

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147471.10 + phase: 0
(532 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
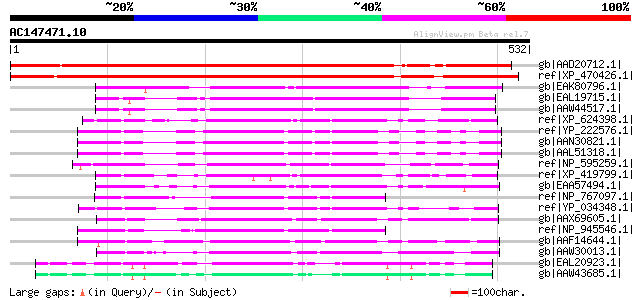
Score E
Sequences producing significant alignments: (bits) Value
gb|AAD20712.1| hypothetical protein [Arabidopsis thaliana] gi|25... 612 e-173
ref|XP_470426.1| putative AFG1-like ATPase [Oryza sativa (japoni... 575 e-162
gb|EAK80796.1| hypothetical protein UM00002.1 [Ustilago maydis 5... 181 7e-44
gb|EAL19715.1| hypothetical protein CNBG3430 [Cryptococcus neofo... 168 4e-40
gb|AAW44517.1| hypothetical protein CNG01350 [Cryptococcus neofo... 167 8e-40
ref|XP_624398.1| PREDICTED: similar to ENSANGP00000014807 [Apis ... 159 3e-37
ref|YP_222576.1| hypothetical protein BruAb1_1905 [Brucella abor... 158 4e-37
gb|AAN30821.1| conserved hypothetical protein [Brucella suis 133... 158 4e-37
gb|AAL51318.1| PUTATIVE ATPASE N2B [Brucella melitensis 16M] gi|... 158 4e-37
ref|NP_595259.1| ATPase [Schizosaccharomyces pombe 972h-] gi|295... 157 8e-37
ref|XP_419799.1| PREDICTED: similar to lactation elevated 1; CG8... 154 5e-36
gb|EAA57494.1| hypothetical protein MG10169.4 [Magnaporthe grise... 150 1e-34
ref|NP_767097.1| hypothetical protein bll0457 [Bradyrhizobium ja... 141 5e-32
ref|YP_034348.1| hypothetical protein BH16580 [Bartonella hensel... 141 6e-32
gb|AAX69605.1| ATPase, putative [Trypanosoma brucei] 139 3e-31
ref|NP_945546.1| AFG1-like ATPase [Rhodopseudomonas palustris CG... 135 4e-30
gb|AAF14644.1| 7138.4 [Leishmania major] 132 3e-29
gb|AAW30013.1| At4g28070 [Arabidopsis thaliana] gi|53850477|gb|A... 132 4e-29
gb|EAL20923.1| hypothetical protein CNBE2840 [Cryptococcus neofo... 114 6e-24
gb|AAW43685.1| conserved hypothetical protein [Cryptococcus neof... 113 1e-23
>gb|AAD20712.1| hypothetical protein [Arabidopsis thaliana] gi|25412329|pir||E84649
hypothetical protein At2g25530 [imported] - Arabidopsis
thaliana gi|15224744|ref|NP_180123.1| AFG1-like ATPase
family protein [Arabidopsis thaliana]
Length = 655
Score = 612 bits (1577), Expect = e-173
Identities = 305/518 (58%), Positives = 385/518 (73%), Gaps = 19/518 (3%)
Query: 1 MEEYHVNLSNWEKKRENERRRILMDEVEKQQNDKDWWK--RLNNKITERWTNSRKRPENV 58
ME+YHV L+ WEKKRE ERR+++++E EK+++D W + K+ RW R R NV
Sbjct: 152 MEDYHVRLAEWEKKREEERRKLMVEEAEKKEDDGMWASVNKQGQKLLGRWVLGR-RQMNV 210
Query: 59 DPGVGKWVSYLKREKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTMLMDMFYSATEGIVK 118
+PGVGKWVSYL RE+KLDS+VG RP PPAPKGLYIYGNVG GKTMLMDMFY AT+GI++
Sbjct: 211 EPGVGKWVSYLNRERKLDSIVGSRPAVPPAPKGLYIYGNVGCGKTMLMDMFYGATDGIIR 270
Query: 119 HRRRYHFHEAMLRINEHMHKTWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKK 178
HR+R+HFHEAML+INE MHK WK+ EKP+Q ISSWIMNLP D K KEWLA EE YK+
Sbjct: 271 HRQRFHFHEAMLKINEQMHKYWKENGAEKPMQYSISSWIMNLPVDEKVKEWLAGEEFYKQ 330
Query: 179 EVQMKHILPDVADKFFLDREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIV 238
++QMKHILP VADKF +D++ +KGA+ILCFDEIQTVDVFAIVALSGI+SRLL++GT++V
Sbjct: 331 QLQMKHILPAVADKFLVDQQSSKKGASILCFDEIQTVDVFAIVALSGIMSRLLTTGTVLV 390
Query: 239 ATSNRAPKDLNEANMVPEFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPI 298
ATSNRAP++LN+ M E F +S LE+HCE + +GSE+DYRR A+ S V+YLWP+
Sbjct: 391 ATSNRAPRELNQDGMQKEIFDKFISKLEKHCEIISIGSEVDYRRVAAKNSAENVHYLWPL 450
Query: 299 ERETINKFEKKWQDATGRFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGA 358
+ +FEK W+ T ++GG++ S T+ VMFGRT+EVPESC GVARFTF+YLCGRP+GA
Sbjct: 451 NDAVLEEFEKMWRQVTDQYGGEITSATLPVMFGRTVEVPESCSGVARFTFEYLCGRPVGA 510
Query: 359 ADYIAVAENYHTVFISDIPMMSMRIRDKTS--LSYFEQQKKKRSLLFRHGDLSLLLTNYT 416
ADYIAVA+NYHT+FISDIP MSM IRDK ++ ++ L+ H L+ + T
Sbjct: 511 ADYIAVAKNYHTIFISDIPAMSMEIRDKARRFITLVDE-------LYNH-HCCLVSSAET 562
Query: 417 IIILVYVAWHPLQLMNYFKELRRAHFLTWREAEGSKLRRDVLAEGNVGSGGTPVGITSIL 476
I ++ L +L F T E E S+LRRDVLAEG++ + G+P I S+L
Sbjct: 563 AIDELFQGTAEGTLF----DLESFQFET--ETEDSRLRRDVLAEGSISAAGSPSSIVSML 616
Query: 477 SGQEELFTFQRAVSRLIEMQTQLYLDGVSNVHPYFQTQ 514
SG+EE+F F RA SRLIEMQT LYL+GV +HPYF Q
Sbjct: 617 SGEEEMFAFARAASRLIEMQTPLYLEGVRFLHPYFHQQ 654
