

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147482.6 + phase: 2 /pseudo
(675 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
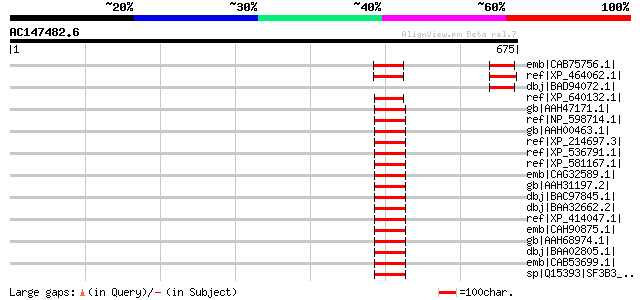
Score E
Sequences producing significant alignments: (bits) Value
emb|CAB75756.1| spliceosomal-like protein [Arabidopsis thaliana]... 87 2e-15
ref|XP_464062.1| putative splicing factor 3b, subunit 3, 130kDa ... 80 2e-13
dbj|BAD94072.1| spliceosomal - like protein [Arabidopsis thaliana] 64 2e-08
ref|XP_640132.1| hypothetical protein DDB0204844 [Dictyostelium ... 62 6e-08
gb|AAH47171.1| Zgc:55440 [Danio rerio] gi|47087273|ref|NP_998668... 59 7e-07
ref|NP_598714.1| splicing factor 3b, subunit 3 [Mus musculus] gi... 58 1e-06
gb|AAH00463.1| SF3B3 protein [Homo sapiens] gi|13111947|gb|AAH03... 58 1e-06
ref|XP_214697.3| PREDICTED: similar to Splicing factor 3b, subun... 58 1e-06
ref|XP_536791.1| PREDICTED: similar to KIAA0017 protein [Canis f... 58 1e-06
ref|XP_581167.1| PREDICTED: similar to KIAA0017 protein, partial... 58 1e-06
emb|CAG32589.1| hypothetical protein [Gallus gallus] 58 1e-06
gb|AAH31197.2| Sf3b3 protein [Mus musculus] 58 1e-06
dbj|BAC97845.1| mKIAA0017 protein [Mus musculus] 58 1e-06
dbj|BAA32662.2| KIAA0017 protein [Homo sapiens] 58 1e-06
ref|XP_414047.1| PREDICTED: similar to KIAA0017 protein [Gallus ... 58 1e-06
emb|CAH90875.1| hypothetical protein [Pongo pygmaeus] 58 1e-06
gb|AAH68974.1| Splicing factor 3b, subunit 3, 130kDa [Homo sapiens] 58 1e-06
dbj|BAA02805.1| KIAA0017 [Homo sapiens] 58 1e-06
emb|CAB53699.1| hypothetical protein [Homo sapiens] gi|7512688|p... 58 1e-06
sp|Q15393|SF3B3_HUMAN Splicing factor 3B subunit 3 (Spliceosome ... 58 1e-06
>emb|CAB75756.1| spliceosomal-like protein [Arabidopsis thaliana]
gi|7019653|emb|CAB75754.1| spliceosomal-like protein
[Arabidopsis thaliana] gi|18410226|ref|NP_567016.1|
splicing factor, putative [Arabidopsis thaliana]
gi|18410222|ref|NP_567015.1| splicing factor, putative
[Arabidopsis thaliana] gi|11358854|pir||T47659
spliceosomal-like protein - Arabidopsis thaliana
Length = 1214
Score = 86.7 bits (213), Expect = 2e-15
Identities = 37/40 (92%), Positives = 39/40 (97%)
Query: 485 CKYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG 524
CKYR DENQLYIFADDCVPRWLTAS+H+DFDTMAGADKFG
Sbjct: 1010 CKYRRDENQLYIFADDCVPRWLTASHHVDFDTMAGADKFG 1049
Score = 63.5 bits (153), Expect = 2e-08
Identities = 30/33 (90%), Positives = 31/33 (93%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEE 672
LCEQFPTLPMDLQRKIADELDRT EILKKLE+
Sbjct: 1176 LCEQFPTLPMDLQRKIADELDRTPAEILKKLED 1208
>ref|XP_464062.1| putative splicing factor 3b, subunit 3, 130kDa [Oryza sativa
(japonica cultivar-group)] gi|42409258|dbj|BAD10521.1|
putative splicing factor 3b, subunit 3, 130kDa [Oryza
sativa (japonica cultivar-group)]
gi|42409127|dbj|BAD10377.1| putative splicing factor 3b,
subunit 3, 130kDa [Oryza sativa (japonica
cultivar-group)]
Length = 1234
Score = 80.5 bits (197), Expect = 2e-13
Identities = 36/40 (90%), Positives = 37/40 (92%)
