

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147483.13 - phase: 0
(208 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
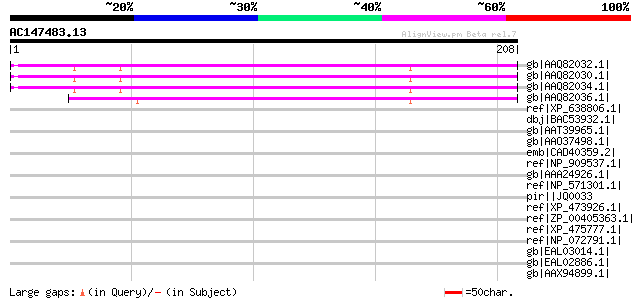
Score E
Sequences producing significant alignments: (bits) Value
gb|AAQ82032.1| unknown [Pisum sativum] 147 2e-34
gb|AAQ82030.1| unknown [Pisum sativum] 146 3e-34
gb|AAQ82034.1| unknown [Pisum sativum] 143 4e-33
gb|AAQ82036.1| unknown [Pisum sativum] 136 3e-31
ref|XP_638806.1| hypothetical protein DDB0218555 [Dictyostelium ... 40 0.042
dbj|BAC53932.1| hypothetical protein [Nicotiana tabacum] 40 0.055
gb|AAT39965.1| hypothetical protein [Solanum demissum] 36 0.61
gb|AAO37498.1| retrotransposon protein, putative, Ty3-gypsy sub-... 36 0.79
emb|CAD40359.2| OSJNBa0093P23.5 [Oryza sativa (japonica cultivar... 34 3.0
ref|NP_909537.1| Putative mutator-like transposase [Oryza sativa... 34 3.0
gb|AAA24926.1| methylase (MFokI) gi|127451|sp|P14871|MTF1_FLAOK ... 33 3.9
ref|NP_571301.1| retinal homeobox gene 2 [Danio rerio] gi|182020... 33 3.9
pir||JQ0033 site-specific DNA-methyltransferase (adenine-specifi... 33 3.9
ref|XP_473926.1| OSJNBb0085C12.3 [Oryza sativa (japonica cultiva... 33 5.1
ref|ZP_00405363.1| COG1264: Phosphotransferase system IIB compon... 33 6.7
ref|XP_475777.1| unknown protein [Oryza sativa (japonica cultiva... 33 6.7
ref|NP_072791.1| hypothetical protein MG129 [Mycoplasma genitali... 33 6.7
gb|EAL03014.1| hypothetical protein CaO19.1725 [Candida albicans... 33 6.7
gb|EAL02886.1| hypothetical protein CaO19.9293 [Candida albicans... 33 6.7
gb|AAX94899.1| transposon protein, putative, mutator sub-class [... 33 6.7
>gb|AAQ82032.1| unknown [Pisum sativum]
Length = 562
Score = 147 bits (370), Expect = 2e-34
Identities = 91/217 (41%), Positives = 122/217 (55%), Gaps = 10/217 (4%)
Query: 1 MHKFFSCIHRRIGQIKNFVNITGMK-KTKSYKFKEVDLVSLRELA--LKVKNQTGFRLQY 57
MH S +H R + + +K KT SY F L SL EL+ + NQ GF QY
Sbjct: 1 MH-IISQVHSRACSLIEMATVPELKRKTCSYSFHREPLTSLIELSNLMTSGNQKGFVDQY 59
Query: 58 GGLLTILRTNVEEKLVHTLVQFYDPSFRCFTFPDFQVVPTLEAYSHLLDSPTAEKTPFTG 117
G +LT+L+ V+ + TL+QFYDP CFTF D+Q+ PTLE YS LL P + PF
Sbjct: 60 GDILTLLKMVVDPVPLQTLLQFYDPELHCFTFQDYQLAPTLEEYSILLSVPVRYQVPFLD 119
Query: 118 PGTSLTPLVVAKDLYLKTSDVSNHLTTKSHIRGFTSKYLLEQATCQ------DALEAILA 171
+ VVA+ L+L +VS+ + + G K+LL A + +A A LA
Sbjct: 120 VPKEVDFRVVARALHLGIKEVSDSWKSSGDVVGLPLKFLLRVAREEAEKGSWEAFHAQLA 179
Query: 172 LLIYGLILFPSLDNFVDMNAIKIFHSKNPVPTLLADT 208
++IYG++LFPS+ NFVD A+ IF NPVPTLLADT