>ref|XP_470426.1| putative AFG1-like ATPase [Oryza sativa (japonica cultivar-group)]
gi|27573346|gb|AAO20064.1| putative AFG1-like ATPase
[Oryza sativa (japonica cultivar-group)]
Length = 740
Score = 575 bits (1481), Expect = e-162
Identities = 290/523 (55%), Positives = 369/523 (70%), Gaps = 18/523 (3%)
Query: 1 MEEYHVNLSNWEKKRENERRRILMDEVEKQQNDKDWWKRLNNKITERWTNSRKRPENVDP 60
ME+YH LS WE RE +RRR+L+ E E +Q D W + + SRKR N++P
Sbjct: 224 MEDYHARLSMWENTREKQRRRLLVQEAEDKQRDGVWIDEKRGFLDK--LVSRKRRGNIEP 281
Query: 61 GVGKWVSYLKREKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTMLMDMFYSATEGIVKHR 120
GVGKWVSYL REKKLD+LVG++P AP APKG+Y+YGNVGSGKTMLMDMFY ATEG++KHR
Sbjct: 282 GVGKWVSYLNREKKLDTLVGQKPVAPIAPKGIYLYGNVGSGKTMLMDMFYGATEGLIKHR 341
Query: 121 RRYHFHEAMLRINEHMHKTWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKKEV 180
RR+HFHEAML I++HMH WK++ E+K ++S SWI +LPFD K KEWL EE+YK+
Sbjct: 342 RRFHFHEAMLEIHDHMHDVWKRRDEDKSIESSAFSWISSLPFDGKIKEWLIGEEKYKQNT 401
Query: 181 QMKHILPDVADKFFLDREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVAT 240
Q HIL VADKF +DR+ + GA+ILCFDEIQT+DVFA+VALSGILSRLLS+GT++V+T
Sbjct: 402 QQNHILLAVADKFLVDRQANKSGASILCFDEIQTIDVFAVVALSGILSRLLSTGTVLVST 461
Query: 241 SNRAPKDLNEANMVPEFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIER 300
SN+AP+DLN+ M E F +LLS L+E+C K+LVG+E DYRR I +++Y WP+
Sbjct: 462 SNKAPEDLNQDGMQREIFLDLLSKLDENCNKILVGTETDYRRLIPTDGLTQIHYFWPLTS 521
Query: 301 ETINKFEKKWQDATGRFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAAD 360
+ + +E W D T + GG +IS TI VMFGR LE+P+SC GVARF F+YLCGRP+GAAD
Sbjct: 522 DIRSMYEAMWHDITRQTGGNIISVTIPVMFGRYLEIPKSCNGVARFDFEYLCGRPVGAAD 581
Query: 361 YIAVAENYHTVFISDIPMMSMRIRDKTS--LSYFEQQKKKRSLLFRHGDLSLLLTNYTII 418
YIA+A NYHT+FISDIP MSM+IRDK ++ ++ L H L L +
Sbjct: 582 YIAIARNYHTIFISDIPAMSMKIRDKARRFITLIDE------LYNHHCRLVCLAASSIDD 635
Query: 419 ILVYVAWHPLQLMNYFKELRRAHFLTWREAEGSKLRRDVLAEGNVGSGGTPVGITSILSG 478
+ PL + F+ EAEG+KLRRDVLAEGNVG+ +P G+ +ILSG
Sbjct: 636 LFQGTDEGPLFDLESFQ--------FEGEAEGAKLRRDVLAEGNVGAAPSPTGLVAILSG 687
Query: 479 QEELFTFQRAVSRLIEMQTQLYLDGVSNVHPYFQTQHKKFQKN 521
QEE+F F+RA+SRLIEMQT LYL+ V VH Q Q K+
Sbjct: 688 QEEMFAFRRAISRLIEMQTSLYLERVERVHSSLQQQSSVLTKS 730
>gb|EAK80796.1| hypothetical protein UM00002.1 [Ustilago maydis 521]
gi|49066654|ref|XP_397617.1| hypothetical protein
UM00002.1 [Ustilago maydis 521]
Length = 550
Score = 181 bits (458), Expect = 7e-44
Identities = 128/425 (30%), Positives = 202/425 (47%), Gaps = 61/425 (14%)
Query: 89 PKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAMLRINEHMH----KTWKKQM 144
PKGLY+YG+VG+GK+MLMD+FY + +RR HFH+ M+ ++ H KT K
Sbjct: 169 PKGLYLYGDVGTGKSMLMDLFYDTLPSNITSKRRIHFHQFMIEAHKRAHFYKSKTHKPSG 228
Query: 145 EEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILPDVADKFFLDREGEEKGA 204
+ SG SS + A E A E +E+ H
Sbjct: 229 IVMMMSSGSSSSASSAGGAASAGEESDAIEAVAREMARNH-------------------- 268
Query: 205 NILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNEANMVPEFFQNLLSN 264
++LCFDE Q D+ + L G+L R+L+ G ++V TSNR P +L + + + F +
Sbjct: 269 SVLCFDEFQVTDIADAMILRGLLERMLAYGVVMVMTSNRHPDELYKNGIQRQSFLPCIDL 328
Query: 265 LEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEKKWQDATGRFGGKVISN 324
L+ + S DYR+ R+ ++V Y P++ +F+K + T V+
Sbjct: 329 LKSRLGVTDLNSGTDYRK--VPRALSKV-YFSPLDDANTREFDKLFDAMTSDPHDPVVEK 385
Query: 325 TISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYHTVFISDIPMMSMRIR 384
++GRTL+VP S + VARFTFD LCGRP AADYI + N+ T+FI DIP M + R
Sbjct: 386 RPLKIWGRTLQVPRSTQRVARFTFDELCGRPRSAADYIEICNNFSTIFIDDIPKMGLNQR 445
Query: 385 D--KTSLSYFEQQKKKRSLLFRHGDLSLLLTNYTIIILVYVAWHPLQLMNYFKELRRAHF 442
D + +++ + + ++ L ++ + LQ+ +
Sbjct: 446 DLARRFITFIDAAYESKTKLLASSEVPI-----------------LQIFS---------- 478
Query: 443 LTWREAEGSKLRRDVLAE--GNVGSGGTPVGITSILSGQEELFTFQRAVSRLIEMQTQLY 500
+A +K D + ++G +G + I +G EELF F R +SRL EM ++ Y
Sbjct: 479 ---GDAGDAKPTADQMRALMDDLGLTMDDLGGSPIFTGDEELFAFARVISRLTEMGSRQY 535
Query: 501 LDGVS 505
+ S
Sbjct: 536 AETAS 540
>gb|EAL19715.1| hypothetical protein CNBG3430 [Cryptococcus neoformans var.
