

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147406.5 - phase: 0 /pseudo
(99 letters)
Database: nr
2,540,612 sequences; 863,360,394 total letters
Searching..................................................done
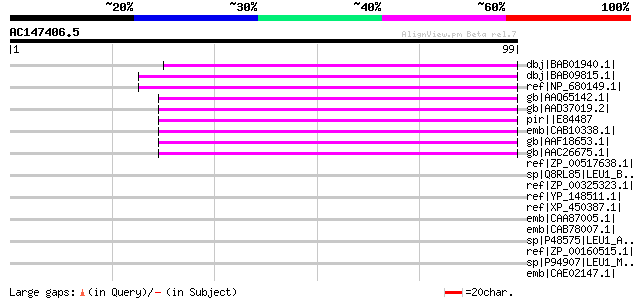
Score E
Sequences producing significant alignments: (bits) Value
dbj|BAB01940.1| non-LTR retrolelement reverse transcriptase-like... 59 4e-08
dbj|BAB09815.1| non-LTR retroelement reverse transcriptase-like ... 59 4e-08
ref|NP_680149.1| reverse transcriptase-related [Arabidopsis thal... 59 4e-08
gb|AAQ65142.1| At5g10850 [Arabidopsis thaliana] gi|8979719|emb|C... 58 7e-08
gb|AAD37019.2| putative non-LTR retrolelement reverse transcript... 57 1e-07
pir||E84487 hypothetical protein At2g07670 [imported] - Arabidop... 57 1e-07
emb|CAB10338.1| hypothetical protein [Arabidopsis thaliana] gi|7... 56 2e-07
gb|AAF18653.1| hypothetical protein [Arabidopsis thaliana] gi|15... 52 3e-06
gb|AAC26675.1| putative Ta11-like non-LTR retroelement protein [... 52 3e-06
ref|ZP_00517638.1| 2-isopropylmalate synthase [Crocosphaera wats... 38 0.071
sp|Q8RL85|LEU1_BACST 2-isopropylmalate synthase (Alpha-isopropyl... 37 0.092
ref|ZP_00325323.1| COG0119: Isopropylmalate/homocitrate/citramal... 36 0.21
ref|YP_148511.1| 2-isopropylmalate synthase [Geobacillus kaustop... 36 0.21
ref|XP_450387.1| hypothetical protein [Oryza sativa (japonica cu... 35 0.35
emb|CAA87005.1| alpha-isopropylmalate synthase [Nostoc sp. PCC 7... 35 0.46
emb|CAB78007.1| putative protein [Arabidopsis thaliana] gi|73210... 35 0.46
sp|P48575|LEU1_ANASP 2-isopropylmalate synthase (Alpha-isopropyl... 35 0.46
ref|ZP_00160515.1| COG0119: Isopropylmalate/homocitrate/citramal... 35 0.46
sp|P94907|LEU1_MICAE 2-isopropylmalate synthase (Alpha-isopropyl... 33 1.3
emb|CAE02147.1| OSJNBa0081G05.2 [Oryza sativa (japonica cultivar... 33 1.7
>dbj|BAB01940.1| non-LTR retrolelement reverse transcriptase-like protein
[Arabidopsis thaliana]
Length = 591
Score = 58.5 bits (140), Expect = 4e-08
Identities = 24/69 (34%), Positives = 40/69 (57%)
Query: 31 VKLLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVITGGPWLIFD 90
VK+LG+ + A+ RL++ W+ G ++D F+MV+F++ + +TGGPW F
Sbjct: 83 VKVLGRKVSLAAVSHRLREMWKPSGAMYVLDLPRHFFMVRFEKEEEFLTALTGGPWRAFG 142
Query: 91 HCLAVTHWT 99
CL V W+
Sbjct: 143 SCLLVKAWS 151
>dbj|BAB09815.1| non-LTR retroelement reverse transcriptase-like [Arabidopsis
thaliana]
Length = 676
Score = 58.5 bits (140), Expect = 4e-08
Identities = 24/74 (32%), Positives = 44/74 (59%)
Query: 26 RMQFVVKLLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVITGGP 85
++ VVK+LG+++ A+ +L++ W+ +GG ++D F++V+FD D +TGGP
Sbjct: 92 KLCMVVKVLGRHVSIAALSRKLRELWKPRGGMYVLDLPRQFFVVRFDVEEDYLAAVTGGP 151
Query: 86 WLIFDHCLAVTHWT 99
W +F L W+
Sbjct: 152 WRLFGSVLMGQAWS 165
>ref|NP_680149.1| reverse transcriptase-related [Arabidopsis thaliana]
Length = 594
Score = 58.5 bits (140), Expect = 4e-08
Identities = 24/74 (32%), Positives = 44/74 (59%)
Query: 26 RMQFVVKLLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVITGGP 85
++ VVK+LG+++ A+ +L++ W+ +GG ++D F++V+FD D +TGGP
Sbjct: 92 KLCMVVKVLGRHVSIAALSRKLRELWKPRGGMYVLDLPRQFFVVRFDVEEDYLAAVTGGP 151
