGDVLLKTQNDDVRLNCIALLTMLAQRQLLREEPAYDTGNMSLS 266
G F KL L +L Q ++++NC+ LL RQLL+ YD +
Sbjct: 224 SGH-----FREKLFPLFLSILDGAQTKELQINCLGLL-----RQLLK----YDLFVSMIM 269
Query: 267 GEVNFKETEDVPEGASLVNLFAEAIKGPLLSSVSQVQIGALDLLFHYLSSVGTSGNQIQV 326
+ E+ EG+S +K LLS +Q+ + + L V +
Sbjct: 270 NQDGLGESAKNIEGSSGNTSLPLVLKKLLLSRDETLQVASAHCITAVL--VHSPAKHASA 327
Query: 327 LVEENIADYLFEILRLS 343
+ +I ++LFE L S
Sbjct: 328 FIHADIPEFLFEHLSSS 344
>gb|AAH25400.1| MGC40042 protein [Homo sapiens]
Length = 582
Score = 48.1 bits (113), Expect = 7e-04
Identities = 70/317 (22%), Positives = 129/317 (40%), Gaps = 32/317 (10%)
Query: 36 ICLVCFSNLISNPSSPTIHVSYALSQLSRSLS-----IPQFLHSIVTFHPHFLVSPLVAA 90
+CL C L+ +P + + LS +L + Q + HF +S L
Sbjct: 56 LCLACALELLPDPGVSLVRKKHMLSCFQDALVRHTSLVTQLVSQDQRVCIHF-ISVLFGL 114
Query: 91 LSSFDDEPIAEQLVHLILNLSASSDPSVCREFVARVSDRVSSGALGWSSRQ--LHSLHCL 148
L S +D + + + +++ ++ + + + D S + L +L L
Sbjct: 115 LCSMEDGSVTDLCIEVLIQITTQLK---LEQTIRCLLDECHKELCNMPSMRGSLATLTLL 171
Query: 149 GVLLNC--EKEDDLHVHIKDVYSLISVLVTGLQLPSEEIRGEVLFVLYKLFALHSTSDEG 206
G L++ D+L + + +L+ L+ GL PSE I+ V ++ KL++ ++
Sbjct: 172 GKLVDAIPALADEL---VMEHGNLMEHLLRGLVYPSEGIQASVCYLYGKLYSSPVAAEML 228
Query: 207 DGSDMLIPFCPKLLYLLGDVLLKTQNDDVRLNCIALLTMLAQRQLLREEPAYDTGNMSLS 266
G F KL L +L Q ++++NC+ LL RQLL+ YD +
Sbjct: 229 SGH-----FREKLFPLFLSILDGAQTKELQINCLGLL-----RQLLK----YDLFVSMIM 274
Query: 267 GEVNFKETEDVPEGASLVNLFAEAIKGPLLSSVSQVQIGALDLLFHYLSSVGTSGNQIQV 326
+ E+ EG+S +K LLS +Q+ + + L V +
Sbjct: 275 NQDGLGESAKNIEGSSGNTSLPLVLKKLLLSRDETLQVASAHCITAVL--VHSPAKHASA 332
Query: 327 LVEENIADYLFEILRLS 343
+ +I ++LFE L S
Sbjct: 333 FIHADIPEFLFEHLSSS 349
>ref|XP_525611.1| PREDICTED: similar to RIKEN cDNA 2310042L06 [Pan troglodytes]
Length = 1760
Score = 43.1 bits (100), Expect = 0.021
Identities = 45/175 (25%), Positives = 77/175 (43%), Gaps = 16/175 (9%)
Query: 169 SLISVLVTGLQLPSEEIRGEVLFVLYKLFALHSTSDEGDGSDMLIPFCPKLLYLLGDVLL 228
+L+ L+ GL PSE I+ V ++ KL++ ++ G F KL L +L
Sbjct: 34 NLMEHLLRGLVYPSEGIQASVCYLYGKLYSSPVAAEMLSGH-----FREKLFPLFLSILD 88
Query: 229 KTQNDDVRLNCIALLTMLAQRQLLREEPAYDTGNMSLSGEVNFKETEDVPEGASLVNLFA 288
Q ++++NC+ LL RQLL+ YD + + +E+ EG+S
Sbjct: 89 GAQTKELQINCLGLL-----RQLLK----YDLFVSMIMNQDGMEESAKNIEGSSGNTSLP 139
Query: 289 EAIKGPLLSSVSQVQIGALDLLFHYLSSVGTSGNQIQVLVEENIADYLFEILRLS 343
+K LLS +Q+ + + L V + + +I ++LFE L S
Sbjct: 140 LVLKKLLLSRDETLQVASAHCITAVL--VHSPAKHAPAFIHADIPEFLFEHLSSS 192
>gb|EAL18573.1| hypothetical protein CNBJ2140 [Cryptococcus neoformans var.