neoformans B-3501A]
Length = 521
Score = 168 bits (425), Expect = 4e-40
Identities = 131/424 (30%), Positives = 190/424 (43%), Gaps = 88/424 (20%)
Query: 89 PKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRR----------RYHFHEAMLRINEHMHK 138
PKGLY+YG+VG+GKTMLMD+F+S I K R R HFH ML + + HK
Sbjct: 160 PKGLYLYGSVGTGKTMLMDLFHST---IPKQFRPTSQGGYGSIRIHFHAFMLDVLQRQHK 216
Query: 139 TWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILPDVADKFFLDRE 198
E K + K +LP+VA L E
Sbjct: 217 L--------------------------------VVEYEKAGLGKKDVLPEVARS--LANE 242
Query: 199 GEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNEANMVPEFF 258
G +LCFDE Q D+ + L G+L RL+S G + + TSNR P +L + + F
Sbjct: 243 GR-----VLCFDEFQVTDIVTAMILRGLLERLMSFGVVCIMTSNRHPDELYINGIQRQSF 297
Query: 259 QNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEKKWQDATGRFG 318
+ ++E E V + S DYR+ S+ N L P + INK + +T
Sbjct: 298 IPAIELIKERFEVVDLDSGTDYRKIPRALSKVYYNPLSPTVKSEINKLFDSFA-STDPVS 356
Query: 319 GKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYHTVFISDIPM 378
+V+ N ++GR L VPES VA+FTF LC +PL AADY+ V + TVF+ DIP
Sbjct: 357 SEVVHNRKVHLWGRELNVPESSGSVAKFTFADLCNKPLSAADYLEVTSKFGTVFVEDIPR 416
Query: 379 MSMRIRDKTS--LSYFEQQKKKRSLLFRHGDLSLLLTNYTIIILVYVAWHPLQLMNYFKE 436
M + RD+ +++ + + ++ LF ++ +
Sbjct: 417 MGLSERDQARRFITFIDACYENKTKLFCSSEVPI-------------------------- 450
Query: 437 LRRAHFLTWREAEGSKLRRDVLAE--GNVGSGGTPVGITSILSGQEELFTFQRAVSRLIE 494
F + + GS + E +G + VG +S+ SG EELF F R VSRL +
Sbjct: 451 -----FQVFSDKHGSAAEDAHMQEVMDELGLDPSAVGSSSLFSGDEELFAFARCVSRLSQ 505
Query: 495 MQTQ 498
M T+
Sbjct: 506 MGTK 509
>gb|AAW44517.1| hypothetical protein CNG01350 [Cryptococcus neoformans var.
neoformans JEC21] gi|58269336|ref|XP_571824.1|
hypothetical protein CNG01350 [Cryptococcus neoformans
var. neoformans JEC21]
Length = 521
Score = 167 bits (423), Expect = 8e-40
Identities = 131/424 (30%), Positives = 189/424 (43%), Gaps = 88/424 (20%)
Query: 89 PKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRR----------RYHFHEAMLRINEHMHK 138
PKGLY+YG+VG+GKTMLMD+F+S I K R R HFH ML + + HK
Sbjct: 160 PKGLYLYGSVGTGKTMLMDLFHST---IPKQFRPTSQGGYGSIRIHFHAFMLDVLQRQHK 216
Query: 139 TWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILPDVADKFFLDRE 198
E K + K +LP+VA L E
Sbjct: 217 L--------------------------------VVEYEKAGLGKKDVLPEVARS--LANE 242
Query: 199 GEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNEANMVPEFF 258
G +LCFDE Q D+ + L G+L RL+S G + + TSNR P +L + + F
Sbjct: 243 GR-----VLCFDEFQVTDIVTAMILRGLLERLMSFGVVCIMTSNRHPDELYINGIQRQSF 297
Query: 259 QNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEKKWQDATGRFG 318
+ ++E E V + S DYR S+ N L P + INK + +T
Sbjct: 298 IPAIELIKERFEVVDLDSGTDYREIPRALSKVYYNPLSPTVKSEINKLFDSFA-STDPVS 356
Query: 319 GKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYHTVFISDIPM 378
+V+ N ++GR L VPES VA+FTF LC +PL AADY+ V + TVF+ DIP
Sbjct: 357 SEVVHNRKVHLWGRELNVPESSGSVAKFTFADLCNKPLSAADYLEVTSKFGTVFVEDIPR 416
Query: 379 MSMRIRDKTS--LSYFEQQKKKRSLLFRHGDLSLLLTNYTIIILVYVAWHPLQLMNYFKE 436
M + RD+ +++ + + ++ LF ++ +
Sbjct: 417 MGLSERDQARRFITFIDACYENKTKLFCSSEVPI-------------------------- 450
Query: 437 LRRAHFLTWREAEGSKLRRDVLAE--GNVGSGGTPVGITSILSGQEELFTFQRAVSRLIE 494
F + + GS + E +G + VG +S+ SG EELF F R VSRL +
Sbjct: 451 -----FQVFSDKHGSAAEDAHMQEVMDELGLDPSAVGSSSLFSGDEELFAFARCVSRLSQ 505
Query: 495 MQTQ 498
M T+
Sbjct: 506 MGTK 509
>ref|XP_624398.1| PREDICTED: similar to ENSANGP00000014807 [Apis mellifera]
Length = 436
Score = 159 bits (401), Expect = 3e-37
Identities = 121/426 (28%), Positives = 205/426 (47%), Gaps = 70/426 (16%)
Query: 75 LDSLVGRRPTAPPAPKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAMLRINE 134
LD +GR+ PP KGLY+YG VG GKTMLMD+FY + +K+++R HFH ML ++
Sbjct: 73 LDKWLGRKRKQPP--KGLYLYGAVGGGKTMLMDLFYQCCQ--IKNKKRVHFHSFMLDVHN 128
Query: 135 HMHKTWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILPDVADKFF 194
+H+ +K + +SS + PFD +P VA
Sbjct: 129 KVHEV------KKNIVRDVSSTKLQ-PFDP---------------------IPPVARSI- 159
Query: 195 LDREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNEANMV 254
+ A +LCFDE Q D+ + L + + L ++G I++ATSNRAP DL + +
Sbjct: 160 ------TEEAWLLCFDEFQVTDIADAMILKRLFTELFNNGVIVIATSNRAPDDLYKNGLQ 213
Query: 255 PEFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEKKWQDAT 314
F + L+ +C + S IDYR +E ++ ++ ++ I+ +K ++ +
Sbjct: 214 RGNFIPFIQVLKNYCIINSLDSGIDYRLKTGLGNE-KIYFI--KGKDAISDVDKVFKYLS 270
Query: 315 GRFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYHTVFIS 374
+ V S TI + GR + ++C + TF+ LC RPLGA+DY+ +++ +HTV I
Sbjct: 271 SKENDVVRSRTICIR-GRNVTFKKTCGQILDSTFEELCDRPLGASDYLELSQAFHTVIIR 329
Query: 375 DIPMMSMRIRDKTSLSYFEQQKKKRSLLFRHGDLSLLLTNYTIIILVYVAWHPLQLMNYF 434
D+P ++ R++ +T +R + + L N +++ H
Sbjct: 330 DVPQLNFRLKSQT----------RRFITL----IDTLYDNKVRVVISAAVPH-------- 367
Query: 435 KELRRAHFLTWREAEGSKLRRDVLAEGNVGSGGTPVGITSILSGQEELFTFQRAVSRLIE 494
F+ E + +R ++ + + G +++ +G+EELF F R +SRL E
Sbjct: 368 ----TKLFVPEGNNEYTDEKRMLMDDLKISHGSDDYK-SNLFTGEEELFAFDRTISRLSE 422
Query: 495 MQTQLY 500
MQT Y
Sbjct: 423 MQTTQY 428
>ref|YP_222576.1| hypothetical protein BruAb1_1905 [Brucella abortus biovar 1 str.