Query: 86 WLIFDHCLAVTHWT 99
W +F L W+
Sbjct: 152 WRLFGSVLMGQAWS 165
>gb|AAQ65142.1| At5g10850 [Arabidopsis thaliana] gi|8979719|emb|CAB96840.1|
putative protein [Arabidopsis thaliana]
gi|15238283|ref|NP_196646.1| expressed protein
[Arabidopsis thaliana] gi|51970522|dbj|BAD43953.1|
putative protein [Arabidopsis thaliana]
gi|11358289|pir||T50794 hypothetical protein T30N20_120
- Arabidopsis thaliana
Length = 181
Score = 57.8 bits (138), Expect = 7e-08
Identities = 22/70 (31%), Positives = 41/70 (58%)
Query: 30 VVKLLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVITGGPWLIF 89
+VK+ G+N+ A+ +L++ W+ +G ++D F+M++F+ + +TGGPW F
Sbjct: 98 IVKVPGRNMTISALSRKLREMWKPRGAMSVVDLPRKFFMIRFEAEEEYMAALTGGPWRAF 157
Query: 90 DHCLAVTHWT 99
D L V W+
Sbjct: 158 DRYLMVQAWS 167
>gb|AAD37019.2| putative non-LTR retrolelement reverse transcriptase [Arabidopsis
thaliana]
Length = 855
Score = 56.6 bits (135), Expect = 1e-07
Identities = 24/70 (34%), Positives = 40/70 (56%)
Query: 30 VVKLLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVITGGPWLIF 89
VVK+LG+N+ + RL++ W+ QG ++D F+M++F+ + +TGGPW F
Sbjct: 57 VVKVLGRNISIANLNRRLREMWKPQGAMFVVDLPCQFFMIRFEREDEYLSALTGGPWRAF 116
Query: 90 DHCLAVTHWT 99
L V W+
Sbjct: 117 GSYLLVQAWS 126
>pir||E84487 hypothetical protein At2g07670 [imported] - Arabidopsis thaliana
Length = 913
Score = 56.6 bits (135), Expect = 1e-07
Identities = 24/70 (34%), Positives = 40/70 (56%)
Query: 30 VVKLLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVITGGPWLIF 89
VVK+LG+N+ + RL++ W+ QG ++D F+M++F+ + +TGGPW F
Sbjct: 57 VVKVLGRNISIANLNRRLREMWKPQGAMFVVDLPCQFFMIRFEREDEYLSALTGGPWRAF 116
Query: 90 DHCLAVTHWT 99
L V W+
Sbjct: 117 GSYLLVQAWS 126
>emb|CAB10338.1| hypothetical protein [Arabidopsis thaliana]
gi|7268308|emb|CAB78602.1| hypothetical protein
[Arabidopsis thaliana] gi|7485201|pir||H71420
hypothetical protein - Arabidopsis thaliana
Length = 655
Score = 56.2 bits (134), Expect = 2e-07
Identities = 23/70 (32%), Positives = 38/70 (53%)
Query: 30 VVKLLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVITGGPWLIF 89
+VK+LG++ + +L+ WR GG ++D F+MV+F+ D +TGGPW +
Sbjct: 97 LVKILGRHTTIEVLSRKLRDLWRPTGGMSVLDLPRQFFMVRFEVEEDYMMALTGGPWRVL 156
Query: 90 DHCLAVTHWT 99
L V W+
Sbjct: 157 GSILMVQAWS 166
>gb|AAF18653.1| hypothetical protein [Arabidopsis thaliana]
gi|15226168|ref|NP_178215.1| hypothetical protein
[Arabidopsis thaliana] gi|25410873|pir||A84420
hypothetical protein At2g01050 [imported] - Arabidopsis
thaliana
Length = 515
Score = 52.4 bits (124), Expect = 3e-06
Identities = 20/70 (28%), Positives = 38/70 (53%)
Query: 30 VVKLLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVITGGPWLIF 89
+VK+LG + + +L++ W+ G +MD F+M++F+ + +TGGPW +
Sbjct: 82 IVKVLGSQIPISVLNRKLRELWKPSGVMTVMDLPRQFFMIRFELEEEYMAALTGGPWRVL 141
Query: 90 DHCLAVTHWT 99
+ L V W+
Sbjct: 142 GNYLLVQDWS 151
>gb|AAC26675.1| putative Ta11-like non-LTR retroelement protein [Arabidopsis
thaliana] gi|15226160|ref|NP_178817.1| zinc knuckle
(CCHC-type) family protein [Arabidopsis thaliana]
gi|25371846|pir||D84488 probable Ta11-like non-LTR
retroelement protein [imported] - Arabidopsis thaliana
Length = 409
Score = 52.4 bits (124), Expect = 3e-06