Query: 485 CKYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG 524
CKYR DENQLYIFADD VPRWLTA+ HIDFDTMAGADKFG
Sbjct: 1030 CKYRRDENQLYIFADDSVPRWLTAANHIDFDTMAGADKFG 1069
Score = 61.6 bits (148), Expect = 8e-08
Identities = 28/35 (80%), Positives = 32/35 (91%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQFP+LP D+QRKIADELDRT GEILKKLE+ +
Sbjct: 1196 LCEQFPSLPADMQRKIADELDRTPGEILKKLEDIR 1230
>dbj|BAD94072.1| spliceosomal - like protein [Arabidopsis thaliana]
Length = 165
Score = 63.5 bits (153), Expect = 2e-08
Identities = 30/33 (90%), Positives = 31/33 (93%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEE 672
LCEQFPTLPMDLQRKIADELDRT EILKKLE+
Sbjct: 127 LCEQFPTLPMDLQRKIADELDRTPAEILKKLED 159
>ref|XP_640132.1| hypothetical protein DDB0204844 [Dictyostelium discoideum]
gi|60468134|gb|EAL66144.1| hypothetical protein
DDB0204844 [Dictyostelium discoideum]
Length = 1256
Score = 62.0 bits (149), Expect = 6e-08
Identities = 25/39 (64%), Positives = 32/39 (81%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG 524
KY+ EN LY+FADD PRW+T+S +D+DT+AGADKFG
Sbjct: 1052 KYKRSENMLYVFADDLAPRWMTSSVMLDYDTVAGADKFG 1090
Score = 39.3 bits (90), Expect = 0.42
Identities = 18/35 (51%), Positives = 24/35 (68%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF TL Q I++EL R+ E++KKLEE +
Sbjct: 1217 LCEQFSTLNYQKQLSISEELSRSPSEVIKKLEEIR 1251
>gb|AAH47171.1| Zgc:55440 [Danio rerio] gi|47087273|ref|NP_998668.1| zgc:55440 [Danio
rerio]
Length = 1217
Score = 58.5 bits (140), Expect = 7e-07
Identities = 27/42 (64%), Positives = 33/42 (78%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+YR +ENQL IFADD PRW+T + +D+DTMA ADKFG IC
Sbjct: 1013 RYRRNENQLIIFADDTYPRWITTACLLDYDTMASADKFGNIC 1054
Score = 37.7 bits (86), Expect = 1.2
Identities = 16/35 (45%), Positives = 25/35 (70%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ Q+ +++ELDRT E+ KKLE+ +
Sbjct: 1178 LCEQFNSMDPHKQKSVSEELDRTPPEVSKKLEDIR 1212
>ref|NP_598714.1| splicing factor 3b, subunit 3 [Mus musculus]
gi|27503728|gb|AAH42580.1| Splicing factor 3b, subunit 3,
130kDa [Mus musculus] gi|15030278|gb|AAH11412.1| Splicing
factor 3b, subunit 3 [Mus musculus]
gi|26353236|dbj|BAC40248.1| unnamed protein product [Mus
musculus]
Length = 1217
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 1013 RYKRNENQLIIFADDTYPRWVTTASLLDYDTVAGADKFGNIC 1054
Score = 38.5 bits (88), Expect = 0.72
Identities = 16/35 (45%), Positives = 26/35 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ + Q+ +++ELDRT E+ KKLE+ +
Sbjct: 1178 LCEQFNSMEPNKQKNVSEELDRTPPEVSKKLEDIR 1212
>gb|AAH00463.1| SF3B3 protein [Homo sapiens] gi|13111947|gb|AAH03146.1| SF3B3
protein [Homo sapiens]
Length = 399
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 195 RYKRNENQLIIFADDTYPRWVTTASLLDYDTVAGADKFGNIC 236
Score = 38.5 bits (88), Expect = 0.72
Identities = 16/35 (45%), Positives = 26/35 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ + Q+ +++ELDRT E+ KKLE+ +
Sbjct: 360 LCEQFNSMEPNKQKNVSEELDRTPPEVSKKLEDIR 394
>ref|XP_214697.3| PREDICTED: similar to Splicing factor 3b, subunit 3, 130kDa [Rattus