9-941] gi|62196915|gb|AAX75215.1| conserved hypothetical
protein [Brucella abortus biovar 1 str. 9-941]
Length = 387
Score = 158 bits (400), Expect = 4e-37
Identities = 127/436 (29%), Positives = 191/436 (43%), Gaps = 107/436 (24%)
Query: 70 KREKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAM 129
++ L L G+R KGLY++G VG GKTMLMDMF+ V+ +RR HF++ M
Sbjct: 53 RKSSALGWLFGKRRETAATIKGLYVHGEVGRGKTMLMDMFFQLLP--VERKRRAHFNDFM 110
Query: 130 LRINEHMHKTWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKK-EVQMKHILPD 188
++E ++ A + +K+ E + +P
Sbjct: 111 ADVHERIY---------------------------------AHRQAHKRGETKQDDPIPP 137
Query: 189 VADKFFLDREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDL 248
VA E + A +LCFDE D+ + LS + S L S G ++VATSN AP +L
Sbjct: 138 VA-------EALSQQAWVLCFDEFAVTDIADAMILSRLFSALFSRGVVLVATSNVAPDNL 190
Query: 249 NEANMVPEFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEK 308
+ + F + L+ H + + + S DYR ++ + + YL P+ ET + +
Sbjct: 191 YRDGLNRQLFLPFIDILKSHVDVINLDSRTDYR---LEKLDRQPVYLSPLGPETERRMDA 247
Query: 309 KWQDATGRFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENY 368
W A + G + + I + GR +EVP + G ARF+FD LC RPLGA+DYIA+A Y
Sbjct: 248 AW--AAHKNGAEEKPDVIHIK-GRDIEVPRAAAGAARFSFDDLCARPLGASDYIAIANRY 304
Query: 369 HTVFISDIPMMSMRIRDKTSLSYFEQQKKKRSLLFRHGDLSLLLTNYTIIILVYVAWHPL 428
T+FI ++P+ L Y + + KR I+L+ V +
Sbjct: 305 PTLFIDNVPV----------LDYSRRNEAKR-----------------FILLIDVLYD-- 335
Query: 429 QLMNYFKELRRAHFLTWREAEGSKLRRDVLAEGNVGSGGTPVGITSILSGQEELFTFQRA 488
A EA+ KL + + GT E F F R
Sbjct: 336 ---------HHARLFVSAEAQPEKL--------YIANSGT------------EAFEFDRT 366
Query: 489 VSRLIEMQTQLYLDGV 504
SRL EMQ+ YLDG+
Sbjct: 367 ASRLFEMQSAEYLDGI 382
>gb|AAN30821.1| conserved hypothetical protein [Brucella suis 1330]
gi|23502779|ref|NP_698906.1| hypothetical protein BR1929
[Brucella suis 1330]
Length = 387
Score = 158 bits (400), Expect = 4e-37
Identities = 127/436 (29%), Positives = 191/436 (43%), Gaps = 107/436 (24%)
Query: 70 KREKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAM 129
++ L L G+R KGLY++G VG GKTMLMDMF+ V+ +RR HF++ M
Sbjct: 53 RKSSALGWLFGKRRETAATIKGLYVHGEVGRGKTMLMDMFFQLLP--VERKRRAHFNDFM 110
Query: 130 LRINEHMHKTWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKK-EVQMKHILPD 188
++E ++ A + +K+ E + +P
Sbjct: 111 ADVHERIY---------------------------------AHRQAHKRGETKQDDPIPP 137
Query: 189 VADKFFLDREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDL 248
VA E + A +LCFDE D+ + LS + S L S G ++VATSN AP +L
Sbjct: 138 VA-------EALSQQAWVLCFDEFTVTDIADAMILSRLFSALFSRGVVLVATSNVAPDNL 190
Query: 249 NEANMVPEFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEK 308
+ + F + L+ H + + + S DYR ++ + + YL P+ ET + +
Sbjct: 191 YRDGLNRQLFLPFIDILKSHVDVINLDSRTDYR---LEKLDRQPVYLSPLGPETERRMDA 247
Query: 309 KWQDATGRFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENY 368
W A + G + + I + GR +EVP + G ARF+FD LC RPLGA+DYIA+A Y
Sbjct: 248 AW--AAHKNGAEEKPDVIHIK-GRDIEVPRAAAGAARFSFDDLCARPLGASDYIAIANRY 304
Query: 369 HTVFISDIPMMSMRIRDKTSLSYFEQQKKKRSLLFRHGDLSLLLTNYTIIILVYVAWHPL 428
T+FI ++P+ L Y + + KR I+L+ V +
Sbjct: 305 PTLFIDNVPV----------LDYSRRNEAKR-----------------FILLIDVLYD-- 335
Query: 429 QLMNYFKELRRAHFLTWREAEGSKLRRDVLAEGNVGSGGTPVGITSILSGQEELFTFQRA 488
A EA+ KL + + GT E F F R
Sbjct: 336 ---------HHARLFVSAEAQPEKL--------YIANSGT------------EAFEFDRT 366
Query: 489 VSRLIEMQTQLYLDGV 504
SRL EMQ+ YLDG+
Sbjct: 367 ASRLFEMQSAEYLDGI 382
>gb|AAL51318.1| PUTATIVE ATPASE N2B [Brucella melitensis 16M]
gi|17986420|ref|NP_539054.1| PUTATIVE ATPASE N2B
[Brucella melitensis 16M] gi|25355785|pir||AC3269
probable ATPase N2B BMEI0136 [imported] - Brucella
melitensis (strain 16M)
Length = 403
Score = 158 bits (400), Expect = 4e-37
Identities = 127/436 (29%), Positives = 191/436 (43%), Gaps = 107/436 (24%)
Query: 70 KREKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAM 129
++ L L G+R KGLY++G VG GKTMLMDMF+ V+ +RR HF++ M
Sbjct: 69 RKSSALGWLFGKRRETAATIKGLYVHGEVGRGKTMLMDMFFQLLP--VERKRRAHFNDFM 126
Query: 130 LRINEHMHKTWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKK-EVQMKHILPD 188
++E ++ A + +K+ E + +P
Sbjct: 127 ADVHERIY---------------------------------AHRQAHKRGETKQDDPIPP 153
Query: 189 VADKFFLDREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDL 248
VA E + A +LCFDE D+ + LS + S L S G ++VATSN AP +L
Sbjct: 154 VA-------EALSQQAWVLCFDEFTVTDIADAMILSRLFSALFSRGVVLVATSNVAPDNL 206
Query: 249 NEANMVPEFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEK 308
+ + F + L+ H + + + S DYR ++ + + YL P+ ET + +
Sbjct: 207 YRDGLNRQLFLPFIDILKSHVDVINLDSRTDYR---LEKLDRQPVYLSPLGPETERRMDA 263
Query: 309 KWQDATGRFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENY 368
W A + G + + I + GR +EVP + G ARF+FD LC RPLGA+DYIA+A Y
Sbjct: 264 AW--AAHKNGAEEKPDVIHIK-GRDIEVPRAAAGAARFSFDDLCARPLGASDYIAIANRY 320
Query: 369 HTVFISDIPMMSMRIRDKTSLSYFEQQKKKRSLLFRHGDLSLLLTNYTIIILVYVAWHPL 428
T+FI ++P+ L Y + + KR I+L+ V +
Sbjct: 321 PTLFIDNVPV----------LDYSRRNEAKR-----------------FILLIDVLYD-- 351