neoformans B-3501A] gi|57229560|gb|AAW45993.1|
hypothetical protein CNJ01330 [Cryptococcus neoformans
var. neoformans JEC21] gi|58260200|ref|XP_567510.1|
hypothetical protein CNJ01330 [Cryptococcus neoformans
var. neoformans JEC21]
Length = 864
Score = 40.8 bits (94), Expect = 0.11
Identities = 37/125 (29%), Positives = 57/125 (45%), Gaps = 15/125 (12%)
Query: 102 QLVHLILNLSASSDPS--VCREFVARVSDRVSSGALGWSSRQLHSLHCLGVLLNCEKEDD 159
QLVH L+ ++D + +C F V V+ L +L +L LG + E +
Sbjct: 349 QLVHATLDRETTADDTDLLCEVFKG-VDSMVTQEILAAGMERLATL--LGHAIAAESQQQ 405
Query: 160 LHVHIKDVYSLISVLVTGLQLPSEEI-RGEVLFVLYKLFAL---------HSTSDEGDGS 209
L HIK + L+ +L + LPS+ + VL +KL L + S DGS
Sbjct: 406 LETHIKSIDQLLRILDSMNNLPSQSVTEPSVLNARHKLLDLLALALQTVERNLSTTRDGS 465
Query: 210 DMLIP 214
D+L+P
Sbjct: 466 DLLLP 470
>ref|NP_614318.1| Lhr-like Superfamily II helicase [Methanopyrus kandleri AV19]
gi|19887567|gb|AAM02248.1| Lhr-like Superfamily II
helicase [Methanopyrus kandleri AV19]
Length = 910
Score = 37.7 bits (86), Expect = 0.90
Identities = 27/97 (27%), Positives = 46/97 (46%), Gaps = 12/97 (12%)
Query: 267 GEVNFKETEDVPEGASLVNLFAEAIKGPLLSSVSQVQIGALDLLFHYLSSVGTSGNQIQV 326
G V +D+ E A + A+KG L +++ G LD+L H L + G+Q+ V
Sbjct: 345 GLVITSNPDDLAEAAVICR---RALKGSL--EPTEIPEGCLDVLAHQLVGLCLDGHQVTV 399
Query: 327 LVEENIADYLFEILRLSVPFHPVQYETLKLIYECISE 363
DY E+ R + P+ + ETL+ + E + +
Sbjct: 400 -------DYALEVFRRAYPYRHLDAETLREVAEFLDD 429
>gb|EAL89138.1| spindle pole body component, putative [Aspergillus fumigatus Af293]
Length = 911
Score = 36.6 bits (83), Expect = 2.0
Identities = 44/196 (22%), Positives = 84/196 (42%), Gaps = 11/196 (5%)
Query: 203 SDEGDGS----DMLIPF--CPKLLYLLGDVLLKTQNDDVRLNCIALLTMLAQRQLLREEP 256
+DE DG D+ P PK+ + L ++ D + AL T L ++ ++
Sbjct: 268 ADEDDGEVDAFDLAEPVDVFPKIPKDFNEQLASSKWKDRKETLDALYTALNVPRI--KDG 325
Query: 257 AYDTGNMSLSGEVNFKETEDVPEGASLVNLFAEAIKGPLLSSVSQVQIGALDLLFHYLSS 316
+D SL+ + V A+ ++L A+ ++G S + ++ L +
Sbjct: 326 PFDDIVRSLAKSMKDANVAVVTVAANCIDLLAKGLRGAFAKHRSTIMPPIMERLKEKKQT 385
Query: 317 VGTSGNQI--QVLVEENIADYLFEILRLSVPFHP-VQYETLKLIYECISECPGAVSTSQM 373
V + Q V N++D L EIL +P V+ ETLK + C+ S +++
Sbjct: 386 VADALGQALDAVFASTNLSDCLEEILEFLKHKNPQVKQETLKFLIRCLRTTRDVPSKAEV 445
Query: 374 EELVLVLIKMLTKHSD 389
+ + K+LT+ ++
Sbjct: 446 KAIADAATKLLTESTE 461
>ref|XP_666992.1| hypothetical protein Chro.20292 [Cryptosporidium hominis]
gi|54658073|gb|EAL36759.1| hypothetical protein
Chro.20292 [Cryptosporidium hominis]