Identities = 22/70 (31%), Positives = 37/70 (52%)
Query: 30 VVKLLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVITGGPWLIF 89
VVK+LG+++ + RL++ W+ GG +++ F+MVKFD D G PW +
Sbjct: 34 VVKVLGRHITIADLNRRLRELWKPNGGMSVLNLPRSFFMVKFDLEEDYLAAAIGDPWRVL 93
Query: 90 DHCLAVTHWT 99
L + W+
Sbjct: 94 GSILMIRAWS 103
>ref|ZP_00517638.1| 2-isopropylmalate synthase [Crocosphaera watsonii WH 8501]
gi|67853951|gb|EAM49269.1| 2-isopropylmalate synthase
[Crocosphaera watsonii WH 8501]
Length = 537
Score = 37.7 bits (86), Expect = 0.071
Identities = 22/50 (44%), Positives = 29/50 (58%), Gaps = 5/50 (10%)
Query: 33 LLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVIT 82
+LGK G +A R RL + GFE+ DND ++F E ADK+K IT
Sbjct: 344 VLGKLSGRNAFRTRLSEL-----GFELSDNDLNNAFLRFKEVADKKKEIT 388
>sp|Q8RL85|LEU1_BACST 2-isopropylmalate synthase (Alpha-isopropylmalate synthase)
(Alpha-IPM synthetase) gi|19918935|gb|AAL99359.1|
2-isopropylmalate synthase [Geobacillus
stearothermophilus]
Length = 515
Score = 37.4 bits (85), Expect = 0.092
Identities = 20/50 (40%), Positives = 30/50 (60%), Gaps = 5/50 (10%)
Query: 33 LLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVIT 82
+LGKN G HA+R R+++ G+ + D + V+F E ADK+K IT
Sbjct: 326 VLGKNSGRHALRYRVEEL-----GYTLSDEEINQLFVRFKELADKKKDIT 370
>ref|ZP_00325323.1| COG0119: Isopropylmalate/homocitrate/citramalate synthases
[Trichodesmium erythraeum IMS101]
Length = 541
Score = 36.2 bits (82), Expect = 0.21
Identities = 20/50 (40%), Positives = 29/50 (58%), Gaps = 5/50 (10%)
Query: 33 LLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVIT 82
+LGK G HA + RLQ+ GF++ D + +KF + ADK+K IT
Sbjct: 344 VLGKLSGRHAFQSRLQEL-----GFKLSDEEINKAFIKFKDLADKKKEIT 388
>ref|YP_148511.1| 2-isopropylmalate synthase [Geobacillus kaustophilus HTA426]
gi|56381035|dbj|BAD76943.1| 2-isopropylmalate synthase
[Geobacillus kaustophilus HTA426]
Length = 515
Score = 36.2 bits (82), Expect = 0.21
Identities = 19/50 (38%), Positives = 30/50 (60%), Gaps = 5/50 (10%)
Query: 33 LLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVIT 82
+LGK+ G HA+R R+++ G+ + D + V+F E ADK+K IT
Sbjct: 326 VLGKHSGRHALRNRVEEL-----GYTLSDEEINQLFVRFKELADKKKDIT 370
>ref|XP_450387.1| hypothetical protein [Oryza sativa (japonica cultivar-group)]
gi|49389024|dbj|BAD26267.1| hypothetical protein [Oryza
sativa (japonica cultivar-group)]
Length = 417
Score = 35.4 bits (80), Expect = 0.35
Identities = 13/45 (28%), Positives = 24/45 (52%)
Query: 42 AMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVITGGPW 86
++R +L + W G +G ++++F E AD +V+ G PW
Sbjct: 64 SLRTKLLRDWNPSGAVNFWRATDGAFLIQFYERADLARVLDGAPW 108
>emb|CAA87005.1| alpha-isopropylmalate synthase [Nostoc sp. PCC 7120]
Length = 531
Score = 35.0 bits (79), Expect = 0.46
Identities = 20/50 (40%), Positives = 30/50 (60%), Gaps = 5/50 (10%)
Query: 33 LLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVIT 82
+LGK+ G +A R RL++ GFE+ + + VKF E ADK+K I+
Sbjct: 344 VLGKHSGRNAFRTRLKEL-----GFELSETELNKAFVKFKEVADKKKEIS 388
>emb|CAB78007.1| putative protein [Arabidopsis thaliana]
gi|7321071|emb|CAB82118.1| putative protein
[Arabidopsis thaliana] gi|15236612|ref|NP_192622.1|
hypothetical protein [Arabidopsis thaliana]