Query: 429 QLMNYFKELRRAHFLTWREAEGSKLRRDVLAEGNVGSGGTPVGITSILSGQEELFTFQRA 488
A EA+ KL + + GT E F F R
Sbjct: 352 ---------HHARLFVSAEAQPEKL--------YIANSGT------------EAFEFDRT 382
Query: 489 VSRLIEMQTQLYLDGV 504
SRL EMQ+ YLDG+
Sbjct: 383 ASRLFEMQSAEYLDGI 398
>ref|NP_595259.1| ATPase [Schizosaccharomyces pombe 972h-] gi|2956750|emb|CAA17914.1|
SPBC115.02c [Schizosaccharomyces pombe]
gi|7492423|pir||T39297 probable atpase - fission yeast
(Schizosaccharomyces pombe)
Length = 454
Score = 157 bits (397), Expect = 8e-37
Identities = 127/441 (28%), Positives = 186/441 (41%), Gaps = 85/441 (19%)
Query: 65 WVSYLKR---EKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRR 121
W+S LK+ KK +L P P PKG+Y+YG+VG GKT LMD+FY V +
Sbjct: 92 WISPLKKMFSRKKSPTLTSSLPV-PGMPKGIYLYGDVGCGKTALMDLFYHNLPPNVTRSQ 150
Query: 122 RYHFHEAMLRINEHMHKTWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKKEVQ 181
R HFH M++++ H ++RY E+
Sbjct: 151 RIHFHAFMMQVHRTSHDL---------------------------------QDRYGFEID 177
Query: 182 -MKHILPDVADKFFLDREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVAT 240
+ HI +A K +LCFDE+Q DV + L + L+ G +I T
Sbjct: 178 FIDHIASGIA-----------KETTVLCFDELQVTDVADALLLRRLFEALMKYGVVIFIT 226
Query: 241 SNRAPKDLNEANMVPEFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIER 300
SNRAP DL + + E F + LE + + + S DYRR +S+ YL+P
Sbjct: 227 SNRAPSDLYKNGIQRESFIPCIKLLEHRLQVICLDSPNDYRRL---KSKTEDTYLYPANS 283
Query: 301 ETINKFEKKWQDATGRFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAAD 360
+ K + W + + V FGR + VP++ VA FTF+ LCG P AAD
Sbjct: 284 PEVKKALENWFLCYADEKDPAHQDEVEV-FGRKIIVPKASGNVAWFTFEQLCGEPKSAAD 342
Query: 361 YIAVAENYHTVFISDIPMMSMRIRDKTSLSYFEQQKKKRSLLFRHGDLSLLLTNYTIIIL 420
Y+++A YH +SDIP +S+ +D L+ T I
Sbjct: 343 YLSLASRYHVFIVSDIPKLSIESKD------------------------LIHRFITFIDA 378
Query: 421 VYVAWHPLQLMNYFKELRRAHFLTWREAEGSKLRRDVLAEGNVGSGGTPVGITSILSGQE 480
+Y L L + E+ +E D A+G + S G E
Sbjct: 379 LYDTHGKLILSS---EVPVQEIYPTAPSEVLSSTADPAAKGKIES-----HYHGAFGGIE 430
Query: 481 ELFTFQRAVSRLIEMQTQLYL 501
E+FTF R +SRL EM+ Q ++
Sbjct: 431 EVFTFTRCLSRLSEMKKQSWI 451
>ref|XP_419799.1| PREDICTED: similar to lactation elevated 1; CG8520 gene product;
lactation elevated-1 [Gallus gallus]
Length = 1272
Score = 154 bits (390), Expect = 5e-36
Identities = 128/426 (30%), Positives = 204/426 (47%), Gaps = 83/426 (19%)
Query: 89 PKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAMLRINEHMHKTWKKQMEEKP 148
PKGLY+YG+VG+GKTM+MDMFYS + V+ ++R HFH ML +++ +H+ + + KP
Sbjct: 319 PKGLYVYGDVGTGKTMVMDMFYSHLK--VERKKRVHFHGFMLDVHQRIHRLKQSLPKRKP 376
Query: 149 LQSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILPDVADKFFLDREGEEKGANILC 208
F K+ + +A VA++ + A +LC
Sbjct: 377 ------------GFMAKSYDPIAP----------------VAEEI-------SEEACLLC 401
Query: 209 FDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDL-----NEANMVPEFFQNLLS 263
FDE Q D+ + L + L +G ++VATSNR P+DL AN VP L
Sbjct: 402 FDEFQVTDIADAMILKQLFENLFQNGVVVVATSNRPPEDLYKNGLQRANFVPFIAVLKLL 461
Query: 264 NLE--------EHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEKKWQDATG 315
L ++C V + S IDYR+ + + ++ YL + K D
Sbjct: 462 LLGGCVTASSCKYCSTVQLDSGIDYRKRVLPAA-GKLYYL--TSEADVEAVMDKLFDELA 518
Query: 316 RFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYHTVFISD 375
+ + I + GR L + ++C +A FTF+ LC RPLGA+DY+ +++++ TVF+ D
Sbjct: 519 QKQNDLTRPRILKVQGRELRLNKACGTIADFTFEELCDRPLGASDYLEISKHFDTVFVRD 578
Query: 376 IPMMSMRIRDKTSLSYFEQQKKKRSLLFRHGDLSLLLTNYTIII-LVYVAWHPLQLMNYF 434
IP ++M K+ ++ F ++L+ T Y + ++ A PLQ + +
Sbjct: 579 IPPLTMA-------------KRTQARRF----ITLIDTFYEHKVRIICSAATPLQSL-FV 620
Query: 435 KELRRAHFLTWREAEGSKLRRDVLAEGNVGSGGTPVGITSILSGQEELFTFQRAVSRLIE 494
E E E S++ D L + G S+ +G+EE+F FQR +SRL E
Sbjct: 621 VEAGSI------ELEDSRVLMDDLDLSQDSAKGL-----SMFTGEEEIFAFQRTISRLTE 669
Query: 495 MQTQLY 500
MQT+ Y
Sbjct: 670 MQTEQY 675
>gb|EAA57494.1| hypothetical protein MG10169.4 [Magnaporthe grisea 70-15]
gi|39969117|ref|XP_365949.1| hypothetical protein
MG10169.4 [Magnaporthe grisea 70-15]
Length = 721
Score = 150 bits (379), Expect = 1e-34
Identities = 129/422 (30%), Positives = 195/422 (45%), Gaps = 72/422 (17%)
Query: 89 PKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAMLRINEHMHKTWKKQMEEKP 148
P+GLY+YG+VGSGKTMLMD+FY VK + R HFH M +++ MHK
Sbjct: 335 PRGLYLYGDVGSGKTMLMDLFYDTLPPSVKSKTRIHFHNFMQDVHKRMHKM--------K 386
Query: 149 LQSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILPDVADKFFLDREGEEKGANILC 208
+Q G D A +AA D+A++ N+LC
Sbjct: 387 MQHGN---------DLDAVPLIAA---------------DIAEQ-----------GNVLC 411
Query: 209 FDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNEANMVPEFFQNLLSNLEEH 268
FDE Q DV + L +L L+S G ++V TSNR P DL + + E F + L+
Sbjct: 412 FDEFQCTDVADAMILRRLLEALMSHGVVLVTTSNRHPDDLYKNGVQRESFIPAIELLKSR 471
Query: 269 CEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEKKWQDATGRFGGKVISNTISV 328
+ + S DYR+ R + V Y P+++ + E KW G T +V
Sbjct: 472 LHVINLDSPTDYRKI--PRPPSGV-YHTPLDKHAQSHAE-KWFAFLGDASDPGHPETQTV 527