Length = 681
Score = 36.2 bits (82), Expect = 2.6
Identities = 58/245 (23%), Positives = 103/245 (41%), Gaps = 41/245 (16%)
Query: 248 QRQLLREEPAYDTGNMSLSGEVNFKETEDVPEGASLVN----LFAEAIKGPLLSSVSQV- 302
QR+++ + P+ D +S+S V + V EG +N + + AI LL+S+SQ+
Sbjct: 302 QREVVIQHPSID--QVSISIGVKLEICPLVDEGEIFMNAVDTIISNAIHTKLLNSLSQLA 359
Query: 303 ---QIGALDLLFHYLSSVGTSGNQIQVLVEENIADYLFEILRLSVPFHPVQYETLKLIYE 359
++ ++ F+ S N ++ +E +F + I +
Sbjct: 360 CDEELTSISWDFYNSSRENCGWNTFSIVSDEKNWRAVFRLG----------------IQQ 403
Query: 360 CISECPGAVSTSQMEELVLVLIKMLTKHSDGEMGMIPETFIMACSVFVALMRSPSCNGAL 419
+S C +S + EE+VL+ I K ++ E+G P+ V L+ C G++
Sbjct: 404 IVSICNTKMSMEEFEEIVLITISDYKKSAEEELGDDPKL------VLDGLIDDWLC-GSI 456
Query: 420 DLSKSIEEAVKQAILACLYVSERNINQILQC-FYLLKEAYAYSHDENSTHISKLELRSGI 478
LSK KQ V +R +I+QC L Y DEN H +K +G
Sbjct: 457 PLSK------KQEYQLFCKVVDRINPEIIQCRCKALFSHIIYYFDEND-HYNKESSFTGC 509
Query: 479 LDICR 483
+ + +
Sbjct: 510 IFVSK 514
>ref|YP_146260.1| hypothetical protein GK0407 [Geobacillus kaustophilus HTA426]
gi|56378784|dbj|BAD74692.1| hypothetical conserved
protein [Geobacillus kaustophilus HTA426]
Length = 217
Score = 35.4 bits (80), Expect = 4.5
Identities = 30/163 (18%), Positives = 63/163 (38%), Gaps = 29/163 (17%)
Query: 50 SPTIHVSYALSQLSRSLSIPQFLHS----------------IVTFHPHFLVSPLVAALSS 93
+P + L ++ R++ +P F + I+ H L+ +
Sbjct: 32 TPVTYACLTLGEMGRNMGVPPFANRETLPHIRKQELEEACRILGIHDLRLLGYRDKTVEF 91
Query: 94 FDDEPIAEQLVHLILNLSASSDPSVCREFVARVSDRVSSGALG---------WSSRQLHS 144
D+E +A+Q+ ++ A ++PS+ F S A G W + +
Sbjct: 92 EDEEELADQIAAIV----AETNPSLVITFYPGYSVHPDHDACGAAVIRALKRWPKEERPT 147
Query: 145 LHCLGVLLNCEKEDDLHVHIKDVYSLISVLVTGLQLPSEEIRG 187
+HC+ NCE++ ++DV S+I + ++ + G
Sbjct: 148 VHCVAFAKNCEQDLGQPDVVRDVSSVIDTKLAAIRAHRSQTEG 190
>ref|XP_319838.2| ENSANGP00000016531 [Anopheles gambiae str. PEST]
gi|55235423|gb|EAA15054.2| ENSANGP00000016531 [Anopheles
gambiae str. PEST]
Length = 544
Score = 35.0 bits (79), Expect = 5.8
Identities = 41/176 (23%), Positives = 77/176 (43%), Gaps = 30/176 (17%)
Query: 346 FHPVQYETLKLIYECISECPGAVSTSQMEELVLVLIKMLTKHSDGEMGMIPETFIMACSV 405
F+P Q ++ E ++ G V S + V ++++ + G++ +P++F+
Sbjct: 79 FYPSQTSYSSIVGETLAAGLGVVGFSWICSPVCTELEVIMMNWLGQLLNLPKSFL----- 133
Query: 406 FVALMRSPSCNGALDLSKSIEEAVKQAILACLYVSERNINQILQCFYLLKEA------YA 459
NG + S E++ A+LA E+ + ++ L EA A
Sbjct: 134 -----NCDEGNGGGIIQGSASESILVAVLAA---REQAVRRLRTQHPELTEADIRGRLVA 185
Query: 460 YSHDENSTHISKLELRSGILDICRTHLLPWLATGIMNG-------EEELFLGLLEI 508