Sbjct: 180 VMIYGIVLFPSMPNFVDYAAVSIFIGGNPVPTLLADT 216
>gb|AAQ82030.1| unknown [Pisum sativum]
Length = 562
Score = 146 bits (369), Expect = 3e-34
Identities = 91/217 (41%), Positives = 123/217 (55%), Gaps = 10/217 (4%)
Query: 1 MHKFFSCIHRRIGQIKNFVNITGMK-KTKSYKFKEVDLVSLRELA--LKVKNQTGFRLQY 57
MH S +H R + + +K KT SY F L SL EL+ + NQ GF QY
Sbjct: 1 MH-IISQVHSRACSLIEMAIVPELKRKTCSYSFHREPLTSLIELSNLMTSGNQKGFVDQY 59
Query: 58 GGLLTILRTNVEEKLVHTLVQFYDPSFRCFTFPDFQVVPTLEAYSHLLDSPTAEKTPFTG 117
G +LT+L+ V+ + TL+QFYDP CFTF D+Q+ PTLE YS LL P + PF
Sbjct: 60 GDILTLLKMVVDPVPLQTLLQFYDPELHCFTFQDYQLAPTLEEYSILLSVPVRYQVPFLD 119
Query: 118 PGTSLTPLVVAKDLYLKTSDVSNHLTTKSHIRGFTSKYLLEQATCQ------DALEAILA 171
+ VVA+ L+L +VS++ + + G K+LL A + +A A LA
Sbjct: 120 VPKEVDFRVVARALHLGIKEVSDNWKSSGDVVGLPLKFLLRVAREEAEKGSWEAFHAQLA 179
Query: 172 LLIYGLILFPSLDNFVDMNAIKIFHSKNPVPTLLADT 208
++IYG++LFPS+ NFVD A+ IF NPVPTLLADT
Sbjct: 180 VMIYGIVLFPSMPNFVDYAAVSIFIGGNPVPTLLADT 216
>gb|AAQ82034.1| unknown [Pisum sativum]
Length = 562
Score = 143 bits (360), Expect = 4e-33
Identities = 90/217 (41%), Positives = 121/217 (55%), Gaps = 10/217 (4%)
Query: 1 MHKFFSCIHRRIGQIKNFVNITGMK-KTKSYKFKEVDLVSLRELA--LKVKNQTGFRLQY 57
MH S +H R + + +K KT SY F L SL EL+ + NQ GF Q+
Sbjct: 1 MH-IISQVHSRACSLIEMATVPELKRKTCSYSFHREPLTSLIELSNLMTSGNQKGFVDQH 59
Query: 58 GGLLTILRTNVEEKLVHTLVQFYDPSFRCFTFPDFQVVPTLEAYSHLLDSPTAEKTPFTG 117
G +LT+L+ V+ + TL+QFYDP CFTF D+Q+ PTLE YS LL P + PF
Sbjct: 60 GDILTLLKMVVDPVPLQTLLQFYDPELHCFTFQDYQLAPTLEEYSILLSVPVRYQVPFLD 119
Query: 118 PGTSLTPLVVAKDLYLKTSDVSNHLTTKSHIRGFTSKYLLEQATCQ------DALEAILA 171
+ VVA+ L+L +VS+ + G K+LL A + +A A LA
Sbjct: 120 VPKEVDFGVVARALHLGIKEVSDSWKYSGDVVGLPLKFLLRVAREEAEKGSWEAFRAQLA 179
Query: 172 LLIYGLILFPSLDNFVDMNAIKIFHSKNPVPTLLADT 208
++IYG++LFPS+ NFVD A+ IF NPVPTLLADT
Sbjct: 180 VMIYGIVLFPSMPNFVDYAAVSIFIGGNPVPTLLADT 216
>gb|AAQ82036.1| unknown [Pisum sativum]
Length = 546
Score = 136 bits (343), Expect = 3e-31
Identities = 79/192 (41%), Positives = 109/192 (56%), Gaps = 8/192 (4%)
Query: 25 KKTKSYKFKEVDLVSLRELALKVKNQT--GFRLQYGGLLTILRTNVEEKLVHTLVQFYDP 82
+KT SY F L SL EL + + GF Q+G +LT+L+T V+ + TL+QFYDP
Sbjct: 9 RKTCSYSFYREPLTSLIELGSLMPSDQLKGFVGQFGDILTLLKTVVDPVPLQTLLQFYDP 68
Query: 83 SFRCFTFPDFQVVPTLEAYSHLLDSPTAEKTPFTGPGTSLTPLVVAKDLYLKTSDVSNHL 142
RCFTF D+Q+ PTLE YS L+ P + PF + V+A+ L + +V ++
Sbjct: 69 ELRCFTFQDYQLAPTLEEYSILMSIPIQHQVPFLDVPKEVDFRVIARALRMSVKEVCDNW 128
Query: 143 TTKSHIRGFTSKYLLEQATCQ------DALEAILALLIYGLILFPSLDNFVDMNAIKIFH 196