Query: 329 MFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYHTVFISDIPMMSMRIRDKTS 388
+GR + VP A FTFD L GRP GAADYI + +Y ++D+P M+ R RD
Sbjct: 528 -WGRKIHVPRVSGRCACFTFDELIGRPTGAADYIELVRSYDAFVVTDVPGMTYRQRDLA- 585
Query: 389 LSYFEQQKKKRSLLFRHGDLSLLLTNYTIIILVYVAWHPL-QLMNYFKELRRAHFLTWRE 447
+R + F + + ++ ++L A PL +L +E+R + T ++
Sbjct: 586 ---------RRFITF----IDAVYESHAKLVLTTAA--PLGELFVSREEMRESLAATRKK 630
Query: 448 AEGSKLRRDVLAEGNVG-------SGGTPVGITSILSGQEELFTFQRAVSRLIEMQTQLY 500
G + D EG +G S + +++ SG EE F F RA+SRL M ++ +
Sbjct: 631 DAGREEPDDGDVEGAMGHMMEDLDSNVDKLRNSNLFSGDEEAFAFARALSRLSHMGSKEW 690
Query: 501 LD 502
++
Sbjct: 691 VE 692
>ref|NP_767097.1| hypothetical protein bll0457 [Bradyrhizobium japonicum USDA 110]
gi|27348705|dbj|BAC45722.1| bll0457 [Bradyrhizobium
japonicum USDA 110]
Length = 394
Score = 141 bits (356), Expect = 5e-32
Identities = 89/298 (29%), Positives = 142/298 (46%), Gaps = 47/298 (15%)
Query: 88 APKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAMLRINEHMHKTWKKQMEEK 147
AP GLYI+G VG GKTMLMD+F+ + V+H+ R HFHE M ++E ++ + +
Sbjct: 65 APHGLYIHGEVGRGKTMLMDLFFQHSS--VEHKHRAHFHEFMADVHERIYDYRQSIARGE 122
Query: 148 PLQSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILPDVADKFFLDREGEEKGANIL 207
+ + N F+ + WL L
Sbjct: 123 IADGDVIALTANAIFE---ESWL------------------------------------L 143
Query: 208 CFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNEANMVPEFFQNLLSNLEE 267
CFDE D+ + L + ++L GT++VATSN AP+DL + + F + + +
Sbjct: 144 CFDEFHVTDIADAMILGRLFAKLFELGTVVVATSNVAPEDLYKGGLNRSLFLPFIKQITD 203
Query: 268 HCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEKKWQDATGRFGGKVISNTIS 327
H + + + D+R ++ + +L P + + ++ W +G K S IS
Sbjct: 204 HMDVARLDARTDFR---LEKLQGVPMWLTPADGDADAVLDRAWSRMSG--SAKCKSRDIS 258
Query: 328 VMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYHTVFISDIPMMSMRIRD 385
+ GR L VP S GVARF+F LC +PLGA+DY+ +A +YHT+ + IP+M R+
Sbjct: 259 IK-GRILHVPCSAHGVARFSFTDLCEKPLGASDYLRLAHDYHTILVDHIPVMDFSQRN 315
>ref|YP_034348.1| hypothetical protein BH16580 [Bartonella henselae str. Houston-1]
gi|49239115|emb|CAF28419.1| hypothetical protein
[Bartonella henselae str. Houston-1]
Length = 401
Score = 141 bits (355), Expect = 6e-32
Identities = 115/431 (26%), Positives = 187/431 (42%), Gaps = 105/431 (24%)
Query: 71 REKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAML 130
+ KK +S+ ++ A P +GLYIYG VG GKTMLMD+F+S K ++R HF++ M
Sbjct: 54 KRKKQNSVAKQKSAANPF-QGLYIYGEVGRGKTMLMDLFFSCLPQ--KRKKRAHFNDFMA 110
Query: 131 RINEHMHKTWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILPDVA 190
++E ++ + + K Q + + ++ D+A
Sbjct: 111 DVHERINIYRQASKDGKTGQ----------------------------KTPILAVVEDLA 142
Query: 191 DKFFLDREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNE 250
+ A +LCFDE D+ + L ++S L G VATSN AP +L
Sbjct: 143 QE-----------AQVLCFDEFSVTDIADAMVLGRLISALFDKGIFFVATSNVAPDNLYY 191
Query: 251 ANMVPEFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEKKW 310
+ E F + L+ H + +G++ DYR ++S + Y+ P+ E + ++ W
Sbjct: 192 NGLNRELFLPFIQVLKAHVHVINLGAKTDYR---LEKSNFQQVYITPLGLEANQRMDQAW 248
Query: 311 QDATGRFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYHT 370
G K S+ S+ GR + +P S G ARF + LC +PL A +Y+A+ E YHT
Sbjct: 249 MLVLK--GQKETSDEFSIK-GRVIHIPRSGVGCARFDYQDLCAKPLAAVEYLALGERYHT 305
Query: 371 VFISDIPMMSMRIRDKTSLSYFEQQKKKRSLLFRHGDLSLLLTNYTIIILVYVAWHPLQL 430
+F+ ++P+M R++T KR +LF + +L Y + +
Sbjct: 306 IFVDNVPVMDDTCRNET----------KRFILF----IDVLYERYIRLFM---------- 341
Query: 431 MNYFKELRRAHFLTWREAEGSKLRRDVLAEGNVGSGGTPVGITSILSGQEELFTFQRAVS 490
+ ++ D L +G + E F FQR S
Sbjct: 342 -------------------SAAVKLDDLYKG--------------YAQTSETFEFQRTQS 368
Query: 491 RLIEMQTQLYL 501
RL EMQ+Q YL
Sbjct: 369 RLFEMQSQDYL 379
>gb|AAX69605.1| ATPase, putative [Trypanosoma brucei]
Length = 490
Score = 139 bits (349), Expect = 3e-31
Identities = 116/415 (27%), Positives = 170/415 (40%), Gaps = 59/415 (14%)
Query: 90 KGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAMLRINEHMHKTWKKQMEEKPL 149
KGLY+YG VG GKTMLMD+ Y +K + R HFH ML + +H
Sbjct: 117 KGLYVYGGVGCGKTMLMDLLYENAPSTIK-KHRVHFHHFMLDVQRTLHTV---------- 165
Query: 150 QSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILPDVADKFFLDREGEEKGANI--L 207
E Q+ PD A F D + +N+ L
Sbjct: 166 ----------------------THESKSDGTQVARRTPDAAINSF-DEVAQRMMSNVELL 202
Query: 208 CFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNEANMVPEFFQNLLSNLEE 267
CFDE+ DV + L + G +++ TSNRAP DL + E F + ++
Sbjct: 203 CFDEVAVTDVAHAMILRRLFHAFYKLGVVVIFTSNRAPDDLYLGGLNREGFLPFIELIKR 262
Query: 268 HCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEKKWQDATGRFGGKVISNTIS 327
CE + S D+R A ++ YL PI F +++ G +
Sbjct: 263 QCEVYDMCSNTDHRLSDAGDAKT---YLAPINEANTATFNEQFLQFCK---GMPAERRVL 316
Query: 328 VMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYHTVFISDIPMMSMRIRDKT 387
+FGR +EVP +C GV RF F +CG L AADY +A+ ++TVFI +P
Sbjct: 317 RVFGRDVEVPAACGGVCRFHFMEICGGELSAADYSVIAKTFNTVFIEGVP---------- 366
Query: 388 SLSYFEQQKKKRSLLFRHGDLSLLLTNYTIIILVYVAWHPLQLMNYFKELRRAHFLTWRE 447
+Y K R LL L L + +++Y + L +E AH +