gi|25371845|pir||G85088 hypothetical protein AT4g08820
[imported] - Arabidopsis thaliana
Length = 404
Score = 35.0 bits (79), Expect = 0.46
Identities = 16/53 (30%), Positives = 28/53 (52%), Gaps = 9/53 (16%)
Query: 56 GFEIMDNDNG---------FYMVKFDEAADKEKVITGGPWLIFDHCLAVTHWT 99
G E+++ NG F+M++F+ + +TGGPW +F + L V W+
Sbjct: 46 GEEVLEAMNGLWKRCMIVKFFMIRFELEEEYLAALTGGPWRVFGNYLMVQDWS 98
>sp|P48575|LEU1_ANASP 2-isopropylmalate synthase (Alpha-isopropylmalate synthase)
(Alpha-IPM synthetase) gi|17133977|dbj|BAB76539.1|
2-isopropylmalate synthase [Nostoc sp. PCC 7120]
gi|17232332|ref|NP_488880.1| 2-isopropylmalate synthase
[Nostoc sp. PCC 7120]
Length = 531
Score = 35.0 bits (79), Expect = 0.46
Identities = 20/50 (40%), Positives = 30/50 (60%), Gaps = 5/50 (10%)
Query: 33 LLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVIT 82
+LGK+ G +A R RL++ GFE+ + + VKF E ADK+K I+
Sbjct: 344 VLGKHSGRNAFRTRLKEL-----GFELSETELNKAFVKFKEVADKKKEIS 388
>ref|ZP_00160515.1| COG0119: Isopropylmalate/homocitrate/citramalate synthases
[Anabaena variabilis ATCC 29413]
Length = 531
Score = 35.0 bits (79), Expect = 0.46
Identities = 20/50 (40%), Positives = 30/50 (60%), Gaps = 5/50 (10%)
Query: 33 LLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVIT 82
+LGK+ G +A R RL++ GFE+ + + VKF E ADK+K I+
Sbjct: 344 VLGKHSGRNAFRTRLKEL-----GFELSETELNKAFVKFKEVADKKKEIS 388
>sp|P94907|LEU1_MICAE 2-isopropylmalate synthase (Alpha-isopropylmalate synthase)
(Alpha-IPM synthetase) gi|1783285|dbj|BAA12849.1|
2-isopropylmalate synthase [Microcystis aeruginosa]
Length = 533
Score = 33.5 bits (75), Expect = 1.3
Identities = 19/50 (38%), Positives = 29/50 (58%), Gaps = 5/50 (10%)
Query: 33 LLGKNLGYHAMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVIT 82
+LGK G +A R RLQ+ GFE+ + + ++F E AD++K IT
Sbjct: 345 VLGKLSGRNAFRSRLQEL-----GFELSETELNNAFIQFKEMADRKKEIT 389
>emb|CAE02147.1| OSJNBa0081G05.2 [Oryza sativa (japonica cultivar-group)]
gi|50923489|ref|XP_472105.1| OSJNBa0081G05.2 [Oryza
sativa (japonica cultivar-group)]
Length = 1711
Score = 33.1 bits (74), Expect = 1.7
Identities = 13/46 (28%), Positives = 23/46 (49%)
Query: 42 AMRERLQKTWRTQGGFEIMDNDNGFYMVKFDEAADKEKVITGGPWL 87
A+ E +++ WR + + ++V F D + V+ GGPWL
Sbjct: 184 ALFEDMRRAWRLRAEMSYKSLKDNLFIVSFSAEGDHKFVLQGGPWL 229
Database: nr
Posted date: Jul 5, 2005 12:34 AM
Number of letters in database: 863,360,394
Number of sequences in database: 2,540,612
Lambda K H
0.327 0.141 0.460
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 183,940,965
Number of Sequences: 2540612
Number of extensions: 6866841
Number of successful extensions: 15844
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 26
Number of HSP's successfully gapped in prelim test: 17
Number of HSP's that attempted gapping in prelim test: 15812
Number of HSP's gapped (non-prelim): 43
length of query: 99
length of database: 863,360,394
effective HSP length: 75
effective length of query: 24
effective length of database: 672,814,494
effective search space: 16147547856
effective search space used: 16147547856
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 68 (30.8 bits)
Medicago: description of AC147406.5