norvegicus]
Length = 1562
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 1358 RYKRNENQLIIFADDTYPRWVTTASLLDYDTVAGADKFGNIC 1399
Score = 38.5 bits (88), Expect = 0.72
Identities = 16/35 (45%), Positives = 26/35 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ + Q+ +++ELDRT E+ KKLE+ +
Sbjct: 1523 LCEQFNSMEPNKQKNVSEELDRTPPEVSKKLEDIR 1557
>ref|XP_536791.1| PREDICTED: similar to KIAA0017 protein [Canis familiaris]
Length = 1283
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 1079 RYKRNENQLIIFADDTYPRWVTTASLLDYDTVAGADKFGNIC 1120
Score = 38.5 bits (88), Expect = 0.72
Identities = 16/35 (45%), Positives = 26/35 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ + Q+ +++ELDRT E+ KKLE+ +
Sbjct: 1244 LCEQFNSMEPNKQKNVSEELDRTPPEVSKKLEDIR 1278
>ref|XP_581167.1| PREDICTED: similar to KIAA0017 protein, partial [Bos taurus]
Length = 1022
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 899 RYKRNENQLIIFADDTYPRWVTTASLLDYDTVAGADKFGNIC 940
>emb|CAG32589.1| hypothetical protein [Gallus gallus]
Length = 503
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 299 RYKRNENQLIIFADDTYPRWVTTATLLDYDTVAGADKFGNIC 340
Score = 39.7 bits (91), Expect = 0.32
Identities = 17/35 (48%), Positives = 26/35 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ + Q+ +A+ELDRT E+ KKLE+ +
Sbjct: 464 LCEQFNSMEPNKQKNVAEELDRTPPEVSKKLEDIR 498
>gb|AAH31197.2| Sf3b3 protein [Mus musculus]
Length = 494
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 290 RYKRNENQLIIFADDTYPRWVTTASLLDYDTVAGADKFGNIC 331
Score = 38.5 bits (88), Expect = 0.72
Identities = 16/35 (45%), Positives = 26/35 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ + Q+ +++ELDRT E+ KKLE+ +
Sbjct: 455 LCEQFNSMEPNKQKNVSEELDRTPPEVSKKLEDIR 489
>dbj|BAC97845.1| mKIAA0017 protein [Mus musculus]
Length = 1122
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 918 RYKRNENQLIIFADDTYPRWVTTASLLDYDTVAGADKFGNIC 959
Score = 38.5 bits (88), Expect = 0.72
Identities = 16/35 (45%), Positives = 26/35 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ + Q+ +++ELDRT E+ KKLE+ +
Sbjct: 1083 LCEQFNSMEPNKQKNVSEELDRTPPEVSKKLEDIR 1117
>dbj|BAA32662.2| KIAA0017 protein [Homo sapiens]
Length = 1253
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 1049 RYKRNENQLIIFADDTYPRWVTTASLLDYDTVAGADKFGNIC 1090
Score = 38.5 bits (88), Expect = 0.72
Identities = 16/35 (45%), Positives = 26/35 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ + Q+ +++ELDRT E+ KKLE+ +
Sbjct: 1214 LCEQFNSMEPNKQKNVSEELDRTPPEVSKKLEDIR 1248
>ref|XP_414047.1| PREDICTED: similar to KIAA0017 protein [Gallus gallus]
Length = 1273
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 1069 RYKRNENQLIIFADDTYPRWVTTATLLDYDTVAGADKFGNIC 1110
Score = 39.7 bits (91), Expect = 0.32
Identities = 17/35 (48%), Positives = 26/35 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ + Q+ +A+ELDRT E+ KKLE+ +
Sbjct: 1234 LCEQFNSMEPNKQKNVAEELDRTPPEVSKKLEDIR 1268
>emb|CAH90875.1| hypothetical protein [Pongo pygmaeus]
Length = 1217