Sbjct: 367 RFTYNSTDVKHRFLL-----LIDELYEHRCKVVIYAQVEIMLLQESKEEFEAAHISSGTT 421
Query: 448 AEGSKLRRDVLAE-GNVGSGGTPVGITSILSGQEELFTFQRAVSRLIEMQTQLYL 501
A + + E + +G S+L + F +R +SRL EM+TQ YL
Sbjct: 422 AATEAEVKPITQEFARLSEFEREIG-RSLLDHTDSAFQMERCLSRLCEMRTQQYL 475
>ref|NP_945546.1| AFG1-like ATPase [Rhodopseudomonas palustris CGA009]
gi|39652895|emb|CAE25637.1| AFG1-like ATPase
[Rhodopseudomonas palustris CGA009]
Length = 394
Score = 135 bits (339), Expect = 4e-30
Identities = 93/316 (29%), Positives = 145/316 (45%), Gaps = 47/316 (14%)
Query: 70 KREKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAM 129
K++ L G P+GLYI+G VG GKTMLMD+F+ A+ +K RR HFHE M
Sbjct: 47 KKQGFFGRLFGGGDAGQAPPQGLYIHGEVGRGKTMLMDLFFDASPIALK--RRSHFHEFM 104
Query: 130 LRINEHMHKTWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILPDV 189
++ ++ +++ ++ I PD
Sbjct: 105 AETHDRING-------------------------------------FRQAIKRGDI-PD- 125
Query: 190 ADKFFLDREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLN 249
AD L A +LCFDE D+ + L + ++L GT++VATSN AP DL
Sbjct: 126 ADVMELTAASILDEAWLLCFDEFHVTDIADAMILGRLFTKLFELGTVVVATSNVAPDDLY 185
Query: 250 EANMVPEFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEKK 309
+ + F + ++ H + + + DYR ++ +L P + E ++
Sbjct: 186 KGGLNRSLFMPFIGQVKRHMRVIRLDARTDYR---LEKFAGMKVWLAPDDAEATATIDRA 242
Query: 310 WQDATGRFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYH 369
W TG G+ +I GR L VP++ VARF+F LC +PLGA+DY+ +A YH
Sbjct: 243 WHRITGTTKGEPRDISIK---GRILHVPQADHHVARFSFADLCQKPLGASDYLRLAHEYH 299
Query: 370 TVFISDIPMMSMRIRD 385
T+ I +P+M R+
Sbjct: 300 TLMIDHVPVMDYADRN 315
>gb|AAF14644.1| 7138.4 [Leishmania major]
Length = 478
Score = 132 bits (332), Expect = 3e-29
Identities = 121/438 (27%), Positives = 188/438 (42%), Gaps = 59/438 (13%)
Query: 70 KREKKLDSLVGRRPTAPPAP----KGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHF 125
++EKK++ + A P KGLY++G VG GKTMLMD+ Y ++ +RR HF
Sbjct: 82 EQEKKVEQALSDDNKAVYHPLSQVKGLYVWGGVGCGKTMLMDLLYDNAPPEIR-KRRLHF 140
Query: 126 HEAMLRINEHMHKTWKKQMEEKPLQSGISSWIMNLPFD-TKAKEWLAAEERYKKEVQMKH 184
H+ ML + + + K EE M P + T + ++ R + +
Sbjct: 141 HQFMLDMQKTSNSIRYKSKEE-----------MQDPANRTNMVRYNTSDNRRRTPDAEIN 189
Query: 185 ILPDVADKFFLDREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRA 244
+ +VA + D E +LCFDE+ DV + L + G +++ TSNR
Sbjct: 190 LFDEVAQRMISDVE-------LLCFDEVAVSDVAHAMILKRLFHSFYKIGLVVIFTSNRP 242
Query: 245 PKDLNEANMVPEFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETIN 304
+DL + + F + +++ C + S +D+R Q YL P+ E +
Sbjct: 243 LEDLYKDGLNRGGFIPFIDLVKKQCIIHHMKSNVDHRLLGHQAD----TYLTPMNSENNS 298
Query: 305 KFEKKWQDATGRFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAV 364
K EK + + + +FGR + VP +C GV F F LCG AADY +
Sbjct: 299 KLEKLFLEMCKAMPA---TERKLEVFGRDVIVPRACGGVCYFDFYELCGGEKSAADYEVI 355
Query: 365 AENYHTVFISDIPMMSMRIRDKTSLSYFEQQKKKRSLLFRHGDLSLLLTNYTIIILVYVA 424
A+ +HT+FI+ +P Y K R LL L L + ++++ A
Sbjct: 356 AKTFHTIFINGVP----------QFPYENSDVKSRFLL-----LIDTLYGHRCKVMIHAA 400
Query: 425 WHPLQLMNYFKELRRAHFLTWREAEGSKLRRDVLAEGNVGSGGTPVGITSILSGQEELFT 484
P QL +E EG R D L+E SG V + + F
Sbjct: 401 VEPPQLQAPKEEAA-------GRIEGDAQRVDQLSEFERESGNRLVDV------DDSAFQ 447
Query: 485 FQRAVSRLIEMQTQLYLD 502
R VSRL EM+T+ YL+
Sbjct: 448 MDRCVSRLFEMRTKEYLE 465
>gb|AAW30013.1| At4g28070 [Arabidopsis thaliana] gi|53850477|gb|AAU95415.1|
At4g28070 [Arabidopsis thaliana]
Length = 473
Score = 132 bits (331), Expect = 4e-29
Identities = 109/413 (26%), Positives = 178/413 (42%), Gaps = 86/413 (20%)
Query: 90 KGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAMLRINEHMHKTWKKQMEEKPL 149
KGLY+YG VG+GKTMLMD+F+ + +R HFH ML ++ + K K +E+ PL
Sbjct: 134 KGLYLYGGVGTGKTMLMDLFFHQLPASWR-TQRIHFHNFMLSVHSRLQK--HKGLED-PL 189
Query: 150 QSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILPDVADKFFLDREGEEKGANILCF 209
+ ++ L ++AD+ A +LC
Sbjct: 190 E------VVGL---------------------------EIADE-----------AILLCL 205
Query: 210 DEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNEANMVPEFFQNLLSNLEEHC 269
DE DV + L+ + L ++G I+VATSNRAP +L E + + F +S L+E C
Sbjct: 206 DEFMVNDVADALILNRLFRHLFNNGIILVATSNRAPDNLYEGGLQRDLFLPFISTLKERC 265
Query: 270 EKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEKKWQDATGRFGGKVISNTISVM 329
+GS +DYR+ + + I ++ ++K+Q G + V+
Sbjct: 266 VVREIGSSVDYRKLTSAEEG-----FYFIGKDISGLLKQKFQLLVG--DQPAGPQVVEVV 318
Query: 330 FGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYHTVFISDIPMMSMRIRDKTSL 389
GR L+VP + +G A F F+ LC RPLGAADY+ + + +HT+ + +P +
Sbjct: 319 MGRKLQVPLAADGCAYFLFEELCDRPLGAADYLGLFKKFHTLALEGVPFFGLH------- 371
Query: 390 SYFEQQKKKRSLLFRHGDLSLLLTNYTIIILVYVAWHPLQLMNYFKELRRAHFLTWREAE 449
R+ +R L ++ +L PL+L+ + A + R +
Sbjct: 372 --------NRTAAYRFVTLVDVMYENKARLLCTAEGSPLELLERIVTISDAQQIAPRTSS 423
Query: 450 GSKLRRDVLAEGNVGSGGTPVGITSILSGQEELFTFQRAVSRLIEMQTQLYLD 502
S+ D + E F R +SRL EM ++ YL+
Sbjct: 424 RSRKSDD----------------PDLCVDNELGFAKDRTISRLTEMNSKEYLE 460
>gb|EAL20923.1| hypothetical protein CNBE2840 [Cryptococcus neoformans var.