Y+ D++++ + K SGIL + LLP TGI+ G EE++ GL +
Sbjct: 186 YTSDQSNSAVEK----SGILGAIKMRLLPADETGILRGSTFIQAVEEDVAKGLFPV 237
>ref|XP_330731.1| hypothetical protein [Neurospora crassa] gi|28925993|gb|EAA34985.1|
hypothetical protein [Neurospora crassa]
Length = 560
Score = 35.0 bits (79), Expect = 5.8
Identities = 20/53 (37%), Positives = 30/53 (55%), Gaps = 1/53 (1%)
Query: 469 ISKLELRSGILDICRTHLLPWLATGIMNGEEELFLGLLEIFHSILLLHCSIIL 521
++ L + S IL + W ATG +NGE + FLG+L FH I L C++ +
Sbjct: 209 LTHLNVVSNILQMADVDGRYWSATGGLNGEGDKFLGVLPFFH-IYGLTCALFM 260
>ref|XP_283292.3| PREDICTED: similar to MGC40042 protein [Mus musculus]
Length = 1374
Score = 35.0 bits (79), Expect = 5.8
Identities = 22/83 (26%), Positives = 41/83 (48%), Gaps = 5/83 (6%)
Query: 169 SLISVLVTGLQLPSEEIRGEVLFVLYKLFALHSTSDEGDGSDMLIPFCPKLLYLLGDVLL 228
+L+ L+ GL P+E ++ + ++ KL++ + ++ G F KL L L
Sbjct: 1221 NLMEHLLRGLVYPNEGVQASICYLYGKLYSSPTAAEMLSGH-----FREKLCALFLSTLD 1275
Query: 229 KTQNDDVRLNCIALLTMLAQRQL 251
Q D+++NC+ LL L + L
Sbjct: 1276 SAQTKDLQINCLGLLRQLLKYDL 1298
>ref|NP_438398.1| branched-chain amino acid transport system II carrier protein
[Haemophilus influenzae Rd KW20]
gi|1573191|gb|AAC21896.1| branched chain amino acid
transport system II carrier protein (brnQ) [Haemophilus
influenzae Rd KW20] gi|1075007|pir||D64056
branched-chain amino acid transport protein braB homolog
- Haemophilus influenzae (strain Rd KW20)
gi|3023408|sp|P71345|BRNQ_HAEIN Branched-chain amino
acid transport system carrier protein (Branched-chain
amino acid uptake carrier)
Length = 436
Score = 34.7 bits (78), Expect = 7.6
Identities = 17/72 (23%), Positives = 32/72 (43%)
Query: 336 LFEILRLSVPFHPVQYETLKLIYECISECPGAVSTSQMEELVLVLIKMLTKHSDGEMGMI 395
L + LR +P Y + L+ C S C + + E + L+K S+G ++
Sbjct: 356 LLQFLRKKLPSIKFTYNSTLLVTVCFSLCDSLNNVKMLPESINSLLKHFPLSSEGMAWLV 415
Query: 396 PETFIMACSVFV 407
P ++ S+F+
Sbjct: 416 PTLVMLVASIFI 427
>ref|ZP_00156068.2| COG1114: Branched-chain amino acid permeases [Haemophilus
influenzae R2866]
Length = 436
Score = 34.7 bits (78), Expect = 7.6
Identities = 17/72 (23%), Positives = 32/72 (43%)
Query: 336 LFEILRLSVPFHPVQYETLKLIYECISECPGAVSTSQMEELVLVLIKMLTKHSDGEMGMI 395
L + LR +P Y + L+ C S C + + E + L+K S+G ++
Sbjct: 356 LLQFLRKKLPSIKFTYNSTLLVTVCFSLCDSLNNVKMLPESINSLLKHFPLSSEGMAWLV 415
Query: 396 PETFIMACSVFV 407
P ++ S+F+
Sbjct: 416 PTLVMLVASIFI 427
>ref|ZP_00155224.1| COG1114: Branched-chain amino acid permeases [Haemophilus