+ G K+LL A + + A LA +IYG++LFPS+ NF+D AI IF
Sbjct: 129 KPSGEVVGMPLKFLLRVAREEAEKGNWEVFHAQLAAMIYGIVLFPSMPNFIDHAAISIFI 188
Query: 197 SKNPVPTLLADT 208
NPVPTLLADT
Sbjct: 189 GGNPVPTLLADT 200
>ref|XP_638806.1| hypothetical protein DDB0218555 [Dictyostelium discoideum]
gi|60467463|gb|EAL65486.1| hypothetical protein
DDB0218555 [Dictyostelium discoideum]
Length = 803
Score = 40.0 bits (92), Expect = 0.042
Identities = 24/94 (25%), Positives = 45/94 (47%), Gaps = 5/94 (5%)
Query: 2 HKFFS--CIHRRIGQIKNFVNITGMKKTKSYKFKEV--DLVSLRELALKVKNQTGFRL-Q 56
HKF S C ++ + ++N+TG F ++ + + + +L K N GF + +
Sbjct: 250 HKFLSTNCSFQQHHDLLVYLNLTGFHSFSKLNFTKICKEFLEINQLEFKKVNLEGFSIFE 309
Query: 57 YGGLLTILRTNVEEKLVHTLVQFYDPSFRCFTFP 90
+ TI N++ K +H +V DP++ FP
Sbjct: 310 INEIFTIALKNLDFKFLHLMVNCMDPNYFSVDFP 343
>dbj|BAC53932.1| hypothetical protein [Nicotiana tabacum]
Length = 464
Score = 39.7 bits (91), Expect = 0.055
Identities = 17/42 (40%), Positives = 24/42 (56%)
Query: 58 GGLLTILRTNVEEKLVHTLVQFYDPSFRCFTFPDFQVVPTLE 99
G L I++ + L+ LV F+DP F F DF++ PTLE
Sbjct: 39 GALTDIMKIKPRDDLIEALVTFWDPVHNVFCFSDFELTPTLE 80
>gb|AAT39965.1| hypothetical protein [Solanum demissum]
Length = 607
Score = 36.2 bits (82), Expect = 0.61
Identities = 17/42 (40%), Positives = 23/42 (54%)
Query: 58 GGLLTILRTNVEEKLVHTLVQFYDPSFRCFTFPDFQVVPTLE 99
G L ++L L+ LV F+DP F F DF++ PTLE
Sbjct: 49 GFLRSLLYVEPRRDLIEALVHFWDPIKNVFHFSDFELTPTLE 90
>gb|AAO37498.1| retrotransposon protein, putative, Ty3-gypsy sub-class [Oryza
sativa (japonica cultivar-group)]
gi|50916509|ref|XP_468651.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 966
Score = 35.8 bits (81), Expect = 0.79
Identities = 41/138 (29%), Positives = 61/138 (43%), Gaps = 32/138 (23%)
Query: 65 RTNVEEKLVHTLVQFYDPSFRCFTFPDFQVVPTLEAYSHLLDSPTAEKTPFTGPGTS--- 121
R NV+ L+ LV + P F P +V PTL+ S+LL P A GP T+
Sbjct: 402 RWNVDRSLLAALVDRWRPETHTFHLPCGEVAPTLQDVSYLLGLPLAGDA--VGPVTTGVD 459
Query: 122 --------LTPLVVAKDLYLKTSDVSNHLTTKSHIRGFTSKYLLE------QATCQD--- 164
TP V + +L +++H T G T ++LL+ QA +
Sbjct: 460 WKDDLTARFTP--VHRAPHLPLQPLAHHRNT-----GPTKRWLLQFIVEQLQAEADEYSY 512
Query: 165 --ALEAILALLIYGLILF 180
LEA L LL++G ++F
Sbjct: 513 SRCLEAYL-LLLFGWVMF 529
>emb|CAD40359.2| OSJNBa0093P23.5 [Oryza sativa (japonica cultivar-group)]
gi|50922621|ref|XP_471671.1| OSJNBa0041M21.10 [Oryza
sativa (japonica cultivar-group)]
gi|38343986|emb|CAD40452.2| OSJNBa0041M21.10 [Oryza
sativa (japonica cultivar-group)]
Length = 2074
Score = 33.9 bits (76), Expect = 3.0