neoformans B-3501A]
Length = 709
Score = 114 bits (286), Expect = 6e-24
Identities = 128/505 (25%), Positives = 204/505 (40%), Gaps = 110/505 (21%)
Query: 27 VEKQQNDKDWWKRLNNKITERWTNSRKRPENVDPGVGKWVSYLKREKKLDSLVGRRPTAP 86
+ +Q+ WWK N + E S+ E + V L E++L +L
Sbjct: 139 IAQQKEQTSWWKG-NRNMGEHLGFSKTNGEVEK----RLVKVLSGEEELANL-------- 185
Query: 87 PAPKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYH------FHEAMLRINEHM---H 137
PKG+ + G GSGK++L+ +FY I K R YH + E +N
Sbjct: 186 KTPKGILLTGPPGSGKSLLLSLFYQLLP-ISKKRIHYHAFTLALYKEVFSEMNRRKSSDE 244
Query: 138 KTWKKQMEEKPL--QSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILPDVADKFFL 195
+ W+++ K L + G S +D + KE L K+E + VA +
Sbjct: 245 EEWERKARNKELAGRKGWKSVFAGGRWDEEGKERLVWA---KEEGMAFNSTSLVASQ--- 298
Query: 196 DREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNEANMVP 255
CFDE Q +D + + +LS G +IV SNR P+DL +
Sbjct: 299 ------------CFDEFQLIDASSAALIRDVLSWYWRLGGVIVTCSNRVPEDLYHHGVQK 346
Query: 256 EFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEKKWQDATG 315
E L L+ CE V V D+RR I + + R W ++ + +FE+ W A
Sbjct: 347 ERLAGFLDALKARCEVVQVDGGRDWRREIERGDQIR----WFHSKQQL-EFERAWSAA-- 399
Query: 316 RFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYHTVFISD 375
LE ES RFTF LC LG ADY+A+ + T FI +
Sbjct: 400 ------------------LETQES-GNTCRFTFSQLCEEALGPADYLALVSTFSTFFIDE 440
Query: 376 IPMMSMRIRD-------------KTSLSYFEQQKKKRSLLFRHGDLSL------------ 410
+P + +R ++ ++ F + S LF L+L
Sbjct: 441 VPTLYLRHKNEARRLINLIDALYESRCQVFLRSPSTPSTLFFPDALNLSSQEEETLTNER 500
Query: 411 LLTNYTIIILVYVAWHPLQLMNYFKELRRAHFLTWREAEGSKLRRDVLAEGNVGSGGTPV 470
+++ ++ + V + P ++Y+ L A R+ E ++ +R GT
Sbjct: 501 MMSAESLSATIQVPYRP--NVSYYNNLSPAQ----RDREATEEKRK----------GTSF 544
Query: 471 GITSILSGQEELFTFQRAVSRLIEM 495
+ I +G++E F ++RAVSRLIEM
Sbjct: 545 SVLGIWTGEDEKFAYKRAVSRLIEM 569
>gb|AAW43685.1| conserved hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21] gi|57227225|gb|AAW43684.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21] gi|58267672|ref|XP_570992.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21] gi|58267670|ref|XP_570991.1| conserved
hypothetical protein [Cryptococcus neoformans var.
neoformans JEC21]
Length = 709
Score = 113 bits (283), Expect = 1e-23
Identities = 128/505 (25%), Positives = 203/505 (39%), Gaps = 110/505 (21%)
Query: 27 VEKQQNDKDWWKRLNNKITERWTNSRKRPENVDPGVGKWVSYLKREKKLDSLVGRRPTAP 86
+ +Q+ WWK N + E S+ E + V L E++L +L
Sbjct: 139 IAQQKEQTSWWKG-NRNMGEHLGFSKTNGEVEK----RLVKVLSGEEELANL-------- 185
Query: 87 PAPKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYH------FHEAMLRINEHM---H 137
PKG+ + G GSGK++L+ +FY I K R YH + E +N
Sbjct: 186 KTPKGILLTGPPGSGKSLLLSLFYQLLP-ISKKRIHYHAFTLALYKEVFSEMNRRKSSDE 244
Query: 138 KTWKKQMEEKPL--QSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILPDVADKFFL 195
+ W+++ K L + G S +D + KE L + KE M + FL
Sbjct: 245 EEWERKARNKELAGKKGWKSVFAGGRWDEEGKERLV----WAKEEGMAF------NSTFL 294
Query: 196 DREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNEANMVP 255
CFDE Q +D + + +LS G +IV SNR P+DL +
Sbjct: 295 VASQ--------CFDEFQLIDASSAALIRDVLSWYWRLGGVIVTCSNRVPEDLYHHGVQK 346
Query: 256 EFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEKKWQDATG 315
E L L+ CE V V D+RR I + + R W ++ ++ FE+ W
Sbjct: 347 ERLAGFLDALKARCEVVQVDGGRDWRREIERGDQIR----WFHSKQQLD-FERAW----- 396
Query: 316 RFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYHTVFISD 375
LE ES RFTF LC LG ADY+A+ + T FI +
Sbjct: 397 ---------------SAVLETQES-GNTCRFTFSQLCEEALGPADYLALVSTFSTFFIDE 440
Query: 376 IPMMSMRIRD-------------KTSLSYFEQQKKKRSLLFRHGDLSL------------ 410
+P + +R ++ ++ F + S LF L+L
Sbjct: 441 VPTLYLRHKNEARRLINLIDALYESRCQVFLRSPSTPSTLFFPDALNLSSQEEETLTNER 500
Query: 411 LLTNYTIIILVYVAWHPLQLMNYFKELRRAHFLTWREAEGSKLRRDVLAEGNVGSGGTPV 470
+++ ++ + V + P ++Y+ L A R+ E ++ +R GT
Sbjct: 501 MMSAESLSATIQVPYRP--NVSYYNNLSPAQ----RDREATEEKRK----------GTSF 544
Query: 471 GITSILSGQEELFTFQRAVSRLIEM 495
+ I +G++E F ++RAVSRLIEM
Sbjct: 545 SVLGIWTGEDEKFAYKRAVSRLIEM 569
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.319 0.135 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 921,520,823
Number of Sequences: 2540612
Number of extensions: 40110262
Number of successful extensions: 142342
Number of sequences better than 10.0: 216
Number of HSP's better than 10.0 without gapping: 186
Number of HSP's successfully gapped in prelim test: 30
Number of HSP's that attempted gapping in prelim test: 141492
Number of HSP's gapped (non-prelim): 581
length of query: 532
length of database: 863,360,394
effective HSP length: 133
effective length of query: 399
effective length of database: 525,458,998
effective search space: 209658140202
effective search space used: 209658140202
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 78 (34.7 bits)
Medicago: description of AC147471.10