influenzae R2846]
Length = 436
Score = 34.7 bits (78), Expect = 7.6
Identities = 17/72 (23%), Positives = 32/72 (43%)
Query: 336 LFEILRLSVPFHPVQYETLKLIYECISECPGAVSTSQMEELVLVLIKMLTKHSDGEMGMI 395
L + LR +P Y + L+ C S C + + E + L+K S+G ++
Sbjct: 356 LLQFLRKKLPSIKFTYNSTLLVTVCFSLCDSLNNVKMLPESINSLLKHFPLSSEGMAWLV 415
Query: 396 PETFIMACSVFV 407
P ++ S+F+
Sbjct: 416 PTLVMLVASIFI 427
>gb|EAK99674.1| hypothetical protein CaO19.10025 [Candida albicans SC5314]
gi|46440278|gb|EAK99586.1| hypothetical protein
CaO19.2489 [Candida albicans SC5314]
Length = 1109
Score = 34.3 bits (77), Expect = 10.0
Identities = 35/133 (26%), Positives = 63/133 (47%), Gaps = 20/133 (15%)
Query: 170 LISVLVTGLQLPSEEIRGEVLFVLYKLFALHSTSDEGDGSDMLIPFCPKLLYLLGDVLLK 229
L+ L + +Q R FVL+ L + +IP LL L G +L
Sbjct: 124 LLPTLFSAVQNTDVHTREVGTFVLFALLESQIAT--------VIPHISDLLSLFGTLLND 175
Query: 230 TQNDDVRLNCIALLTMLAQRQLLREEPAYDTGNMSLSGEVNFKETEDVPEGASLVNLFAE 289
+++ +VR+N I L +++ Q++ E+ + + L+G+ F+ T VP S++N+F +
Sbjct: 176 SESKEVRINSIMSLDVIS--QIIEED---EERTVELAGK--FQST--VP---SMINIFKD 223
Query: 290 AIKGPLLSSVSQV 302
I G + S V
Sbjct: 224 VISGDDVESAKSV 236
>ref|XP_656480.1| Rab GTPase activating protein, putative [Entamoeba histolytica
HM-1:IMSS] gi|56473652|gb|EAL51067.1| Rab GTPase
activating protein, putative [Entamoeba histolytica
HM-1:IMSS]
Length = 447
Score = 34.3 bits (77), Expect = 10.0
Identities = 18/68 (26%), Positives = 38/68 (55%)
Query: 151 LLNCEKEDDLHVHIKDVYSLISVLVTGLQLPSEEIRGEVLFVLYKLFALHSTSDEGDGSD 210
+L E E+++ H + ++ LV ++ + E++ VLY +FALHS + + +G++
Sbjct: 196 ILKIENENNIVNHRNVIKRILYCLVAIDKIKYTQGMNELVSVLYYVFALHSNNQDFEGAE 255
Query: 211 MLIPFCPK 218
+ +C K
Sbjct: 256 VSSYYCMK 263
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.321 0.137 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 859,700,556
Number of Sequences: 2540612
Number of extensions: 35227678
Number of successful extensions: 72638
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 14
Number of HSP's that attempted gapping in prelim test: 72609
Number of HSP's gapped (non-prelim): 25
length of query: 526
length of database: 863,360,394
effective HSP length: 133
effective length of query: 393
effective length of database: 525,458,998
effective search space: 206505386214
effective search space used: 206505386214
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 77 (34.3 bits)
Medicago: description of AC147484.3