Identities = 34/148 (22%), Positives = 53/148 (34%), Gaps = 8/148 (5%)
Query: 57 YGGLLTILRTNVEEKLVHTLVQFYDPSFRCFTFPDFQVVPTLEAYSHLLDSPTAEKTPFT 116
+GGLL I R NV L + +D F + T HLLD P+ + F
Sbjct: 493 FGGLLKIHRINVHRILCMWIANQFDTKAEAFKIQGSYLSLTNRDAEHLLDLPSQGEEIFE 552
Query: 117 GPGTSLTPLVVAKDLYLKTSDVSNHLTTKSHIRGFTSKYLLEQATCQDALEAILALLIYG 176
P T L D + TS H+ T+K + + D L + G
Sbjct: 553 PPQTKNRDLF---DEFKTTSKQGAHIKLSLQEYFQTNKGIHD-----DNFIHRFVLFVIG 604
Query: 177 LILFPSLDNFVDMNAIKIFHSKNPVPTL 204
+ L P+ +V + + N + +
Sbjct: 605 VFLCPTTQRYVSSAYLNLVEDVNAISAI 632
>ref|NP_909537.1| Putative mutator-like transposase [Oryza sativa (japonica
cultivar-group)] gi|19697447|gb|AAL93082.1| Putative
mutator-like transposase [Oryza sativa (japonica
cultivar-group)]
Length = 1530
Score = 33.9 bits (76), Expect = 3.0
Identities = 22/66 (33%), Positives = 32/66 (48%), Gaps = 8/66 (12%)
Query: 65 RTNVEEKLVHTLVQFYDPSFRCFTFPDFQVVPTLEAYSHLLDSPTAEKTPFTGPGTSLTP 124
R +V+ L+ TLV + P F P +V PTL+ S+LL P A G ++ P
Sbjct: 962 RWSVDRSLLATLVDRWRPETHTFHLPCGEVAPTLQDVSYLLGLPLA--------GDAVGP 1013
Query: 125 LVVAKD 130
+ A D
Sbjct: 1014 VTTAVD 1019
>gb|AAA24926.1| methylase (MFokI) gi|127451|sp|P14871|MTF1_FLAOK Modification
methylase FokI (Adenine-specific methyltransferase FokI)
(M.FokI)
Length = 647
Score = 33.5 bits (75), Expect = 3.9
Identities = 22/71 (30%), Positives = 33/71 (45%), Gaps = 3/71 (4%)
Query: 103 HLLDSPTAEKTPFTGPGTSLTPLVVAKDLYLKTSDVS---NHLTTKSHIRGFTSKYLLEQ 159
HLL++ P T L P V +K LY + +V NHL K++ R Y E
Sbjct: 231 HLLETIALYDYPEIYGKTGLRPYVESKSLYCQKKEVGNAFNHLIEKANFRHILVSYSSEG 290
Query: 160 ATCQDALEAIL 170
++ +E+IL
Sbjct: 291 LLLEEEIESIL 301
>ref|NP_571301.1| retinal homeobox gene 2 [Danio rerio]
gi|18202035|sp|O42357|RX2_BRARE Retinal homeobox protein
Rx2 gi|2240028|gb|AAB62326.1| retinal homeobox protein
[Danio rerio]
Length = 327
Score = 33.5 bits (75), Expect = 3.9
Identities = 34/113 (30%), Positives = 48/113 (42%), Gaps = 15/113 (13%)
Query: 36 DLVSLRELALKVKNQTGFRLQYGGLLTILRTNVEEKLVHTLVQFYDPSFRCFTFPDFQVV 95
D+ S ELA+KV N R+Q + +EK+ ++ +D R F P +
Sbjct: 161 DVYSREELAMKV-NLPEVRVQVWFQNRRAKWRRQEKMDTGTMKLHDSPIRSFNRPP--MA 217
Query: 96 PTLEAYSHLL------DSPTAEKTP------FTGPGTSLTPLVVAKDLYLKTS 136
P + S+ L SP + TP F GPG SL P A +L TS
Sbjct: 218 PNVGPMSNSLPLDPWLSSPLSSATPMHSIPGFMGPGQSLQPTYTAHPGFLNTS 270
>pir||JQ0033 site-specific DNA-methyltransferase (adenine-specific) (EC
2.1.1.72) - Flavobacterium sp gi|148724|gb|AAA24933.1|
methyltransferase
Length = 647
Score = 33.5 bits (75), Expect = 3.9