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 1013 RYKRNENQLIIFADDTYPRWVTTASLLDYDTVAGADKFGNIC 1054
Score = 38.9 bits (89), Expect = 0.55
Identities = 16/35 (45%), Positives = 26/35 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ + Q+ +++ELDRT E+ KKLE+ +
Sbjct: 1178 LCEQFNSMEPNKQKNVSEELDRTPPEVTKKLEDIR 1212
>gb|AAH68974.1| Splicing factor 3b, subunit 3, 130kDa [Homo sapiens]
Length = 1217
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 1013 RYKRNENQLIIFADDTYPRWVTTASLLDYDTVAGADKFGNIC 1054
Score = 38.5 bits (88), Expect = 0.72
Identities = 16/35 (45%), Positives = 26/35 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ + Q+ +++ELDRT E+ KKLE+ +
Sbjct: 1178 LCEQFNSMEPNKQKNVSEELDRTPPEVSKKLEDIR 1212
>dbj|BAA02805.1| KIAA0017 [Homo sapiens]
Length = 399
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 195 RYKRNENQLIIFADDTYPRWVTTASLLDYDTVAGADKFGNIC 236
Score = 38.5 bits (88), Expect = 0.72
Identities = 16/35 (45%), Positives = 26/35 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ + Q+ +++ELDRT E+ KKLE+ +
Sbjct: 360 LCEQFNSMEPNKQKNVSEELDRTPPEVSKKLEDIR 394
>emb|CAB53699.1| hypothetical protein [Homo sapiens] gi|7512688|pir||T14779
hypothetical protein DKFZp434P041.1 - human (fragment)
Length = 215
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 11 RYKRNENQLIIFADDTYPRWVTTASLLDYDTVAGADKFGNIC 52
Score = 38.5 bits (88), Expect = 0.72
Identities = 16/35 (45%), Positives = 26/35 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ + Q+ +++ELDRT E+ KKLE+ +
Sbjct: 176 LCEQFNSMEPNKQKNVSEELDRTPPEVSKKLEDIR 210
>sp|Q15393|SF3B3_HUMAN Splicing factor 3B subunit 3 (Spliceosome associated protein 130)
(SAP 130) (SF3b130) (Pre-mRNA splicing factor SF3b 130
kDa subunit) (STAF130) gi|6006515|emb|CAB56791.1|
spliceosomal protein SAP 130 [Homo sapiens]
Length = 1217
Score = 57.8 bits (138), Expect = 1e-06
Identities = 26/42 (61%), Positives = 34/42 (80%), Gaps = 1/42 (2%)
Query: 486 KYRWDENQLYIFADDCVPRWLTASYHIDFDTMAGADKFG-IC 526
+Y+ +ENQL IFADD PRW+T + +D+DT+AGADKFG IC
Sbjct: 1013 RYKRNENQLIIFADDTYPRWVTTASLLDYDTVAGADKFGNIC 1054
Score = 38.5 bits (88), Expect = 0.72
Identities = 16/35 (45%), Positives = 26/35 (73%)
Query: 640 LCEQFPTLPMDLQRKIADELDRTRGEILKKLEEYK 674
LCEQF ++ + Q+ +++ELDRT E+ KKLE+ +
Sbjct: 1178 LCEQFNSMEPNKQKNVSEELDRTPPEVSKKLEDIR 1212
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.360 0.160 0.619
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,004,319,785
Number of Sequences: 2540612
Number of extensions: 36577171
Number of successful extensions: 149689
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 51
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 149589
Number of HSP's gapped (non-prelim): 100
length of query: 675
length of database: 863,360,394
effective HSP length: 135
effective length of query: 540
effective length of database: 520,377,774
effective search space: 281003997960
effective search space used: 281003997960
T: 11
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 79 (35.0 bits)
Medicago: description of AC147482.6