Identities = 22/71 (30%), Positives = 33/71 (45%), Gaps = 3/71 (4%)
Query: 103 HLLDSPTAEKTPFTGPGTSLTPLVVAKDLYLKTSDVS---NHLTTKSHIRGFTSKYLLEQ 159
HLL++ P T L P V +K LY + +V NHL K++ R Y E
Sbjct: 231 HLLETIALYDYPEIYGKTGLRPYVESKSLYCQKKEVGNAFNHLIEKANFRHILVSYSSEG 290
Query: 160 ATCQDALEAIL 170
++ +E+IL
Sbjct: 291 LLLEEEIESIL 301
>ref|XP_473926.1| OSJNBb0085C12.3 [Oryza sativa (japonica cultivar-group)]
gi|38345701|emb|CAE01929.2| OSJNBb0085C12.3 [Oryza
sativa (japonica cultivar-group)]
Length = 1356
Score = 33.1 bits (74), Expect = 5.1
Identities = 38/136 (27%), Positives = 58/136 (41%), Gaps = 28/136 (20%)
Query: 65 RTNVEEKLVHTLVQFYDPSFRCFTFPDFQVVPTLEAYSHLLDSPTAEKTPFTGPGTSLTP 124
R V+ LV LV + P F P +V PTL+ S+LL P A GP T+
Sbjct: 732 RWTVDRSLVAALVDRWRPETHTFHLPCGEVAPTLQDVSYLLGLPLAGDA--VGPVTTAVD 789
Query: 125 ---------LVVAKDLYLKTSDVSNHLTTKSHIRGFTSKYLLE------QATCQD----- 164
+V + +L +++H T G T ++LL+ QA +
Sbjct: 790 WQDDLTARFALVQRAPHLPLEPLAHHRNT-----GPTKRWLLQLTVEQLQAEADEYSYSR 844
Query: 165 ALEAILALLIYGLILF 180
LEA L L ++G ++F
Sbjct: 845 CLEAYL-LWLFGWVMF 859
>ref|ZP_00405363.1| COG1264: Phosphotransferase system IIB components [Mycoplasma
genitalium G-37]
Length = 111
Score = 32.7 bits (73), Expect = 6.7
Identities = 23/64 (35%), Positives = 33/64 (50%), Gaps = 5/64 (7%)
Query: 13 GQIKNFVNITGMKKTKSYKFKEVDLVSLREL-ALKV----KNQTGFRLQYGGLLTILRTN 67
G +NFVNI FK+V+ VSL +L AL + KNQ FR G + L+
Sbjct: 47 GGKENFVNIKTTPTQLIVTFKDVNSVSLTKLNALNIKGINKNQNQFRFVLGNFVNELKKK 106
Query: 68 VEEK 71
+E++
Sbjct: 107 IEDE 110
>ref|XP_475777.1| unknown protein [Oryza sativa (japonica cultivar-group)]
gi|47900426|gb|AAT39220.1| unknown protein [Oryza sativa
(japonica cultivar-group)]
Length = 896
Score = 32.7 bits (73), Expect = 6.7
Identities = 36/131 (27%), Positives = 57/131 (43%), Gaps = 18/131 (13%)
Query: 65 RTNVEEKLVHTLVQFYDPSFRCFTFPDFQVVPTLEAYSHLLDSPTAEKTPFTGPGTSLTP 124
R V+ L+ LV + P F P +V PTL+ S+LL P A GP T+
Sbjct: 288 RWTVDRSLLAALVDRWRPETHTFHLPCGEVAPTLQDVSYLLGLPLAGDA--VGPVTTAVD 345
Query: 125 ---------LVVAKDLYLKTSDVSNHLT---TKSHIRGFTSKYLLEQA---TCQDALEAI 169
+V + +L +++H TK + FT + L +A + LEA
Sbjct: 346 WEDDLTARFALVQRAPHLPLEPLAHHRNTGPTKRWLLQFTVEQLQAKADEYSYSRCLEAY 405
Query: 170 LALLIYGLILF 180
L L ++G ++F
Sbjct: 406 L-LWLFGWVMF 415
>ref|NP_072791.1| hypothetical protein MG129 [Mycoplasma genitalium G-37]
gi|3844720|gb|AAC71347.1| conserved hypothetical protein
[Mycoplasma genitalium G-37] gi|1361758|pir||C64214
phosphotransferase system glucose-specific permease
homolog - Mycoplasma genitalium
gi|2496309|sp|Q49397|Y129_MYCGE Hypothetical protein
MG129
Length = 117
Score = 32.7 bits (73), Expect = 6.7
Identities = 23/64 (35%), Positives = 33/64 (50%), Gaps = 5/64 (7%)
Query: 13 GQIKNFVNITGMKKTKSYKFKEVDLVSLREL-ALKV----KNQTGFRLQYGGLLTILRTN 67
G +NFVNI FK+V+ VSL +L AL + KNQ FR G + L+
Sbjct: 53 GGKENFVNIKTTPTQLIVTFKDVNSVSLTKLNALNIKGINKNQNQFRFVLGNFVNELKKK 112
Query: 68 VEEK 71
+E++
Sbjct: 113 IEDE 116
>gb|EAL03014.1| hypothetical protein CaO19.1725 [Candida albicans SC5314]
Length = 725
Score = 32.7 bits (73), Expect = 6.7
Identities = 23/66 (34%), Positives = 32/66 (47%), Gaps = 11/66 (16%)
Query: 59 GLLTILRTNVEEKLVHTLVQFYDPSFRCFTFPDFQVVPTLEAYSHLLDSPTAEKTPFTGP 118
G +T L T+V E+ V T+ T P F V TL ++++ SPT T TG
Sbjct: 481 GPVTTLTTSVPEQSVITI-----------TGPGFTTVFTLTPGTNVITSPTGTATKPTGT 529
Query: 119 GTSLTP 124
G+ TP
Sbjct: 530 GSEATP 535
>gb|EAL02886.1| hypothetical protein CaO19.9293 [Candida albicans SC5314]
Length = 687
Score = 32.7 bits (73), Expect = 6.7
Identities = 23/66 (34%), Positives = 32/66 (47%), Gaps = 11/66 (16%)
Query: 59 GLLTILRTNVEEKLVHTLVQFYDPSFRCFTFPDFQVVPTLEAYSHLLDSPTAEKTPFTGP 118
G +T L T+V E+ V T+ T P F V TL ++++ SPT T TG
Sbjct: 443 GPVTTLTTSVPEQSVITI-----------TGPGFTTVFTLTPGTNVITSPTGTATKPTGT 491
Query: 119 GTSLTP 124
G+ TP
Sbjct: 492 GSEATP 497
>gb|AAX94899.1| transposon protein, putative, mutator sub-class [Oryza sativa
(japonica cultivar-group)]
Length = 932
Score = 32.7 bits (73), Expect = 6.7
Identities = 22/66 (33%), Positives = 30/66 (45%), Gaps = 8/66 (12%)
Query: 65 RTNVEEKLVHTLVQFYDPSFRCFTFPDFQVVPTLEAYSHLLDSPTAEKTPFTGPGTSLTP 124
R V+ LV LV + P F P +V PTL+ S+LL P A G ++ P
Sbjct: 835 RWTVDRSLVAALVDRWRPETHTFHLPCGEVAPTLQDVSYLLGLPLA--------GDAVGP 886
Query: 125 LVVAKD 130
+ A D
Sbjct: 887 VTTAVD 892
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.323 0.139 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 343,737,206
Number of Sequences: 2540612
Number of extensions: 13812902
Number of successful extensions: 28093
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 28066
Number of HSP's gapped (non-prelim): 31
length of query: 208
length of database: 863,360,394
effective HSP length: 122
effective length of query: 86
effective length of database: 553,405,730
effective search space: 47592892780
effective search space used: 47592892780
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 72 (32.3 bits)
Medicago: description of AC147483.13


